Featured
-
-
Article
| Open Access Rapid detection of identity-by-descent tracts for mega-scale datasets
Rapid detection of identity-by-descent tracts for mega-scale datasets
Traditional methods to identify genomic regions identical-by-descent (IBD) do not scale well to biobank-level datasets. Here, the authors describe a new IBD algorithm, iLASH, which uses LocAlity-Sensitive Hashing to provide rapid IBD estimation when applied to the PAGE and UK Biobank datasets.
- Ruhollah Shemirani
- , Gillian M. Belbin
- & José Luis Ambite
-
Article
| Open Access Cell segmentation-free inference of cell types from in situ transcriptomics data
Cell segmentation-free inference of cell types from in situ transcriptomics data
Inaccurate cell segmentation has been the major problem for cell-type identification and tissue characterization of the in situ spatially resolved transcriptomics data. Here we show a robust cell segmentation-free computational framework (SSAM), for identifying cell types and tissue domains in 2D and 3D.
- Jeongbin Park
- , Wonyl Choi
- & Naveed Ishaque
-
Article
| Open Access Hybrid AI-assistive diagnostic model permits rapid TBS classification of cervical liquid-based thin-layer cell smears
Hybrid AI-assistive diagnostic model permits rapid TBS classification of cervical liquid-based thin-layer cell smears
Technical advancements have significantly improved early diagnosis of cervical cancer, but accurate diagnosis is still difficult due to various practical factors. Here, the authors develop an artificial intelligence assistive diagnostic solution to improve cervical liquid-based thin-layer cell smear diagnosis according to clinical TBS criteria in a large multicenter study.
- Xiaohui Zhu
- , Xiaoming Li
- & Yanqing Ding
-
Article
| Open Access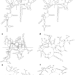 R2DT is a framework for predicting and visualising RNA secondary structure using templates
R2DT is a framework for predicting and visualising RNA secondary structure using templates
Non-coding RNA function is poorly understood, partly due to the challenge of determining RNA secondary (2D) structure. Here, the authors present a framework for the reproducible prediction and visualization of the 2D structure of a wide array of RNAs, which enables linking RNA sequence to function.
- Blake A. Sweeney
- , David Hoksza
- & Anton I. Petrov
-
Article
| Open Access Time trajectories in the transcriptomic response to exercise - a meta-analysis
Time trajectories in the transcriptomic response to exercise - a meta-analysis
Regular exercise promotes overall health and prevents non-communicable diseases, but the adaptation mechanisms are unclear. Here, the authors perform a meta-analysis to reveal time-specific patterns of the acute and long-term exercise response in human skeletal muscle, and identify sex- and age-specific changes.
- David Amar
- , Malene E. Lindholm
- & Euan A. Ashley
-
Article
| Open Access Variant-specific inflation factors for assessing population stratification at the phenotypic variance level
Variant-specific inflation factors for assessing population stratification at the phenotypic variance level
Pooling participant-level genetic data into a single analysis can result in variance stratification, reducing statistical performance. Here, the authors develop variant-specific inflation factors to assess variance stratification and apply this to pooled individual-level data from whole genome sequencing.
- Tamar Sofer
- , Xiuwen Zheng
- & Kenneth M. Rice
-
Article
| Open Access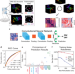 Deep learning connects DNA traces to transcription to reveal predictive features beyond enhancer–promoter contact
Deep learning connects DNA traces to transcription to reveal predictive features beyond enhancer–promoter contact
Recent advances in super-resolution microscopy have made it possible to measure chromatin 3D structure and transcription in thousands of single cells. Here, authors present a deep learning-based approach to characterise how chromatin structure relates to transcriptional state of individual cells and determine which structural features of chromatin regulation are important for gene expression state.
- Aparna R. Rajpurkar
- , Leslie J. Mateo
- & Alistair N. Boettiger
-
Article
| Open Access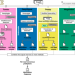 Systematic benchmarking of tools for CpG methylation detection from nanopore sequencing
Systematic benchmarking of tools for CpG methylation detection from nanopore sequencing
Several existing algorithms predict the methylation of DNA using Nanopore sequencing signals, but it is unclear how they compare in performance. Here, the authors benchmark the performance of several such tools, and propose METEORE, a consensus tool that improves prediction accuracy.
- Zaka Wing-Sze Yuen
- , Akanksha Srivastava
- & Eduardo Eyras
-
Article
| Open Access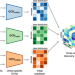 MOGONET integrates multi-omics data using graph convolutional networks allowing patient classification and biomarker identification
MOGONET integrates multi-omics data using graph convolutional networks allowing patient classification and biomarker identification
Our understanding of human disease can be improved by integrating the abundance of high throughput biomedical data. Here, the authors use deep learning methods successfully used on images to integrate various types of omics data to improve patient classification and identify disease biomarkers.
- Tongxin Wang
- , Wei Shao
- & Kun Huang
-
Article
| Open Access Large variation in anti-SARS-CoV-2 antibody prevalence among essential workers in Geneva, Switzerland
Large variation in anti-SARS-CoV-2 antibody prevalence among essential workers in Geneva, Switzerland
Many job sectors classified as ‘essential’ have continued operating with limited restrictions during the COVID-19 pandemic, potentially placing workers at higher risk of infection. Here, the authors show that seropositivity rates in workers vary widely across and between job sectors in Geneva, Switzerland.
- Silvia Stringhini
- , María-Eugenia Zaballa
- & Idris Guessous
-
Article
| Open Access Optimizing vaccine allocation for COVID-19 vaccines shows the potential role of single-dose vaccination
Optimizing vaccine allocation for COVID-19 vaccines shows the potential role of single-dose vaccination
Most COVID-19 vaccines require two doses but a single dose provides partial protection, so it is unclear how best to prioritize vaccine distribution in the context of limited supply. Here, the authors show that campaigns in which some age groups receive one dose while others receive both doses may be optimal.
- Laura Matrajt
- , Julia Eaton
- & Holly Janes
-
Article
| Open Access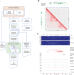 HiC-DC+ enables systematic 3D interaction calls and differential analysis for Hi-C and HiChIP
HiC-DC+ enables systematic 3D interaction calls and differential analysis for Hi-C and HiChIP
The genome-wide investigation of chromatin organization enables insights into global gene expression control. Here, the authors present a computationally efficient method for the analysis of chromatin organization data and use it to recover principles of 3D organization across conditions.
- Merve Sahin
- , Wilfred Wong
- & Christina S. Leslie
-
Article
| Open Access Replicate sequencing libraries are important for quantification of allelic imbalance
Replicate sequencing libraries are important for quantification of allelic imbalance
Allele-specific expression in diploid organisms can be quantified by RNA-seq and it is common practice to rely on a single library. Here, the authors show that the standard approach has variable error rate and present Qllelic as a tool to improve reproducibility of allele-specific RNA-seq analysis.
- Asia Mendelevich
- , Svetlana Vinogradova
- & Alexander A. Gimelbrant
-
Article
| Open Access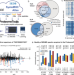 Deep learning boosts sensitivity of mass spectrometry-based immunopeptidomics
Deep learning boosts sensitivity of mass spectrometry-based immunopeptidomics
The identification of HLA peptides by mass spectrometry is non-trivial. Here, the authors extended and used the wealth of data from the ProteomeTools project to improve the prediction of non-tryptic peptides using deep learning, and show their approach enables a variety of immunological discoveries.
- Mathias Wilhelm
- , Daniel P. Zolg
- & Bernhard Kuster
-
Article
| Open Access Quantitative single-cell proteomics as a tool to characterize cellular hierarchies
Quantitative single-cell proteomics as a tool to characterize cellular hierarchies
Single-cell proteomics can provide insights into the molecular basis for cellular heterogeneity. Here, the authors develop a multiplexed single-cell proteomics and computational workflow, and show that their strategy captures the cellular hierarchies in an Acute Myeloid Leukemia culture model.
- Erwin M. Schoof
- , Benjamin Furtwängler
- & Bo T. Porse
-
Article
| Open Access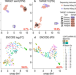 MOCCASIN: a method for correcting for known and unknown confounders in RNA splicing analysis
MOCCASIN: a method for correcting for known and unknown confounders in RNA splicing analysis
Confounding factors on gene expression analysis can be analyzed by several existing tools. Here the authors develop an algorithm called MOCCASIN to correct the effect of known and unknown confounders on RNA splicing quantification.
- Barry Slaff
- , Caleb M. Radens
- & Yoseph Barash
-
Article
| Open Access Systems genetics in diversity outbred mice inform BMD GWAS and identify determinants of bone strength
Systems genetics in diversity outbred mice inform BMD GWAS and identify determinants of bone strength
Osteoporosis GWAS faces two challenges, causal gene discovery and a lack of phenotypic diversity. Here, the authors use the Diversity Outbred mouse population to inform human GWAS using networks and map genetic loci for 55 bone traits, identifying new potential bone strength genes.
- Basel M. Al-Barghouthi
- , Larry D. Mesner
- & Charles R. Farber
-
Article
| Open Access A deep learning approach to identify gene targets of a therapeutic for human splicing disorders
A deep learning approach to identify gene targets of a therapeutic for human splicing disorders
Drugs that modify RNA splicing are promising treatments for many genetic diseases. Here the authors show that deep learning strategies can predict drug targets, strongly supporting the use of in silico approaches to expand the therapeutic potential of drugs that modulate RNA splicing.
- Dadi Gao
- , Elisabetta Morini
- & Susan A. Slaugenhaupt
-
Article
| Open Access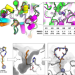 Anchor extension: a structure-guided approach to design cyclic peptides targeting enzyme active sites
Anchor extension: a structure-guided approach to design cyclic peptides targeting enzyme active sites
Cyclic peptides are of particular interest due to their pharmacological properties, but their design for binding to a target protein is challenging. Here, the authors present a computational “anchor extension” methodology for de novo design of cyclic peptides that bind to the target protein with high affinity, and validate the approach by developing cyclic peptides that inhibit histone deacetylases 2 and 6.
- Parisa Hosseinzadeh
- , Paris R. Watson
- & David Baker
-
Article
| Open Access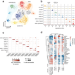 The regulatory landscape of Arabidopsis thaliana roots at single-cell resolution
The regulatory landscape of Arabidopsis thaliana roots at single-cell resolution
Existing studies of the chromatin accessibility, the primary mark of regulatory DNA, in Arabidopsis are based mainly on bulk samples. Here, the authors report the regulatory landscape of Arabidopsis thaliana roots at single-cell resolution.
- Michael W. Dorrity
- , Cristina M. Alexandre
- & Josh T. Cuperus
-
Article
| Open Access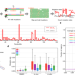 RNA structure probing reveals the structural basis of Dicer binding and cleavage
RNA structure probing reveals the structural basis of Dicer binding and cleavage
Sequencing methods such as icSHAPE were developed to probe RNA structures transcriptome-wide in cells. To probe intact RNA structures, the authors develop icSHAPE-MaP and apply to Dicer-bound substrates showing that distance measuring is important for Dicer cleavage of pre-miRNAs.
- Qing-Jun Luo
- , Jinsong Zhang
- & Qiangfeng Cliff Zhang
-
Article
| Open Access Nuclear compartmentalization of TERT mRNA and TUG1 lncRNA is driven by intron retention
Nuclear compartmentalization of TERT mRNA and TUG1 lncRNA is driven by intron retention
RNA localization plays an important role in transcriptome regulation. The majority of TERT transcripts are detected in the nucleus and TUG1 lncRNAs in both the nucleus and cytoplasm. Here, the authors combine single-cell RNA imaging, antisense oligonucleotides and splicing analyses to show that retention of specific introns drives stable compartmentalization of TERT and TUG1 transcripts in the nucleus, and that splicing of TERT retained introns is mitotically regulated.
- Gabrijela Dumbović
- , Ulrich Braunschweig
- & John L. Rinn
-
Article
| Open Access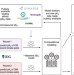 Crowdsourced mapping of unexplored target space of kinase inhibitors
Crowdsourced mapping of unexplored target space of kinase inhibitors
The IDG-DREAM Challenge carried out crowdsourced benchmarking of predictive algorithms for kinase inhibitor activities on unpublished data. This study provides a resource to compare emerging algorithms and prioritize new kinase activities to accelerate drug discovery and repurposing efforts.
- Anna Cichońska
- , Balaguru Ravikumar
- & Tero Aittokallio
-
Article
| Open Access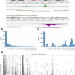 Discovery of widespread transcription initiation at microsatellites predictable by sequence-based deep neural network
Discovery of widespread transcription initiation at microsatellites predictable by sequence-based deep neural network
Mammalian genomes are scattered with repetitive sequences, but their biology remains largely elusive. Here, the authors show that transcription can initiate from short tandem repetitive sequences, and that genetic variants linked to human diseases are preferentially found at repeats with high transcription initiation level.
- Mathys Grapotte
- , Manu Saraswat
- & Charles-Henri Lecellier
-
Article
| Open Access Model-based analysis uncovers mutations altering autophagy selectivity in human cancer
Model-based analysis uncovers mutations altering autophagy selectivity in human cancer
Although autophagy has been linked to tumourigenesis, it is unclear how genomic alterations affect autophagy selectivity in tumours. Here, the authors establish a pipeline that integrates computational and experimental approaches to show that altered autophagy selectivity is frequent in cancer cells and link glycogen autophagy with tumourigenesis.
- Zhu Han
- , Weizhi Zhang
- & Da Jia
-
Article
| Open Access Generative modeling of single-cell time series with PRESCIENT enables prediction of cell trajectories with interventions
Generative modeling of single-cell time series with PRESCIENT enables prediction of cell trajectories with interventions
Single-cell RNA-Seq allows us to observe snapshots of how biological systems change over time at cellular resolution. Here, the authors develop a generative framework that uses time-resolved single-cell data to model how cells change in physical time, including in response to perturbations.
- Grace Hui Ting Yeo
- , Sachit D. Saksena
- & David K. Gifford
-
Article
| Open Access Integrating genomics and metabolomics for scalable non-ribosomal peptide discovery
Integrating genomics and metabolomics for scalable non-ribosomal peptide discovery
Current genome mining methods predict many putative non-ribosomal peptides (NRPs) from their corresponding biosynthetic gene clusters, but it remains unclear which of those exist in nature and how to identify their post-assembly modifications. Here, the authors develop NRPminer, a modification-tolerant tool for the discovery of NRPs from large genomic and mass spectrometry datasets, and use it to find 180 NRPs from different environments.
- Bahar Behsaz
- , Edna Bode
- & Hosein Mohimani
-
Article
| Open Access Enhancing CRISPR-Cas9 gRNA efficiency prediction by data integration and deep learning
Enhancing CRISPR-Cas9 gRNA efficiency prediction by data integration and deep learning
High-quality gRNA activity data is needed for accurate on-target efficiency predictions. Here the authors generate activity data for over 10,000 gRNA and build a deep learning model CRISPRon for improved performance predictions.
- Xi Xiang
- , Giulia I. Corsi
- & Yonglun Luo
-
Article
| Open Access Automated annotation and visualisation of high-resolution spatial proteomic mass spectrometry imaging data using HIT-MAP
Automated annotation and visualisation of high-resolution spatial proteomic mass spectrometry imaging data using HIT-MAP
MALDI-mass spectrometry imaging (MSI) can reveal the distribution of proteins in tissues but tools for protein identification and annotation are sparse. Here, the authors develop an open-source bioinformatic workflow for false discovery rate-controlled protein annotation and spatial mapping from MALDI-MSI data.
- G. Guo
- , M. Papanicolaou
- & A. C. Grey
-
Article
| Open Access Multimodal analysis of cell-free DNA whole-genome sequencing for pediatric cancers with low mutational burden
Multimodal analysis of cell-free DNA whole-genome sequencing for pediatric cancers with low mutational burden
Liquid biopsies enable minimally invasive applications for diagnosis and treatment monitoring. Here the authors analyse fragmentation patterns of circulating tumour DNA on multiple levels and develop a bioinformatic tool, LIQUORICE, to accurately detect and classify paediatric cancers with low mutational burden.
- Peter Peneder
- , Adrian M. Stütz
- & Eleni M. Tomazou
-
Article
| Open Access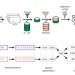 Retention time prediction using neural networks increases identifications in crosslinking mass spectrometry
Retention time prediction using neural networks increases identifications in crosslinking mass spectrometry
Predicting chromatographic retention times (RTs) has proven beneficial in proteomics but has not yet been achieved for crosslinked peptides. Here, the authors develop an RT prediction tool for crosslinked peptides and leverage predicted RTs to increase identifications in crosslinking mass spectrometry studies.
- Sven H. Giese
- , Ludwig R. Sinn
- & Juri Rappsilber
-
Article
| Open Access Impact of DNA methylation on 3D genome structure
Impact of DNA methylation on 3D genome structure
Multi-layered epigenetic regulation in higher eukaryotes makes it challenging to disentangle the individual effects of modifications on chromatin structure and function. Here, the authors expressed mammalian DNA methyltransferases in yeast, which have no DNA methylation, to show that methylation has intrinsic effects on chromatin structure.
- Diana Buitrago
- , Mireia Labrador
- & Modesto Orozco
-
Article
| Open Access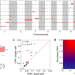 Chromosomal copy number heterogeneity predicts survival rates across cancers
Chromosomal copy number heterogeneity predicts survival rates across cancers
Intratumour heterogeneity (ITH) is associated with worse prognosis in cancer, and efficient frameworks to measure it are needed. Here the authors develop a method to estimate copy number heterogeneity, and propose that it is driven by chromosomal instability and can predict pan-cancer survival.
- Erik van Dijk
- , Tom van den Bosch
- & Daniël M. Miedema
-
Article
| Open Access Structure-based protein function prediction using graph convolutional networks
Structure-based protein function prediction using graph convolutional networks
The rapid increase in the number of proteins in sequence databases and the diversity of their functions challenge computational approaches for automated function prediction. Here, the authors introduce DeepFRI, a Graph Convolutional Network for predicting protein functions by leveraging sequence features extracted from a protein language model and protein structures.
- Vladimir Gligorijević
- , P. Douglas Renfrew
- & Richard Bonneau
-
Article
| Open Access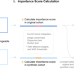 Identification of putative causal loci in whole-genome sequencing data via knockoff statistics
Identification of putative causal loci in whole-genome sequencing data via knockoff statistics
Association analyses that capture rare and noncoding variants in whole genome sequencing data are limited by factors like statistical power. Here, the authors present KnockoffScreen, a statistical method using the knockoff framework to detect, localise and prioritise rare and common risk variants at genome-wide scale.
- Zihuai He
- , Linxi Liu
- & Iuliana Ionita-Laza
-
Article
| Open Access An electronic neuromorphic system for real-time detection of high frequency oscillations (HFO) in intracranial EEG
An electronic neuromorphic system for real-time detection of high frequency oscillations (HFO) in intracranial EEG
A major challenge across a variety of fields is how to process the vast quantities of data produced by sensors without large computation resources. Here, the authors present a neuromorphic chip which can detect a relevant signature of epileptogenic tissue from intracranial recordings in patients.
- Mohammadali Sharifshazileh
- , Karla Burelo
- & Giacomo Indiveri
-
Article
| Open Access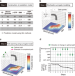 Health improvement framework for actionable treatment planning using a surrogate Bayesian model
Health improvement framework for actionable treatment planning using a surrogate Bayesian model
Clinical decision-making regarding treatments based on personal characteristics leads to effective health improvements. Here, the authors introduce a modeling framework to evaluate the actionability of treatment pathways.
- Kazuki Nakamura
- , Ryosuke Kojima
- & Yasushi Okuno
-
Article
| Open Access Synthetic neural-like computing in microbial consortia for pattern recognition
Synthetic neural-like computing in microbial consortia for pattern recognition
Complex biological systems have individual cells acting collectively to solve complex tasks. Here the authors implement neural network-like computing in a bacterial consortia to recognise patterns.
- Ximing Li
- , Luna Rizik
- & Ramez Daniel
-
Article
| Open Access Detecting and phasing minor single-nucleotide variants from long-read sequencing data
Detecting and phasing minor single-nucleotide variants from long-read sequencing data
Cellular genetic heterogeneity is common across biological conditions, yet application of long-read sequencing to this subject is limited by error rates. Here, the authors present iGDA, a tool for detection and phasing of minor variants from long-read sequencing data, allowing accurate reconstruction of haplotypes.
- Zhixing Feng
- , Jose C. Clemente
- & Eric E. Schadt
-
Article
| Open Access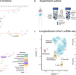 Evolution of core archetypal phenotypes in progressive high grade serous ovarian cancer
Evolution of core archetypal phenotypes in progressive high grade serous ovarian cancer
High-grade serous ovarian cancer (HGSOC) is prone to developing resistance to treatment. Here, the authors use single-cell RNA-seq and an analysis of archetypes, and find that shifts in metabolism and proliferation are associated with the response to treatment and clonal heterogeneity in HGSOC.
- Aritro Nath
- , Patrick A. Cosgrove
- & Andrea H. Bild
-
Article
| Open Access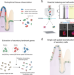 Clump sequencing exposes the spatial expression programs of intestinal secretory cells
Clump sequencing exposes the spatial expression programs of intestinal secretory cells
Combining scRNA-seq with spatial information to enable the reconstruction of spatially-resolved cell atlases is challenging for rare cell types. Here the authors present ClumpSeq, an approach for sequencing small clumps of tissue attached cells, and apply it to establish spatial atlases for all secretory cell types in the small intestine.
- Rita Manco
- , Inna Averbukh
- & Shalev Itzkovitz
-
Article
| Open Access Mining mutation contexts across the cancer genome to map tumor site of origin
Mining mutation contexts across the cancer genome to map tumor site of origin
The vast majority of somatic mutations observed in tumors are rare. Here, the authors show that these large numbers of rare mutations are more predictive of the tissue of origin of a tumor than the information from a few common driver mutations.
- Saptarshi Chakraborty
- , Axel Martin
- & Ronglai Shen
-
Article
| Open Access Permutation-based identification of important biomarkers for complex diseases via machine learning models
Permutation-based identification of important biomarkers for complex diseases via machine learning models
Study of human disease remains challenging due to convoluted disease etiologies and complex molecular mechanisms at genetic, genomic, and proteomic levels. Here, the authors propose a computationally efficient Permutation-based Feature Importance Test to assist interpretation and selection of individual features in complex machine learning models for complex disease analysis.
- Xinlei Mi
- , Baiming Zou
- & Jianhua Hu
-
Article
| Open Access Controlling COVID-19 via test-trace-quarantine
Controlling COVID-19 via test-trace-quarantine
Initial COVID-19 containment in the United States focused on limiting mobility, including school and workplace closures, with enormous societal and economic costs. Here, the authors demonstrate the feasibility of a test-trace-quarantine strategy using an agent-based model and detailed data on the Seattle region.
- Cliff C. Kerr
- , Dina Mistry
- & Daniel J. Klein
-
Article
| Open Access Interpretation of T cell states from single-cell transcriptomics data using reference atlases
Interpretation of T cell states from single-cell transcriptomics data using reference atlases
One challenge of single cell RNA sequencing analysis is how to consistently identify cell subtypes and states across different datasets. Here the authors propose the use of a reference single-cell atlas as a stable system of coordinates to characterize T cell states across studies, diseases and species.
- Massimo Andreatta
- , Jesus Corria-Osorio
- & Santiago J. Carmona
-
Article
| Open Access Gene-level metagenomic architectures across diseases yield high-resolution microbiome diagnostic indicators
Gene-level metagenomic architectures across diseases yield high-resolution microbiome diagnostic indicators
Here, combing the massive gene-universe of the gut microbiome to identify strain-specific, cross-disease, associations across seven human diseases, the authors introduce the concept of microbiome architecture, defined as the complete set of positive and negative associations between microbial genes and human host disease, highlighting microbiome architectures as potential diagnostic indicators.
- Braden T. Tierney
- , Yingxuan Tan
- & Chirag J. Patel
-
Article
| Open Access A global resource for genomic predictions of antimicrobial resistance and surveillance of Salmonella Typhi at pathogenwatch
A global resource for genomic predictions of antimicrobial resistance and surveillance of Salmonella Typhi at pathogenwatch
Whole genome sequencing data are increasingly becoming routinely available but generating actionable insights is challenging. Here, the authors describe Pathogenwatch, a web tool for genomic surveillance of S. Typhi, and demonstrate its use for antimicrobial resistance assignment and strain risk assessment.
- Silvia Argimón
- , Corin A. Yeats
- & David M. Aanensen
-
Article
| Open Access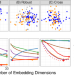 Supervised dimensionality reduction for big data
Supervised dimensionality reduction for big data
Biomedical measurements usually generate high-dimensional data where individual samples are classified in several categories. Vogelstein et al. propose a supervised dimensionality reduction method which estimates the low-dimensional data projection for classification and prediction in big datasets.
- Joshua T. Vogelstein
- , Eric W. Bridgeford
- & Mauro Maggioni
-
Article
| Open Access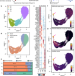 The neutrotime transcriptional signature defines a single continuum of neutrophils across biological compartments
The neutrotime transcriptional signature defines a single continuum of neutrophils across biological compartments
Differentiating neutrophil functional states is difficult. Here the authors show, using single cell RNA-sequencing and trajectory analyses, that mouse neutrophils can be presented as a transcriptome continuum rather than discrete subsets, but are affected by inflammation to express distinct transcriptional states.
- Ricardo Grieshaber-Bouyer
- , Felix A. Radtke
- & Hideyuki Yoshida
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Machine-learning predicts genomic determinants of meiosis-driven structural variation in a eukaryotic pathogen
Machine-learning predicts genomic determinants of meiosis-driven structural variation in a eukaryotic pathogen