Abstract
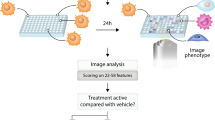
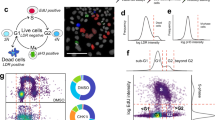
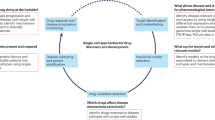
Drug screening is often limited to cell-free assays involving purified enzymes, but it is arguably best applied against systems that represent disease states or complex physiological cellular networks. Here, we describe a high-content, cell-based drug discovery platform based on phosphospecific flow cytometry, or phosphoflow, that enabled screening for inhibitors against multiple endogenous kinase signaling pathways in heterogeneous primary cell populations at the single-cell level. From a library of small-molecule natural products, we identified pathway-selective inhibitors of Jak-Stat and MAP kinase signaling. Dose-response experiments in primary cells confirmed pathway selectivity, but importantly also revealed differential inhibition of cell types and new druggability trends across multiple compounds. Lead compound selectivity was confirmed in vivo in mice. Phosphoflow therefore provides a unique platform that can be applied throughout the drug discovery process, from early compound screening to in vivo testing and clinical monitoring of drug efficacy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Nicholson, R.L., Welch, M., Ladlow, M. & Spring, D.R. Small-molecule screening: advances in microarraying and cell-imaging technologies. ACS Chem. Biol. 2, 24–30 (2007).
Valet, G. Cytomics as a new potential for drug discovery. Drug Discov. Today 11, 785–791 (2006).
Lang, P., Yeow, K., Nichols, A. & Scheer, A. Cellular imaging in drug discovery. Nat. Rev. Drug Discov. 5, 343–356 (2006).
Nolan, G.P. What's wrong with drug screening today. Nat. Chem. Biol. 3, 187–191 (2007).
Chow, S., Patel, H. & Hedley, D.W. Measurement of MAP kinase activation by flow cytometry using phospho-specific antibodies to MEK and ERK: potential for pharmacodynamic monitoring of signal transduction inhibitors. Cytometry 46, 72–78 (2001).
Krutzik, P.O. & Nolan, G.P. Intracellular phospho-protein staining techniques for flow cytometry: monitoring single cell signaling events. Cytometry A 55, 61–70 (2003).
Krutzik, P.O., Clutter, M.R. & Nolan, G.P. Coordinate analysis of murine immune cell surface markers and intracellular phosphoproteins by flow cytometry. J. Immunol. 175, 2357–2365 (2005).
Krutzik, P.O., Hale, M.B. & Nolan, G.P. Characterization of the murine immunological signaling network with phosphospecific flow cytometry. J. Immunol. 175, 2366–2373 (2005).
Krutzik, P.O., Irish, J.M., Nolan, G.P. & Perez, O.D. Analysis of protein phosphorylation and cellular signaling events by flow cytometry: techniques and clinical applications. Clin. Immunol. 110, 206–221 (2004).
Irish, J.M. et al. Single cell profiling of potentiated phospho-protein networks in cancer cells. Cell 118, 217–228 (2004).
Irish, J.M., Kotecha, N. & Nolan, G.P. Mapping normal and cancer cell signalling networks: towards single-cell proteomics. Nat. Rev. Cancer 6, 146–155 (2006).
Jacobberger, J.W. et al. Immunoreactivity of Stat5 phosphorylated on tyrosine as a cell-based measure of Bcr/Abl kinase activity. Cytometry A 54, 75–88 (2003).
Irish, J.M., Czerwinski, D.K., Nolan, G.P. & Levy, R. Kinetics of B cell receptor signaling in human B cell subsets mapped by phosphospecific flow cytometry. J. Immunol. 177, 1581–1589 (2006).
Zell, T. et al. Single-cell analysis of signal transduction in CD4 T cells stimulated by antigen in vivo. Proc. Natl. Acad. Sci. USA 98, 10805–10810 (2001).
Ilangumaran, S., Finan, D. & Rottapel, R. Flow cytometric analysis of cytokine receptor signal transduction. J. Immunol. Methods 278, 221–234 (2003).
Kersh, E.N. et al. TCR signal transduction in antigen-specific memory CD8 T cells. J. Immunol. 170, 5455–5463 (2003).
Krutzik, P.O. & Nolan, G.P. Fluorescent cell barcoding in flow cytometry allows high-throughput drug screening and signaling profiling. Nat. Methods 3, 361–368 (2006).
Sachs, K., Perez, O., Pe'er, D., Lauffenburger, D.A. & Nolan, G.P. Causal protein-signaling networks derived from multiparameter single-cell data. Science 308, 523–529 (2005).
Edwards, B.S., Oprea, T., Prossnitz, E.R. & Sklar, L.A. Flow cytometry for high-throughput, high-content screening. Curr. Opin. Chem. Biol. 8, 392–398 (2004).
Young, S.M. et al. High-throughput screening with HyperCyt flow cytometry to detect small molecule formylpeptide receptor ligands. J. Biomol. Screen. 10, 374–382 (2005).
Jacobberger, J.W. Increasing the power of cytometry. Nat. Methods 3, 343–344 (2006).
Van Meter, M.E. et al. K-RasG12D expression induces hyperproliferation and aberrant signaling in primary hematopoietic stem/progenitor cells. Blood 109, 3945–3952 (2007).
O'Shea, J.J., Pesu, M., Borie, D.C. & Changelian, P.S. A new modality for immunosuppression: targeting the JAK/STAT pathway. Nat. Rev. Drug Discov. 3, 555–564 (2004).
Luo, C. & Laaja, P. Inhibitors of JAKs/STATs and the kinases: a possible new cluster of drugs. Drug Discov. Today 9, 268–275 (2004).
Zhang, J.H., Chung, T.D. & Oldenburg, K.R. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 4, 67–73 (1999).
Quintas-Cardama, A. et al. Phase I/II study of subcutaneous homoharringtonine in patients with chronic myeloid leukemia who have failed prior therapy. Cancer 109, 248–255 (2007).
Chen, R., Gandhi, V. & Plunkett, W. A sequential blockade strategy for the design of combination therapies to overcome oncogene addiction in chronic myelogenous leukemia. Cancer Res. 66, 10959–10966 (2006).
Dessureault, J. & Krause, M.O. The effect of a new antibiotic, cryptosporiopsin, on RNA synthesis in L-cells. Can. J. Biochem. 53, 149–154 (1975).
Burres, N.S., Hunter, J.E. & Wright, A.E. A mammalian cell agar-diffusion assay for the detection of toxic compounds. J. Nat. Prod. 52, 522–527 (1989).
Marumo, H., Kitaura, K., Morimoto, M., Tanaka, H. & Omura, S. The mode of action of nanaomycin A in Gram-positive bacteria. J. Antibiot. (Tokyo) 33, 885–890 (1980).
Suzuki, H. et al. Immunosuppressive activity of streptonigrin in vitro and in vivo. Biosci. Biotechnol. Biochem. 60, 789–793 (1996).
Thompson, J.E. et al. Photochemical preparation of a pyridone containing tetracycle: a Jak protein kinase inhibitor. Bioorg. Med. Chem. Lett. 12, 1219–1223 (2002).
Narla, R.K., Liu, X.P., Myers, D.E. & Uckun, F.M. 4-(3′-Bromo-4′hydroxylphenyl)-amino-6,7-dimethoxyquinazoline: a novel quinazoline derivative with potent cytotoxic activity against human glioblastoma cells. Clin. Cancer Res. 4, 1405–1414 (1998).
Nam, S. et al. Indirubin derivatives inhibit Stat3 signaling and induce apoptosis in human cancer cells. Proc. Natl. Acad. Sci. USA 102, 5998–6003 (2005).
Feng, B.Y., Shelat, A., Doman, T.N., Guy, R.K. & Shoichet, B.K. High-throughput assays for promiscuous inhibitors. Nat. Chem. Biol. 1, 146–148 (2005).
Hopkins, A.L. & Groom, C.R. The druggable genome. Nat. Rev. Drug Discov. 1, 727–730 (2002).
Overington, J.P., Al-Lazikani, B. & Hopkins, A.L. How many drug targets are there? Nat. Rev. Drug Discov. 5, 993–996 (2006).
Cheng, A.C. et al. Structure-based maximal affinity model predicts small-molecule druggability. Nat. Biotechnol. 25, 71–75 (2007).
Kitano, H. A robustness-based approach to systems-oriented drug design. Nat. Rev. Drug Discov. 6, 202–210 (2007).
Clutter, M.R., Krutzik, P.O. & Nolan, G.P. Phospho-specific flow cytometry in drug discovery. Drug Discov. Today Technol. 2, 295–302 (2005).
Darzynkiewicz, Z., Crissman, H. & Jacobberger, J.W. Cytometry of the cell cycle: cycling through history. Cytometry A 58, 21–32 (2004).
Schulz, K.R., Danna, E.A., Krutzik, P.O. & Nolan, G.P. Single-cell phospho-protein analysis by flow cytometry. Curr. Protoc. Immunol. 78, 8.17 (2007).
Acknowledgements
We thank E. Danna, K. Schulz and K. Gibbs for critical reading of this manuscript. We thank BD Pharmingen, in particular G. Gao, R. Campos and B. Balderas, for kindly providing antibody reagents and technical expertise. We also thank the NCI DTP and J. Johnson for providing the natural product library used in this study. P.O.K. was supported in part by a Howard Hughes Medical Institute Predoctoral Fellowship. G.P.N. was supported by US National Heart, Lung and Blood Institute contract N01-HV-28183 and US National Institutes of Health grant AI35304.
Author information
Authors and Affiliations
Contributions
P.O.K. designed the study, performed experiments, analyzed data and wrote the manuscript. J.M.C. performed experiments and analyzed data. M.R.C. performed experiments and helped write the manuscript. G.P.N. helped to design the study and write the manuscript.
Corresponding author
Ethics declarations
Competing interests
G.P.N. and P.O.K. are each paid consultants to Becton Dickinson, a provider of antibodies and flow cytometry-based reagents. G.P.N. consults occasionally with multiple pharmaceutical and biotechnology concerns about flow-based analysis of signaling systems.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Table 1 (PDF 1179 kb)
Rights and permissions
About this article
Cite this article
Krutzik, P., Crane, J., Clutter, M. et al. High-content single-cell drug screening with phosphospecific flow cytometry. Nat Chem Biol 4, 132–142 (2008). https://doi.org/10.1038/nchembio.2007.59
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2007.59
This article is cited by
-
Electron transfer-triggered imaging of EGFR signaling activity
Nature Communications (2022)
-
Functional patient-derived cellular models for neuropsychiatric drug discovery
Translational Psychiatry (2021)
-
Exploring the neuropsychiatric spectrum using high-content functional analysis of single-cell signaling networks
Molecular Psychiatry (2020)
-
Streptonigrin at low concentration promotes heterochromatin formation
Scientific Reports (2020)
-
Nanoscale monitoring of mitochondria and lysosome interactions for drug screening and discovery
Nano Research (2019)