Abstract
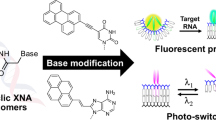
Further development of DNA nanotechnology requires new functional oligonucleotides composed of nucleobases beyond the native four. In this review, we demonstrate new methodology for DNA and RNA functionalization using a base surrogate prepared from d-threoninol (2-amino-1,3-butanediol). Using this nucleobase surrogate, we can introduce functional molecules at any position of the sequence. Our methodology is conceptually similar to the copolymerization of multiple monomers: phosphoramidite monomers corresponding to the base surrogate and natural nucleotides are copolymerized on a solid support to prepare the functional oligonucleotides. Copolymerization allows for stable functional motifs, including wedges, interstrand-wedges, dimers and clusters. By selecting suitable functional molecules and motifs, we can design photofunctional oligonucleotides, such as: (1) photoresponsive DNA that enables reversible formation and dissociation of the duplex by photoirradiation; (2) [2+2] photocycloaddition of stilbene derivatives; (3) orientation-dependent FRET (fluorescence (Förster) resonance energy transfer) systems; (4) sequence-specific fluorescent probe for the detection of DNA and RNA; and (5) functional siRNA for fluorescent labeling of mature RISC (RNA-induced silencing complex).
Similar content being viewed by others
Introduction
In addition to its function as a carrier of genetic information, DNA has also been engineered as a nanomaterial for the design of nanomachines and nanostructures.1, 2, 3 Although various DNA nanoarchitectures have been reported,4, 5, 6 possible functions are limited by only four natural nucleobases. For further evolution of DNA nanotechnology, new methodology that enables incorporation of novel functional molecules into DNA and RNA is highly desired. Conventionally, functional molecules have been introduced by chemical modification of natural nucleotides. For example, the 5-position of the pyrimidine base is a suitable modification site because the resulting functionality does not interfere with canonical Watson–Crick base pairing.7, 8 Similarly, functional molecules have been tethered to the 2′-hydroxyl group on ribose.9, 10, 11 Although these modifications are expected to minimize deviations from B- or A-type duplexes, several drawbacks exist. The synthetic modification of nucleotides is laborious. For example, it takes seven steps to synthesize the phosphoramidite monomer of azobenzene tethered to uridine at the 2′-position when uridine is used as a starting material.12 Four modified monomers, corresponding to A, U(T), G and C, are necessary to incorporate multiple functional molecules at any position of the sequence. Furthermore, since multiple modifications at these positions often destabilize the duplex,13 only minor changes to the duplex are permitted.
Herein, we demonstrate a simple but effective methodology of DNA and RNA modification. We applied the concept of copolymerization, often employed in the field of polymer chemistry, to the synthesis of functionalized DNA and RNA. The newly designed base surrogate d-threoninol (2-amino-1,3-butanediol) is used as a scaffold for the functional molecule (Figure 1).14 This diol is commercially available, or it can be easily obtained by the reduction of commercially available d-threonine. A functional molecule, incorporated at the 2 position of the threoninol via an amide bond, can be easily converted to a phosphoramidite monomer.15 Even positively charged molecules that are difficult to tether to deoxyribose can be introduced.16 Copolymerization of the surrogate monomer with the monomers of four natural nucleotides on a solid support allows for ‘full model change’ of the DNA duplex. Multiple functional base surrogates can be introduced at any position in the sequence. In this review, we show four stable functional motifs designed with these surrogates as well as several photofunctional DNA and RNA oligonucleotides based on these motifs.
Synthetic route of the phosphoramidite monomer of functionalized molecules tethered to threoninol as a scaffold, and the basic design of a sequence with the base surrogate. Note that the counterstrand, with respect to the modified strand, is native DNA or RNA, and the base surrogate in the duplex faces a phosphodiester linkage in the counterstrand.
Basic sequence design: insertion of a base surrogate on d-threoninol
The base surrogate does not replace one of the nucleotides in the sequence but rather is inserted into the duplex, creating a single-bulge (lower panel in Figure 1).15 Note that the counterstrand is a native DNA or RNA oligomer. The use of d-threoninol is crucial, because functional molecules bound to the l-form significantly destabilize the duplex while those on the d-form do not.14 The functional molecule on the threoninol is intercalated between the adjacent base-pairs, which is evidenced by NOESY.17 Figure 2 shows NOESY correlations between the aromatic and imino proton signals of a DNA duplex modified with trans-azobenzene on d- or l-threoninol. In both scaffolds, distinct NOE signals are observed between the imino protons (T10, G4) of the nucleobases and the aromatic protons (H8(H12), H9(H11)) of azobenzene, unambiguously demonstrating the intercalation. On d-threoninol, the NOE signal between G4 and H8(H12) or H9(H11) appears with similar signal intensity to that between T10 and H8(H12) or H9(H11) (Figure 2a), indicating that azobenzene is symmetrically intercalated between the G4–C9 and A3–T10 pairs without distorting the B-type duplex. However, in the NOESY plot of the l-form, the NOE signal between G4 and H8(H12) or H9(H11) is stronger than that between T10 and H8(H12) or H9(H11) (Figure 2b). This asymmetrical NOE suggests that azobenzene is stacked closer to the pair G4–C9 than to T10–A3, indicating distortion of the canonical B-type duplex. Destabilization of the duplex is enhanced when multiple azobenzenes are tethered to the l-form.14
Four functional motifs designed by the copolymerization of natural nucleotides and the base surrogates on d-threoninol
The conjugation of natural nucleotides with the base surrogates on d-threoninol allows for functional copolymers. As mentioned above, this sequence design does not sacrifice natural base-pairs. Furthermore, stacking interactions between the functional molecule and the base-pairs more than compensates for the destabilization caused by insertion of the surrogate, as long as the functional molecule is a planar structure of suitable size.18 Hence, the incorporation of multiple surrogates actually stabilizes the duplex. By copolymerizing these surrogates with natural nucleotides, the four stable functional motifs shown in Figure 3 have been developed.19
Wedge motif
One strand is a copolymer of natural nucleotides and surrogates, whereas its counterstrand is a ‘homo’ polymer of natural nucleotides (Figure 3a).18 Upon duplex formation, two or more nucleotides are inserted between the surrogates to avoid destabilization induced by asymmetrical sequence design. 2D-NOESY revealed that multiple azobenzenes were intercalated between the adjacent base-pairs (Supplementary Figure 1).
Interstrand-wedge motif
Both strands are copolymers of nucleotides and surrogates, and two nucleotides are inserted between the surrogates.20 Hybridization of these two strands results in an alternating copolymer of base-pairs and surrogates (Figure 3b). NMR measurements revealed that all the functional molecules bound to threoninol moieties were intercalated without disturbing the base pairing of the natural nucleotides.21 This motif is remarkably more stable than the wedge motif due to its symmetrical sequence design.
Dimer motif
Both strands are alternating copolymers of nucleotides and surrogates (Figure 3c). As the surrogates are located at base pairing positions in each strand, hybridization results in stacked pairs of surrogates as well as base pairing of nucleotides.22, 23 This stacking interaction between the functional molecules, as well as the symmetrical sequence design, remarkably increases the stability of the duplex.
Cluster motif
The cluster motif is an extension of the dimer motif. Both strands are composed of ABA-type block copolymers, and the surrogates are alternately stacked in the duplex (Figure 3d).24, 25, 26 Consecutive base surrogates are not necessarily arranged at the center of the strands; terminal positions are also possible.27 Interestingly, this motif accepts various functional molecules. Even non-planar molecules, such as cyclohexyl groups, can be assembled, stabilizing the duplex by hydrophobic interactions.28, 29, 30 If the interaction between the surrogates is sufficiently strong, stable hybrid clusters can be prepared without the assistance of natural base-pairs.31
By selecting suitable functional molecules and motifs, functions that are difficult to achieve with natural DNA and RNA can be realized. In the next section, five kinds of photofunctional oligonucleotides that have been developed are introduced.
Photoregulation of the formation and dissociation of DNA duplexes by azobenzene-tethered DNA
Spontaneous hybridization is a key feature that characterizes DNA and RNA as supramolecular assemblies. Almost all of the DNA-based nanomachines or nanoarchitectures, as well as enzymes that catalyze DNA replication or transcription, are based on this supramolecular property. Hence, if the formation and dissociation of DNA duplexes can be controlled via irradiation with light of a specific wavelength, photon-driven nanomachines and the photoregulation of gene expression can be expected. For this purpose, photoswitchable molecules are introduced into DNA or RNA. Among photoresponsive molecules, azobenzene derivatives are suitable for this method because (1) they are photochemically stable and do not decompose in aqueous environments; and (2) trans–cis structural isomerization reversibly occurs by irradiating azobenzene with light of wavelength longer than 300 nm.32 Multiple azobenzene moieties have been tethered to DNA via d-threoninol, and reversible photoregulation of duplex formation has been realized.18 In the trans form (λ>400 nm), the planar azobenzenes are intercalated and stabilize the duplex.17 In the cis form (300 nm<λ<400 nm), however, the non-planar structures interfere with the base pairing of adjacent nucleobases,14, 33 which results in dissociation of the duplex. For the photoregulation of hybridization, all four functional motifs are available. Figure 4a shows a schematic illustration of the photoresponsive hairpin-tethering of six azobenzenes in an Interstrand-wedge motif.20 Visible light irradiation isomerizes azobenzene to the planar trans form, which closes the hairpin. In contrast, ultraviolet (UV) light irradiation induces a random coil transition due to the dissociation of the stem. A gel-shift assay (Figure 4b) revealed that the band appeared only at the hairpin position after visible light irradiation, whereas a new band emerged at the random coil position after UV irradiation.20 Thus, perfect photoregulation of the formation and dissociation of the DNA duplex was attained using an Interstrand-wedge motif with six tethered azobenzenes. Photon-driven nanomachines (Figure 5a),34 photoswitchable DNA enzymes (Figure 5b),35, 36 and photoresponsive nanocapsules (Figure 5c)37 have been designed based on the reversible photoregulation of hybridization with photoresponsive oligonucleotides. Other photoregulated systems have been successfully designed using the azobenzene derivatives.38, 39, 40, 41
(a) Schematic illustration of the formation and dissociation of a photoresponsive hairpin involving six azobenzenes on d-threoninol in an Interstrand-wedge motif. (b) Gel-shift assay of the photoresponsive hairpin after UV or visible light irradiation.20 Lane 1, 30-nt single-stranded DNA; Lane 2, the photoresponsive hairpin after 3 min UV irradiation at 60 °C; Lane 3, the photoresponsive hairpin after 1 min visible light irradiation; Lane 4, the native hairpin DNA without azobenzene moieties. Adapted with permission from Liang et al.20 Copyright (2009) Wiley-VCH Verlag GmbH & Co. KGaA
[2+2] Photocycloaddition of aggregated stilbene derivatives in DNA duplexes
Stilbene is well-known as a photoresponsive molecule that can be subjected to photoisomerization between its trans and cis form. Unlike azobenzene, cis-to-trans photoisomerization of stilbene requires UV light of wavelengths shorter than 300 nm,42 which can also cause the formation of undesired thymine dimers. Furthermore, intercalated trans stilbene is not easily photoisomerized, as isomerization requires rotation of the benzene ring around the C=C bond through disruption of the π-bond,42 which needs sufficiently wide space. Hence, stilbene derivatives cannot be used in place of azobenzenes for the reversible photoregulation of hybridization. However, stilbene derivatives undergo [2+2] photocycloaddition reactions,43, 44, 45 which can be useful for the covalent crosslinking of two strands. We have introduced stilbazoles into duplexes as a dimer motif and applied them to the crosslinking of the two strands via photocycloaddition (Figure 6).46 Before photoirradiation, single species of the stable dimers were confirmed by NMR analysis. UV spectra and TOF-MS analysis revealed that photocycloaddition reactions occurred rapidly following irradiation with UV light at 340 nm, which significantly increased the stability of the duplex. However, both HPLC and NMR analyses showed that two photocrosslinked products were generated. Before photoirradiation, stilbazole can adopt two conformations, between which interconversion is possible, via rotation about sigma bonds. Very fast interconversion resulted in averaged NMR signals, preventing discrimination between the two conformers. After UV irradiation, however, the photocycloaddition reaction yielded two diastereomers, because the two conformers were fixed by the photocycloaddition reaction (Figure 7).
Molecular modeling, as well as NMR analysis, revealed that the stilbene dimer requires a suitable conformation for photocycloaddition in the DNA duplex, which allowed us to investigate the intrinsic reactivity of stilbene derivatives. Photocycloadditions have also been investigated in homogeneous solutions, in which photoreaction follows dimer formation. Accordingly, photocycloaddition obeys second-order reaction kinetics and depends on the association constant of dimer formation as well as the intrinsic photoreactivity. In this copolymerized DNA-based system, the dimer is already formed in the duplex, so the association step has already occurred and only the intrinsic photoreactivity determines the rate. Thus, the photocycloaddition becomes first-order in the DNA duplex at temperatures below the melting temperature, and the rate of reaction reflects only the intrinsic photoreactivity. Yamada et al. previously reported that photocycloaddition reactions of stilbazole proceeded faster in the presence of HCl.47 This acceleration was hypothesized to be due to favorable dimer formation associated with the positive charge on the protonated pyridine ring, but an increase in the intrinsic reactivity by protonation could not be ruled out. Our investigation of the dimer motif revealed that the first-order reaction rate of stilbazole was almost the same as that of methylstilbazolium,46 which supports the above hypothesis. This study demonstrates that our system is suitable for evaluating the intrinsic reactivity of stilbene derivatives. We synthesized various stilbene derivatives, shown in Figure 6, and evaluated the intrinsic reactivities of the resulting homo-dimers.48 We found that the reactivity was closely related to the excitation energy. Recently, we have developed a styrylpyrene moiety (Figure 6) that effects photocycloaddition with visible light irradiation (455 nm).49 Although its quantum yield is not high, due to the low excitation energy, relatively harmless 450 nm light is advantageous for biological applications. Interestingly, this crosslinking reaction was reversible; the cycloreversion (the reverse reaction of cycloaddition) occurred upon irradiation with light of λ=340 nm. Like azobenzene, styrylpyrene can induce crosslinking and un-crosslinking of two strands reversibly, via irradiation with UV (λ=340 nm) or visible light (λ=455 nm).
Compared to the homo-clusters, hetero-clusters are very difficult to prepare. Thus, cross-photocycloaddition between different stilbene derivatives has not yet been achieved. However, using our interstrand clustering method,26, 50 it is easy to prepare hetero-dimers of stilbene derivatives by hybridizing two complementary strands with different chromophores (Figure 6). We investigated the cross-reactivity of stilbene derivatives and found that reactivity was highly correlated with the highest occupied molecular orbital (HOMO)/lowest unoccupied molecular orbital (LUMO) energy gaps between the derivatives.48 Interestingly, non-reactive nitrostilbene was able to cross-react with methylstilbazolium and stilbazole, because these two moieties have HOMO/LUMO energies similar to that of nitrostilbene.
Orientation-dependent FRET in the DNA duplex
Fluorescence (Förster) resonance energy transfer (FRET), nonradiative energy transfer from an excited donor fluorophore to an acceptor chromophore,51, 52 is an effective method for the control of the fluorescent behaviors of chromophore assemblies. FRET is mainly applied to the measurement of the distance between biomolecules labeled with a donor and an acceptor in nanoscale (that is, as a molecular ruler).53 Increases in the apparent Stokes’ shift improve the sensitivity of fluorescent probe.54 Theory predicts that FRET efficiency depends on both the distance between, and orientation of, the donor and acceptor dyes. Practically, most FRET analyses focus on the distance-dependence by assuming that mutual orientation is averaged by free movement. However, if the chromophores are fixed and their movement is restricted, FRET efficiency should be affected by their orientation as well as the distance. Only limited reports have been presented on orientation-dependent FRET.55, 56
The DNA duplex is an ideal scaffold to compare theoretical prediction of FRET with experiments, because the canonical B-type duplex takes a rigid helical structure and its structural parameters have been determined. Accordingly, if the donor and acceptor are fixed in the duplex, and the number of base-pairs between them are systematically changed, both the distance and orientation of FRET pairs are simultaneously controlled in a predictable manner.55, 56 We have introduced a typical FRET pair, pyrene and perylene, into a DNA duplex via d-threoninol as donor and acceptor, respectively (Figure 8a).54, 57 Firmly intercalated chromophores cannot move freely, so FRET efficiency is expected to reflect orientation between the donor and acceptor as well as distance. Figures 8b and c shows static emission intensities of pyrene and perylene as a function of the number of base-pairs between them. As the number of base-pairs increased, a decrease in the perylene emission and an increase in the pyrene emission were synchronously observed, reflecting distance-dependent FRET. However, these changes were not monotonous, but periodic: local maxima and minima of pyrene and perylene emission at 8, 13 and 18 base-pairs were observed, indicating unusual decreases in FRET efficiency.58 Similar local maxima were also observed in the life-time of pyrene at the same number of base-pairs. This cycle of five base-pairs corresponds to a half-turn (180°) of the B-type helix (10.4 base-pairs per turn), clearly reflecting the orientation between the pyrene and perylene moieties.
Orientation-dependent FRET system using a DNA duplex in which d-threoninol-tethered pyrene and perylene function as donor and acceptor, respectively.58 (a) Sequence design of the DNA duplex. (b) Emission intensity of pyrene as a function of inserted number of A–T pairs. (c) Emission intensity of perylene as a function of inserted number of A–T pairs. (d) Comparison of experimentally obtained FRET efficiencies (closed circles, static fluorescence of pyrene; open circles, time-resolved fluorescence spectroscopy) with theoretically determined values (cross marks and solid gray line, cylinder model; broken lines, averaged orientation model). Adapted with permission from Kato et al.58 Copyright (2013) American Chemical Society.
FRET efficiencies experimentally determined from both static fluorescence and life-time measurements of pyrene were plotted against the number of base-pairs (Figure 8d, the ‘FRET plot’). The decline in efficiency at 8, 13 and 18 base-pair separations suggests that the transition dipole moments of pyrene and perylene are perpendicularly oriented. We also compared these experimental values with theoretical prediction by assuming B-type geometry, which typically exhibits the structural parameters of axial rise and rotation angle per residue of 3.2 Å and 36°, respectively. As shown in Figure 8d, the calculated values are in excellent agreement with the experimental values.58 This result clearly demonstrates that this FRET system using donors and acceptors bound to d-threoninol precisely reflects the orientation and distance factors of B-type geometry and that the FRET plots provide duplex structural information. An advantage of this method is that it is not limited to pyrene–perylene; any donor–acceptor combination can be selected. These FRET plots can be applied in the structural analysis of oligonucleotides, even in the case of artificial duplexes in aqueous solution, through selection of a suitable donor–acceptor pair.
In-stem molecular beacons and stemless linear probes that detect DNA and RNA with sequence specificity
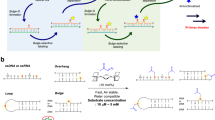
The sequence-specific detection of DNA and RNA is a key biological tool for individualized medicine and for visualization of functional RNAs that play important roles in cells. As hybridization occurs sequence-specifically, even one mismatched pair destabilizes the duplex. However, silent natural bases cannot give rise to visible strand recognition signals. Hence, probe molecules that convert strand recognition into detectable signals are introduced into oligonucleotides. Conventionally, molecular beacons that tether fluorophores and quenchers at either termini of hairpin DNA have been applied to the fluorescent detection of DNA and RNA (Figure 9a).59 In the absence of target DNA or RNA, the fluorophore does not emit light due to the contact with the quencher in the closed hairpin. Hybridization with the target opens the hairpin and lights up the fluorophore that has been separated from the quencher. Accordingly, MBs (molecular beacon) can convert strand recognition into a fluorescent signal. However, MBs have several drawbacks. Only one pair of fluorophore and quencher can be incorporated at the termini; multiple fluorophores are not possible. Additionally, insufficient contact between the fluorophore and quencher in the absence of target DNA, results in background emission due to breathing effects of the terminal pairs in the closed MB, which lowers the sensitivity. Finally, hybridization of the target with the loop region of the MB is followed by the dissociation of the stem, which delays the response speed.
To overcome these drawbacks, we have developed an in-stem molecular beacon (ISMB) that involves multiple fluorophore-quencher pairs bound to d-threoninols at the stem regions as a dimer motif (Figure 9b).60, 61, 62, 63 This design allows for the incorporation of multiple fluorophores and an increase in emission intensity when the ISMB is hybridized with the target. Strong contact between the fluorophore and quencher in the stem remarkably suppresses background emissions in the absence of the target. As a result, the signal-to-background ratio (S/B ratio) is greatly improved. However, incorporation of many fluorophore-quencher pairs requires longer stems, which further delayed the response rate. Without the assistance of a chaperone polymer, the slow response time of the ISMB could not be solved.64
To overcome all of the above drawbacks, we have recently designed a stemless linear probe involving multiple fluorophores in DNA in a wedge motif (Figure 9c).65 In the absence of the target, the flexible linear probe facilitates self-quenching of the fluorophores. Self-quenching is improved greatly when the number of fluorophores increases. Hybridization with the target separates each fluorophore by intercalation and causes all of the fluorophores to fluoresce. Hence, as the number of fluorophores increases, the sensitivity of the probe becomes greater. The absence of a stem structure greatly improved the response rate, and the second-order rate constant of the linear probe is ~80-fold greater than that of the MB targeting the same sequence.65 The introduction of fluorescence quenchers at the termini is also an effective way to minimize background emission. We have designed a stemless linear probe with four perylenes and two anthraquinones as fluorophore and quencher, respectively (Figure 10a). Using this system, the S/B ratio became as large as 1600.66
(a) Fluorescent emission spectra of a linear probe measured at 20 °C in the presence (solid blue line) and absence (dotted red line) of target RNA. (b) Fluorescent imaging of mRNA transcribed from a DsRed-encoding plasmid by the linear probe.66 Upper panel, linear probe only; lower panel, linear probe and plasmid. Adapted with permission from Asanuma et al.66 Copyright (2015) Wiley-VCH Verlag GmbH & Co. KGaA
Our linear probe has another advantage over MBs. The loop regions of MBs are typically composed only of natural DNA, and MBs are easily degraded in cell lysate, causing them to emit light even in the absence of the target.67 This low nuclease resistance prevents the application of MBs in cells. In contrast, a linear probe, in which non-natural base surrogates are intermittently inserted into DNA, has improved nuclease resistance.66 This property is advantageous for fluorescent imaging of RNA in living cells. A linear probe targeting a region of the DsRed gene was designed (Figure 10a) and applied to the fluorescent imaging of mRNA in living cells. As designed, the linear probe exhibited a blue emission from cells transfected with a plasmid encoding DsRed (Figure 10b).66
The linear probe can label not only single-stranded DNA or RNA but also double-stranded DNA (dsDNA), with the aid of unmodified peptide nucleic acid (PNA). As base surrogates in the oligonucleotide interfere with hybridization with PNA, the linear probe does not form a stable duplex with its complementary PNA and remains quenched even in its presence. In contrast, both the linear probe and PNA form stable duplexes with DNA, so that dsDNA can be fluorescently labeled in a strand-invasive manner (Figure 11). We designed a linear probe with a perylene derivative and anthraquinone and achieved strand-invasive labeling of dsDNA in the presence of PNA using heat-shock-treatment-like polymerase chain reaction.68
Double-duplex invasion of a linear probe and unmodified PNA (peptide nucleic acid) for the fluorescent labeling of double-stranded DNA.68
Fluorescent trafficking of siRNA using a fluorophore-quencher pair
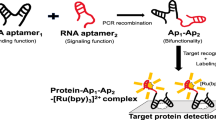
Short interference RNAs (siRNA) are now commonly used to knockout specific genes through an endogenous RNA interference (RNAi) pathway, due to their powerful silencing ability.69, 70, 71, 72 It is widely accepted that the antisense (guide) strand of siRNA is incorporated into the RNAi machinery to form an RNA-induced silencing complex (RISC). However, the RNAi machinery in living cells is not yet fully understood. The development of an siRNA-based molecular probe would thus be a useful tool for tracing the fate of siRNA in cells. For this purpose, we designed functional siRNA (Figure 12) in which a fluorophore (thiazole orange; TO) and a quencher (methyl red; MR) pair was incorporated as a dimer motif near the 5′-terminus of the sense strand.73 The siRNA does not emit fluorescence before incorporation into the RNA machinery due to the close contact of TO with MR, whereas the mature RISC involving the single-stranded antisense sequence exhibits emission from TO. The introduction of the dimer at the fifth position from the 5′-terminus of the sense strand is particularly important because this specific position allows for the selective loading of the antisense strand to RISC without decreasing the RNAi activity.73, 74 Using this functional siRNA, we could successfully monitor the intracellular fate of siRNA in living cells.
Design of functional siRNA (short interference RNA) for fluorescent tracing of its fate.73
Conclusions
In this review, methodology for the novel functionalization of DNA and RNA has been demonstrated. Because the copolymerization concept is simple and easily extended, other applications in addition to those enumerated above have been achieved, including a quantum-dot-like fluorophore assembly75 and new artificial nucleic acids, acyclic threoninol nucleic acid76, 77 and serinol nucleic acid,78, 79 that can form more stable homo-duplexes than DNA or RNA. Due to facile programmability and availability, fields beyond nanotechnology have turned to DNA and RNA. Recent years have seen the development of a new field of ‘molecular robotics,’ in which integrated nano- to microsystems of sensors, logic gates and actuators operate autonomously in response to the environment.80 In this novel field, DNA and RNA are regarded as useful nanomaterials for device design. We believe that the concept of ‘copolymerization,’ together with new XNAs, will contribute to the development of these new fields.
References
Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Yurke, B., Turberfield, A. J., Mills, A. P. Jr, Simmel, F. C. & Neumann, J. L. A DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Rothemund, P. K. W. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Aldaye, F. A., Palmer, A. L. & Sleiman, H. F. Assembling materials with DNA as the guide. Science 321, 1795–1799 (2008).
Seeman, N. C. Nanomaterials based on DNA. Annu. Rev. Biochem. 79, 65–87 (2010).
Tørring, T., Voigt, N. V., Nangreave, J., Yan, H. & Gothelf, K. V. DNA origami: a quantum leap for self-assembly of complex structures. Chem. Soc. Rev. 40, 5636–5646 (2011).
Ahmadian, M., Zhang, P. M. & Bergstrom, D. E. A comparative study of the thermal stability of oligodeoxyribonucleotides containing 5-substituted 2'-deoxyuridines. Nucleic Acids Res. 26, 3127–3135 (1998).
Graham, D., Parkinson, J. A. & Brown, T. DNA duplexes stabilized by modified monomer residues: synthesis and stability. J. Chem. Soc. Perkins Trans. 1, 1131–1138 (1998).
Yamana, K., Nishijima, Y., Ikeda, T., Gokota, T., Ozaki, H., Nakano, H., Sangen, O. & Shimidzu, T. Synthesis and interactive properties of an oligonucleotide with anthraquinone at the sugar fragment. Bioconjugate Chem. 1, 319–324 (1990).
Yamana, K., Iwase, R., Furutani, S., Tsuchida, H., Zako, H., Yamaoka, T. & Murakami, A. 2’-Pyrene modified oligonucleotide provides a highly sensitive fluorescent probe of RNA. Nucleic Acids Res. 27, 2387–2392 (1999).
Umek, R. M., Lin, S. W., Vielmetter, J., Terbrueggen, R. H., Irvine, B., Yu, C. J., Kayyem, J. F., Yowanto, H., Blackburn, G. F., Farkas, D. H. & Chen, Y. P. Electronic detection of nucleic acids: a versatile platform for molecular diagnostics. J. Mol. Diagn. 3, 74–84 (2001).
Asanuma, H., Yoshida, T., Ito, T. & Komiyama, M. Photo-responsive oligonucleotides carrying azobenzene at the 2'-position of the ribonucleotide. Tetrahedron Lett. 40, 7995–7998 (1999).
Zatsepin, T. S., Gait, M. J. & Oretskaya, T. S. 2'-functionalized nucleic acids as structural tools in molecular biology. IUBMB Life 56, 209–214 (2004).
Asanuma, H., Takarada, T., Yoshida, T., Liang, X. G. & Komiyama, M. Enantioselective incorporation of azobenzenes into oligodeoxyribonucleotide for the effective photo-regulation of duplex formation. Angew. Chem. Int. Ed. Engl. 40, 2671–2673 (2001).
Kashida, H., Liang, X. G. & Asanuma, H. Rational design of functional DNA with a non-ribose acyclic scaffold. Curr. Org. Chem. 13, 1065–1084 (2009).
Kashida, H., Ito, H., Fujii, T., Hayashi, T. & Asanuma, H. Positively charged base surrogate for highly stable ‘base pairing’ through electrostatic and stacking interactions. J. Am. Chem. Soc. 131, 9928–9930 (2009).
Liang, X., Asanuma, H., Kashida, H., Takasu, A., Sakamoto, T., Kawai, G. & Komiyama, M. NMR study on the photo-responsive DNA tethering an azobenzene. Assignment of the absolute configuration of two diastereomers and structure determination of their duplexes in the trans-form. J. Am. Chem. Soc. 125, 16408–16415 (2003).
Asanuma, H., Liang, X. G., Nishioka, H., Matsunaga, D., Liu, M. Z. & Komiyama, M. Synthesis of azobenzene-tethered DNA for reversible photo-regulation of DNA. Nat. Protoc. 2, 203–212 (2007).
Asanuma, H., Kashida, H. & Kamiya, Y. De novo design of functional oligonucleotides with acyclic scaffolds. Chem. Rec. 14, 1055–1069 (2014).
Liang, X. G., Mochizuki, T. & Asanuma, H. A supra-photoswitch involving sandwiched DNA base pairs and azobenzenes for light-driven nanostructures and nanodevices. Small 5, 1761–1768 (2009).
Liang, X. G., Nishioka, H., Mochizuki, T. & Asanuma, H. An interstrand-wedged duplex composed of alternating DNA base pairs and covalently attached intercalators. J. Mater. Chem. 20, 575–581 (2010).
Liang, X. G., Mochizuki, T., Fujii, T., Kashida, H. & Asanuma, H. Design of a functional nanomaterial with recognition ability for constructing light-driven nanodevices. LNCS 6518, 112–122 (2011).
Liang, X. G., Mochizuki, T., Fujii, T., Kashida, H. & Asanuma, H. Design of an artificial functional nanomaterial with high recognition ability. Nat. Comput. 11, 231–238 (2012).
Kashida, H., Fujii, T. & Asanuma, H. Threoninol as a scaffold of dyes (threoninol-nucleotide) and their stable interstrand clustering in duplexes. Org. Biomol. Chem. 6, 2892–2899 (2008).
Fujii, T., Kashida, H. & Asanuma, H. Analysis of coherent heteroclustering of different dyes by use of threoninol nucleotides for comparison with the molecular exciton theory. Chem. Eur. J. 15, 10092–10102 (2009).
Fujii, T., Urushihara, M., Kashida, H., Ito, H., Liang, X. G., Yagi-Utsumi, M., Kato, K. & Asanuma, H. Reversed assembling of dyes in RNA duplex compared with those in DNA. Chem. Eur. J. 18, 13304–13313 (2012).
Kashida, H., Hayashi, T., Fujii, T. & Asanuma, H. A cationic dye triplet as a unique ‘glue’ that can connect fully matched termini of DNA duplexes. Chem. Eur. J. 17, 2614–2622 (2011).
Kashida, H., Sekiguchi, K. & Asanuma, H. Insulator base pairs for lighting-up perylenediimide in a DNA duplex. Chem. Eur. J. 16, 11554–11557 (2010).
Kashida, H., Sekiguchi, K., Higashiyama, N., Kato, T. & Asanuma, H. Cyclohexyl ‘base pairs’ stabilize duplexes and intensify pyrene fluorescence by shielding it from natural base pairs. Org. Biomol. Chem. 9, 8313–8320 (2011).
Kashida, H., Higashiyama, N., Kato, T. & Asanuma, H. Evaluation of intrinsic spectroscopic properties of chromophore assemblies by shielding with cyclohexyl base pairs within a DNA duplex. Bioorg. Med. Chem. 21, 6191–6197 (2013).
Doi, T., Sakakibara, T., Kashida, H., Araki, Y., Wada, T. & Asanuma, H. Hetero-selective DNA-like duplex stabilized by donor-acceptor interaction. Chem. Eur. J. 21, 15974–15980 (2015).
Kamiya, Y. & Asanuma, H. Light-driven DNA nanomachine with a photoresponsive molecular engine. Acc. Chem. Res. 47, 1663–1672 (2014).
Liu, M., Asanuma, H. & Komiyama, M. Azobenzene-tethered T7 promoter for efficient photoregulation of transcription. J. Am. Chem. Soc. 128, 1009–1015 (2006).
Liang, X. G., Nishioka, H., Takenaka, N. & Asanuma, H. A DNA nanomachine powered by light irradiation. ChemBioChem 9, 702–705 (2008).
Zhou, M. G., Liang, X. G., Mochizuki, T. & Asanuma, H. A light-driven DNA nanomachine for efficiently photoswitching RNA digestion. Angew. Chem. Int. Ed. Engl. 49, 2167–2170 (2010).
Liang, X. G., Zhou, M. G., Kato, K. & Asanuma, H. Photoswitch nucleic acid catalytic activity by regulating topological structure with a universal supra-photoswitch. ACS Synth. Biol. 2, 194–202 (2013).
Tanaka, F., Mochizuki, T., Liang, X. G., Asanuma, H., Tanaka, S., Suzuki, K., Kitamura, S., Nishikawa, A., Ui-Tei, K. & Hagiya, M. Robust and photo-controllable DNA capsules using azobenzenes. Nano Lett. 10, 3560–3565 (2010).
Liang, X. G., Takenaka, N., Nishioka, H. & Asanuma, H. Molecular design for reversing the photoswitching mode of turning ON and OFF DNA hybridization. Chem. Asian J. 3, 553–560 (2008).
Nishioka, H., Liang, X. G. & Asanuma, H. Effect of the ortho modification of azobenzene on the photoregulatory efficiency of DNA hybridization and thermal stability of its cis-form. Chem. Eur. J. 16, 2054–2062 (2010).
Liang, X. G., Wakuda, R., Fujioka, K. & Asanuma, H. Photoregulation of DNA transcription by using photoresponsive T7 promoters and clarification of its mechanism. FEBS J. 277, 1551–1561 (2010).
Kamiya, Y., Takagi, T., Ooi, H., Ito, H., Liang, X. G. & Asanuma, H. Synthetic gene involving azobenzene-tethered T7 promoter for the photocontrol of gene expression by visible light. ACS Synth. Biol. 4, 365–370 (2015).
Bandara, H. M. D. & Burdette, S. C. Photoisomerization in different classes of azobenzene. Chem. Soc. Rev. 41, 1809–1825 (2012).
Goodall, G. W. & Hayes, W. Advances in cycloaddition polymerizations. Chem. Soc. Rev. 35, 280–312 (2006).
Daku, L. M. L., Linares, J. & Boillot, M.-L. Ab initio static and molecular dynamics study of the absorption spectra of the 4-styrylpyridine photoswitch in its cis and trans forms. Phys. Chem. Chem. Phys. 12, 6107–6123 (2010).
Schraub, M. & Hampp, N. Selective [2+2]-cycloaddition in methacrylic stilbene polymers without interference from E/Z-isomerization. Macromolecules 44, 8755–8762 (2011).
Kashida, H., Doi, T., Sakakibara, T., Hayashi, T. & Asanuma, H. p-Stilbazole moieties as artificial base pairs for photo-cross-linking of DNA duplex. J. Am. Chem. Soc. 135, 7960–7966 (2013).
Yamada, S., Uematsu, N. & Yamashita, K. Role of cation-π interactions in the photodimerization of trans-4-styrylpyridines. J. Am. Chem. Soc., 129, 12100–12101 (2007).
Doi, T., Kashida, H. & Asanuma, H. Efficiency of [2+2] photodimerization of various stilbene derivatives within the DNA duplex scaffold. Org. Biomol. Chem. 13, 4430–4437 (2015).
Doi, T., Kawai, H., Murayama, K., Kashida, H. & Asanuma, H. Visible-light-triggered cross-linking of DNA duplexes by reversible [2++2] photocycloaddition of styrylpyrene. Chem. Eur. J. 22, 10533–10538 (2016).
Kashida, H., Asanuma, H. & Komiyama, M. Alternating hetero H aggregation of different dyes by interstrand stacking from two DNA–dye conjugates. Angew. Chem. Int. Ed. Engl. 43, 6522–6525 (2004).
Jares-Erijman, E. A. & Jovin, T. M. FRET imaging. Nat. Biotechnol. 21, 1387–1395 (2003).
Preus, S. & Wilhelmsson, L. M. Advances in quantitative FRET-based methods for studying nucleic acids. ChemBioChem 13, 1990–2001 (2012).
Sapsford, K. E., Berti, L. & Medintz, I. L. Materials for fluorescence resonance energy transfer analysis: beyond traditional donor-acceptor combinations. Angew. Chem. Int. Ed. Engl. 45, 4562–4588 (2006).
Kashida, H., Takatsu, T., Sekiguchi, K. & Asanuma, H. An efficient fluorescence resonance energy transfer (FRET) between pyrene and perylene assembled in a DNA duplex and its potential for discriminating single-base changes. Chem. Eur. J. 16, 2479–2486 (2010).
Iqbal, A., Arslan, S., Okumus, B., Wilson, T. J., Giraud, G., Norman, D. G., Ha, T. & Lilley, D. M. J. Orientation dependence in fluorescent energy transfer between Cy3 and Cy5 terminally attached to double-stranded nucleic acids. Proc. Natl. Acad. Sci. USA 105, 11176–11181 (2008).
Börjesson, K., Preus, S., El-Sapheer, A. H., Brown, T., Albinsson, B. & Wilhelmsson, L. M. Nucleic acid base analog FRET-pair facilitating detailed structural measurements in nucleic acid containing systems. J. Am. Chem. Soc. 131, 4288–4293 (2009).
Masuko, M., Ouchi, S., Sode, K., Ohtani, H. & Shimadzu, A. Fluorescence resonance energy transfer from pyrene to perylene labels for nucleic acid hybridization assays under homogeneous solution conditions. Nucleic Acids Res. 28, e34 (2000).
Kato, T., Kashida, H., Kishida, H., Yada, H., Okamoto, H. & Asanuma, H. Development of a robust model system of FRET using base-surrogates tethering fluorophores for strict control of their position and orientation within DNA duplex. J. Am. Chem. Soc. 135, 741–750 (2013).
Tyagi, S. & Kramer, F. R. Molecular beacons: probes that fluoresce upon hybridization. Nat. Biotechnol. 14, 303–308 (1996).
Kashida, H., Takatsu, T., Fujii, T., Sekiguchi, K., Liang, X. G., Niwa, K., Takase, T., Yoshida, Y. & Asanuma, H. In-stem molecular beacon containing a pseudo base pair of threoninol nucleotides for the removal of background emission. Angew. Chem. Int. Ed. Engl. 48, 7044–7047 (2009).
Hara, Y., Fujii, T., Kashida, H., Sekiguchi, K., Liang, X. G., Niwa, K., Takase, T., Yoshida, Y. & Asanuma, H. Coherent quenching of a fluorophore for the design of a highly sensitive in-stem molecular beacon. Angew. Chem. Int. Ed. Engl. 49, 5502–5506 (2010).
Fujii, T., Hara, Y., Osawa, T., Kashida, H., Liang, X. G., Yoshida, Y. & Asanuma, H. Bulge-like asymmetric hetero dye-clustering in DNA duplex results in efficient quenching of background emission based on the maximized excitonic interaction. Chem. Eur. J. 18, 10865–10872 (2012).
Kashida, H., Osawa, T., Morimoto, K., Kamiya, Y. & Asanuma, H. Molecular design of Cy3 derivative for highly sensitive in-stem molecular beacon and its application to the wash-free FISH. Bioorg. Med. Chem. 23, 1758–1762 (2015).
Asanuma, H., Osawa, T., Kashida, H., Fujii, T., Liang, X. G., Niwa, K., Yoshida, Y., Shimada, N. & Maruyama, A. Polycation-chaperoned in-stem molecular beacon system. Chem. Commun. 48, 1760–1762 (2012).
Asanuma, H., Akahane, M., Kondo, N., Osawa, T., Kato, T. & Kashida, H. Quencher-free linear probe with multiple fluorophores on acyclic scaffold. Chem. Sci. 3, 3165–3169 (2012).
Asanuma, H., Akahane, M., Niwa, R., Kashida, H. & Kamiya, Y. Highly sensitive and robust linear probe for detection of mRNA in cells. Angew. Chem. Int. Ed. Engl. 54, 4315–4319 (2015).
Murayama, K., Kamiya, Y., Kashida, H. & Asanuma, H. Ultra-sensitive molecular beacon designed with totally serinol nucleic acid (SNA) for monitoring mRNA in cell. ChemBioChem 16, 1298–1301 (2015).
Aasanuma, H., Niwa, R., Akahane, M., Murayama, K., Kashida, H. & Kamiya, Y. Strand-invading linear probe combined with unmodified PNA. Bioorg. Med. Chem. 24, 4129–4137 (2016).
Fire, A., Xu, S., Montgomery, M. K., Kostas, S. A., Driver, S. E. & Mello, C. C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Elbashir, S. M., Harborth, J., Lendeckel, W., Yalcin, A., Weber, K. & Tuschl, T. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494–498 (2001).
Kamiya, Y., Takai, J., Ito, H., Murayama, K., Kashida, H. & Asanuma, H. Enhancement of stability and activity of siRNA by terminal substitution with serinol nucleic acid (SNA). ChemBioChem 15, 2549–2555 (2014).
Kamiya, Y., Iishiba, K., Doi, T., Tsuda, K., Kashida, H. & Asanuma, H. Terminus-free siRNA prepared by photo-crosslinking activated via slicing by Ago2. Biomater. Sci. 3, 1534–1538 (2015).
Kamiya, Y., Ito, A., Ito, H., Urushihara, M., Takai, J., Fujii, T., Liang, X. G., Kashida, H. & Asanuma, H. Selective labeling of mature RISC using siRNA carrying fluorophore-quencher pair. Chem. Sci. 4, 4016–4021 (2013).
Ito, H., Urushihara, M., Liang, X. G. & Asanuma, H. Improvement of RNAi activity and strand-selectivity of RISC formation by modified siRNA involving intercalators near 5'-termini. ChemBioChem 13, 311–315 (2012).
Kashida, H., Sekiguchi, K., Liang, X. G. & Asanuma, H. Accumulation of fluorophores into DNA duplexes to mimic the properties of quantum dots. J. Am. Chem. Soc. 132, 6223–6230 (2010).
Asanuma, H., Toda, T., Murayama, K., Liang, X. G. & Kashida, H. Unexpectedly stable artificial duplex from flexible acyclic threoninol. J. Am. Chem. Soc. 132, 14702–14703 (2010).
Murayama, K., Kashida, H. & Asanuma, H. Acyclic L-threoninol nucleic acid (L-aTNA) with suitable structural rigidity cross-pairs with DNA and RNA. Chem. Commun. 51, 6500–6503 (2015).
Kashida, H., Murayama, K., Toda, T. & Asanuma, H. Control of the chirality and helicity of oligomers of serinol nucleic acid (SNA) by sequence design. Angew. Chem. Int. Ed. Engl. 50, 1285–1288 (2011).
Murayama, K., Tanaka, Y., Toda, T., Kashida, H. & Asanuma, H. Highly stable duplex formation by artificial nucleic acids aTNA and SNA with acyclic scaffolds. Chem. Eur. J. 19, 14151–14158 (2013).
Hagiya, M., Konagaya, A., Kobayashi, S., Saito, H. & Murata, S. Molecular robots with sensors and intelligence. Acc. Chem. Res. 47, 1681–1690 (2014).
Acknowledgements
This work was supported by the Scientific Research on Innovative Areas ‘Molecular Robotics’ (no. 24104005), a Grant-in-Aid for challenging Exploratory Research (no. 16K12522) from the Ministry of Education, Culture, Sports, Science and Technology, Japan. Support by the Asahi Glass Foundation (H.A.) is also acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Polymer Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Asanuma, H., Murayama, K., Kamiya, Y. et al. Design of photofunctional oligonucleotides by copolymerization of natural nucleobases with base surrogates prepared from acyclic scaffolds. Polym J 49, 279–289 (2017). https://doi.org/10.1038/pj.2016.120
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pj.2016.120
This article is cited by
-
Functionalization of acyclic xenonucleic acid with modified nucleobases
Polymer Journal (2023)
-
Nucleic acid-based fluorescent sensor systems: a review
Polymer Journal (2022)
-
Quantitative evaluation of energy migration between identical chromophores enabled by breaking symmetry
Communications Chemistry (2018)