« Prev Next »
Introduction
During embryonic development, specialized cells (e.g., muscle or immune cells) arise from a common stem cell that differentiates via a series of cellular changes triggered by specific gene expression patterns. Scientists can recover these embryonic stem (ES) cells from embryos and manipulate them in vitro to study early development. They can also differentiate ES cells into cell types that are useful for therapeutic purposes, such as transplantation. This technology raises a significant ethical concern because most ES cells arise from human embryos. Some ethical concerns may be circumvented by the discovery that somatic cells can be reprogrammed to a pluripotent state. The reprogrammed cells, called induced pluripotent stem (iPS) cells, exhibit functional similarities to ES cells and present an exciting area of research. The ability to reprogram somatic cells into iPS cells that are pluripotent and can self-renew has transformed the fields of developmental biology and regenerative medicine.
What Are ES Cells?
ES cells were originally derived from the inner cell mass (ICM) or epiblast of mouse embryos. These cells exhibit an almost unlimited proliferative capacity in culture and maintain their pluripotent potential to differentiate into all cell lineages in the body. However, to create ES cell lines, it is necessary to destroy preimplantation embryos at the blastocyst stage of 100–200 cells. Many people believe that life begins at conception and that performing research on ES cells derived from the destruction of human blastocysts is morally unacceptable. Another important issue is that transplanted ES cells for therapeutic purposes can trigger host immune rejection. These issues are largely resolved by using iPS stems cells that arise from somatic cells because they can be recovered from an adult source. Furthermore, if cells are recovered from a human donor who will also later receive the differentiated cells, this method also largely mitigates any immune reaction encountered during transplantation.
Reprogramming Somatic Cells into Pluripotent iPS Cells
How do researchers reprogram somatic cells to a pluripotent state? Earlier experiments in cell fusion and nuclear transfer showed that gene expression in differentiated cells remained dynamic and reversible. Silent genes in a specific cell type can be reactivated by fusing the cells with a different cell type. Subsequently, several studies showed that introduction of defined transcription factors could convert specialized cell types from one lineage to another. When somatic cells were reprogrammed by transferring their nuclei into oocytes or by fusion with ES cells, genome-wide transcriptional activity and DNA methylation patterns were converted from the somatic state to an embryonic state.
In 2006, Kazutoshi Takahashi and Shinya Yamanaka established for the first time murine ES-like cell lines from mouse embryonic fibroblasts (MEFs) and skin fibroblasts by simply expressing four transcription factor genes encoding Oct4, Sox2, Klf4, and c-Myc (Figure 1) (Takahashi & Yamanaka 2006). They called these somatic cell-derived cell lines induced pluripotent stem (iPS) cells. These iPS cell lines exhibit similar morphology and growth properties as ES cells and express ES cell-specific genes. Transplantation of iPS cells into immunodeficient mice resulted in the formation of germ-cell-tumor (teratoma)-containing tissues from all three germ layers, confirming the pluripotent potential of iPS cells. However, there were two problems: the low efficiency of establishing iPS cell lines and some variations in gene expression profiling between iPS cells and ES cells. The latter issue raised the concern that cell reprogramming may be insufficient to restore full pluripotency in somatic cells as exhibited by ES cells.
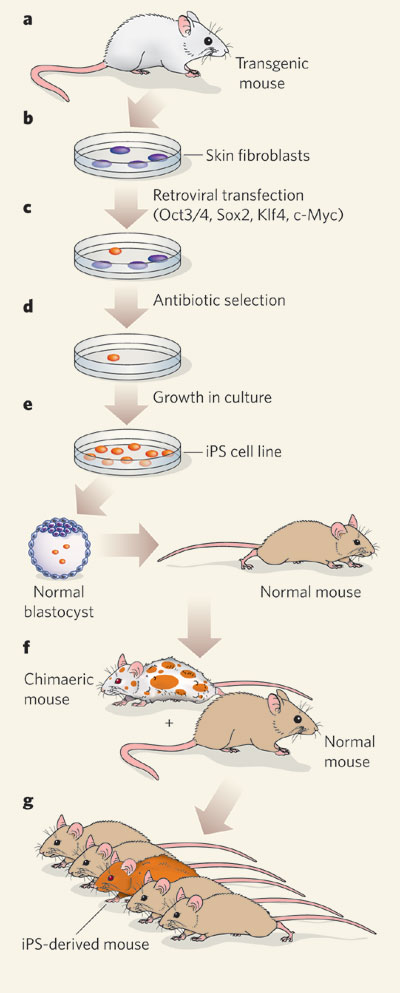
The most stringent test for pluripotency is known as the tetraploid blastocyst complementation assay. The method involves merging the embryonic (ES or iPS cells) and extraembryonic tissue (tetraploid cells) from two different species of animals and then testing to see if the embryonic tissue is sufficient for the normal development to the adult stage. Using this assay, several studies now show the production of fertile adult mice derived entirely from iPS cells, therefore confirming the true pluripotency gained by iPS cells during the reprogramming process.
In the years following Takahashi and Yamanaka's initial success in reprogramming mouse cells, multiple groups used the same strategy to generate human iPS cells, which resemble human ES cells by all measured criteria. The ability to use the same set of transcription factors to reprogram both mouse and human somatic cells reinforces the conserved nature of transcription factor networks in these two species to control self-renewal and pluripotency. Generation of iPS cells from somatic cells from any individual largely solves the two major obstacles mentioned earlier.
What Somatic Cell Types Can Be Reprogrammed into iPS Cells?
Most iPS cell lines established in earlier studies were derived from fibroblasts. Can iPS cells be established from other somatic cell types? If so, at what frequency? These questions were soon answered, although not completely. Using somatic cells harvested from transgenic mouse lines containing the four reprogramming transcription factors, several groups showed that iPS cell lines could be generated from different cell types, such as neuronal progenitor cells, keratinocytes, hepatocytes, B cells, and fibroblasts of mouse tail tips, kidneys, muscles, and adrenal glands. These results demonstrate the capacity of the four transcription factors to mediate reprogramming regardless of developmental origins and epigenetic states of a cell.
The efficiency of cell reprogramming varies among different cell types. For example, human keratinocytes from skin biopsies can be reprogrammed to pluripotency at much higher frequency and faster speed than fibroblasts. This difference is attributed to the finding that keratinocytes express much higher levels of endogenous c-Myc and Klf4 than fibroblasts, which may accelerate the conversion of keratinocytes to iPS cells. Recent studies show that mesenchymal-to-epithelial transition (MET) is a crucial early phase during the reprogramming of fibroblasts into iPS cells. Striking similarities between the initial stage of reprogramming and MET were detected, including up-regulation of epithelial genes such as E-cadherin and Epcam as well as the concomitant down-regulation of mesenchymal genes such as Snail and N-cadherin.
In contrast to keratinocytes, which are an epithelial cell type, fibroblasts may need to undergo an initial MET during the reprogramming process. This additional transition step may result in reduced efficiency and a prolonged duration to reprogram fibroblasts to reach pluripotency. Consistent with this model, hepatocytes and gastric epithelial cells in mice also appear to be more easily reprogrammed than fibroblasts.
The Cell Differentiation Status Matters
Will the differentiation status of a somatic cell affect the efficiency of iPS cell generation? Likely it will. In fact, it seems that for successful reprogramming, differentiation status, which itself may depend on the epigenetic genomic state of a somatic cell, must be reset to a pluripotent state. Furthermore, ectopic expression of Oct4 alone is sufficient to convert adult neural stem cells into iPS cells and therefore appears to support such a model. Compounds that inhibit DNA or histone modifications also generate iPS cells more efficiently. A link between cell differentiation stages and reprogramming efficiency is further supported by the observation that hematopoietic stem and progenitor cells can be reprogrammed into iPS cells much more efficiently than terminally differentiated B and T lymphocytes.
How does the source of iPS cells affect the supply? For practical consideration, acquiring samples from a skin biopsy of a patient to produce fibroblasts for reprogramming represents a convenient option. However, the biopsy procedure itself and the need to expand fibroblasts in culture still represent major hurdles to be overcome before the iPS cell technology can become more broadly applicable. Hematopoietic (blood) cells may represent a more suitable source than skin fibroblasts to derive iPS cells because taking blood samples from patients is common and is less invasive than taking a skin biopsy. Moreover, collecting blood is not as costly as performing a skin biopsy. Several recent studies have made iPS cell lines from patients' blood samples, thereby demonstrating the feasibility of using this accessible resource for cell reprogramming and removing the need to expand these cells in culture.
Modeling Human Diseases with iPS Cells
Can we use patient iPS cells to model human diseases in vitro? Availability of patient-specific iPS cell lines provides unprecedented opportunities to elucidate disease mechanisms in vitro, to carry out drug screening and toxicology studies, and to advance cell replacement therapy in regenerative medicine (Colman & Dreesen 2009). Reprogramming of fibroblasts from patients with Mendelian and complex genetic disorders — such as amyotrophic lateral sclerosis, type 1 diabetes, Parkinson's disease, and Duchenne muscular dystrophy — allows the establishment of disease-specific iPS cell lines. To study the disease mechanism, a key issue is whether the affected cell type derived from iPS cells can recapitulate the disease phenotype.
Two studies demonstrated that iPS-derived cells actually exhibited partial disease phenotypes. The first case involves the iPS cells derived from spinal muscular atrophy (SMA). The major symptom of SMA is selective degeneration of motoneurons, and the SMA iPS cells showed reduced capacity to form mature motoneurons in culture when compared with those derived from normal iPS cells.
The second case involves the iPS cells derived from familial dysautonomia (FD). FD is a fatal peripheral neuropathy characterized by the depletion of autonomic and sensory neurons. FD iPS cells showed marked defects in neurogenesis and reduced migration of iPS cell-derived neural crest precursors. These studies demonstrate the potential promises of using iPS cells to model early-onset human diseases such as SMA and FD. However, the use of iPS cells to model adult-onset diseases such as Parkinson's disease and amyotrophic lateral sclerosis proves to be more elusive. Relevant cell types derived from patients' iPS cells of these diseases have so far failed to exhibit disease-related phenotypes. Exposure of the differentiating cells to stress conditions by increasing the level of nitrogen and oxygen reactive species, proinflammatory factors, or even toxins may be necessary to speed up the emergence of pathological status in relevant cell types derived from iPS cells.
The other limitation in modeling human diseases with iPS cells is that a single cell type may not be sufficient to manifest the full spectrum of pathogenesis. Interaction among different cell types may be important to reconstruct the disease phenotypes faithfully. In this case, we first need to identify the interacting cell types and then work out a protocol for iPS cell differentiation into these cell types so as to recapitulate full disease phenotypes. Ultimately, it may be necessary to transplant iPS-derived cells into immunodeficient mice to reveal disease phenotypes.
The Exciting Potential and Limitations of Reprogrammed iPS Cells
The discovery that somatic cells can be reprogrammed into iPS cells has already had major effects on research in stem cell biology and regenerative medicine. Is this technology mature enough for research and therapy? Many obstacles remain and need to be resolved to take full advantage of this technology in research and therapy (Saha & Jaenisch 2009).
The most noted problem is the use of retroviral and lentiviral vectors to introduce the four transcription factor genes into somatic cells for cell reprogramming. These viral vectors preferentially integrate into active genes and therefore have the potential to activate flanking cellular genes and transform the transplanted cells. In addition, most of the four introduced transcription factors possess oncogenic potentials, and persistent expression of any of them may provide cell growth advantage and increase the chance for cell transformation. Although expression of these four genes for the most part is silent in established iPS cell lines, residual expression or reactivation of their expression in transplanted iPS cells can induce tumors in mice. Thus, although iPS cells derived from this route may be suitable for the study of disease mechanisms or for drug screening and validation, they definitely are not suitable for cell replacement therapy.
Many alternative gene delivery strategies — including the use of episomal vectors, nonintegrating viral vectors, transient DNA transfection, transposons, and protein transduction — can overcome this problem. A general principle common to all these strategies is the transient expression of the four transcription factors at sufficient levels to trigger the initiation of the cell reprogramming event without permanent integration of the four genes into the host genome. Although these strategies work for the most part, the efficiency of generating iPS cell lines is significantly reduced compared with the approach of retroviral and lentiviral vectors.
Use of small molecules to activate the pluripotency program in somatic cells represents perhaps the safest approach to create reprogramming factor-free iPS cells. Several small molecules, when used singly, could substitute for some of the reprogramming factors. However, so far it is not possible to use only small molecules to reprogram somatic cells. High throughput screening of small molecules for cell reprogramming is ongoing in many laboratories, and the ultimate goal would be to establish iPS cells free of any exogenously introduced DNA fragments.
Yet another problem with iPS cells in the study of disease mechanism is that defining a disease-related phenotype is frequently hindered by the intrinsic variability in differentiation potentials observed among different iPS cell lines. This variability makes it less certain that any observed phenotype in cells derived from a single iPS cell line is caused by the defective gene function. Therefore, to ensure that the exhibited phenotypes are not unique to a specific iPS cell line or a particular patient, it is important to evaluate several iPS cell lines generated from the same patient as well as those generated from different patients with the same disease. Alternatively, restoration of the missing gene function in mutant iPS cells provides an ideal isogenic control for any observed phenotype.
Use in Sickle-Cell Anemia
Development of iPS cell technology has already revolutionized the fields of developmental biology and regenerative medicine. This technology has successfully treated sickle-cell anemia in a humanized mouse model. In this model, the human bs sickle globin gene replaces the endogenous mouse b-globin gene, causing sickle-cell anemia (Hanna 2007). Fibroblasts from the tail tip of this mouse model were converted into iPS cells, and gene-specific targeting in vitro corrected the human bs sickle globin gene in the iPS cells. The genetically modified iPS cells were then differentiated into hematopoietic progenitor cells in vitro, followed by transplantation into the mouse model. This procedure rescued the disease phenotype and provided proof of the principle for using iPS cells in gene- and cell-based therapy.
Remaining Challenges
For practical application of this technology in clinics, we need to resolve issues involving the use of viral vectors for iPS cell establishment, the safe strategy to genetically modify iPS cells, differentiation of iPS cells into relevant cell types in vitro for transplantation, and removal of contaminating stem or progenitor cells before transplantation. Study of abnormal phenotypes exhibited in iPS-derived cell lineages, such as in the cases of SMA and FD, opens up the possibility of using this technology to model human diseases in vitro. However, to facilitate the study of underlying disease mechanisms, we need to reveal a full spectrum of disease phenotypes in the affected cell type derived from iPS cells, especially those involved in functionality of the differentiated cells. The iPS cell technology may also contribute to drug validation and screening.
In the case of SMA, administration of valproic acid and tobramycin showed an improvement in the biochemical characteristics of iPS cells, but it remains unclear whether such treatment also improves the phenotypic deficit such as the neuron outgrowth defect. Treatment of FD iPS cells with kinetin, a plant hormone, throughout neuronal differentiation resulted in a significant increase in the percentage of neurons. These studies illustrate the promise of this technology in drug validation and screening. Because drug screening in most cases requires the exhibition of a phenotype in the cell target, developing efficient strategies to generate such cell targets from iPS cells for high throughput assays remains a major challenge. Despite these challenges, the potential of iPS cells remains enormous. Resolution of these challenges will have significant implications in the understanding of human diseases and will have major effects on the therapeutic treatment of these diseases.
References and Recommended Reading
Colman, A. & Dreesen, O. Pluripotent stem cells and disease modeling. Cell Stem Cell 5, 244–247 (2009).
Hanna, J. et al. Treatment of sickle cell anemia mouse model with iPS cells generated from autologous skin. Science 318, 1920–1923 (2007).
Saha, K. & Jaenisch, R. Technical challenges in using human induced pluripotent stem cells to model disease. Cell Stem Cell 5, 584–595 (2009).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).






























