Featured
-
-
Article
| Open Access Gene-level metagenomic architectures across diseases yield high-resolution microbiome diagnostic indicators
Gene-level metagenomic architectures across diseases yield high-resolution microbiome diagnostic indicators
Here, combing the massive gene-universe of the gut microbiome to identify strain-specific, cross-disease, associations across seven human diseases, the authors introduce the concept of microbiome architecture, defined as the complete set of positive and negative associations between microbial genes and human host disease, highlighting microbiome architectures as potential diagnostic indicators.
- Braden T. Tierney
- , Yingxuan Tan
- & Chirag J. Patel
-
Article
| Open Access A global resource for genomic predictions of antimicrobial resistance and surveillance of Salmonella Typhi at pathogenwatch
A global resource for genomic predictions of antimicrobial resistance and surveillance of Salmonella Typhi at pathogenwatch
Whole genome sequencing data are increasingly becoming routinely available but generating actionable insights is challenging. Here, the authors describe Pathogenwatch, a web tool for genomic surveillance of S. Typhi, and demonstrate its use for antimicrobial resistance assignment and strain risk assessment.
- Silvia Argimón
- , Corin A. Yeats
- & David M. Aanensen
-
Article
| Open Access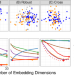 Supervised dimensionality reduction for big data
Supervised dimensionality reduction for big data
Biomedical measurements usually generate high-dimensional data where individual samples are classified in several categories. Vogelstein et al. propose a supervised dimensionality reduction method which estimates the low-dimensional data projection for classification and prediction in big datasets.
- Joshua T. Vogelstein
- , Eric W. Bridgeford
- & Mauro Maggioni
-
Article
| Open Access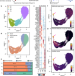 The neutrotime transcriptional signature defines a single continuum of neutrophils across biological compartments
The neutrotime transcriptional signature defines a single continuum of neutrophils across biological compartments
Differentiating neutrophil functional states is difficult. Here the authors show, using single cell RNA-sequencing and trajectory analyses, that mouse neutrophils can be presented as a transcriptome continuum rather than discrete subsets, but are affected by inflammation to express distinct transcriptional states.
- Ricardo Grieshaber-Bouyer
- , Felix A. Radtke
- & Hideyuki Yoshida
-
Article
| Open Access Connectivity characterization of the mouse basolateral amygdalar complex
Connectivity characterization of the mouse basolateral amygdalar complex
The basolateral amygdala is implicated in several behavior-related states including anxiety, autism, and addiction. The authors apply circuit-level pathway tracing methods combined with computational techniques to provide a comprehensive connectivity atlas of the mouse basolateral amygdala complex.
- Houri Hintiryan
- , Ian Bowman
- & Hong-Wei Dong
-
Article
| Open Access AutoSpill is a principled framework that simplifies the analysis of multichromatic flow cytometry data
AutoSpill is a principled framework that simplifies the analysis of multichromatic flow cytometry data
Flow cytometry allows the simultaneous quantification of many markers in and on a cell, but the analysis of such data is complicated. Here, the authors propose AutoSpill, a framework that facilitates the analysis of such data by automating parts of the analysis and requiring fewer controls.
- Carlos P. Roca
- , Oliver T. Burton
- & Adrian Liston
-
Article
| Open Access Hierarchical progressive learning of cell identities in single-cell data
Hierarchical progressive learning of cell identities in single-cell data
Classification methods for scRNA-seq data are limited in their ability to learn from multiple datasets simultaneously. Here the authors present scHPL, a hierarchical progressive learning method that automatically finds relationships between cell populations across multiple datasets and constructs a classification tree.
- Lieke Michielsen
- , Marcel J. T. Reinders
- & Ahmed Mahfouz
-
Article
| Open Access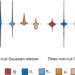 Total genetic contribution assessment across the human genome
Total genetic contribution assessment across the human genome
Quantifying the effects of individual loci on the human phenome is a challenging task. Here, the authors introduce a modelling technique, TGCA, that assesses total genetic contribution per locus and apply this to UK Biobank phenotype domains, revealing top loci and links to tissue-specific gene expression.
- Ting Li
- , Zheng Ning
- & Xia Shen
-
Article
| Open Access Using mobile phone data to reveal risk flow networks underlying the HIV epidemic in Namibia
Using mobile phone data to reveal risk flow networks underlying the HIV epidemic in Namibia
Human mobility influences the spatial distribution of infectious diseases such as HIV. Here, the authors use call data records from mobile phones to model HIV networks in Namibia and estimate that ~40% of the risk of HIV acquisition is driven by mobility.
- Eugenio Valdano
- , Justin T. Okano
- & Sally Blower
-
Article
| Open Access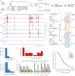 Tissue context determines the penetrance of regulatory DNA variation
Tissue context determines the penetrance of regulatory DNA variation
The functional consequences of variation in human regulatory DNA depend on the local chromatin environment and the cell/tissue context. Here the authors use highly diverged hybrid mice to study genetic effects on DNA accessibility in vivo across multiple cell and tissue types.
- Jessica M. Halow
- , Rachel Byron
- & Matthew T. Maurano
-
Article
| Open Access Pairing a high-resolution statistical potential with a nucleobase-centric sampling algorithm for improving RNA model refinement
Pairing a high-resolution statistical potential with a nucleobase-centric sampling algorithm for improving RNA model refinement
Predicting RNA structure from sequence is challenging due to the relative sparsity of experimentally-determined RNA 3D structures for model training. Here, the authors propose a way to incorporate knowledge on interactions at the atomic and base–base level to refine the prediction of RNA structures.
- Peng Xiong
- , Ruibo Wu
- & Yaoqi Zhou
-
Article
| Open Access Landscape of allele-specific transcription factor binding in the human genome
Landscape of allele-specific transcription factor binding in the human genome
Single-nucleotide variants in enhancers or promoters may affect gene transcription by altering transcription factor binding sites. Here the authors present a meta-analysis empowered by a new statistical method covering thousands of ChIP-Seq experiments resulting in the identification of more than 500 thousand allele-specific binding (ASB) events in the human genome.
- Sergey Abramov
- , Alexandr Boytsov
- & Ivan V. Kulakovskiy
-
Article
| Open Access Distinct axial and lateral interactions within homologous filaments dictate the signaling specificity and order of the AIM2-ASC inflammasome
Distinct axial and lateral interactions within homologous filaments dictate the signaling specificity and order of the AIM2-ASC inflammasome
AIM2-ASC inflammasomes are filamentous signalling platforms that play a central role in host innate defence. Here, the authors present the filament cryo-EM structure of the inflammasome receptor AIM2, which is very similar to the adaptor ASC filament structure. By employing Rosetta and Molecular Dynamics simulations the authors provide further insights into the directionality and recognition mechanisms of the individual AIM2 and ASC filaments, which is further validated with biochemical and cellular experiments.
- Mariusz Matyszewski
- , Weili Zheng
- & Jungsan Sohn
-
Article
| Open Access Estimating COVID-19 mortality in Italy early in the COVID-19 pandemic
Estimating COVID-19 mortality in Italy early in the COVID-19 pandemic
Estimates of COVID-19-related mortality are limited by incomplete testing. Here, the authors perform counterfactual analyses and estimate that there were 59,000–62,000 deaths from COVID-19 in Italy until 9th September 2020, approximately 1.5 times higher than official statistics.
- Chirag Modi
- , Vanessa Böhm
- & Uroš Seljak
-
Article
| Open Access The epidemicity index of recurrent SARS-CoV-2 infections
The epidemicity index of recurrent SARS-CoV-2 infections
Several prognostic indices are available to predict the long-term fate of emerging infectious diseases and the effect of their containment measures, including a variety of reproduction numbers. Here, the authors introduce the epidemicity index, a complementary index to evaluate the potential for transient increases of SARS-Cov-2 epidemics.
- Lorenzo Mari
- , Renato Casagrandi
- & Marino Gatto
-
Article
| Open Access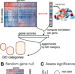 Overcoming false-positive gene-category enrichment in the analysis of spatially resolved transcriptomic brain atlas data
Overcoming false-positive gene-category enrichment in the analysis of spatially resolved transcriptomic brain atlas data
Identifying enriched gene sets in transcriptomic data is routine analysis. Here, the authors show that conventional gene category enrichment analysis (GCEA) applied to brain-wide atlas data yields biased results and develop a flexible ensemble-based null model framework to enable appropriate inference in GCEA.
- Ben D. Fulcher
- , Aurina Arnatkeviciute
- & Alex Fornito
-
Article
| Open Access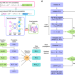 Neural network aided approximation and parameter inference of non-Markovian models of gene expression
Neural network aided approximation and parameter inference of non-Markovian models of gene expression
Cells are complex systems that make decisions biologists struggle to understand. Here, the authors use neural networks to approximate the solution of mathematical models that capture the history and randomness of biochemical processes in order to understand the principles of transcription control.
- Qingchao Jiang
- , Xiaoming Fu
- & Ramon Grima
-
Article
| Open Access Integration of machine learning and genome-scale metabolic modeling identifies multi-omics biomarkers for radiation resistance
Integration of machine learning and genome-scale metabolic modeling identifies multi-omics biomarkers for radiation resistance
Personalized prediction of tumor radiosensitivity would facilitate development of precision medicine workflows for cancer treatment. Here, the authors integrate machine learning and genome-scale metabolic modeling approaches to identify multi-omics biomarkers predictive of radiation response.
- Joshua E. Lewis
- & Melissa L. Kemp
-
Article
| Open Access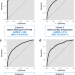 A machine learning model for identifying patients at risk for wild-type transthyretin amyloid cardiomyopathy
A machine learning model for identifying patients at risk for wild-type transthyretin amyloid cardiomyopathy
Transthyretin amyloid cardiomyopathy is a treatable but often unrecognized cause of heart failure. We derived and validated a machine learning model based on medical diagnostic codes that identifies heart failure patients at risk for wild-type transthyretin amyloid cardiomyopathy.
- Ahsan Huda
- , Adam Castaño
- & Sanjiv J. Shah
-
Article
| Open Access The RNA landscape of the human placenta in health and disease
The RNA landscape of the human placenta in health and disease
Placental dysfunction can have catastrophic or barely discernible effects ranging from miscarriage to apparently normal birth. Here the authors present a comprehensive analysis of the human placental transcriptome and identify circular RNAs and piRNAs.
- Sungsam Gong
- , Francesca Gaccioli
- & D. Stephen Charnock-Jones
-
Article
| Open Access SARS-CoV-2 gene content and COVID-19 mutation impact by comparing 44 Sarbecovirus genomes
SARS-CoV-2 gene content and COVID-19 mutation impact by comparing 44 Sarbecovirus genomes
The SARS-CoV-2 gene set remains unresolved, hindering dissection of COVID-19 biology. Comparing 44 Sarbecovirus genomes provides a high-confidence protein-coding gene set. The study characterizes protein-level and nucleotide-level evolutionary constraints, and prioritizes functional mutations from the ongoing COVID-19 pandemic.
- Irwin Jungreis
- , Rachel Sealfon
- & Manolis Kellis
-
Article
| Open Access Artificial intelligence-enabled fully automated detection of cardiac amyloidosis using electrocardiograms and echocardiograms
Artificial intelligence-enabled fully automated detection of cardiac amyloidosis using electrocardiograms and echocardiograms
Cardiac amyloidosis is difficult to identify, given low prevalence and similarity of the symptoms to more prevalent disorders. Here the authors present a multi-modality, artificial intelligence-enabled pipeline, that enables automated detection of cardiac amyloidosis from inexpensive and accessible measures.
- Shinichi Goto
- , Keitaro Mahara
- & Rahul C. Deo
-
Article
| Open Access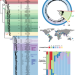 Global population structure and genotyping framework for genomic surveillance of the major dysentery pathogen, Shigella sonnei
Global population structure and genotyping framework for genomic surveillance of the major dysentery pathogen, Shigella sonnei
Whole genome sequencing is increasingly being adopted for Shigella sonnei outbreak investigation and surveillance, but there is no global classification standard. Here, the authors develop and validate a genomic framework implemented using open-source software, and demonstrate its application using surveillance data.
- Jane Hawkey
- , Kalani Paranagama
- & Kathryn E. Holt
-
Article
| Open Access A coordinated progression of progenitor cell states initiates urinary tract development
A coordinated progression of progenitor cell states initiates urinary tract development
Nephric duct (ND)-derived ureteric buds (UB) form the kidney collecting duct system, while ureteric tips promote nephron formation. Here the authors use single-cell RNA-seq and introduce Cluster RNA-seq to identify four progenitor populations in developing ND/UB regulated by the transcription factors Tfap2a/b and Gata3.
- Oraly Sanchez-Ferras
- , Alain Pacis
- & Maxime Bouchard
-
Article
| Open Access Comprehensive cell type decomposition of circulating cell-free DNA with CelFiE
Comprehensive cell type decomposition of circulating cell-free DNA with CelFiE
Tissue damage and turnover lead to the release of DNA in the blood and can be used to monitor changes in tissue state. Here, the authors developed a tool to accurately estimate the proportion of cell types contributing to cell-free DNA in the blood, with an application to pregnant women and ALS patients.
- Christa Caggiano
- , Barbara Celona
- & Noah Zaitlen
-
Article
| Open Access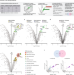 Genoppi is an open-source software for robust and standardized integration of proteomic and genetic data
Genoppi is an open-source software for robust and standardized integration of proteomic and genetic data
Genetic variation can impact protein complexes and interaction networks, but reconciling genetic and proteomic information remains challenging. To address this need, the authors develop Genoppi —a computational tool for integrating genetics and cell-type-specific proteomics data.
- Greta Pintacuda
- , Frederik H. Lassen
- & Kasper Lage
-
Article
| Open Access Deep learning-based predictive identification of neural stem cell differentiation
Deep learning-based predictive identification of neural stem cell differentiation
The differentiation of neural stem cells (NSCs) into neurons is a critical part in devising potential cell-based therapeutic strategies for central nervous system diseases but NSCs fate determination and prediction is problematic. Here, the authors present a deep neural network model for predictable reliable identification of NSCs fate.
- Yanjing Zhu
- , Ruiqi Huang
- & Rongrong Zhu
-
Article
| Open Access Systematic inference and comparison of multi-scale chromatin sub-compartments connects spatial organization to cell phenotypes
Systematic inference and comparison of multi-scale chromatin sub-compartments connects spatial organization to cell phenotypes
Computational algorithms to infer chromatin sub-compartments and compartment domains require high-resolution Hi-C maps. Here the authors present Calder, an algorithm that can infer sub-compartments and compartment domains with variable resolution Hi-C data, and they apply it to more than a hundred Hi-C experiments to study sub-compartment repositioning.
- Yuanlong Liu
- , Luca Nanni
- & Giovanni Ciriello
-
Article
| Open Access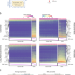 Decoupling epithelial-mesenchymal transitions from stromal profiles by integrative expression analysis
Decoupling epithelial-mesenchymal transitions from stromal profiles by integrative expression analysis
Epithelial cancer cells can transition into a mesenchymal phenotype to enable invasion and metastasis. Here, the authors use previously published single-cell and bulk RNA sequencing datasets to decouple the mesenchymal expression profiles of cancer and stromal cells.
- Michael Tyler
- & Itay Tirosh
-
Article
| Open Access Regression plane concept for analysing continuous cellular processes with machine learning
Regression plane concept for analysing continuous cellular processes with machine learning
High-content screening prompted the development of software enabling discrete phenotypic analysis of single cells. Here, the authors show that supervised continuous machine learning can drive novel discoveries in diverse imaging experiments and present the Regression Plane module of Advanced Cell Classifier.
- Abel Szkalisity
- , Filippo Piccinini
- & Peter Horvath
-
Article
| Open Access CopulaNet: Learning residue co-evolution directly from multiple sequence alignment for protein structure prediction
CopulaNet: Learning residue co-evolution directly from multiple sequence alignment for protein structure prediction
Protein structure prediction is a challenge. A new deep learning framework, CopulaNet, is a major step forward toward end-to-end prediction of inter-residue distances and protein tertiary structures with improved accuracy and efficiency.
- Fusong Ju
- , Jianwei Zhu
- & Dongbo Bu
-
Article
| Open Access Deep generative model embedding of single-cell RNA-Seq profiles on hyperspheres and hyperbolic spaces
Deep generative model embedding of single-cell RNA-Seq profiles on hyperspheres and hyperbolic spaces
Single-cell RNA-seq allows the study of tissues at cellular resolution. Here, the authors demonstrate how deep learning can be used to gain biological insight from such data by accounting for biological and technical variability. Data exploration is improved by accurately visualizing cells on an interactive 3D surface.
- Jiarui Ding
- & Aviv Regev
-
Article
| Open Access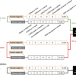 Learning a genome-wide score of human–mouse conservation at the functional genomics level
Learning a genome-wide score of human–mouse conservation at the functional genomics level
Understanding conserved functional genomic properties between human and mouse provides important context for mouse model studies. Here, the authors present a genome-wide conservation score integrating epigenomic, transcription factor binding, and transcriptomic data from mouse and human genomes.
- Soo Bin Kwon
- & Jason Ernst
-
Article
| Open Access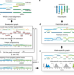 Integrative reconstruction of cancer genome karyotypes using InfoGenomeR
Integrative reconstruction of cancer genome karyotypes using InfoGenomeR
Karyotyping of cancer genomes at the base-level is technically challenging. Here, the authors introduce InfoGenomeR, an algorithm that can infer cancer genome karyotypes from whole-genome sequencing data, and test their model on breast, ovarian and brain cancer samples; and identify private and shared mutations between primary and metastatic cancer samples.
- Yeonghun Lee
- & Hyunju Lee
-
Article
| Open Access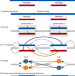 Extended haplotype-phasing of long-read de novo genome assemblies using Hi-C
Extended haplotype-phasing of long-read de novo genome assemblies using Hi-C
Methods to produce haplotype-resolved genome assemblies often rely on access to family trios. The authors present FALCON-Phase, a tool that combines ultra-long range Hi-C chromatin interaction data with a long read de novo assembly to extend haplotype phasing to the contig or scaffold level.
- Zev N. Kronenberg
- , Arang Rhie
- & Sarah B. Kingan
-
Article
| Open Access Genomic insights into the conservation status of the world’s last remaining Sumatran rhinoceros populations
Genomic insights into the conservation status of the world’s last remaining Sumatran rhinoceros populations
Highly endangered species like the Sumatran rhinoceros are at risk from inbreeding. Five historical and 16 modern genomes from across the species range show mutational load, but little evidence for local adaptation, suggesting that future inbreeding depression could be mitigated by assisted gene flow among populations.
- Johanna von Seth
- , Nicolas Dussex
- & Love Dalén
-
Article
| Open Access Protein design and variant prediction using autoregressive generative models
Protein design and variant prediction using autoregressive generative models
The ability to design functional sequences is central to protein engineering and biotherapeutics. Here the authors introduce a deep generative alignment-free model for sequence design applied to highly variable regions and design and test a diverse nanobody library with improved properties for selection experiments.
- Jung-Eun Shin
- , Adam J. Riesselman
- & Debora S. Marks
-
Article
| Open Access The VRNetzer platform enables interactive network analysis in Virtual Reality
The VRNetzer platform enables interactive network analysis in Virtual Reality
Data-rich networks can be difficult to interpret beyond a certain size. Here, the authors introduce a platform that uses virtual reality to allow the visual exploration of large networks, while interfacing with data repositories and other analytical methods to improve the interpretation of big data.
- Sebastian Pirch
- , Felix Müller
- & Jörg Menche
-
Article
| Open Access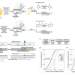 CRISPR-Cas9 cytidine and adenosine base editing of splice-sites mediates highly-efficient disruption of proteins in primary and immortalized cells
CRISPR-Cas9 cytidine and adenosine base editing of splice-sites mediates highly-efficient disruption of proteins in primary and immortalized cells
Base editors can inactivate splice sites or introduce stop codons into a gene sequence. Here the authors present SpliceR to design, rank, and test sgRNAs for efficient gene disruption in T cells.
- Mitchell G. Kluesner
- , Walker S. Lahr
- & Branden S. Moriarity
-
Article
| Open Access Leveraging community mortality indicators to infer COVID-19 mortality and transmission dynamics in Damascus, Syria
Leveraging community mortality indicators to infer COVID-19 mortality and transmission dynamics in Damascus, Syria
Reported COVID-19 mortality rates have been relatively low in Syria, but there has been concern about overwhelmed health systems. Here, the authors use community mortality indicators and estimate that <3% of COVID-19 deaths in Damascus were reported as of 2 September 2020.
- Oliver J. Watson
- , Mervat Alhaffar
- & Patrick Walker
-
Article
| Open Access Machine learning guided aptamer refinement and discovery
Machine learning guided aptamer refinement and discovery
Current aptamer discovery approaches are unable to probe the complete space of possible sequences. Here, the authors use machine learning to facilitate the development of DNA aptamers with improved binding affinities, and truncate them without significantly compromising binding affinity.
- Ali Bashir
- , Qin Yang
- & B. Scott Ferguson
-
Article
| Open Access An integrative analysis of the age-associated multi-omic landscape across cancers
An integrative analysis of the age-associated multi-omic landscape across cancers
Our understanding of the age-related molecular alterations in cancer is still limited. Here, the authors perform a pan-cancer analysis of age-associated genomic, transcriptomic, and epigenetic alterations, linking age-related gene expression changes to age-related DNA methylation alterations
- Kasit Chatsirisupachai
- , Tom Lesluyes
- & João Pedro de Magalhães
-
Article
| Open Access Genomic architecture and prediction of censored time-to-event phenotypes with a Bayesian genome-wide analysis
Genomic architecture and prediction of censored time-to-event phenotypes with a Bayesian genome-wide analysis
Few genome-wide association studies have explored the genetic architecture of age-of-onset for traits and diseases. Here, the authors develop a Bayesian approach to improve prediction in timing-related phenotypes and perform age-of-onset analyses across complex traits in the UK Biobank.
- Sven E. Ojavee
- , Athanasios Kousathanas
- & Matthew R. Robinson
-
Article
| Open Access Spatially interacting phosphorylation sites and mutations in cancer
Spatially interacting phosphorylation sites and mutations in cancer
Dysregulated phosphorylation is well-known in cancers, but it has largely been studied in isolation from mutations. Here the authors introduce HotPho, a tool that can discover spatial interactions between phosphosites and mutations, which are associated with activating mutation and genetic dependencies in cancer.
- Kuan-lin Huang
- , Adam D. Scott
- & Li Ding
-
Article
| Open Access Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Functional RNA secondary structure is important for the pre-mRNA processing including splicing, cleavage and polyadenylation, and RNA editing. Here the authors present a catalog of conserved long-range RNA structures in the human transcriptome by defining pairs of conserved complementary regions (PCCR) in pre-aligned evolutionarily conserved regions.
- Svetlana Kalmykova
- , Marina Kalinina
- & Dmitri Pervouchine
-
Article
| Open Access Detecting protein and DNA/RNA structures in cryo-EM maps of intermediate resolution using deep learning
Detecting protein and DNA/RNA structures in cryo-EM maps of intermediate resolution using deep learning
It is challenging to extract structural information from EM density maps at intermediate or low resolutions. Here, the authors present Emap2sec+, a program for detecting nucleotides and protein secondary structures in EM density maps at 5 to 10 Å resolution.
- Xiao Wang
- , Eman Alnabati
- & Daisuke Kihara
-
Article
| Open Access multiSLIDE is a web server for exploring connected elements of biological pathways in multi-omics data
multiSLIDE is a web server for exploring connected elements of biological pathways in multi-omics data
The integration and interpretation of different omics data types is an ongoing challenge for biologists. Here, the authors present a web-based, interactive tool called multiSLIDE for the visualization of protein, phosphoprotein, and RNA data presented as interlinked heatmaps.
- Soumita Ghosh
- , Abhik Datta
- & Hyungwon Choi
-
Article
| Open Access Design of multi-scale protein complexes by hierarchical building block fusion
Design of multi-scale protein complexes by hierarchical building block fusion
De novo design of self-assembling protein nanostructures and materials is of significant interest, however design of complex, multi-component assemblies is challenging. Here, the authors present a stepwise hierarchical approach to build such assemblies using helical repeat and helical bundle proteins as building blocks, and provide an in-depth structural characterization of the resulting assemblies.
- Yang Hsia
- , Rubul Mout
- & David Baker
-
Article
| Open Access An integrated multi-omics analysis identifies prognostic molecular subtypes of non-muscle-invasive bladder cancer
An integrated multi-omics analysis identifies prognostic molecular subtypes of non-muscle-invasive bladder cancer
Multiple molecular profiling methods are required to study urothelial non-muscle-invasive bladder cancer (NMIBC) due to its heterogeneity. Here the authors integrate multi-omics data of 834 NMIBC patients, identifying a molecular subgroup associated with multiple alterations and worse outcomes.
- Sia Viborg Lindskrog
- , Frederik Prip
- & Lars Dyrskjøt
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Interpretation of T cell states from single-cell transcriptomics data using reference atlases
Interpretation of T cell states from single-cell transcriptomics data using reference atlases