« Prev Next »
In the majority of cases, two copies of each chromosome — and therefore, two copies of each gene — arrive in every fertilized egg. However, in some cases, one of the genes within a pair is somehow switched "off"; this occurs because of the process known as imprinting. Whether or not particular genes are inactivated in this way depends on which parent the genes were inherited from. In such cases, a gene is only expressed when it comes from the designated or "correct" parent; thus, if a person receives this particular gene from his or her other parent, the gene is permanently turned off. Another way in which one gene copy can be turned off is by inactivation of the entire X chromosome. Although they sometimes yield similar results, imprinting and X inactivation are regulated by completely different mechanisms.
Discovery of Imprinting
Genomic imprinting, also referred to as gametic or parental imprinting, was discovered almost simultaneously by two laboratories in 1984. Prior to the publications of McGrath and Solter (1984) and Surani et al. (1984), conflicting results existed about whether both parental genomes were required for normal development. Most laboratories were unable to generate viable embryos from activated oocytes. This lack of successful parthenogenesis was supportive of the requirement of both parental genomes, but direct evidence of this requirement first came from Surani et al. (1984).
Working with mice, Surani and colleagues performed an experiment at the earliest stages of development. Because activated oocytes did not survive into development, these researchers hypothesized that normal development required both a maternal and a paternal genome. To test this idea, the scientists relied on their ability to independently manipulate the maternal and paternal genomes through the pronuclei of sperm and egg cells. Thus, they were able to collect nuclei from oocytes and nuclei from sperm cells and manually inject these nuclei into regular oocytes. Surani et al. used this technique to look at several types of embryos. Specifically, the team first examined embryos created from eggs to which the nucleus from another oocyte had been added (thus creating a case of maternal uniparental disomy). Next, the team considered embryos created from eggs to which the nucleus of a sperm cell had been added.
Upon examining their results, Surani et al. noted that activated oocytes alone were not viable; similarly, adding the nucleus from an egg to a regular oocyte did not produce a healthy embryo. The parthenogenic, activated oocytes were rescued only when the scientists added the paternal contribution (Table 1). Although the rescue rate was not 100% (only 9 of 24 pregnancies resulted in live mouse pups), the level of success was still dramatic considering that none of the embryos survived when the entire genetic contribution came from the mother. These results supported the idea that both maternal and paternal contributions are required for normal development.
| Genetic Component | Number of Total Pregnancies | Number of Viable Embryos |
| Activated oocyte + egg pronucleus | 48 | 0 |
| Activated oocyte + sperm pronucleus | 24 | 9 |
| Table 1: Summary of results from experiments examining the differential role of egg or sperm pronuclei in mouse development | ||
But why are both types of contributions required, and how does a fertilized egg differentiate between the two? Surani et al. speculated that during production of eggs or sperm, some of the cells' genes are imprinted with an indication of the gamete in which the genes began. It was not until 1991, however, that this suspicion was confirmed via the identification of the first paternally imprinted (IgfII) and maternally imprinted (Igfr2, H19) genes (DeChiara et al., 1991; Barlow et al., 1991; Bartolomei et al., 1991). All three of these genes play a critical role in embryonic development (Barlow, 1995).
Imprinting in Somatic Cells
Imprinting results in expression patterns that are different from classical Mendelian inheritance patterns, because even though both parents contribute equally to the genetic content of their offspring, the imprinted genes contributed from the father and mother are not equally expressed. In particular, when the gene at a maternally imprinted locus is expressed, the copy of the imprinted gene from the mother is always turned "off," whereas the copy from the father is always turned "on." The opposite is true of a paternally imprinted gene.
So how, then, are imprinted genes marked to distinguish between the maternal and paternal contributions? It turns out that some genes have stretches of alternating cytosine and guanine nucleotides, which are known as CpG islands. These sequences are domains that can be methylated. Methylation is a type of chemical modification of DNA that can be copied immediately after replication, making it epigenetically inherited. With proper signals, DNA methylation can be subsequently removed without changing the original DNA sequence. Thus, imprinted genes can be marked by parental-specific methylation of CpG-rich domains. After that marking is present, the DNA is compacted via binding of specific proteins that recognize the 5-methyl cytosines. An imprint can be erased and then reestablished during gametogenesis to match the sex of the gamete-producing parent, thus playing an important role in maintenance of allele-specific gene expression (Ariel et al., 1995). De novo methylation establishes these patterns for different tissues, which are then maintained during somatic cell divisions.
In mammals, most imprinted genes are found in clusters on the genome, and these clusters often contain the sequences for noncoding RNAs. The relationship between the expression of noncoding RNAs and the silencing of linked protein-coding genes was studied by Sleutels et al. (2002), who used expression of a truncated Air RNA on mouse chromosome 7 to study its involvement in silencing of the IGf2r/Slc22a2/Slc22a3 gene cluster, which is imprinted and maternally expressed. Their results provided evidence that noncoding RNAs are also an important factor in genomic imprinting.
For imprinting to work, the epigenetic signal must be consistent in the mature gamete and the cells originating from the zygote that this gamete forms; however, this signal also has to be reversible in the next generation, depending on the sex of the offspring. During embryonic development, somatic cells maintain monoallelic expression of imprinted genes, but germ cells need to be imprinted to reflect the sex of the embryo. For example, if a paternally imprinted gene is inherited by a female, when she produces eggs, the paternal imprints must be erased and replaced with the maternal version.
X Inactivation
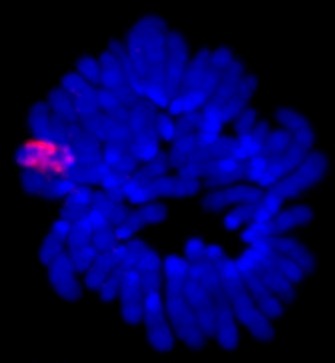
Ariel et al. (1995) used polymerase chain reaction to study the extent of methylation on the noncoding RNA gene called Xist in early mice embryos and reported that methylation of that gene was an important part of the X-imprinting process. In placental mammals, noncoding RNA is also believed to play a role in random imprinting of the X chromosome (Ng et al., 2007). The Xist gene, found on the X chromosome itself, produces noncoding RNA (siRNA) that associates with the X chromosome, coating its entire length (Figure 1). In adults, the Xist gene is expressed only on the inactive X chromosome (Xi). It also appears that the RNA might direct DNA methylation to the silent X, further compacting it. During X chromosome inactivation, another gene for noncoding RNA, Tsix, on Xi, is downregulated, but the Xist and Tsix genes need to interact with each other for X chromosome inactivation to take place. These interactions are mediated by a region of DNA called the X-pairing region (Xpr), which is found about 200 kilobases upstream of Xist. Research indicates that trans interactions between Xpr regions result in higher Xist RNA levels (Figure 1).
In females, differential methylation patterns are erased at some point and reestablished based on the random selection of one of the X chromosomes in the late blastocyst (Ariel et al., 1995; Ng et al., 2007). The mechanism for this process involves transcription of the genes Tsix and Xite and the involvement of a chromatin insulator, called Ctcf, in the X-inactivation center (Xu et al., 2007).
Despite what we know about gene imprinting, X chromosome inactivation, DNA methylation, and the various transcription factors involved, much needs to be discovered before we can fully explain any of these processes and how the message to express a specific gene based on the parental source is transmitted. Although imprinting and X inactivation both result in decreased gene expression, it is important to note that the mechanisms utilized are completely distinct, revealing the diversity of cellular processes that act as "switches" to keep gene expression in the "off" position for specific loci. Gaining a more detailed understanding of these switches will open doors for medical research on the determinants of various diseases and the causes of anomalies in embryonic development, in addition to leading to advances in stem-cell and cloning research.
References and Recommended Reading
Ariel, M., et al. Gamete-specific methylation correlates with imprinting of the murine Xist gene. Nature Genetics 9, 312–315 (1995) doi:10.1038/ng0395-312 (link to article)
Barlow, D. Gametic imprinting in mammals. Science 270, 1610–1613 (1995) doi:10.1126/science.270.5242.1610
Barlow, D. P., et al. The mouse insulin-like growth factor type-2 receptor is imprinted and closely linked to the Tme locus. Nature 349, 84–87(1991) doi:10.1038/349084a0 (link to article)
Bartolomei, M. S., et al. Parental imprinting of the mouse H19 gene. Nature 351, 153–155 (1991). doi:10.1038/351153a0 (link to article)
DeChiara, T. M., et al. Parental imprinting of the mouse insulin-like growth factor II gene. Cell 64, 849–859 (1991)
McGrath, J., & Solter, D. Completion of mouse embryogenesis requires both the maternal and paternal genomes. Cell 37, 179–183 (1984)
Ng, K., et al. Xist and the order of silencing. EMBO Reports 8, 34–39 (2007) doi:10.1038/sj.embor.7400871
Sleutels, F., et al. The non-coding Air RNA is required for silencing autosomal imprinted genes. Nature 415, 810–813 (2002) doi:10.1038/415810a (link to article)
Surani, M. A. H., et al. Development of reconstituted mouse eggs suggests imprinting of the genome during gametogenesis. Nature 308, 548–550 (1984) doi:10.1038/308548a0 (link to article)
Xu, N., et al. Evidence that homologous X-chromosome pairing requires transcription and Ctcf protein. Nature Genetics 39, 1390–1396 (2007) doi:10.1038/ng.2007.5 (link to article)






























