Featured
-
-
Article
| Open Access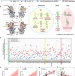 Tracing genetic diversity captures the molecular basis of misfolding disease
Tracing genetic diversity captures the molecular basis of misfolding disease
Pei et al. applied Gaussian process-based machine learning to capture dynamic spatial covariance relationships managed by proteostasis to mediate cooperative folding on a residue basis as a standard model for precision disease management.
- Pei Zhao
- , Chao Wang
- & William E. Balch
-
Article
| Open Access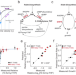 The genetic landscape of a metabolic interaction
The genetic landscape of a metabolic interaction
Reynolds and colleagues examine a biochemically-mediated epistatic interaction between metabolic enzymes involved in folate metabolism and show that biochemical coupling shapes the range of enzyme activities sufficient to rescue cell growth.
- Thuy N. Nguyen
- , Christine Ingle
- & Kimberly A. Reynolds
-
Article
| Open Access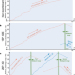 Data-driven recombination detection in viral genomes
Data-driven recombination detection in viral genomes
Here, the authors present RecombinHunt, a computational method based on big data analysis, that enhances community-based detection of recombinant viral lineages.
- Tommaso Alfonsi
- , Anna Bernasconi
- & Stefano Ceri
-
Article
| Open Access Phylogenomic profiles of whole-genome duplications in Poaceae and landscape of differential duplicate retention and losses among major Poaceae lineages
Phylogenomic profiles of whole-genome duplications in Poaceae and landscape of differential duplicate retention and losses among major Poaceae lineages
Grasses share a whole-genome duplication called rho, but the adaptive implications are unclear. Here, the authors conduct phylogenomic and phylotranscriptomic analyses of 363 grasses, identifying additional whole-genome duplications and finding that duplicates are implicated in environmental adaptations or morphogenesis.
- Taikui Zhang
- , Weichen Huang
- & Hong Ma
-
Article
| Open Access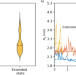 Tertiary structure and conformational dynamics of the anti-amyloidogenic chaperone DNAJB6b at atomistic resolution
Tertiary structure and conformational dynamics of the anti-amyloidogenic chaperone DNAJB6b at atomistic resolution
Adupa et al show how the anti-amyloidogenic molecular chaperone DNAJB6 adopts three conformational states that determine the accessibility of its substrate binding domain. In all states, interactions with HSP70 are shielded, suggesting that functional interactions only may occur upon substrate binding.
- Vasista Adupa
- , Elizaveta Ustyantseva
- & Patrick R. Onck
-
Article
| Open Access Microenvironmental reorganization in brain tumors following radiotherapy and recurrence revealed by hyperplexed immunofluorescence imaging
Microenvironmental reorganization in brain tumors following radiotherapy and recurrence revealed by hyperplexed immunofluorescence imaging
Improved imaging techniques are required to help advance our understanding of the complex role of the tumour microenvironment (TME). Here, the authors develop a high-throughput, highly multiplexed tissue visualisation workflow and demonstrate its utility by characterising the response of the TME to radiotherapy in preclinical models of glioblastoma.
- Spencer S. Watson
- , Benoit Duc
- & Johanna A. Joyce
-
Article
| Open Access One substrate many enzymes virtual screening uncovers missing genes of carnitine biosynthesis in human and mouse
One substrate many enzymes virtual screening uncovers missing genes of carnitine biosynthesis in human and mouse
With structural models now available on a proteome scale, Malatesta et al. show that structure-based screening can help identify proteins catalyzing orphan reactions in metabolic pathways, offering functional insights beyond sequence-based approaches.
- Marco Malatesta
- , Emanuele Fornasier
- & Riccardo Percudani
-
Article
| Open Access Deep learning predictions of TCR-epitope interactions reveal epitope-specific chains in dual alpha T cells
Deep learning predictions of TCR-epitope interactions reveal epitope-specific chains in dual alpha T cells
Prediction of the specificity of a T cell receptor from amino acid sequence has been performed using different methods and approaches. Here the authors use TCRab sequences with known specificity to develop a deep learning TCR-epitope interaction predictor and use this method to predict specificity of dual alpha chain TCRs and TCRs specific for different antigens.
- Giancarlo Croce
- , Sara Bobisse
- & David Gfeller
-
Article
| Open Access Integrated proteogenomic and metabolomic characterization of papillary thyroid cancer with different recurrence risks
Integrated proteogenomic and metabolomic characterization of papillary thyroid cancer with different recurrence risks
Papillary thyroid cancers (PTC) generally have good prognosis, but their recurrence rate remains high. Here, the authors use proteogenomics and metabolomics to identify molecular features in PTC tumours and determine PTC subtypes that are associated with prognosis and potential targeted therapies.
- Ning Qu
- , Di Chen
- & Rongliang Shi
-
Article
| Open Access Systematic investigation of chemo-immunotherapy synergism to shift anti-PD-1 resistance in cancer
Systematic investigation of chemo-immunotherapy synergism to shift anti-PD-1 resistance in cancer
The design of new combinatorial regimens represents an opportunity to improve response to immune checkpoint inhibitors in patients with cancer. Here the authors computationally model the interaction between chemotherapy and immunotherapy by studying treatment-induced expression changes associated with response to anti-PD-1, identifying chemotherapeutic drugs or small molecule inhibitors that can overcome resistance to anti-PD-1.
- Yue Wang
- , Dhamotharan Pattarayan
- & Da Yang
-
Article
| Open Access Cell-type-specific mRNA transcription and degradation kinetics in zebrafish embryogenesis from metabolically labeled single-cell RNA-seq
Cell-type-specific mRNA transcription and degradation kinetics in zebrafish embryogenesis from metabolically labeled single-cell RNA-seq
This study analyzes the embryonic replacement of maternally contributed mRNA with new mRNA in single cells and shows dynamic spatio-temporal regulation of maternal mRNA decay and cell-type specific retention within the earliest specified cell types in zebrafish embryos.
- Lior Fishman
- , Avani Modak
- & Michal Rabani
-
Article
| Open Access Allopolyploid origin and diversification of the Hawaiian endemic mints
Allopolyploid origin and diversification of the Hawaiian endemic mints
Hawaiian endemic mints represent the second largest plant radiation in the archipelago. Here, the authors present a reference genome and numerous resequenced individuals to uncover evidence for polyploidy, geographic speciation and localized hybridization underlying diversification in this lineage
- Crystal M. Tomlin
- , Sitaram Rajaraman
- & Charlotte Lindqvist
-
Article
| Open Access Teacher-student collaborated multiple instance learning for pan-cancer PDL1 expression prediction from histopathology slides
Teacher-student collaborated multiple instance learning for pan-cancer PDL1 expression prediction from histopathology slides
PDL1 expression is a common biomarker for immunotherapy response in cancer, and it is usually quantified using immunohistochemistry. Here, the authors develop a weakly supervised learning approach combining multiple instance learning and a teacher-student framework to predict PDL1 expression from histopathological imaging.
- Darui Jin
- , Shangying Liang
- & Xiangzhi Bai
-
Article
| Open Access Nonlinear DNA methylation trajectories in aging male mice
Nonlinear DNA methylation trajectories in aging male mice
DNA methylation is an age biomarker, but nonlinear aspects of its age-related dynamics are not well characterized. Here, the authors identify loci that undergo sudden methylation changes at specific life stages in the aging colon of male mice.
- Maja Olecka
- , Alena van Bömmel
- & Steve Hoffmann
-
Article
| Open Access Accurately clustering biological sequences in linear time by relatedness sorting
Accurately clustering biological sequences in linear time by relatedness sorting
Accurately clustering biological sequences is an increasingly important task but is challenging for large datasets. This study introduces a new approach called ‘relatedness sorting’ to accurately cluster sequences with linear-time scalability.
- Erik Wright
-
Article
| Open Access Comparative characterization of the infant gut microbiome and their maternal lineage by a multi-omics approach
Comparative characterization of the infant gut microbiome and their maternal lineage by a multi-omics approach
Here, the authors employ multi-omics on a cohort comprising three generations of family members, showing that fecal microbiota populations, functions, and metabolome of infants vary greatly from their maternal lineage, exhibiting a less diverse microbiota and differences in various metabolite classes including short- and branched-chain fatty acids.
- Tomás Clive Barker-Tejeda
- , Elisa Zubeldia-Varela
- & Alma Villaseñor
-
Article
| Open Access Single-cell multiomics reveals the interplay of clonal evolution and cellular plasticity in hepatoblastoma
Single-cell multiomics reveals the interplay of clonal evolution and cellular plasticity in hepatoblastoma
Hepatoblastoma (HB) is the most frequent paediatric liver tumour with heterogeneous cellular phenotypes that influence clinical outcomes. Here, the authors integrate bulk, single-cell, and spatial multi-omics to characterise HB cells, and find that clonal evolution and epigenetic plasticity shape response to therapy.
- Amélie Roehrig
- , Theo Z. Hirsch
- & Eric Letouzé
-
Article
| Open Access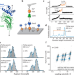 Engineering an artificial catch bond using mechanical anisotropy
Engineering an artificial catch bond using mechanical anisotropy
Catch bonds are unique protein-protein interactions where the bond lifetime increases under external pulling forces. Here, the authors engineer an artificial catch bond based on a non-catch bonding human gut bacterial adhesion protein complex.
- Zhaowei Liu
- , Haipei Liu
- & Michael A. Nash
-
Article
| Open Access scButterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders
scButterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders
Technical limitations of simultaneously multi-omics profiling lead to highly noisy multi-modal data and substantial costs. Here, authors proposed a versatile framework and data augmentation schemes, capable of single-cell cross-modality translation and multiple extensive applications.
- Yichuan Cao
- , Xiamiao Zhao
- & Shengquan Chen
-
Article
| Open Access Systematic HOIP interactome profiling reveals critical roles of linear ubiquitination in tissue homeostasis
Systematic HOIP interactome profiling reveals critical roles of linear ubiquitination in tissue homeostasis
Authors perform an in vivo mass spectrometry-based interactome analysis of HOIL-1-interacting protein, a key component of linear ubiquitination assembly complex.
- Yesheng Fu
- , Lei Li
- & Lingqiang Zhang
-
Article
| Open Access De novo diploid genome assembly using long noisy reads
De novo diploid genome assembly using long noisy reads
Most existing assemblers failed to generate high-quality phased assemblies using long noisy reads. Here, the authors present PECAT, a Phased Error Correction and Assembly Tool, for reconstructing diploid genomes from long noisy reads.
- Fan Nie
- , Peng Ni
- & Jianxin Wang
-
Article
| Open Access DeepDOF-SE: affordable deep-learning microscopy platform for slide-free histology
DeepDOF-SE: affordable deep-learning microscopy platform for slide-free histology
Histopathology can be limited by the time-consuming and labour-intensive preparation of slides from resected tissue. Here, the authors report DeepDOF-SE, a deep-learning-enabled microscope to rapidly scan intact tissue at cellular resolution without the need for physical sectioning.
- Lingbo Jin
- , Yubo Tang
- & Ashok Veeraraghavan
-
Article
| Open Access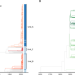 The recent rapid expansion of multidrug resistant Ural lineage Mycobacterium tuberculosis in Moldova
The recent rapid expansion of multidrug resistant Ural lineage Mycobacterium tuberculosis in Moldova
Chitwood et al. report on the rapid expansion of a Ural-lineage multidrug resistant strain of Mycobacterium tuberculosis in Moldova. This strain has an estimated reproduction number more than two times greater than otherwise similar drug susceptible strains.
- Melanie H. Chitwood
- , Caroline Colijn
- & Benjamin Sobkowiak
-
Article
| Open Access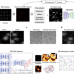 Context-aware deep learning enables high-efficacy localization of high concentration microbubbles for super-resolution ultrasound localization microscopy
Context-aware deep learning enables high-efficacy localization of high concentration microbubbles for super-resolution ultrasound localization microscopy
Ultrasound localisation microscopy enables deep tissue microvascular imaging. Here, authors introduce LOCA-ULM, a deep learning pipeline enhancing localisation accuracy in high microbubble concentrations. LOCA-ULM reveals dense cerebrovascular networks and enhances the sensitivity of functional ULM.
- YiRang Shin
- , Matthew R. Lowerison
- & Pengfei Song
-
Article
| Open Access Genomic language model predicts protein co-regulation and function
Genomic language model predicts protein co-regulation and function
A gene’s function is governed by its sequence, structure and context. Here, the authors develop a genomic language model that learns contextualized functional representations from diverse and large-scale metagenomic datasets.
- Yunha Hwang
- , Andre L. Cornman
- & Peter R. Girguis
-
Article
| Open Access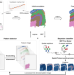 Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Recognising spatial spots’ biological identity in spatial transcriptomics remains a challenge. Here, authors introduce Pianno, a tool that helps annotate the biological structures or cell-type constructions across diverse tissues, offering new perspectives on understanding spatial transcriptomics.
- Yuqiu Zhou
- , Wei He
- & Ying Zhu
-
Article
| Open Access Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks
Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks
Rise of new viral strains is a major public health challenge, demanding advanced detection and forecasting methods. This study shows how examining communities within networks of viral mutations enables early detection of emerging strains.
- Fatemeh Mohebbi
- , Alex Zelikovsky
- & Pavel Skums
-
Article
| Open Access Reconstructing the evolution history of networked complex systems
Reconstructing the evolution history of networked complex systems
Evolution processes of complex networked systems in biology and social sciences, and their underlying mechanisms, still need better understanding. The authors propose a machine learning approach to reconstruct the evolution history of complex networks.
- Junya Wang
- , Yi-Jiao Zhang
- & Yanqing Hu
-
Article
| Open Access Data-driven identification of predictive risk biomarkers for subgroups of osteoarthritis using interpretable machine learning
Data-driven identification of predictive risk biomarkers for subgroups of osteoarthritis using interpretable machine learning
Osteoarthritis can be caused by multiple biological mechanisms but the drivers of disease risk are not well understood. Here, the authors use data from UK Biobank in machine learning models to identify clinical and biological markers associated with development of osteoarthritis and identify sub-groups with different risk profiles.
- Rikke Linnemann Nielsen
- , Thomas Monfeuga
- & Ramneek Gupta
-
Article
| Open Access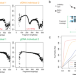 FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA
FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA
DNA methylation from cell-free DNA (cfDNA) can be profiled using whole genome bisulfite sequencing (WGBS). Here, the authors develop a computational method, FinaleMe, that predicts DNA methylation and tissues of-origin in cfDNA and validate its performance using paired deep and shallow-coverage whole-genome sequencing (WGS) and WGBS data.
- Yaping Liu
- , Sarah C. Reed
- & Manolis Kellis
-
Article
| Open Access PLMSearch: Protein language model powers accurate and fast sequence search for remote homology
PLMSearch: Protein language model powers accurate and fast sequence search for remote homology
Homologous protein search is one of the most commonly used methods for protein analysis. Here, authors propose PLMSearch, a search method that takes only sequences as input and can search millions of protein pairs in seconds while maintaining sensitivity comparable to SOTA structure search methods.
- Wei Liu
- , Ziye Wang
- & Shanfeng Zhu
-
Article
| Open Access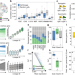 Gene-expression memory-based prediction of cell lineages from scRNA-seq datasets
Gene-expression memory-based prediction of cell lineages from scRNA-seq datasets
Combining experimental lineage tracing with single cell transcriptomics is technically demanding. Here, authors present GEMLI, a computational tool to annotate cell lineages in single cell RNA sequencing data solely based on gene expression.
- A. S. Eisele
- , M. Tarbier
- & D. M. Suter
-
Article
| Open Access DELVE: feature selection for preserving biological trajectories in single-cell data
DELVE: feature selection for preserving biological trajectories in single-cell data
Characteristic genes or proteins driving continuous biological processes are difficult to uncover from noisy single-cell data. Here, authors present DELVE, an unsupervised feature selection method to identify core molecular features driving cell fate decisions.
- Jolene S. Ranek
- , Wayne Stallaert
- & Jeremy E. Purvis
-
Article
| Open Access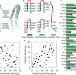 Hairpin trimer transition state of amyloid fibril
Hairpin trimer transition state of amyloid fibril
Amyloid fibrils are ordered protein assemblies implicated in neurodegenerative disease. Here the authors show that hairpin trimers can be transition states of fibril nucleation, explaining how different fibril isoforms may arise from alternative nucleation sites.
- Levent Sari
- , Sofia Bali
- & Milo M. Lin
-
Article
| Open Access Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY
Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY
Placenta histopathology for maternal and newborn health is highly specialised and time consuming. Here, authors present a deep learning pipeline for quantifying cells and tissues in placenta whole slide images, revealing biological and clinical insights.
- Claudia Vanea
- , Jelisaveta Džigurski
- & Christoffer Nellåker
-
Article
| Open Access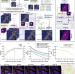 SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
Imaging mass cytometry (IMC) is a powerful single-cell resolution platform for targeted spatial proteomics, but it can be constrained by imaging noise and resolution. Here, the authors propose SpiDe-Sr, a super-resolution network embedded with a denoising module for IMC spatial resolution enhancement.
- Rui Chen
- , Jiasu Xu
- & Xianting Ding
-
Article
| Open Access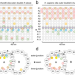 Uncovering structural themes across cilia microtubule inner proteins with implications for human cilia function
Uncovering structural themes across cilia microtubule inner proteins with implications for human cilia function
The inside surface of microtubules contains so-called microtubule inner proteins, but little is known about their identity. Here the authors use bioinformatics to identify structural motifs within this class of proteins and potential new members.
- Jens S. Andersen
- , Aaran Vijayakumaran
- & Kenneth Bødtker Schou
-
Article
| Open Access High-throughput prediction of protein conformational distributions with subsampled AlphaFold2
High-throughput prediction of protein conformational distributions with subsampled AlphaFold2
Protein dynamics, crucial for life, are difficult and expensive to predict. This study shows that AI-based structure prediction methods can be modified for rapidly predicting the conformational landscapes of proteins, with strong correlations with experimentally-measured relative state populations.
- Gabriel Monteiro da Silva
- , Jennifer Y. Cui
- & Brenda M. Rubenstein
-
Article
| Open Access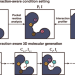 3D molecular generative framework for interaction-guided drug design
3D molecular generative framework for interaction-guided drug design
Designing a molecule that favorably binds to a protein pocket is a keystone of drug discovery. Zhung et al. devise DeepICL, which leverages the generalizable features of non-covalent protein-ligand interactions on a 3D molecular generative model, improving the quality of AI-designed molecules.
- Wonho Zhung
- , Hyeongwoo Kim
- & Woo Youn Kim
-
Article
| Open Access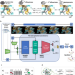 A dual diffusion model enables 3D molecule generation and lead optimization based on target pockets
A dual diffusion model enables 3D molecule generation and lead optimization based on target pockets
Structure-based generative chemistry is crucial in computer-aided drug discovery. Here, authors propose PMDM, a conditional generative model for 3D molecule generation tailored to specific targets. Extensive experiments demonstrate that PMDM can effectively generate rational bioactive molecules
- Lei Huang
- , Tingyang Xu
- & Hengtong Zhang
-
Article
| Open Access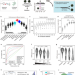 Precise prediction of phase-separation key residues by machine learning
Precise prediction of phase-separation key residues by machine learning
Understanding intracellular phase separation is essential for transcriptional control, cell fate, and disease. Here the authors report PSPHunter which accurately predicts key residues, aiding in disease-associated protein identification and mechanistic insights.
- Jun Sun
- , Jiale Qu
- & Junjun Ding
-
Article
| Open Access Predicting and improving complex beer flavor through machine learning
Predicting and improving complex beer flavor through machine learning
Perception and appreciation of food flavour depends on many factors, posing a challenge for effective prediction. Here, the authors combine extensive chemical and sensory analyses of 250 commercial Belgian beers to train machine learning models that enable flavour and consumer appreciation prediction.
- Michiel Schreurs
- , Supinya Piampongsant
- & Kevin J. Verstrepen
-
Article
| Open Access Diverging co-translational protein complex assembly pathways are governed by interface energy distribution
Diverging co-translational protein complex assembly pathways are governed by interface energy distribution
Protein complex assembly can occur co-translationally. Here, the authors uncover diverging assembly pathways and hotspot disruptions in N-terminal acetyltransferases, enzymes implicated in neurodegenerative diseases. Their model predicts co-translational assembly based on interface energy distribution.
- Johannes Venezian
- , Hagit Bar-Yosef
- & Ayala Shiber
-
Article
| Open Access Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
DNA double-strand breaks (DSBs) are repaired by a hierarchically regulated network of pathways. Here, authors develop ICP for deciphering somatic DSB repair patterns in multicellular organisms and discover developmental regulation in flies and mosquitoes, enabling tracking of mutant alleles and interhomolog copying of gene cassettes.
- Zhiqian Li
- , Lang You
- & Ethan Bier
-
Article
| Open Access Multi-omic integration of microbiome data for identifying disease-associated modules
Multi-omic integration of microbiome data for identifying disease-associated modules
Here, Muller et al. introduce MintTea, a method for analyzing multi-omic microbiome data and identifying disease-associated modules comprising mixed sets of features that collectively shift in disease, offering insights into microbiome-disease interactions.
- Efrat Muller
- , Itamar Shiryan
- & Elhanan Borenstein
-
Article
| Open Access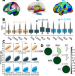 The genetic architecture of multimodal human brain age
The genetic architecture of multimodal human brain age
The biological basis of brain aging is not well understood, but it has implications for human health. Here, the authors explore the genetic basis of human brain aging, finding genetic variants, genes and potential causal relationships with disease.
- Junhao Wen
- , Bingxin Zhao
- & Christos Davatzikos
-
Article
| Open Access InPACT: a computational method for accurate characterization of intronic polyadenylation from RNA sequencing data
InPACT: a computational method for accurate characterization of intronic polyadenylation from RNA sequencing data
Intronic polyadenylation (IPA) can produce transcripts with truncated coding regions and has been implicated in diverse biological processes and diseases. Here, the authors present a computational method for the accurate delineation of IPA events using RNA-sequencing data.
- Xiaochuan Liu
- , Hao Chen
- & Yang Yang
-
Article
| Open Access Predicting nuclear G-quadruplex RNA-binding proteins with roles in transcription and phase separation
Predicting nuclear G-quadruplex RNA-binding proteins with roles in transcription and phase separation
RNA G-quadruplexes are important regulatory elements, yet our knowledge of their structure-based interactions is at present limited. Here the authors combine experimental and computational methods to develop a predictive tool, G4-FUNNIES, to estimate proteins’ RNA G4-binding propensities.
- Johanna Luige
- , Alexandros Armaos
- & Ulf Andersson Vang Ørom
-
Article
| Open Access Allele-specific transcriptional effects of subclonal copy number alterations enable genotype-phenotype mapping in cancer cells
Allele-specific transcriptional effects of subclonal copy number alterations enable genotype-phenotype mapping in cancer cells
Quantifying the impact of copy-number alterations (CNAs) on gene expression at the subclone level in cancer remains a challenge. Here, the authors develop TreeAlign, a method that integrates sample-matched single-cell DNA and RNA sequencing data to infer the impact of CNAs on subclonal gene expression.
- Hongyu Shi
- , Marc J. Williams
- & Sohrab P. Shah
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Detecting m6A at single-molecular resolution via direct RNA sequencing and realistic training data
Detecting m6A at single-molecular resolution via direct RNA sequencing and realistic training data