Abstract
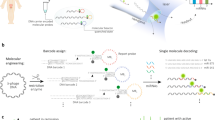
The ability to count biomolecules such as cancer-biomarker miRNAs with the naked eye is seemingly impossible in molecular diagnostics. Here, we show an ultrasensitive naked-eye-counting strategy for quantifying miRNAs by employing T7 phage—a bacteria-specific virus nanoparticle—as a surrogate. The phage is genetically engineered to become fluorescent and capable of binding a miRNA-capturing gold nanoparticle (GNP) in a one-to-one manner. Target miRNAs crosslink the resultant phage–GNP couple and miRNA-capturing magnetic microparticles, forming a sandwich complex containing equimolar phage and miRNA. The phage is then released from the complex and developed into one macroscopic fluorescent plaque in a Petri dish by plating it in a host bacterial medium. Counting the plaques by the naked eye enables the quantification of miRNAs with detection limits of ∼3 and ∼5 aM for single-target and two-target miRNAs, respectively. This approach offers ultrasensitive and convenient quantification of disease biomarkers by the naked eye.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hamburg, M. A. & Collins, F. S. The path to personalized medicine. New Engl. J. Med. 363, 301–304 (2010).
Taton, T. A., Mirkin, C. A. & Letsinger, R. L. Scanometric DNA array detection with nanoparticle probes. Science 289, 1757–1760 (2000).
Valentini, P. et al. Gold-nanoparticle-based colorimetric discrimination of cancer-related point mutations with picomolar sensitivity. ACS Nano 7, 5530–5538 (2013).
Cao, Y. C., Jin, R. & Mirkin, C. A. Nanoparticles with Raman spectroscopic fingerprints for DNA and RNA detection. Science 297, 1536–1540 (2002).
Qiu, F., Jiang, D., Ding, Y., Zhu, J. & Huang, L. L. Monolayer-barcoded nanoparticles for on-chip DNA hybridization assay. Angew. Chem. Int. Ed. 47, 5009–5012 (2008).
Zhou, X., Cao, P., Tian, Y. & Zhu, J. Expressed peptide assay for DNA detection. J. Am. Chem. Soc. 132, 4161–4168 (2010).
Nam, J.-M., Stoeva, S. I. & Mirkin, C. A. Bio-bar-code-based DNA detection with PCR-like sensitivity. J. Am. Chem. Soc. 126, 5932–5933 (2004).
Stoeva, S. I., Lee, J.-S., Smith, J. E., Rosen, S. T. & Mirkin, C. A. Multiplexed detection of protein cancer markers with biobarcoded nanoparticle probes. J. Am. Chem. Soc. 128, 8378–8379 (2006).
Mao, C. et al. Viral assembly of oriented quantum dot nanowires. Proc. Natl Acad. Sci. USA 100, 6946–6951 (2003).
Mao, C. et al. Virus-based toolkit for the directed synthesis of magnetic and semiconducting nanowires. Science 303, 213–217 (2004).
Liu, A., Abbineni, G. & Mao, C. Nanocomposite films assembled from genetically engineered filamentous viruses and gold nanoparticles: Nanoarchitecture- and humidity-tunable surface plasmon resonance spectra. Adv. Mater. 21, 1001–1005 (2009).
Mao, C., Wang, F. & Cao, B. Controlling nanostructures of mesoporous silica fibers by supramolecular assembly of genetically modifiable bacteriophages. Angew. Chem. Int. Ed. 51, 6411–6415 (2012).
Wang, F., Nimmo, S., Cao, B. & Mao, C. B. Oxide formation on biological nanostructures via a structure-directing agent: Towards an understanding of precise transcription. Chem. Sci. 3, 2639–2645 (2012).
Eber, F. J., Eiben, S., Jeske, H. & Wege, C. Bottom-up-assembled nanostar colloids of gold cores and tubes derived from tobacco mosaic virus. Angew. Chem. Int. Ed. 52, 7203–7207 (2013).
Cao, B., Zhu, Y., Wang, L. & Mao, C. B. Controlled alignment of filamentous supramolecular assemblies of biomolecules into centimeter-scale highly ordered patterns by using nature-inspired magnetic guidance. Angew. Chem. Int. Ed. 52, 11750–11754 (2013).
Mao, C., Liu, A. & Cao, B. Virus-based chemical and biological sensing. Angew. Chem. Int. Ed. 48, 6790–6810 (2009).
Mohan, K., Donavan, K. C., Arter, J. A., Penner, R. M. & Weiss, G. A. Sub-nanomolar detection of prostate-specific membrane antigen in synthetic urine by synergistic, dual-ligand phage. J. Am. Chem. Soc. 135, 7761–7767 (2013).
Cuervo, A. et al. Structural characterization of the bacteriophage T7 tail machinery. J. Biol. Chem. 288, 26290–26299 (2013).
Chan, L. Y., Kosuri, S. & Endy, D. Refactoring bacteriophage T7. Mol. Syst. Biol. 1, 2005.0018 (2005).
Slootweg, E. J. et al. Fluorescent T7 display phages obtained by translational frameshift. Nucleic Acids Res. 34, e137 (2006).
Tsuboyama, M. & Maeda, I. Combinatorial parallel display of polypeptides using bacteriophage T7 for development of fluorescent nano-bioprobes. J. Biosci. Bioeng. 116, 28–33 (2013).
Huang, Y. et al. Programmable assembly of nanoarchitectures using genetically engineered viruses. Nano Lett. 5, 1429–1434 (2005).
Garcia-Doval, C. & Raaij, M. J. V. Structure of the receptor-binding carboxy-terminal domain of bacteriophage T7 tail fibers. Proc. Natl Acad. Sci. USA 109, 9390–9395 (2012).
Dunn, J. J. & Studier, F. W. Complete nucleotide sequence of bacteriophage T7 DNA and the locations of T7 genetic elements. J. Mol. Biol. 166, 477–535 (1983).
Johnson, S. M. et al. RAS is regulated by the let-7 microRNA family. Cell 120, 635–647 (2005).
Esquela-Kerscher, A. & Slack, F. J. Oncomirs—microRNAs with a role in cancer. Nature Rev. Cancer 6, 259–269 (2006).
Lin, P. Y., Yu, S. L. & Yang, P. C. MicroRNA in lung cancer. Br. J Cancer 103, 1144–1148 (2010).
Bousquet, M., Harris, M. H., Zhou, B. & Lodish, H. F. MicroRNA miR-125b causes leukemia. Proc. Natl Acad. Sci. USA 107, 21558–21563 (2010).
Monya, B. qPCR: Quicker and easier but don’t be sloppy. Nature Methods 8, 207–212 (2011).
Kumar, M. S. et al. Suppression of non-small cell lung tumor development by the let-7 microRNA family. Proc. Natl Acad. Sci. USA 105, 3903–3908 (2008).
Serwer, P. & Pichler, M. E. Electrophoresis of bacteriophage T7 and T7 capsids in agarose gels. J. Virol. 28, 917–928 (1978).
Xu, T. et al. MicroRNA-195 suppresses tumorigenicity and regulates G1/S transition of human hepatocellular carcinoma cells. Hepatology 50, 113–121 (2009).
Jusufović, E. et al. let-7b and miR-126 are down-regulated in tumor tissue and correlate with microvessel density and survival outcomes in non-small-cell lung cancer. PLoS ONE 7, e45577 (2012).
Liu, Z. L., Wang, H., Liu, J. & Wang, Z. X. MicroRNA-21 (miR-21) expression promotes growth, metastasis, and chemo- or radioresistance in non-small cell lung cancer cells by targeting PTEN. Mol. Cell Biochem. 372, 35–45 (2013).
Thellin, O. et al. Housekeeping genes as internal standards: Use and limits. J. Biotechnol. 75, 291–295 (1999).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, research0034 (2002).
Dheda, K. et al. Validation of housekeeping genes for normalizing RNA expression in real-time PCR. BioTechniques 37, 112–119 (2004).
Acknowledgements
This work was supported by the National Key Basic Research Program of China (2011CB933503), the Special Funds of the National Natural Science Foundation of China for Basic Research Projects of Scientific Instruments (61127002), the Basic Research Program of Jiangsu Province (BK2011036) and the Jiangsu Province Funds for Distinguished Young Scientists (BK20140049). Y.Z. and C.M. are grateful for financial support from the National Institutes of Health (EB015190 and CA200504), the National Science Foundation (CMMI-1234957), the Department of Defense Congressionally Directed Medical Research Programs, the Oklahoma Center for Adult Stem Cell Research (434003) and the Oklahoma Center for the Advancement of Science and Technology (HR14-160).
Author information
Authors and Affiliations
Contributions
X.Z. and C.M conceived the experiments. X.Z., P.C., C.M. and Y.Z. performed the experiments. W.L. assisted with the experiments, X.Z., Y.Z. and C.M. wrote the manuscript and analysed the data. N.G. and C.M. designed and supervised the project. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 4632 kb)
Rights and permissions
About this article
Cite this article
Zhou, X., Cao, P., Zhu, Y. et al. Phage-mediated counting by the naked eye of miRNA molecules at attomolar concentrations in a Petri dish. Nature Mater 14, 1058–1064 (2015). https://doi.org/10.1038/nmat4377
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat4377
This article is cited by
-
Advanced graphene oxide-based paper sensor for colorimetric detection of miRNA
Microchimica Acta (2022)
-
Intermolecular distance measurement with TNT suppressor on the M13 bacteriophage-based Förster resonance energy transfer system
Scientific Reports (2019)
-
DNA origami-based shape IDs for single-molecule nanomechanical genotyping
Nature Communications (2017)
-
Naked-eye sensitive ELISA-like assay based on gold-enhanced peroxidase-like immunogold activity
Analytical and Bioanalytical Chemistry (2016)