Abstract
X-ray crystallography at X-ray free-electron laser sources is a powerful method for studying macromolecules at biologically relevant temperatures. Moreover, when combined with complementary techniques like X-ray emission spectroscopy, both global structures and chemical properties of metalloenzymes can be obtained concurrently, providing insights into the interplay between the protein structure and dynamics and the chemistry at an active site. The implementation of such a multimodal approach can be compromised by conflicting requirements to optimize each individual method. In particular, the method used for sample delivery greatly affects the data quality. We present here a robust way of delivering controlled sample amounts on demand using acoustic droplet ejection coupled with a conveyor belt drive that is optimized for crystallography and spectroscopy measurements of photochemical and chemical reactions over a wide range of time scales. Studies with photosystem II, the phytochrome photoreceptor, and ribonucleotide reductase R2 illustrate the power and versatility of this method.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Emma, P. et al. First lasing and operation of an Ångstrom-wavelength free-electron laser. Nat. Photonics 4, 641–647 (2010).
Chapman, H.N. et al. Femtosecond X-ray protein nanocrystallography. Nature 470, 73–77 (2011).
Boutet, S. et al. High-resolution protein structure determination by serial femtosecond crystallography. Science 337, 362–364 (2012).
Kern, J. et al. Simultaneous femtosecond X-ray spectroscopy and diffraction of photosystem II at room temperature. Science 340, 491–495 (2013).
Benkovic, S.J. & Hammes-Schiffer, S. A perspective on enzyme catalysis. Science 301, 1196–1202 (2003).
Levantino, M., Yorke, B.A., Monteiro, D.C., Cammarata, M. & Pearson, A.R. Using synchrotrons and XFELs for time-resolved X-ray crystallography and solution scattering experiments on biomolecules. Curr. Opin. Struct. Biol. 35, 41–48 (2015).
DePonte, D.P. et al. Gas dynamic virtual nozzle for generation of microscopic droplet streams. J. Phys. D Appl. Phys. 41, 195505 (2008).
Sierra, R.G. et al. Nanoflow electrospinning serial femtosecond crystallography. Acta Crystallogr. D Biol. Crystallogr. 68, 1584–1587 (2012).
Weierstall, U. et al. Lipidic cubic-phase injector facilitates membrane protein serial femtosecond crystallography. Nat. Commun. 5, 3309 (2014).
Sugahara, M. et al. Grease matrix as a versatile carrier of proteins for serial crystallography. Nat. Methods 12, 61–63 (2015).
Hunter, M.S. et al. Fixed-target protein serial microcrystallography with an X-ray free-electron laser. Sci. Rep. 4, 6026 (2014).
Baxter, E.L. et al. High-density grids for efficient data collection from multiple crystals. Acta Crystallogr. D Struct. Biol. 72, 2–11 (2016).
Roedig, P. et al. Room-temperature macromolecular crystallography using a micropatterned silicon chip with minimal background scattering. J. Appl. Crystallogr. 49, 968–975 (2016).
Oghbaey, S. et al. Fixed target combined with spectral mapping: approaching 100% hit rates for serial crystallography. Acta Crystallogr. D Struct. Biol. 72, 944–955 (2016).
Roessler, C.G. et al. Acoustic injectors for drop-on-demand serial femtosecond crystallography. Structure 24, 631–640 (2016).
Roessler, C.G. et al. Acoustic methods for high-throughput protein crystal mounting at next-generation macromolecular crystallographic beamlines. J. Synchrotron Radiat. 20, 805–808 (2013).
Griese, J.J., Srinivas, V. & Högbom, M. Assembly of nonheme Mn–Fe active sites in heterodinuclear metalloproteins. J. Biol. Inorg. Chem. 19, 759–774 (2014).
Jiang, W., Hoffart, L.M., Krebs, C. & Bollinger, J.M. Jr. A manganese(IV) –iron(IV) intermediate in assembly of the manganese(IV) –iron(III) cofactor of Chlamydia trachomatis ribonucleotide reductase. Biochemistry 46, 8709–8716 (2007).
Jiang, W. et al. A manganese(IV) –iron(III) cofactor in Chlamydia trachomatis ribonucleotide reductase. Science 316, 1188–1191 (2007).
Kern, J. et al. Taking snapshots of photosynthetic water oxidation using femtosecond X-ray diffraction and spectroscopy. Nat. Commun. 5, 4371 (2014).
Young, I.D. et al. Structure of photosystem II and substrate binding at room temperature. Nature 540, 453–457 (2016).
Kok, B., Forbush, B. & McGloin, M. Cooperation of charges in photosynthetic O2 evolution—I. a linear four-step mechanism. Photochem. Photobiol. 11, 457–475 (1970).
Renger, G. Primary Processes of Photosynthesis: Principles and Apparatus (Royal Society of Chemistry, Cambridge, UK, 2008).
de Wijn, R. & van Gorkom, H.J. Kinetics of electron transfer from QA to QB in photosystem II. Biochemistry 40, 11912–11922 (2001).
Alonso-Mori, R. et al. Toward characterization of photo-excited electron transfer and catalysis in natural and artificial systems using XFELs. Faraday Discuss. 194, 621–638 (2016).
Messinger, J. et al. Absence of Mn-centered oxidation in the S2 → S3 transition: implications for the mechanism of photosynthetic water oxidation. J. Am. Chem. Soc. 123, 7804–7820 (2001).
Glatzel, P. & Bergmann, U. High-resolution 1s core hole X-ray spectroscopy in 3d transition metal complexes—electronic and structural information. Coord. Chem. Rev. 249, 65–95 (2005).
Yano, J. & Yachandra, V. Mn4Ca cluster in photosynthesis: where and how water is oxidized to dioxygen. Chem. Rev. 114, 4175–4205 (2014).
Alonso-Mori, R. et al. A multicrystal wavelength-dispersive X-ray spectrometer. Rev. Sci. Instrum. 83, 073114 (2012).
Alonso-Mori, R. et al. Energy-dispersive X-ray emission spectroscopy using an X-ray free-electron laser in a shot-by-shot mode. Proc. Natl. Acad. Sci. USA 109, 19103–19107 (2012).
Alonso-Mori, R. et al. Photon-in photon-out hard X-ray spectroscopy at the Linac Coherent Light Source. J. Synchrotron Radiat. 22, 612–620 (2015).
Vankó, G. et al. Probing the 3d spin momentum with X-ray emission spectroscopy: the case of molecular-spin transitions. J. Phys. Chem. B 110, 11647–11653 (2006).
Svyazhin, A., Kurmaev, E., Shreder, E., Shamin, S. & Sahle, C.J. Local moments and electronic correlations in Fe-based Heusler alloys: Kα X-ray emission spectra measurements. J. Alloys Compd. 679, 268–276 (2016).
Burgie, E.S. et al. Crystallographic and electron microscopic analyses of a bacterial phytochrome reveal local and global rearrangements during photoconversion. J. Biol. Chem. 289, 24573–24587 (2014).
Edlund, P. et al. The room temperature crystal structure of a bacterial phytochrome determined by serial femtosecond crystallography. Sci. Rep. 6, 35279 (2016).
Bhattacharya, S., Auldridge, M.E., Lehtivuori, H., Ihalainen, J.A. & Forest, K.T. Origins of fluorescence in evolved bacteriophytochromes. J. Biol. Chem. 289, 32144–32152 (2014).
Zhang, C.F., Farrens, D.L., Bjorling, S.C., Song, P.S. & Kliger, D.S. Time-resolved absorption studies of native etiolated oat phytochrome. J. Am. Chem. Soc. 114, 4569–4580 (1992).
Mroginski, M.A., Murgida, D.H. & Hildebrandt, P. The chromophore structural changes during the photocycle of phytochrome: a combined resonance Raman and quantum chemical approach. Acc. Chem. Res. 40, 258–266 (2007).
Braslavsky, S.E., Gärtner, W. & Schaffner, K. Phytochrome photoconversion. Plant Cell Environ. 20, 700–706 (1997).
Li, F. et al. X-ray radiation induces deprotonation of the bilin chromophore in crystalline D. radiodurans phytochrome. J. Am. Chem. Soc. 137, 2792–2795 (2015).
Lundin, D., Berggren, G., Logan, D.T. & Sjöberg, B.M. The origin and evolution of ribonucleotide reduction. Life (Basel) 5, 604–636 (2015).
Hellmich, J. et al. Native-like photosystem II superstructure at 2.44 Å resolution through detergent extraction from the protein crystal. Structure 22, 1607–1615 (2014).
Ibrahim, M. et al. Improvements in serial femtosecond crystallography of photosystem II by optimizing crystal uniformity using microseeding procedures. Struct. Dyn. 2, 041705 (2015).
Kern, J. et al. Purification, characterization and crystallization of photosystem II from Thermosynechococcus elongatus cultivated in a new type of photobioreactor. Biochim. Biophys. Acta 1706, 147–157 (2005).
Wagner, J.R., Zhang, J., Brunzelle, J.S., Vierstra, R.D. & Forest, K.T. High-resolution structure of Deinococcus bacteriophytochrome yields new insights into phytochrome architecture and evolution. J. Biol. Chem. 282, 12298–12309 (2007).
Kutin, Y. et al. Divergent assembly mechanisms of the manganese–iron cofactors in R2lox and R2c proteins. J. Inorg. Biochem. 162, 164–177 (2016).
Dassama, L.M.K. et al. O2-evolving chlorite dismutase as a tool for studying O2-utilizing enzymes. Biochemistry 51, 1607–1616 (2012).
Fuller, F.D. et al. Droplet on tape: protocol. Nat. Protoc. http://dx.doi.org/10.1038/protex.2017.017 (2017).
Chollet, M. et al. The X-ray pump–probe instrument at the Linac coherent light source. J. Synchrotron Radiat. 22, 503–507 (2015).
Herrmann, S. et al. CSPAD-140k: a versatile detector for LCLS experiments. Nucl. Instrum. Methods Phys. Res. A 718, 550–553 (2013).
Carini, G. et al. in 2014 IEEE Nuclear Science Symposium and Medical Imaging Conference (NSS/MIC) 1–3 (IEEE, Seattle, WA, USA; 2014).
Sauter, N.K., Hattne, J., Grosse-Kunstleve, R.W. & Echols, N. New Python-based methods for data processing. Acta Crystallogr. D Biol. Crystallogr. 69, 1274–1282 (2013).
Hattne, J. et al. Accurate macromolecular structures using minimal measurements from X-ray free-electron lasers. Nat. Methods 11, 545–548 (2014).
Sauter, N.K., Grosse-Kunstleve, R.W. & Adams, P.D. Robust indexing for automatic data collection. J. Appl. Crystallogr. 37, 399–409 (2004).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Högbom, M. et al. The radical site in chlamydial ribonucleotide reductase defines a new R2 subclass. Science 305, 245–248 (2004).
Acknowledgements
We thank T. Rendahl from LCLS for help with controls and B. Martins, S. Myers, M. Cowan, G. Shea-McCarthy and C. Whalen (Brookhaven National Laboratory) for help with engineering and controls, R. Ellson, J. Olechno, R. Stearns and B. Hadimioglu (LABCYTE) for sharing their experience with the acoustic transducers and related issues, C. Saracini (LBNL) for his help with the preliminary testing of the DOT system, P. Glatzel (ESRF) for discussion of the Fe Kα emission and F. Houle (LBNL) for sharing her reaction–diffusion simulation software Kinetiscope and for useful discussions. This work was supported by the Director, Office of Science, Office of Basic Energy Sciences (OBES), Division of Chemical Sciences, Geosciences and Biosciences (CSGB) of the US Department of Energy (DOE) under contract DE-AC02-05CH11231 (J.Y. and V.K.Y.) for X-ray methodology and instrumentation, by the US National Institutes of Health (NIH) grants GM110501 (J.Y.) for instrumentation development for XFEL experiments, GM102520 (N.K.S.) and GM117126 (N.K.S.) for development of computational protocols for XFEL data and GM055302 (V.K.Y.) for PS II biochemistry, structure and mechanism, a Ruth L. Kirschstein National Research Service Award (5 F32 GM116423-02; F.D.F.), the Human Frontiers Science Project award no. RGP0063/2013 310 (J.Y., U.B. and A.Z.), and the Science and Technology Center program of the US National Science Foundation (NSF) through BioXFEL under agreement no. 1231306 (J.A.C., M.D.M. and G.N.P.). R.D.V. is supported by NSF grant MCB-1329956. J.A.C. was supported by a training fellowship from the Gulf Coast Consortia on the Houston Area Molecular Biophysics Program (NIHGMS grant no. T32GM008280). C.J.P. was supported by NIH NRSA grant GM113389-01. Portions of this work were supported by Brookhaven National Laboratory (BNL)-US DOE, Laboratory Directed Research and Development grant 11-008 (C.G.R., M. Allaire., A.M.O.), NIH–NCRR grant 2-P41-RR012408 (A.M.O. and M. Allaire), NIH–National Institute of General Medical Sciences (NIGMS) grants 8P41GM103473-16 (A.M.O.) and Y1GM008003 (M. Allaire) and US DOE, Office of Biological and Environmental Research (OBER) grant FWP BO-70 (A.M.O. and B.A.). A.M.O., P.A. and P.T.D. were supported in part by Diamond Light Source, and A.M.O. acknowledges support from a Strategic Award from the Wellcome Trust and the Biotechnology and Biological Sciences Research Council (grant 102593). P.B. was supported by a Wellcome Trust DPhil studentship. M.H. received support from the Knut and Alice Wallenberg Foundation, the Swedish Cancer Society, the Wenner–Gren foundations and the Swedish Research Council (grants 2013-541 and 2013-5884). C.A.S. acknowledges support from the US DOE, Office of Science, Division of CSGB. A.Z. acknowledges support from the DFG-Cluster of Excellence 'UniCat', coordinated by the Technische Universität Berlin, and Sfb1078 (Humboldt Universität Berlin), TP A5. J.M. acknowledges support from the Solar Fuels Strong Research Environment (Umeå University), the Artificial Leaf Project (K&A Wallenberg Foundation 2011.0055) and Energimyndigheten (36648-1). This research work used resources from the National Energy Research Scientific Computing Center, a DOE Office of Science User Facility supported by the Office of Science, DOE, under contract no. DE-AC02-05CH11231. Testing of crystals and various parts of the setup were carried out at synchrotron facilities that were provided by the Advanced Light Source (ALS) in Berkeley and the Stanford Synchrotron Radiation Light Source (SSRL) in Stanford, which were funded by the DOE OBES. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research, and by the NIH (grant P41GM103393). Use of the Linac Coherent Light Source (LCLS) and SSRL, SLAC National Accelerator Laboratory, is supported by the US DOE, Office of Science, OBES under contract no. DE-AC02-76SF00515. BNL's contribution to data collection at the LCLS is supported by the Life Science and Biomedical Technology Research (LSBR) program at the National Synchrotron Light Source II (NSLS-II), which operates under a DOE BER contract (DE-SC0012704) and DOE BES contract (DE-AC02-98CH10886), and is supported by NIH–NIGMS grant P41GM111244.
Author information
Authors and Affiliations
Contributions
A.M.O., J.K., V.K.Y. and J.Y. conceived the experiment; F.D.F., S.G., J.K., V.K.Y. and J.Y. designed the experiment; E.S.B., R.C., C.J.P., J.A.C., M.I., R.H., A.Z., H. Lebrette, V.S., M.Z., S.K., J.M., A.K.B., J.M.B., C.K., M.H., G.N.P. and R.D.V. prepared, characterized and provided the phytochrome, PS II and RNR samples; F.D.F., J.K., S.G., C.G.R. and A.M.O. designed the acoustic injectors; F.D.F., S.G., B.A., E.P., C.d.L., C.A.S., C.G.R., R.G.S., T.K., M.K., S.K., P.T.D., U.B., G.N.P., J.K., V.K.Y., A.M.O. and J.Y. performed the SFX and XES experiments; J.M.G., C.A.S., S.N., J.E.K., D.Z., M.C., S.S., H. Lemke, D.S., M.L. and R.A.-M. set up the beam lines; A.S.B., I.D.Y., T.M.-C., P.A., P.B., L.L., M.D.M., T.K., M. Amin, M. Allaire, F.D.F., J.K., E.S.B., T.F., C.W. and N.K.S. performed XRD and XES data analysis; F.D.F., J.K., E.S.B., A.M.O., V.K.Y. and J.Y. wrote the paper with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
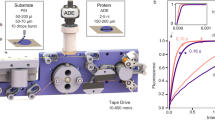
Supplementary Figure 1 Photograph of the DOT system and details of the optical pump setup.
a. A photograph of the drop-on-tape (DOT) device mounted at the X-ray Pump Probe (XPP) endstation of the Linac Coherent Light Source (LCLS) with various important components labeled. b. Schematic of the fiber optics setup for sample illumination. c. Example output of the feedback system to ensure that drop deposition is in phase with laser pump pulses. The arrival time of each drop over one of the IR “gates” is detected by changes in the IR transmission (yellow and pink traces). In addition, the laser transmission of the sample at each of the three illumination points on the tape for each individual drop is measured (blue and green traces shown here are for the pump #2 and #3) and delays are tuned to bring the IR and pump signals in phase. This information can also be used after the experiment to reject signal from drops that did not receive the correct illumination.
Supplementary Figure 2 Treatment of background originating from the Kapton belt.
a. Diffraction image showing the belt background on the XRD CCD detector with the maximum absorption of Kapton highlighted in red and the minimum in blue. b. A simplified geometry of the conveyor belt and the shadow it casts on the CCD. c. Illustrations of the parameters used in the Kapton absorption correction from left to right: on beam axis view, zoomed in beam axis view, and top view.
Supplementary Figure 3 Crystalline samples used in this study.
a. Crystal images of Phytochrome PAS-GAF region (~ 50 μm), b. Phytochrome PSM (~ 100 μm), c. RNR (20 - 30 μm), and d. PS II (20 - 50 μm).
Supplementary Figure 4 Statistics for three example diffraction experiments.
Shown is the indexing rate over time for suspensions of PAS-GAF (blue), PSM (red), PS II (turquoise) and RNR (black) crystals in the DOT setup. Data were collected at 10 Hz and the % of X-ray laser shots that yielded an indexable diffraction pattern are given as a function of total sample run time. Fluctuations in the indexing rate are largely a function of crystal density in the syringe, and also arise from the adjustment of the beam height in terms of the belt.
Supplementary Figure 5 Bilin binding modes in DrBphP.
Four representative DrBphP structures were superposed. The positions of Asp207 and Tyr263 of the PSM (green, PDB code 4Q0J) and our room temperature, SFX PAS-GAF structure (yellow, PDB code 5MG0) were highly congruent, and indicate hydrogen bonding between the two residues. The positions of these residues differed from those found in a separate SFX structure collected at LCLS (magenta, PDB code 5L8M)1, and the photochemically compromised Asp207-Ala mutant (cyan, PDB code 4Q0I), which both show an outward splay of the Tyr263 side chain from residue 207, and absence of hydrogen bonding (e.g., a 3.9 Å inter-residue distance for 5L8M). The D-ring of our PAS-GAF structure appears to be in a position between that found for the PSM and the Asp207-Ala mutant. Pyrrole rings A and D are indicated.
1. Edlund, P. et al. The room temperature crystal structure of a bacterial phytochrome determined by serial femtosecond crystallography. Sci. Rep. 6, 35279 (2016).
Supplementary Figure 6 Omit Fo-Fc electron density maps for the PAS-GAF and PSM constructs of DrBphP.
The two orthogonal views of the PAS-GAF maps (a and b) are calculated to 2.0 Å resolution and contoured at +3 σ (green) and -3 σ (red). Two orthogonal views of the PSM maps (c and d) calculated to 3.2 Å resolution and contoured at +3 σ (green) and -3 σ (red).
Supplementary Figure 7 XES data processing.
a. Raw image (minus outlier pixels), with wide ROI shown in red. b. Smooth polynomial background fit to data outside the wide ROI and extrapolated into the ROI (intensified by 4x for illustrative purposes). c. Integration of the wide ROI for both the raw and background. d. Corrected image with tight ROI based on Gaussian fit. e. Final corrected spectrum.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–2 (PDF 1574 kb)
Supplementary Protocol
Supplementary Protocol (PDF 3362 kb)
Rights and permissions
About this article
Cite this article
Fuller, F., Gul, S., Chatterjee, R. et al. Drop-on-demand sample delivery for studying biocatalysts in action at X-ray free-electron lasers. Nat Methods 14, 443–449 (2017). https://doi.org/10.1038/nmeth.4195
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4195
This article is cited by
-
Stimulated X-ray emission spectroscopy
Photosynthesis Research (2024)
-
In situ X-ray spectroscopies beyond conventional X-ray absorption spectroscopy on deciphering dynamic configuration of electrocatalysts
Nature Communications (2023)
-
Growing and making nano- and microcrystals
Nature Protocols (2023)
-
Structural evidence for intermediates during O2 formation in photosystem II
Nature (2023)
-
Serial femtosecond crystallography
Nature Reviews Methods Primers (2022)