« Prev Next »
In the early part of the twentieth century, scientists knew what genes did, but they did not know what they were. Francis Crick, one of the codiscoverers of the three-dimensional double helical structure of DNA, was among the first to propose that a gene was a linear sequence of nucleotides and that each gene encoded a single protein. Crick called this proposal the sequence hypothesis (Crick, 1958); other scientists have since referred to it as the genes-on-a-string hypothesis. In Crick's words, this hypothesis "assumes that the specificity of a piece of nucleic acid is expressed solely by the sequence of its bases, and this sequence is a (simple) code for the amino acid sequence of a particular protein." Crick freely admitted that his hypothesis was just that: a hypothesis "for which proof is completely lacking." However, in an effort to rationalize his speculation, Crick cited some experimental work with bacteriophages that had been conducted by American molecular biologist Seymour Benzer. Benzer's work demonstrated that, in Crick's words, "a functional gene consists of many sites arranged strictly in a linear order" (Crick, 1958; italics original).
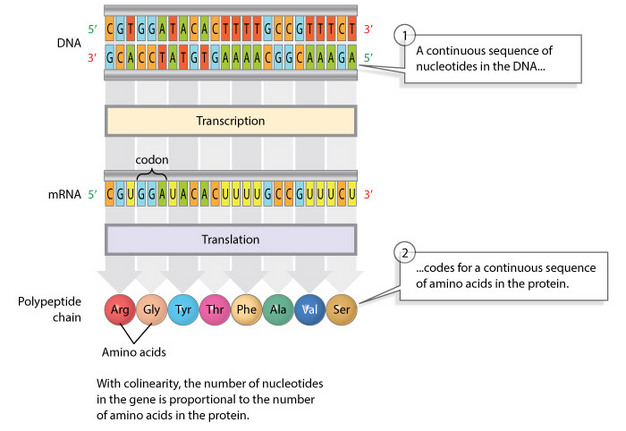
Today, scientists no longer speak of the sequence hypothesis. Instead, the notion that nucleotide sequences (genes) directly dictate amino acid sequences is known as colinearity (Figure 1). Scientists have confirmed that colinearity is a regular occurrence among many viruses, like the ones Benzer studied, as well as among bacteria. However, it turns out that colinearity is the exception, not the rule, in eukaryotic genomes.
Alternatives to Colinearity
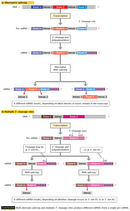
One of the first clues that the colinearity of DNA and amino acid sequences is not as simple as what Crick had proposed was the discovery of RNA splicing in the 1970s. Using common cold viruses as their experimental systems, English molecular chemist Richard Roberts and American molecular biologist Philip Sharp independently discovered that genes can be split into several segments along the genome (Berget et al., 1977; Chow et al., 1977). Then, using electron microscopy, both scientists observed that a single messenger RNA (mRNA) molecule hybridized not to a single stretch of DNA but to as many as four or more discontinuous DNA segments (Figure 2).
Roberts and Sharp also noted that the genetic material actually breaks apart and then re-forms itself at certain points in protein synthesis. Specifically, the sections of DNA that encode protein production are known as exons, and the noncoding sections interspersed among the exons are known as introns. During splicing, which occurs after transcription (i.e., the synthesis of RNA from a DNA template), the introns are removed and the exons are joined, or spliced together.
Roberts's and Sharp's findings not only raised serious doubts about the concept of a gene as a continuous, clearly demarcated segment of DNA, but they also led to a flurry of research activity, with scientists curious about whether the same was true in other species. As other researchers were quick to discover, discontinuous gene structure and splicing during RNA processing are the norm, not the exception, in most eukaryotes. Some vertebrate genes contain as many as 50 exons, and exons often make up only a small portion of the transcribed region of a gene. For example, in one early splicing study that involved examination of the intron-exon pattern of a chicken ovalbumin gene, Stein et al. (1980) measured eight exons ranging in length from 20 to 181 base pairs and seven introns ranging in length from 264 to 1,150 base pairs. Since that study, scientists have detected introns as long as 50,000 base pairs or more in some species.
The final protein products encoded by any given intron-exon sequence also vary in structure, depending on which exons are spliced back together during RNA processing. This so-called "alternative splicing" is illustrated in Figure 3. Scientists have also since learned that eukaryotic cells have evolved another "alternative" mRNA processing pathway: the use of multiple 3' cleavage sites in a single exon. (Every intron has a 5' and 3' splice site.) As illustrated in Figure 3, the end result is the same as with alternative splicing: different mRNA molecules are produced from a single protein-coding gene. Clearly, contrary to the conventional notion of a single gene encoding a single protein, a single continuous stretch of DNA can encode multiple mRNA molecules and, ultimately, multiple protein products.
Transcription Units Instead of Genes
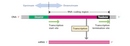
Given the vast quantity of DNA that appears to have little protein-encoding power and the fact that so much of this DNA resides right in the middle of functional genes (as introns), some scientists prefer to think in terms of "transcription units" rather than "genes." A transcription unit is a linear sequence of DNA that extends from a transcription start site to a transcription stop site (Figure 4).
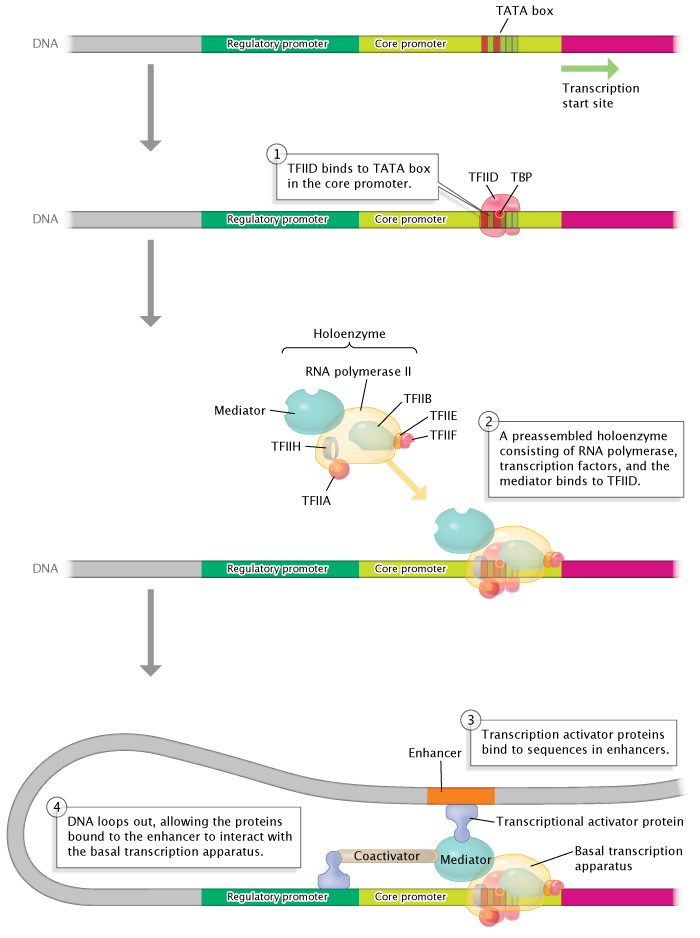
The promoter, a DNA sequence that lies upstream of the RNA coding region, serves as an indicator of where and in which direction transcription should proceed. The promoter is not actually transcribed; its role is purely regulatory. While promoters vary tremendously among eukaryotes, there are some common features. For example, most promoters lie immediately upstream of the transcription unit (transcription proceeds in an upstream to downstream direction), and most contain what is known as a TATA box; this is a sequence that is recognized and bound by a so-called TATA binding protein. The TATA binding protein helps position the RNA polymerase machinery and initiates transcription. Some promoters work in concert with other types of regulatory sequences known as enhancers, which sometimes lie several kilobases further upstream or downstream from the coding sequence itself, or even within introns. These two sequences are able to interact because of the way DNA molecules bend in space, enabling sections that would otherwise be very far from each other to interact (via DNA-binding proteins). Enhancer regions serve as binding sites for proteins known as activators (Figure 5). The proteins that bind to promoters to regulate transcription are called transcription factors. The RNA coding region, the main component of the transcription unit, contains the actual exons and introns. The terminator, a sequence of nucleotides at the end of the transcription unit, is transcribed along with the RNA coding region. The terminator serves as a speed bump of sorts; transcription stops only after this region has been transcribed.
Scientists have recently discovered that some mRNA molecules are coded by exons from multiple transcription units through a process known as trans-splicing. In fact, in 2005, a European group of researchers estimated that about 4% to 5% of tandem transcription units (i.e., distinct but adjacent transcription units) in humans are transcribed together to create single "chimeric" mRNA molecules (Parra et al., 2005). Scientists are not sure how this occurs. Some speculate that transcription overrides the first transcription terminator and doesn't stop until it reaches the second termination site; others suspect that both transcripts are formed independently and then spliced together to form the chimeric mRNA molecule.
Delineating Gene Regions
It seems that the more scientists learn about the genome and gene expression, the less they seem to be able to identify the point along a stretch of nucleotides at which a single gene actually begins and ends; indeed, it appears to be increasingly more difficult to determine whether there are even actual discrete nucleotide start and stop points for genes. This complexity continues to make it difficult for scientists to agree on exactly what a gene is. At the very least, scientists now know that Crick's original sequence hypothesis was overly simplistic, at least for eukaryotes. Genes are not linear sequences of DNA that directly correspond one-to-one with their protein counterparts.
Moreover, scientists now know that not all transcribed RNA molecules, or transcripts, end up being translated into protein products. For example, in a study of the mouse genome, researchers found that as much as 63% of the genome is transcribed but only about 1% to 2% is translated into a functional protein product (FANTOM Consortium et al., 2005). So not only is the notion of colinearity overly simplistic, but so too is the notion that all genes encode proteins. Many code other types of molecules, like tRNA and rRNA, that have important known cellular functions. Other non-protein-coding RNAs work to regulate gene expression at multiple levels, and still other transcripts have unknown functions.
References and Recommended Reading
Beadle, G. W., & Tatus, E. L. Genetic control of biochemical reactions in Neurospora. Proceedings of the National Academy of Sciences 27, 499–506 (1941)
Berget, S. M., et al. Spliced segments at the 5' terminus of adenovirus 2 late mRNA. Proceedings of the National Academy of Sciences 74, 3171–3175 (1977)
Chow, L. T., et al. An amazing sequence arrangement at the 5' ends of adenovirus 2 messenger RNA. Cell 12, 1–8 (1977)
Crick, F. On protein synthesis. The Biological Replication of Macromolecules: Symposium for the Society of Experimental Biology 12, 138–162 (1958)
FANTOM Consortium, et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005) doi:10.1126/science.1112014
Parra, G., et al. Tandem chimerism as a means to increase protein complexity in the human genome. Genome Research 16, 37–44 (2006)
Stein, J. P., et al. Ovomucoid intervening sequences specify functional domains and generate protein polymorphism. Cell 21, 681–687 (1980)



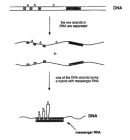
 Figure 2
Figure 2