Featured
-
-
Article
| Open Access Reduction in mobility and COVID-19 transmission
Reduction in mobility and COVID-19 transmission
Social distancing policies aiming to reduce COVID-19 transmission have been reflected in reductions in human mobility. Here, the authors show that reduced mobility is correlated with decreased transmission, but that this relationship weakened over time as social distancing measures were relaxed.
- Pierre Nouvellet
- , Sangeeta Bhatia
- & Christl A. Donnelly
-
Article
| Open Access Biological and therapeutic implications of a unique subtype of NPM1 mutated AML
Biological and therapeutic implications of a unique subtype of NPM1 mutated AML
Molecular heterogeneity of acute myeloid leukaemia (AML) across patients is a major challenge for prognosis and therapy. Here, the authors show that NPM1 mutated AML is a heterogeneous class, consisting of two subtypes which exhibit distinct molecular characteristics, differentiation state, patient survival and drug response.
- Arvind Singh Mer
- , Emily M. Heath
- & Benjamin Haibe-Kains
-
Article
| Open Access Loop competition and extrusion model predicts CTCF interaction specificity
Loop competition and extrusion model predicts CTCF interaction specificity
Boundaries of topologically associated domains in genomes are marked by CTCF and cohesin binding. Here the authors predict CTCF interaction specificity by building a simple mathematical model with features including loop competition and extrusion.
- Wang Xi
- & Michael A. Beer
-
Article
| Open Access A scalable physician-level deep learning algorithm detects universal trauma on pelvic radiographs
A scalable physician-level deep learning algorithm detects universal trauma on pelvic radiographs
Pelvic radiographs (PXRs) are essential for detecting proximal femur and pelvis injuries in trauma patients, but none of the currently available algorithms can detect all kinds of trauma-related radiographic findings. Here, the authors develop a multiscale deep learning algorithm trained with weakly supervised point annotation.
- Chi-Tung Cheng
- , Yirui Wang
- & Le Lu
-
Article
| Open Access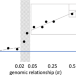 Phenotypic covariance across the entire spectrum of relatedness for 86 billion pairs of individuals
Phenotypic covariance across the entire spectrum of relatedness for 86 billion pairs of individuals
Assigning inter-individual similarities to genetic and non-genetic factors is central to quantitative genetics. Here, the authors look at phenotypic covariance among pairs of individuals for 32 traits across the UK Biobank, from nominally unrelated pairs through to monozygotic twins.
- Kathryn E. Kemper
- , Loic Yengo
- & Peter M. Visscher
-
Article
| Open Access Predicting mammalian hosts in which novel coronaviruses can be generated
Predicting mammalian hosts in which novel coronaviruses can be generated
Homologous recombination between co-infecting coronaviruses can produce novel pathogens. Here, Wardeh et al. develop a machine learning approach to predict associations between mammals and multiple coronaviruses and hence estimate the potential for generation of novel coronaviruses by recombination.
- Maya Wardeh
- , Matthew Baylis
- & Marcus S. C. Blagrove
-
Article
| Open Access Modelling safe protocols for reopening schools during the COVID-19 pandemic in France
Modelling safe protocols for reopening schools during the COVID-19 pandemic in France
The role of children in the spread of COVID-19 is not fully understood, and the circumstances under which schools should be opened are therefore debated. Here, the authors demonstrate protocols by which schools in France can be safely opened without overwhelming the healthcare system.
- Laura Di Domenico
- , Giulia Pullano
- & Vittoria Colizza
-
Article
| Open Access Analysis of metagenome-assembled viral genomes from the human gut reveals diverse putative CrAss-like phages with unique genomic features
Analysis of metagenome-assembled viral genomes from the human gut reveals diverse putative CrAss-like phages with unique genomic features
Here, the authors analyze 4907 Circular Metagenome Assembled Genomes from human microbiomes and identify and characterize nearly 600 diverse genomes of crAss-like phages, finding two putative families with unusual genomic features, including high density of self-splicing introns and inteins.
- Natalya Yutin
- , Sean Benler
- & Eugene V. Koonin
-
Article
| Open Access Causal network models of SARS-CoV-2 expression and aging to identify candidates for drug repurposing
Causal network models of SARS-CoV-2 expression and aging to identify candidates for drug repurposing
Given the severity of the SARS-CoV-2 pandemic, a major challenge is to rapidly repurpose existing approved drugs for clinical interventions. Here, the authors identify robust druggable protein targets within a principled causal framework that makes use of multiple data modalities and integrates aging signatures.
- Anastasiya Belyaeva
- , Louis Cammarata
- & Caroline Uhler
-
Article
| Open Access Fast and precise single-cell data analysis using a hierarchical autoencoder
Fast and precise single-cell data analysis using a hierarchical autoencoder
Accurate analysis of single-cell RNA sequencing (scRNA-seq) data is affected by issues including technical noise and high dropout rate. Here, the authors develop a hierarchical autoencoder, scDHA, which outperforms existing methods in scRNA-seq analyses such as cell segregation and classification.
- Duc Tran
- , Hung Nguyen
- & Tin Nguyen
-
Article
| Open Access PrimeDesign software for rapid and simplified design of prime editing guide RNAs
PrimeDesign software for rapid and simplified design of prime editing guide RNAs
Prime editing guide RNA design is more complex than for standard CRISPR-based nucleases or base editors. Here the authors present PrimeDesign and PrimeVar for the rapid and simplified design of pegRNA and ngRNA combinations.
- Jonathan Y. Hsu
- , Julian Grünewald
- & Luca Pinello
-
Article
| Open Access Machine learning identifies candidates for drug repurposing in Alzheimer’s disease
Machine learning identifies candidates for drug repurposing in Alzheimer’s disease
Clinical trials of novel therapeutics for Alzheimer’s Disease (AD) have provided largely negative results, so far. Here, the authors present a machine learning framework that quantifies potential associations between the pathology of AD severity and gene-based molecular mechanisms to enable drug repurposing.
- Steve Rodriguez
- , Clemens Hug
- & Artem Sokolov
-
Article
| Open Access Rare genetic variants affecting urine metabolite levels link population variation to inborn errors of metabolism
Rare genetic variants affecting urine metabolite levels link population variation to inborn errors of metabolism
Metabolites are indicators of health and disease; genetic studies can reveal variants influencing their levels. Here, the authors investigate the contribution of rare, exonic variants on the levels of urine metabolites and generate predictions on metabolic consequences underlying metabolic disease.
- Yurong Cheng
- , Pascal Schlosser
- & Anna Köttgen
-
Article
| Open Access RNA secondary structure prediction using deep learning with thermodynamic integration
RNA secondary structure prediction using deep learning with thermodynamic integration
Accurately predicting the secondary structure of non-coding RNAs can help unravel their function. Here the authors propose a method integrating thermodynamic information and deep learning to improve the robustness of RNA secondary structure prediction compared to several existing algorithms.
- Kengo Sato
- , Manato Akiyama
- & Yasubumi Sakakibara
-
Article
| Open Access Automatic deep learning-driven label-free image-guided patch clamp system
Automatic deep learning-driven label-free image-guided patch clamp system
Patch clamp recording of neurons is slow and labor-intensive. Here the authors present a method for automated deep learning driven label-free image guided patch clamp physiology to perform measurements on hundreds of human and rodent neurons.
- Krisztian Koos
- , Gáspár Oláh
- & Peter Horvath
-
Article
| Open Access LETR1 is a lymphatic endothelial-specific lncRNA governing cell proliferation and migration through KLF4 and SEMA3C
LETR1 is a lymphatic endothelial-specific lncRNA governing cell proliferation and migration through KLF4 and SEMA3C
Long noncoding RNAs regulate tissue-specific gene expression. Here the authors profile lineage-specific lncRNAs in human dermal lymphatic and blood vascular endothelial cells (LECs and BECs) and show a role of LEC-specific lncRNA, LETR1, in cell proliferation and migration.
- Luca Ducoli
- , Saumya Agrawal
- & Michael Detmar
-
Article
| Open Access Replication dynamics of recombination-dependent replication forks
Replication dynamics of recombination-dependent replication forks
Replication forks that are stalled at obstacles on the DNA template can be restarted by homologous recombination. Here, the authors show replication dynamics during homologous recombination-dependent replication fork restart by combining polymerase usage sequencing and a Monte Carlo mathematical model.
- Karel Naiman
- , Eduard Campillo-Funollet
- & Antony M. Carr
-
Article
| Open Access Forecasting influenza activity using machine-learned mobility map
Forecasting influenza activity using machine-learned mobility map
Human mobility plays a central role in the spread of infectious diseases and can help in forecasting incidence. Here the authors show a comparison of multiple mobility benchmarks in forecasting influenza, and demonstrate the value of a machine-learned mobility map with global coverage at multiple spatial scales.
- Srinivasan Venkatramanan
- , Adam Sadilek
- & Madhav Marathe
-
Article
| Open Access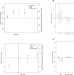 Genetic predictors of participation in optional components of UK Biobank
Genetic predictors of participation in optional components of UK Biobank
Large BioBank studies are commonly used in GWAS, but may be biased by factors affecting participation and dropout. Here the authors show that some of the factors affecting participation may have underlying genetic components.
- Jessica Tyrrell
- , Jie Zheng
- & Kate Tilling
-
Article
| Open Access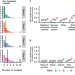 Quantifying population contact patterns in the United States during the COVID-19 pandemic
Quantifying population contact patterns in the United States during the COVID-19 pandemic
Physical distancing measures have been widely adopted to reduce the spread of COVID-19. This study quantifies changes in interpersonal contact patterns in the US and finds an 82% reduction in contacts during early lockdowns in March and steady increases thereafter.
- Dennis M. Feehan
- & Ayesha S. Mahmud
-
Article
| Open Access Genome-wide fine-mapping identifies pleiotropic and functional variants that predict many traits across global cattle populations
Genome-wide fine-mapping identifies pleiotropic and functional variants that predict many traits across global cattle populations
Genomic prediction of phenotype may be improved by using DNA mutations with functional, evolutionary, and pleiotropic consequences. Here the authors describe a method for genome-wide fine-mapping of QTLs and develop a genotyping array for improved prediction of genetic values for cattle traits.
- Ruidong Xiang
- , Iona M. MacLeod
- & Michael E. Goddard
-
Article
| Open Access Assessing the influence of climate on wintertime SARS-CoV-2 outbreaks
Assessing the influence of climate on wintertime SARS-CoV-2 outbreaks
Spread of SARS-CoV-2 in the early phase of the pandemic has been driven by high population susceptibility, but virus sensitivity to climate may play a role in future outbreaks. Here, the authors simulate SARS-CoV-2 dynamics in winter assuming climate dependence is similar to an endemic coronavirus strain.
- Rachel E. Baker
- , Wenchang Yang
- & Bryan T. Grenfell
-
Article
| Open Access Individualized interactomes for network-based precision medicine in hypertrophic cardiomyopathy with implications for other clinical pathophenotypes
Individualized interactomes for network-based precision medicine in hypertrophic cardiomyopathy with implications for other clinical pathophenotypes
Understanding patient-specific pathobiological pathways is a critical step for advancing precision medicine. Here the authors show that individualized protein-protein interaction networks provide key insight on patient-level pathobiology and clinically relevant pathophenotypic characteristics in a complex disease.
- Bradley A. Maron
- , Rui-Sheng Wang
- & Joseph Loscalzo
-
Article
| Open Access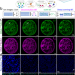 Nanoscopic subcellular imaging enabled by ion beam tomography
Nanoscopic subcellular imaging enabled by ion beam tomography
Secondary ion beam mass spectrometry (SIMS) is a method to obtain a chemical snapshot of biological tissue, but the spatial resolution is low. Here, the authors develop a computational and technology pipeline to localise a chemical signal in SIMS in 3D and sub-25 nm accuracy, called Ion Beam Tomography
- Ahmet F. Coskun
- , Guojun Han
- & Garry P. Nolan
-
Article
| Open Access The molecular basis of socially mediated phenotypic plasticity in a eusocial paper wasp
The molecular basis of socially mediated phenotypic plasticity in a eusocial paper wasp
Connecting genotypes to complex social behaviour is challenging. Taylor et al. use machine learning to show a strong response of caste-associated gene expression to queen loss, wherein individual wasp’s expression profiles become intermediate between queen and worker states, even in the absence of behavioural changes.
- Benjamin A. Taylor
- , Alessandro Cini
- & Seirian Sumner
-
Article
| Open Access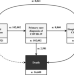 The natural history of symptomatic COVID-19 during the first wave in Catalonia
The natural history of symptomatic COVID-19 during the first wave in Catalonia
Establishing the natural history of COVID-19 requires longitudinal data from population-based cohorts. Here, the authors use linked primary care, testing, and hospital data to describe the disease in ~100,000 individuals with a COVID-19 diagnosis among a population of ~5.5 million in Catalonia, Spain.
- Edward Burn
- , Cristian Tebé
- & Talita Duarte-Salles
-
Article
| Open Access A fast and efficient colocalization algorithm for identifying shared genetic risk factors across multiple traits
A fast and efficient colocalization algorithm for identifying shared genetic risk factors across multiple traits
Statistical colocalisation is a method to identify causal genes and shared genetic aetiology across traits. Here, the authors describe HyPrColoc, an efficient Bayesian divisive clustering algorithm which integrates summary statistics from genome-wide association studies to detect clusters of colocalised traits from large numbers of traits.
- Christopher N. Foley
- , James R. Staley
- & Joanna M. M. Howson
-
Article
| Open Access A practical solution to pseudoreplication bias in single-cell studies
A practical solution to pseudoreplication bias in single-cell studies
Single cell genomics uses cells from the same individual, or pseudoreplicates, that can introduce biases and inflate type I error rates. Here the authors apply generalized linear mixed models with a random effect for individual, to properly account for both zero inflation and the correlation structure among cells within an individual.
- Kip D. Zimmerman
- , Mark A. Espeland
- & Carl D. Langefeld
-
Article
| Open Access PopDel identifies medium-size deletions simultaneously in tens of thousands of genomes
PopDel identifies medium-size deletions simultaneously in tens of thousands of genomes
Identifying structural variants (SVs) from whole genome sequence data has been a significant bioinformatic challenge. Here, the authors describe PopDel, which uses a joint SV detection approach to reliably and efficiently identify 500-10,000 bp deletions across large population cohorts.
- Sebastian Niehus
- , Hákon Jónsson
- & Birte Kehr
-
Article
| Open Access Identification and analysis of splicing quantitative trait loci across multiple tissues in the human genome
Identification and analysis of splicing quantitative trait loci across multiple tissues in the human genome
The profiling of genetic variants affecting splicing can give insight into disease mechanisms. Here, the authors develop a pipeline for discovery of variants affecting splicing (sQTLs) and with application to the GTEx dataset they generate a catalog of human sQTLs.
- Diego Garrido-Martín
- , Beatrice Borsari
- & Roderic Guigó
-
Article
| Open Access Artificial intelligence in sepsis early prediction and diagnosis using unstructured data in healthcare
Artificial intelligence in sepsis early prediction and diagnosis using unstructured data in healthcare
Early prediction and diagnosis of sepsis, which is critical in reducing mortality, is challenging as many of its signs and symptoms are similar to other less critical conditions. Here, the authors develop an artificial intelligence algorithm which uses both structured data and unstructured clinical notes to predict sepsis.
- Kim Huat Goh
- , Le Wang
- & Gamaliel Yu Heng Tan
-
Article
| Open Access Splicing-associated chromatin signatures: a combinatorial and position-dependent role for histone marks in splicing definition
Splicing-associated chromatin signatures: a combinatorial and position-dependent role for histone marks in splicing definition
Chromatin is known to regulate splicing by modulating recruitment of splicing factors. Using machine learning approaches, the authors have underlined a chromatin code for alternative splicing regulation that is conserved amongst cell lines.
- E. Agirre
- , A. J. Oldfield
- & R. F. Luco
-
Article
| Open Access Backbone-independent NMR resonance assignments of methyl probes in large proteins
Backbone-independent NMR resonance assignments of methyl probes in large proteins
Here, the authors present Methyl Assignments Using Satisfiability (MAUS), a method for the assignment of methyl groups using raw NOE data. They use eight proteins in the 10–45 kDa size range as test cases and show that MAUS yields 100% accurate assignments at high completeness levels.
- Santrupti Nerli
- , Viviane S. De Paula
- & Nikolaos G. Sgourakis
-
Article
| Open Access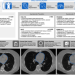 Deep convolutional neural networks to predict cardiovascular risk from computed tomography
Deep convolutional neural networks to predict cardiovascular risk from computed tomography
Coronary artery calcium is an accurate predictor of cardiovascular events but this information is not routinely quantified. Here the authors show a robust and time-efficient deep learning system to automatically quantify coronary calcium on CT scans and predict cardiovascular events in a large, multicentre study.
- Roman Zeleznik
- , Borek Foldyna
- & Hugo J. W. L. Aerts
-
Article
| Open Access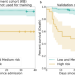 Integrating deep learning CT-scan model, biological and clinical variables to predict severity of COVID-19 patients
Integrating deep learning CT-scan model, biological and clinical variables to predict severity of COVID-19 patients
The SARS-COV-2 pandemic has put pressure on intensive care units, so that predicting severe deterioration early is a priority. Here, the authors develop a multimodal severity score including clinical and imaging features that has significantly improved prognostic performance in two validation datasets compared to previous scores.
- Nathalie Lassau
- , Samy Ammari
- & Michael G. B. Blum
-
Article
| Open Access RUNX1/RUNX1T1 mediates alternative splicing and reorganises the transcriptional landscape in leukemia
RUNX1/RUNX1T1 mediates alternative splicing and reorganises the transcriptional landscape in leukemia
The fusion gene RUNX1/RUNX1T1 is oncogenic in acute myeloid leukemia. Here, the authors show that the fusion gene alters the transcriptional landscape of the cells by changing the structure of the 5’UTR, altering isoform expression, and controlling the expression of splicing factors.
- Vasily V. Grinev
- , Farnaz Barneh
- & Olaf Heidenreich
-
Article
| Open Access Detection of aberrant splicing events in RNA-seq data using FRASER
Detection of aberrant splicing events in RNA-seq data using FRASER
Aberrant splicing is a major contributor to rare disease, but detection accuracy using current methods is limited. Here, the authors develop an algorithm that detects aberrant splicing and intron retention events from RNA-seq data and apply it to diagnosis in mitochondrial disease.
- Christian Mertes
- , Ines F. Scheller
- & Julien Gagneur
-
Article
| Open Access An integrative multiomic network model links lipid metabolism to glucose regulation in coronary artery disease
An integrative multiomic network model links lipid metabolism to glucose regulation in coronary artery disease
Some cholesterol-lowering drugs can increase the risk of type 2 diabetes, but the mechanism behind this is not fully understood. Here the authors show that there is a single genetic regulatory module that influences both cholesterol levels and glucose levels, providing a link between cholesterol levels and diabetes.
- Ariella T. Cohain
- , William T. Barrington
- & Eric E. Schadt
-
Article
| Open Access Sarcoma classification by DNA methylation profiling
Sarcoma classification by DNA methylation profiling
Sarcomas are morphologically heterogeneous tumours rendering their classification challenging. Here the authors developed a classifier using DNA methylation data from several soft tissue and bone sarcoma subtypes, which has the potential to improve classification for research and clinical purposes.
- Christian Koelsche
- , Daniel Schrimpf
- & Andreas von Deimling
-
Article
| Open Access MVP predicts the pathogenicity of missense variants by deep learning
MVP predicts the pathogenicity of missense variants by deep learning
Accurate prediction of variant pathogenicity is essential to understanding genetic risks in disease. Here, the authors present a deep neural network method for prediction of missense variant pathogenicity, MVP, and demonstrate its utility in prioritizing de novo variants contributing to developmental disorders.
- Hongjian Qi
- , Haicang Zhang
- & Yufeng Shen
-
Article
| Open Access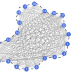 A network-based framework for shape analysis enables accurate characterization of leaf epidermal cells
A network-based framework for shape analysis enables accurate characterization of leaf epidermal cells
While cell shape is crucial for function and development of organisms, versatile frameworks for cell shape quantification, comparison, and classification remain underdeveloped. Here, the authors use a network-based framework for Arabidopsis leaf epidermal cell shape characterization and classification.
- Jacqueline Nowak
- , Ryan Christopher Eng
- & Zoran Nikoloski
-
Article
| Open Access Crystal structure of steroid reductase SRD5A reveals conserved steroid reduction mechanism
Crystal structure of steroid reductase SRD5A reveals conserved steroid reduction mechanism
Steroid 5α-reductase 2 (SRD5A2), a testosterone metabolism enzyme, is implicated in human disease. Structural and biochemical analyses of PbSRD5A, a bacterial homolog, reveal SRD5A2 substrate binding pocket and provide framework for the design of new drugs targeting this enzyme.
- Yufei Han
- , Qian Zhuang
- & Ruobing Ren
-
Article
| Open Access A spatially resolved brain region- and cell type-specific isoform atlas of the postnatal mouse brain
A spatially resolved brain region- and cell type-specific isoform atlas of the postnatal mouse brain
Alternative RNA splicing varies across the brain. Its mapping at single cell resolution is unclear. Here, the authors provide a spatial and single-cell splicing atlas reporting brain region- and cell type-specific expression of different isoforms in the postnatal mouse brain.
- Anoushka Joglekar
- , Andrey Prjibelski
- & Hagen U. Tilgner
-
Article
| Open Access Modelling the global burden of drug-resistant tuberculosis avertable by a post-exposure vaccine
Modelling the global burden of drug-resistant tuberculosis avertable by a post-exposure vaccine
Vaccines preventing tuberculosis disease progression have shown promising results in recent trials. Here, the authors use mathematical modelling to estimate that this type of vaccine could avert 10% of cases of rifampicin-resistant tuberculosis and 7% of deaths from 2020-2035.
- Han Fu
- , Joseph A. Lewnard
- & Nimalan Arinaminpathy
-
Article
| Open Access Mathematical model of COVID-19 intervention scenarios for São Paulo—Brazil
Mathematical model of COVID-19 intervention scenarios for São Paulo—Brazil
Incidence of COVID-19 has been high in parts of South America including Brazil, and information on effective intervention strategies is needed. Here, the authors use mathematical modelling to show that reductions in social distancing should be made gradually to avoid a severe second peak of cases.
- Osmar Pinto Neto
- , Deanna M. Kennedy
- & Renato Amaro Zângaro
-
Article
| Open Access Cellular Heterogeneity–Adjusted cLonal Methylation (CHALM) improves prediction of gene expression
Cellular Heterogeneity–Adjusted cLonal Methylation (CHALM) improves prediction of gene expression
Here, the authors introduce Cell Heterogeneity–Adjusted cLonal Methylation (CHALM) as a methylation quantification method that considers the heterogeneity of sequenced bulk cells. They apply CHALM to methylation datasets to detect differentially methylated genes that exhibit distinct biological functions supporting underlying mechanisms.
- Jianfeng Xu
- , Jiejun Shi
- & Wei Li
-
Article
| Open Access Heme-binding enables allosteric modulation in an ancient TIM-barrel glycosidase
Heme-binding enables allosteric modulation in an ancient TIM-barrel glycosidase
Family 1 glycosidases (GH1) are present in the three domains of life and share classical TIM-barrel fold. Structural and biochemical analyses of a resurrected ancestral GH1 enzyme reveal heme binding, not known in its modern descendants. Heme rigidifies the TIM-barrel and allosterically enhances catalysis.
- Gloria Gamiz-Arco
- , Luis I. Gutierrez-Rus
- & Jose M. Sanchez-Ruiz
-
Article
| Open Access Two distinct catalytic pathways for GH43 xylanolytic enzymes unveiled by X-ray and QM/MM simulations
Two distinct catalytic pathways for GH43 xylanolytic enzymes unveiled by X-ray and QM/MM simulations
Family 43 glycoside hydrolases (GH43) are involved in the breakdown of hemicellulose. Functional, structural and computational characterization of a GH43 enzyme, including a snapshot of an active Michaelis complex, reveal the hydrolysis mechanism and suggest two possible reaction pathways.
- Mariana A. B. Morais
- , Joan Coines
- & Mario T. Murakami
-
Article
| Open Access Deep learning encodes robust discriminative neuroimaging representations to outperform standard machine learning
Deep learning encodes robust discriminative neuroimaging representations to outperform standard machine learning
Recent critical commentaries unfavorably compare deep learning (DL) with standard machine learning (SML) for brain imaging data analysis. Here, the authors show that if trained following prevalent DL practices, DL methods substantially improve compared to SML methods by encoding robust discriminative brain representations.
- Anees Abrol
- , Zening Fu
- & Vince Calhoun
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Aquila enables reference-assisted diploid personal genome assembly and comprehensive variant detection based on linked reads
Aquila enables reference-assisted diploid personal genome assembly and comprehensive variant detection based on linked reads