Abstract
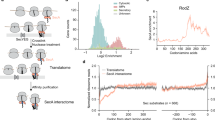
Signal-recognition particle (SRP)-dependent targeting of translating ribosomes to membranes is a multistep quality-control process. Ribosomes that are translating weakly hydrophobic signal sequences can be rejected from the targeting reaction even after they are bound to the SRP. Here we show that the early complex, formed by Escherichia coli SRP and its receptor FtsY with ribosomes translating the incorrect cargo EspP, is unstable and rearranges inefficiently into subsequent conformational states, such that FtsY dissociation is favored over successful targeting. The N-terminal extension of EspP is responsible for these defects in the early targeting complex. The cryo-electron microscopy structure of this 'false' early complex with EspP revealed an ordered M domain of SRP protein Ffh making two ribosomal contacts, and the NG domains of Ffh and FtsY forming a distorted, flexible heterodimer. Our results provide a structural basis for SRP-mediated signal-sequence selection during recruitment of the SRP receptor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Gilmore, R., Walter, P. & Blobel, G. Protein translocation across the endoplasmic reticulum. II. Isolation and characterization of the signal recognition particle receptor. J. Cell Biol. 95, 470–477 (1982).
Keenan, R.J., Freymann, D.M., Stroud, R.M. & Walter, P. The signal recognition particle. Annu. Rev. Biochem. 70, 755–775 (2001).
Nagai, K. et al. Structure, function and evolution of the signal recognition particle. EMBO J. 22, 3479–3485 (2003).
Doudna, J.A. & Batey, R.T. Structural insights into the signal recognition particle. Annu. Rev. Biochem. 73, 539–557 (2004).
Poritz, M.A. et al. An E. coli ribonucleoprotein containing 4.5S RNA resembles mammalian signal recognition particle. Science 250, 1111–1117 (1990).
Freymann, D.M., Keenan, R.J., Stroud, R.M. & Walter, P. Structure of the conserved GTPase domain of the signal recognition particle. Nature 385, 361–364 (1997).
Batey, R.T., Rambo, R.P., Lucast, L., Rha, B. & Doudna, J.A. Crystal structure of the ribonucleoprotein core of the signal recognition particle. Science 287, 1232–1239 (2000).
Janda, C.Y. et al. Recognition of a signal peptide by the signal recognition particle. Nature 465, 507–510 (2010).
Hainzl, T., Huang, S., Merilainen, G., Brannstrom, K. & Sauer-Eriksson, A.E. Structural basis of signal-sequence recognition by the signal recognition particle. Nat. Struct. Mol. Biol. 18, 389–391 (2011).
Montoya, G., Svensson, C., Luirink, J. & Sinning, I. Crystal structure of the NG domain from the signal-recognition particle receptor FtsY. Nature 385, 365–368 (1997).
Angelini, S., Deitermann, S. & Koch, H.G. FtsY, the bacterial signal-recognition particle receptor, interacts functionally and physically with the SecYEG translocon. EMBO Rep. 6, 476–481 (2005).
Weiche, B. et al. A cleavable N-terminal membrane anchor is involved in membrane binding of the Escherichia coli SRP receptor. J. Mol. Biol. 377, 761–773 (2008).
Egea, P.F. et al. Substrate twinning activates the signal recognition particle and its receptor. Nature 427, 215–221 (2004).
Focia, P.J., Shepotinovskaya, I.V., Seidler, J.A. & Freymann, D.M. Heterodimeric GTPase core of the SRP targeting complex. Science 303, 373–377 (2004).
Zhang, X., Rashid, R., Wang, K. & Shan, S.O. Sequential checkpoints govern substrate selection during cotranslational protein targeting. Science 328, 757–760 (2010).
Holtkamp, W. et al. Dynamic switch of the signal recognition particle from scanning to targeting. Nat. Struct. Mol. Biol. 19, 1332–1337 (2012).
Halic, M. et al. Following the signal sequence from ribosomal tunnel exit to signal recognition particle. Nature 444, 507–511 (2006).
Peluso, P., Shan, S.O., Nock, S., Herschlag, D. & Walter, P. Role of SRP RNA in the GTPase cycles of Ffh and FtsY. Biochemistry 40, 15224–15233 (2001).
Zhang, X., Schaffitzel, C., Ban, N. & Shan, S.O. Multiple conformational switches in a GTPase complex control co-translational protein targeting. Proc. Natl. Acad. Sci. USA 106, 1754–1759 (2009).
Estrozi, L.F., Boehringer, D., Shan, S.O., Ban, N. & Schaffitzel, C. Structure of the E. coli co-translational targeting complex in the stable early conformation. Nat. Struct. Mol. Biol. 18, 88–90 (2011).
Lam, V.Q., Akopian, D., Rome, M., Henningsen, D. & Shan, S.O. Lipid activation of the signal recognition particle receptor provides spatial coordination of protein targeting. J. Cell Biol. 190, 623–635 (2010).
Shen, K., Arslan, S., Akopian, D., Ha, T. & Shan, S.O. Activated GTPase movement on an RNA scaffold drives co-translational protein targeting. Nature 492, 271–275 (2012).
Ataide, S.F. et al. The crystal structure of the signal recognition particle in complex with its receptor. Science 331, 881–886 (2011).
Connolly, T., Rapiejko, P. & Gilmore, R. Requirement of GTP hydrolysis for dissociation of the signal recognition particle from its receptor. Science 252, 1171–1173 (1991).
Tian, H., Boyd, D. & Beckwith, J. A mutant hunt for defects in membrane protein assembly yields mutations affecting the bacterial signal recognition particle and Sec machinery. Proc. Natl. Acad. Sci. USA 97, 4730–4735 (2000).
Valent, Q.A. et al. Early events in preprotein recognition in E. coli: interaction of SRP and trigger factor with nascent polypeptides. EMBO J. 14, 5494–5505 (1995).
Lee, H.C. & Bernstein, H.D. The targeting pathway of Escherichia coli presecretory and integral membrane proteins is specified by the hydrophobicity of the targeting signal. Proc. Natl. Acad. Sci. USA 98, 3471–3476 (2001).
Peterson, J.H., Szabady, R.L. & Bernstein, H.D. An unusual signal peptide extension inhibits the binding of bacterial presecretory proteins to the signal recognition particle, trigger factor, and the SecYEG complex. J. Biol. Chem. 281, 9038–9048 (2006).
Szabady, R.L., Peterson, J.H., Skillman, K.M. & Bernstein, H.D. An unusual signal peptide facilitates late steps in the biogenesis of a bacterial autotransporter. Proc. Natl. Acad. Sci. USA 102, 221–226 (2005).
Schuwirth, B.S. et al. Structures of the bacterial ribosome at 3.5 A resolution. Science 310, 827–834 (2005).
Rosendal, K.R., Wild, K., Montoya, G. & Sinning, I. Crystal structure of the complete core of archaeal signal recognition particle and implications for interdomain communication. Proc. Natl. Acad. Sci. USA 100, 14701–14706 (2003).
Schaffitzel, C. et al. Structure of the E. coli signal recognition particle bound to a translating ribosome. Nature 444, 503–506 (2006).
Powers, T. & Walter, P. Co-translational protein targeting catalyzed by the Escherichia coli signal recognition particle and its receptor. EMBO J. 16, 4880–4886 (1997).
Shan, S.O., Chandrasekar, S. & Walter, P. Conformational changes in the GTPase modules of the signal reception particle and its receptor drive initiation of protein translocation. J. Cell Biol. 178, 611–620 (2007).
Zhang, X., Kung, S. & Shan, S.O. Demonstration of a multistep mechanism for assembly of the SRP x SRP receptor complex: implications for the catalytic role of SRP RNA. J. Mol. Biol. 381, 581–593 (2008).
Braig, D. et al. Signal sequence-independent SRP-SR complex formation at the membrane suggests an alternative targeting pathway within the SRP cycle. Mol. Biol. Cell 22, 2309–2323 (2011).
Shen, K. & Shan, S.O. Transient tether between the SRP RNA and SRP receptor ensures efficient cargo delivery during cotranslational protein targeting. Proc. Natl. Acad. Sci. USA 107, 7698–7703 (2010).
Scheres, S.H. et al. Disentangling conformational states of macromolecules in 3D-EM through likelihood optimization. Nat. Methods 4, 27–29 (2007).
Shaikh, T.R. et al. SPIDER image processing for single-particle reconstruction of biological macromolecules from electron micrographs. Nat. Protoc. 3, 1941–1974 (2008).
Zhang, X. et al. Direct visualization reveals dynamics of a transient intermediate during protein assembly. Proc. Natl. Acad. Sci. USA 108, 6450–6455 (2011).
Knoops, K., Schoehn, G. & Schaffitzel, C. Cryo-electron microscopy of ribosomal complexes in cotranslational folding, targeting, and translocation. Wiley Interdiscip. Rev. RNA 3, 429–441 (2012).
Cross, B.C., Sinning, I., Luirink, J. & High, S. Delivering proteins for export from the cytosol. Nat. Rev. Mol. Cell Biol. 10, 255–264 (2009).
Bradshaw, N., Neher, S.B., Booth, D.S. & Walter, P. Signal sequences activate the catalytic switch of SRP RNA. Science 323, 127–130 (2009).
Peterson, J.H., Woolhead, C.A. & Bernstein, H.D. The conformation of a nascent polypeptide inside the ribosome tunnel affects protein targeting and protein folding. Mol. Microbiol. 78, 203–217 (2010).
Raine, A. et al. Targeting and insertion of heterologous membrane proteins in E. coli. Biochimie 85, 659–668 (2003).
Flanagan, J.J. et al. Signal recognition particle binds to ribosome-bound signal sequences with fluorescence-detected subnanomolar affinity that does not diminish as the nascent chain lengthens. J. Biol. Chem. 278, 18628–18637 (2003).
Miyazaki, E., Kida, Y., Mihara, K. & Sakaguchi, M. Switching the sorting mode of membrane proteins from cotranslational endoplasmic reticulum targeting to posttranslational mitochondrial import. Mol. Biol. Cell 16, 1788–1799 (2005).
del Alamo, M. et al. Defining the specificity of cotranslationally acting chaperones by systematic analysis of mRNAs associated with ribosome-nascent chain complexes. PLoS Biol. 9, e1001100 (2011).
Lauring, B., Sakai, H., Kreibich, G. & Wiedmann, M. Nascent polypeptide-associated complex protein prevents mistargeting of nascent chains to the endoplasmic reticulum. Proc. Natl. Acad. Sci. USA 92, 5411–5415 (1995).
Schaffitzel, C. & Ban, N. Generation of ribosome nascent chain complexes for structural and functional studies. J. Struct. Biol. 158, 463–471 (2007).
Heymann, J.B. & Belnap, D.M. Bsoft: image processing and molecular modeling for electron microscopy. J. Struct. Biol. 157, 3–18 (2007).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Valle, M. et al. Incorporation of aminoacyl-tRNA into the ribosome as seen by cryo-electron microscopy. Nat. Struct. Biol. 10, 899–906 (2003).
Valle, M. et al. Locking and unlocking of ribosomal motions. Cell 114, 123–134 (2003).
Fischer, N., Konevega, A.L., Wintermeyer, W., Rodnina, M.V. & Stark, H. Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy. Nature 466, 329–333 (2010).
Rosenthal, P.B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Scheres, S.H., Nunez-Ramirez, R., Sorzano, C.O., Carazo, J.M. & Marabini, R. Image processing for electron microscopy single-particle analysis using XMIPP. Nat. Protoc. 3, 977–990 (2008).
Penczek, P.A., Kimmel, M. & Spahn, C.M. Identifying conformational states of macromolecules by eigen-analysis of resampled cryo-EM images. Structure 19, 1582–1590 (2011).
Pettersen, E.F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Acknowledgements
We thank T. Shaikh for advice with Spider refinement, P. Penczek for assistance with SPARX, M. Bacia for excellent technical assistance and members of the protein expression facility at European Molecular Biology Laboratory Heidelberg as well as the Partnership for Structural Biology in Grenoble for support. The Polara microscope is part of the Structural Biology and Dynamics Groupement d'intérêt scientifique–Infrastrutures en Biologie Sante et Agronomie platform of the Institut de Biologie Structurale. C.S. acknowledges support by the Agence Nationale de la Recherche (ANR-09-JCJC-0044), the region Rhône-Alpes (CIBLE_1976) and the European Research Council Starting grant (project 281331). K.K. was supported by a postdoctoral European Molecular Biology Organization fellowship. We thank I. Saraogi in the Shan laboratory for sharing unpublished results and for critical reading of the manuscript. S.S. is supported by US National Institutes of Health grant R01 GM078024, and the Fellowship for science and engineering from the David and Lucile Packard foundation. A.A. was supported by the US National Institute of General Medical Sciences Ruth L. Kirschstein National Research Service Award (F31GM095294) and the National Institutes of Health National Research Service Award Training grant 5T32GM07616. X.Z. is supported by the Howard Hughes Medical Institute Fellowship of the Helen Hay Whitney Foundation.
Author information
Authors and Affiliations
Contributions
C.S., I.B., X.Z. and S.S. designed experiments; C.S., K.H., O.v.L., A.A. and X.Z. prepared samples; A.A. and X.Z. carried out biochemical experiments; K.K., G.S. and M.K. performed the electron microscopy; O.v.L., M.K. and C.S. performed image analysis and model building; C.S., O.v.L., A.A., X.Z. and S.S. prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Table 1 (PDF 3052 kb)
Rights and permissions
About this article
Cite this article
von Loeffelholz, O., Knoops, K., Ariosa, A. et al. Structural basis of signal sequence surveillance and selection by the SRP–FtsY complex. Nat Struct Mol Biol 20, 604–610 (2013). https://doi.org/10.1038/nsmb.2546
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2546
This article is cited by
-
Protein Transport Across the Bacterial Plasma Membrane by the Sec Pathway
The Protein Journal (2019)