Abstract
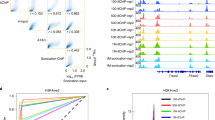
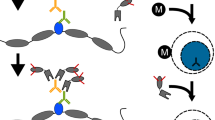
In gene regulation, proteins function as members of protein complexes to recognize chromosomal target DNA loci. In dissecting the pluripotent state in mouse embryonic stem (mES) cells, we have used in vivo biotinylation of critical transcription factors for affinity purification of protein complexes and chromatin immunoprecipitation (ChIP)-on-chip for target identification, respectively. Here, we describe detailed procedures for such studies to dissect protein–protein and protein–DNA interactions in mES cells. Specifically, the following three procedures will be described: (i) in vivo biotinylation system setup in mES cells; (ii) affinity purification of multiprotein complexes by one-step streptavidin capture and tandem anti-FLAG/streptavidin affinity purification; (iii) biotin-mediated ChIP (bioChIP). The system setup takes ∼50 d to complete, and it takes another ∼15 d and ∼3 d to perform affinity purification of protein complexes and bioChIP, respectively.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Shevchenko, A., Schaft, D., Roguev, A., Pijnappel, W.W. & Stewart, A.F. Deciphering protein complexes and protein interaction networks by tandem affinity purification and mass spectrometry: analytical perspective. Mol. Cell Proteomics 1, 204–212 (2002).
de Boer, E. et al. Efficient biotinylation and single-step purification of tagged transcription factors in mammalian cells and transgenic mice. Proc. Natl. Acad. Sci. USA 100, 7480–7485 (2003).
Wang, J. et al. A protein interaction network for pluripotency of embryonic stem cells. Nature 444, 364–368 (2006).
Laniel, M.A., Beliveau, A. & Guerin, S.L. Electrophoretic mobility shift assays for the analysis of DNA-protein interactions. Methods Mol. Biol. 148, 13–30 (2001).
Molloy, P.L. Electrophoretic mobility shift assays. Methods Mol. Biol. 130, 235–246 (2000).
Klug, S.J. & Famulok, M. All you wanted to know about SELEX. Mol. Biol. Rep. 20, 97–107 (1994).
Collas, P. & Dahl, J.A. Chop it, ChIP it, check it: the current status of chromatin immunoprecipitation. Front Biosci. 13, 929–943 (2008).
Turner, F.B., Cheung, W.L. & Cheung, P. Chromatin immunoprecipitation assay for mammalian tissues. Methods Mol. Biol. 325, 261–272 (2006).
Boyer, L.A. et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441, 349–353 (2006).
Valouev, A. et al. Genome-wide analysis of transcription factor binding sites based on ChIP-Seq data. Nat. Methods 5, 829–834 (2008).
Schatz, P.J. Use of peptide libraries to map the substrate specificity of a peptide-modifying enzyme: a 13 residue consensus peptide specifies biotinylation in Escherichia coli. Biotechnology 11, 1138–1143 (1993).
Kim, J., Chu, J., Shen, X., Wang, J. & Orkin, S.H. An extended transcriptional network for pluripotency of embryonic stem cells. Cell 132, 1049–1061 (2008).
Lee, T.I., Johnstone, S.E. & Young, R.A. Chromatin immunoprecipitation and microarray-based analysis of protein location. Nat. Protoc. 1, 729–748 (2006).
Conner, D.A. Mouse embryo fibroblast (MEF) feeder cell preparation. In Current Protocols in Molecular Biology (eds. Frederick, M., Ausubel, et al.) Chapter 23, Unit 23.2 (2001).
Siu, F.K.Y., Lee, L.T.O. & Chow, B.K.C. Southwestern blotting in investigating transcriptional regulation. Nat. Protoc. 3, 51–58 (2008).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J.V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Schmittgen, T.D. & Livak, K.J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108 (2008).
Robertson, G. et al. Genome-wide profiles of STAT1 DNA association using chromatin immunoprecipitation and massively parallel sequencing. Nat. Methods 4, 651–657 (2007).
Acknowledgements
We acknowledge Tyler Moran for his technical assistance in developing these protocols. This work is supported by Seed Grant from the Harvard Stem Cell Institute Cell Reprogramming Program to J.W. J.K. is a Howard Hughes Medical Institute research associate. A.B.C. is supported by NIH Grant R01 HL075705. S.H.O. is an Investigator of Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Kim, J., Cantor, A., Orkin, S. et al. Use of in vivo biotinylation to study protein–protein and protein–DNA interactions in mouse embryonic stem cells. Nat Protoc 4, 506–517 (2009). https://doi.org/10.1038/nprot.2009.23
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2009.23
This article is cited by
-
The mismatch-repair proteins MSH2 and MSH6 interact with the imprinting control regions through the ZFP57-KAP1 complex
Epigenetics & Chromatin (2022)
-
A MYC-ZNF148-ID1/3 regulatory axis modulating cancer stem cell traits in aggressive breast cancer
Oncogenesis (2022)
-
Inner nuclear protein Matrin-3 coordinates cell differentiation by stabilizing chromatin architecture
Nature Communications (2021)
-
Motif-driven interactions between RNA and PRC2 are rheostats that regulate transcription elongation
Nature Structural & Molecular Biology (2021)
-
OCT4 cooperates with distinct ATP-dependent chromatin remodelers in naïve and primed pluripotent states in human
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.