Abstract
Angiogenic sprouting needs to be tightly controlled. It has been suggested that the Notch ligand dll4 expressed in leading tip cells restricts angiogenesis by activating Notch signalling in trailing stalk cells. Here, we show using live imaging in zebrafish that activation of Notch signalling is rather required in tip cells. Notch activation initially triggers expression of the chemokine receptor cxcr4a. This allows for proper tip cell migration and connection to the pre-existing arterial circulation, ultimately establishing functional arterial–venous blood flow patterns. Subsequently, Notch signalling reduces cxcr4a expression, thereby preventing excessive blood vessel growth. Finally, we find that Notch signalling is dispensable for limiting blood vessel growth during venous plexus formation that does not generate arteries. Together, these findings link the role of Notch signalling in limiting angiogenesis to its role during artery formation and provide a framework for our understanding of the mechanisms underlying blood vessel network expansion and maturation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Benedito, R. & Hellstrom, M. Notch as a hub for signaling in angiogenesis. Exp. Cell Res. 319, 1281–1288 (2013).
Phng, L. K. & Gerhardt, H. Angiogenesis: a team effort coordinated by notch. Dev. Cell 16, 196–208 (2009).
Siekmann, A. F., Affolter, M. & Belting, H. G. The tip cell concept 10 years after: new players tune in for a common theme. Exp. Cell Res. 319, 1255–1263 (2013).
Jakobsson, L. et al. Endothelial cells dynamically compete for the tip cell position during angiogenic sprouting. Nat. Cell Biol. 12, 943–953 (2010).
Lobov, I. B. et al. Delta-like ligand 4 (Dll4) is induced by VEGF as a negative regulator of angiogenic sprouting. Proc. Natl Acad. Sci. USA 104, 3219–3224 (2007).
Ubezio, B. et al. Synchronization of endothelial Dll4-Notch dynamics switch blood vessels from branching to expansion. eLife 5, e12167 (2016).
Hellstrom, M. et al. Dll4 signalling through Notch1 regulates formation of tip cells during angiogenesis. Nature 445, 776–780 (2007).
Leslie, J. D. et al. Endothelial signalling by the Notch ligand Delta-like 4 restricts angiogenesis. Development 134, 839–844 (2007).
Siekmann, A. F. & Lawson, N. D. Notch signalling limits angiogenic cell behaviour in developing zebrafish arteries. Nature 445, 781–784 (2007).
Suchting, S. et al. The Notch ligand Delta-like 4 negatively regulates endothelial tip cell formation and vessel branching. Proc. Natl Acad. Sci. USA 104, 3225–3230 (2007).
Lawson, N. D. et al. Notch signaling is required for arterial-venous differentiation during embryonic vascular development. Development 128, 3675–3683 (2001).
Lawson, N. D., Vogel, A. M. & Weinstein, B. M. sonic hedgehog and vascular endothelial growth factor act upstream of the Notch pathway during arterial endothelial differentiation. Dev. Cell 3, 127–136 (2002).
Duarte, A. et al. Dosage-sensitive requirement for mouse Dll4 in artery development. Genes Dev. 18, 2474–2478 (2004).
Gale, N. W. et al. Haploinsufficiency of delta-like 4 ligand results in embryonic lethality due to major defects in arterial and vascular development. Proc. Natl Acad. Sci. USA 101, 15949–15954 (2004).
Krebs, L. T. et al. Haploinsufficient lethality and formation of arteriovenous malformations in Notch pathway mutants. Genes Dev. 18, 2469–2473 (2004).
Bussmann, J., Wolfe, S. A. & Siekmann, A. F. Arterial-venous network formation during brain vascularization involves hemodynamic regulation of chemokine signaling. Development 138, 1717–1726 (2011).
Strasser, G. A., Kaminker, J. S. & Tessier-Lavigne, M. Microarray analysis of retinal endothelial tip cells identifies CXCR4 as a mediator of tip cell morphology and branching. Blood 115, 5102–5110 (2010).
del Toro, R. et al. Identification and functional analysis of endothelial tip cell-enriched genes. Blood 116, 4025–4033 (2010).
Kidoya, H. et al. APJ regulates parallel alignment of arteries and veins in the skin. Dev. Cell 33, 247–259 (2015).
Xu, C. et al. Arteries are formed by vein-derived endothelial tip cells. Nat. Commun. 5, 5758 (2014).
Kiefer, F. & Siekmann, A. F. The role of chemokines and their receptors in angiogenesis. Cell. Mol. Life Sci. 68, 2811–2830 (2011).
Nagasawa, T. CXC chemokine ligand 12 (CXCL12) and its receptor CXCR4. J. Mol. Med. (Berl) 92, 433–439 (2014).
Ara, T., Tokoyoda, K., Okamoto, R., Koni, P. A. & Nagasawa, T. The role of CXCL12 in the organ-specific process of artery formation. Blood 105, 3155–3161 (2005).
Tachibana, K. et al. The chemokine receptor CXCR4 is essential for vascularization of the gastrointestinal tract. Nature 393, 591–594 (1998).
Fujita, M. et al. Assembly and patterning of the vascular network of the vertebrate hindbrain. Development 138, 1705–1715 (2011).
Siekmann, A. F., Standley, C., Fogarty, K. E., Wolfe, S. A. & Lawson, N. D. Chemokine signaling guides regional patterning of the first embryonic artery. Gene Dev. 23, 2272–2277 (2009).
Harrison, M. R. et al. Chemokine-guided angiogenesis directs coronary vasculature formation in zebrafish. Dev. Cell 33, 442–454 (2015).
Ivins, S. et al. The CXCL12/CXCR4 axis plays a critical role in coronary artery development. Dev. Cell 33, 455–468 (2015).
Li, W. et al. Peripheral nerve-derived CXCL12 and VEGF-A regulate the patterning of arterial vessel branching in developing limb skin. Dev. Cell 24, 359–371 (2013).
Isogai, S., Horiguchi, M. & Weinstein, B. M. The vascular anatomy of the developing zebrafish: an atlas of embryonic and early larval development. Dev. Biol. 230, 278–301 (2001).
Kaufman, R. et al. Development and origins of zebrafish ocular vasculature. BMC Dev. Biol. 15, 18 (2015).
Kochhan, E. et al. Blood flow changes coincide with cellular rearrangements during blood vessel pruning in zebrafish embryos. PLoS ONE 8, e75060 (2013).
Claxton, S. & Fruttiger, M. Periodic Delta-like 4 expression in developing retinal arteries. Gene Exp. Patterns 5, 123–127 (2004).
Hofmann, J. J. & Luisa Iruela-Arispe, M. Notch expression patterns in the retina: an eye on receptor-ligand distribution during angiogenesis. Gene Exp. Patterns 7, 461–470 (2007).
Quillien, A. et al. Distinct Notch signaling outputs pattern the developing arterial system. Development 141, 1544–1552 (2014).
Ninov, N., Borius, M. & Stainier, D. Y. Different levels of Notch signaling regulate quiescence, renewal and differentiation in pancreatic endocrine progenitors. Development 139, 1557–1567 (2012).
Li, X. et al. Generation of destabilized green fluorescent protein as a transcription reporter. J. Biol. Chem. 273, 34970–34975 (1998).
Bray, S. J. Notch signalling in context. Nat. Rev. Mol. Cell Biol. 17, 722–735 (2016).
Coumailleau, F., Furthauer, M., Knoblich, J. A. & Gonzalez-Gaitan, M. Directional Delta and Notch trafficking in Sara endosomes during asymmetric cell division. Nature 458, 1051–1055 (2009).
Costa, G. et al. Asymmetric division coordinates collective cell migration in angiogenesis. Nat. Cell Biol. 18, 1292–1301 (2016).
Itoh, M. et al. Mind bomb is a ubiquitin ligase that is essential for efficient activation of Notch signaling by Delta. Dev. Cell 4, 67–82 (2003).
Sainson, R. C. et al. Cell-autonomous notch signaling regulates endothelial cell branching and proliferation during vascular tubulogenesis. FASEB J. 19, 1027–1029 (2005).
Packham, I. M. et al. Microarray profiling reveals CXCR4a is downregulated by blood flow in vivo and mediates collateral formation in zebrafish embryos. Physiol. Genomics 38, 319–327 (2009).
Isogai, S., Lawson, N. D., Torrealday, S., Horiguchi, M. & Weinstein, B. M. Angiogenic network formation in the developing vertebrate trunk. Development 130, 5281–5290 (2003).
De Bock, K. et al. Role of PFKFB3-driven glycolysis in vessel sprouting. Cell 154, 651–663 (2013).
Williams, C. K. et al. Regulation of CXCR4 by the Notch ligand delta-like 4 in endothelial cells. Cancer Res. 68, 1889–1895 (2008).
le Noble, F. et al. Flow regulates arterial-venous differentiation in the chick embryo yolk sac. Development 131, 361–375 (2004).
Serra, H. et al. PTEN mediates Notch-dependent stalk cell arrest in angiogenesis. Nat. Commun. 6, 7935 (2015).
Pitulescu, M. E. Dll4 and Notch signalling couples sprouting angiogenesis and artery formation. Nat. Cell Biol. 19, http://dx.doi.org/10.1038/ncb3555 (2017).
Noguera-Troise, I. et al. Blockade of Dll4 inhibits tumour growth by promoting non-productive angiogenesis. Nature 444, 1032–1037 (2006).
Westerfield, M. The Zebrafish Book: A Guide for the Laboratory use of Zebrafish (Brachydanio rerio) (Univ. Oregon Press, 1993).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Wang, Y. et al. Moesin1 and Ve-cadherin are required in endothelial cells during in vivo tubulogenesis. Development 137, 3119–3128 (2010).
Asakawa, K. et al. Genetic dissection of neural circuits by Tol2 transposon-mediated Gal4 gene and enhancer trapping in zebrafish. Proc. Natl Acad. Sci. USA 105, 1255–1260 (2008).
Parsons, M. J. Notch-responsive cells initiate the secondary transition in larval zebrafish pancreas. Mech. Dev. 126, 898–912 (2009).
Qian, F. et al. Microarray analysis of zebrafish cloche mutant using amplified cDNA and identification of potential downstream target genes. Dev. Dyn. 233, 1163–1172 (2005).
Roman, B. L. et al. Disruption of acvrl1 increases endothelial cell number in zebrafish cranial vessels. Development 129, 3009–3019 (2002).
Hogan, B. M. et al. ccbe1 is required for embryonic lymphangiogenesis and venous sprouting. Nat. Genetics 41, 396–398 (2009).
Lawson, N. D. & Weinstein, B. M. In vivo imaging of embryonic vascular development using transgenic zebrafish. Dev. Biol. 248, 307–318 (2002).
Bussmann, J. et al. Arteries provide essential guidance cues for lymphatic endothelial cells in the zebrafish trunk. Development 137, 2653–2657 (2010).
Traver, D. et al. Transplantation and in vivo imaging of multilineage engraftment in zebrafish bloodless mutants. Nat. Immunol. 4, 1238–1246 (2003).
Jiang, Y. J. et al. Mutations affecting neurogenesis and brain morphology in the zebrafish, Danio rerio. Development 123, 205–216 (1996).
Scheer, N. & Campos-Ortega, J. A. Use of the Gal4-UAS technique for targeted gene expression in the zebrafish. Mech. Dev. 80, 153–158 (1999).
Bussmann, J. & Schulte-Merker, S. Rapid BAC selection for tol2-mediated transgenesis in zebrafish. Development 138, 4327–4332 (2011).
Hermkens, D. M. et al. Sox7 controls arterial specification in conjunction with hey2 and efnb2 function. Development 142, 1695–1704 (2015).
Bahary, N. et al. Duplicate VegfA genes and orthologues of the KDR receptor tyrosine kinase family mediate vascular development in the zebrafish. Blood 110, 3627–3636 (2007).
Kochhan, E. & Siekmann, A. F. in Chemokines: Methods and Protocols (eds Cardona, E. A. & Ubogu, E. E.) 145–159 (Humana Press, 2013).
Brend, T. & Holley, S. A. Zebrafish whole mount high-resolution double fluorescent in situ hybridization. J. Vis. Exp. http://dx.doi.org/10.3791/1229 (2009).
Moore, J. C. et al. Post-transcriptional mechanisms contribute to Etv2 repression during vascular development. Dev. Biol. 384, 128–140 (2013).
Ulrich, F., Ma, L. H., Baker, R. G. & Torres-Vazquez, J. Neurovascular development in the embryonic zebrafish hindbrain. Dev. Biol. 357, 134–151 (2011).
Kronstein, R. et al. Caveolin-1 opens endothelial cell junctions by targeting catenins. Cardiovasc. Res. 93, 130–140 (2012).
Acknowledgements
This work was funded by the Max Planck Society (A.F.S.), the Deutsche Forschungsgemeinschaft (DFG SI-1374/3-1; DFG SI-1374/3-2; DFG SI-1374/4-1; DFG SI-1374/5-1; A.F.S.; DFG SCHN 43076-2/663; H.S.) and an ERC starting grant (260794-ZebrafishAngio; A.F.S.). This work was supported by the Deutsche Forschungsgemeinschaft (DFG) Cells-in-Motion Cluster of Excellence (EXC 1003-CIM), University of Münster, Germany (Flexible funds FF-2014-15 to A.F.S. and H.S.) and the National Institutes of Health (R01 HL093467, N.D.L.). We would like to thank W. Sugden for critical reading of the manuscript and J. Bussmann for generating the Tg(dll4:gal4)mu106 zebrafish line.
Author information
Authors and Affiliations
Contributions
S.S.H. and A.F.S. designed the experiments, performed experiments, analysed the data and wrote the manuscript. M.L. performed in situ hybridization and analysed data. R.T. performed HUAEC experiments, qPCR analysis and ChIP. L.W. analysed data. K.W. analysed data. J.C.M. and N.D.L. provided the fli1b antibody. H.S. provided HUAECs.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Validation and activation of dll4 transgenic reporter line.
(a) Double FISH of dll4 mRNA together with GFP mRNA and an antibody staining to detect GFP protein at different time points in Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a double transgenic embryos to mark ECs with dll4 expression. Inset indicates zoomed in NCA region below (n = 8 embryos for each time point, 2 independent experiments). (b) Double FISH of dll4 mRNA together with GFP mRNA and an antibody staining to detect GFP protein in the trunk region of a zebrafish embryo at 24 hpf. Insets mark zoomed in region. Arrowheads mark dll4 mRNA (red), GFP mRNA (green), GFP protein (white) or overlay of channels (n = 8 embryos, 2 independent experiments). (c) Still images of video 2 at indicated time points of NCA sprouting from the PMBC and connecting to the CrDI in Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a, marking cells expressing dll4 in green, and Tg(kdrl:NLS-mcherry)is4, which marks all EC nuclei in red, triple transgenic embryos. White arrowheads with numbers mark individual ECs. Note onset of GFP expression in cell number 1 at the 2:36 h time point, followed by GFP expression in cells 2 and 3. (d) Still images of video 3 at indicated time points of NCA sprouting from the PMBC and connecting to the CrDI in Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a; Tg(kdrl:NLS-mcherry)is4 triple transgenic embryos. White arrowheads with numbers mark individual ECs. Note onset of GFP expression in cell number 1 and 2 at the 5:00 h time point, followed by GFP expression in cell number 3. (e) Quantification of fluorescence intensity of the green Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a signal normalized to red Tg(kdrl:NLS-mcherry)is4 signal over time in video 2. (f) Quantification of fluorescence intensity of the green Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a signal normalized to red Tg(kdrl:NLS-mcherry)is4 signal over time in video 3. (n = 3 embryos, 3 independent experiments). t0 = 26 hpf. Scale bars, 50 μm. A.U.-arbitrary units; CrDI-cranial division of the internal carotid artery; EC-endothelial cell; h-hour; hpf-hours post fertilization; ISV-intersegmental blood vessel; NCA-nasal ciliary artery; PMBC-primordial midbrain channel; FISH-Fluorescent in situ hybridization.
Supplementary Figure 2 Notch signalling activity in newly forming arteries.
(a) Living triple transgenic Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a; Tg(kdrl:Hsa.HRAS-mcherry)s916zebrafish embryo at 50 hpf. Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a double transgenic embryos mark ECs with dll4 expression in green, while Tg(kdrl:Hsa.HRAS-mcherry)s916 marks EC membranes in red. Note EGFP expression in NCA (n = 10 embryos, 3 independent experiments). (b) Living double transgenic Tg(TP1:GFP)um14; Tg(kdrl:Hsa.HRAS-mcherry)s916 zebrafish embryo. Tg(TP1:GFP)um14 marks cells with Notch pathway activation in green (n = 10 embryos, 3 independent experiments). (c) Transplantation scheme. Kdrl knockdown cells (green) were transferred into wt hosts (red). (d) Representative embryo, showing sprouting NCA cells at 26 hpf time point. Note absence of green fluorescent donor ECs in sprouting NCA (n = 9 embryos, 3 independent experiments). (e) dll4 expression in sprouting NCA cells in control embryo at 30 hpf after DMSO treatment for 4 h and experimental embryo at 30 hpf after SU5416 (VEGF inhibitor) treatment for 4 h (n = 10 embryos, 2 independent experiments). (f) dll4 expression in sprouting NCA cells in control embryo at 32 hpf after DMSO treatment for 4 h and experimental embryo at 30 hpf after SU5416 (VEGF inhibitor) treatment for 4 h (n = 10 embryos, 2 independent experiments). Arrowheads indicate dll4 expressing ECs, marked by Tg(kdrl:EGFP)s843. In experimental embryos, while ECs are present in the position of the NCA, these do not express dll4 (n = 10 embryos). (g,h) Still images of g video 2 and h video 3 at indicated time points of NCA sprouting in Tg(TP1:GFP)um14; Tg(kdrl:NLS-mcherry)is4 double transgenic embryos. White arrowheads with numbers indicate individual ECs. Note onset of GFP expression in cell number 1 at the 4:35 h time point, followed by GFP expression in cells 3 and 4 in video 2. Note onset of GFP expression in cell number 1 at the 3:28 h time point, followed by division of this cell, in video 3. GFP expression can be subsequently detected in cell 2. (i,j) Quantification of fluorescence intensity of green Tg(TP1:GFP)um14 signal normalized to red Tg(kdrl:NLS-mcherry)is4 signal over time in i video 2 and j video 3. (n = 3 embryos, 3 independent experiments). t0 = 26 hpf. Scale bars, 50 μm. A.U.-arbitrary units; CrDI-Cranial division of the internal carotid artery; PMBC-primordial midbrain channel; NCA-nasal ciliary artery; h-hour; hpf-hours post fertilization.
Supplementary Figure 3 Activation of Notch signalling in transgenic reporter zebrafish line expressing destabilized Venus protein in sprouting eye blood vessels.
(a) Still images of video 1 at indicated time points of NCA sprouting in Tg(TP1:Venus-PEST)s940; Tg(kdrl:NLS-mcherry)is4 double transgenic embryos. Tg(TP1:Venus-PEST)s940 marks cells with Notch pathway activation in green. Tg(kdrl:NLS-mcherry)is4 marks EC nuclei in red. White arrowheads with numbers mark individual ECs. Note onset of Venus-PEST expression in cell number 1.1 at the 3:28 h time point, followed by Venus-PEST expression in cells 1.2 and 3. (b) Still images of video 2 at indicated time points of NCA sprouting in Tg(TP1:Venus-PEST)s940; Tg(kdrl:NLS-mcherry)is4 double transgenic embryos. White arrowheads with numbers mark individual ECs. Note onset of GFP expression in cell number 1.1 and 1.2 at the 7:37 h time point. (c) Still images of video 3 at indicated time points of NCA sprouting in Tg(TP1:Venus-PEST)s940; Tg(kdrl:NLS-mcherry)is4 double transgenic embryos. White arrowheads with numbers mark individual ECs. Note onset of GFP expression in cell number 1 at the 6:36 h time point. (d) Quantification of fluorescence intensity of green Tg(TP1:Venus-PEST)s940 signal normalized to red Tg(kdrl:NLS-mcherry)is4 signal over time in video 1. (e) Quantification of fluorescence intensity of green Tg(TP1:Venus-PEST)s940 signal normalized to red Tg(kdrl:NLS-mcherry)is4 signal over time in video 2. (f) Quantification of fluorescence intensity of green Tg(TP1:Venus-PEST)s940 signal normalized to red Tg(kdrl:NLS-mcherry)is4 signal over time in video 3. (n = 3 embryos, 3 independent experiments). t0 = 26 hpf. Scale bars, 50 μm. A.U.-arbitrary units; CrDI-Cranial division of the internal carotid artery; PMBC-primordial midbrain channel; NCA-nasal ciliary artery; hpf-hours post fertilization.
Supplementary Figure 4 Simultaneous detection of dll4 expression and Notch pathway activation during NCA formation.
Still images at indicated time points of NCA sprouting in Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a; Tg(TP1:H2B-mcherry)s939 triple transgenic embryos to detect dll4 expression and Notch pathway activation simultaneously. Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a marks cells with dll4 expression in green, while Tg(TP1:H2B-mcherry)s939marks cells with Notch pathway activation in red. White arrowheads with numbers mark individual ECs. (a) Video 1: Note onset of dll4 expression in cell number 1 at the 2:15 h time point, followed by Notch activation at 5:10 h time point. (b) Video 2: Note onset of dll4 expression in cell number 1 at the 1:16 h time point, followed by Notch activation at 5:34 h time point. (c) Video 3: Note onset of dll4 expression in cell number 1 at the 2:16 h time point, followed by Notch activation at 7:21 h time point. (n = 3 embryos, 3 independent experiments). t0 = 26 hpf. Scale bars, 50 μm. CrDI-Cranial division of the internal carotid artery; EC-endothelial cell; NCA-nasal ciliary artery; h-hour; hpf-hours post fertilization.
Supplementary Figure 5 Absence of Notch activation in dll4 knockdown embryos during NCA formation.
(a,b) Still images of a video 2 and b video 3 at indicated time points of NCA sprouting in Tg(TP1:GFP)um14; Tg(kdrl:NLS-mcherry)is4 double transgenic embryo injected with dll4 MO. Tg(TP1:GFP)um14 marks cells with Notch pathway activation in green, while Tg(kdrl:NLS-mcherry)is4 marks EC nuclei in red. White arrowheads with numbers indicate individual ECs. (c,d) Quantification of fluorescence intensity of green Tg(TP1:GFP)um14 signal normalized to red Tg(kdrl:NLS-mcherry)is4 signal over time in c video 2 and d video 3. t0 = 26 hpf. (n = 3 embryos, 3 independent experiments). (e) Transplantation scheme. Transferring dll4 knockdown cells from Tg(kdrl:Hsa.HRAS-mcherry)s916; Tg(TP1:GFP)um14 into wt hosts allows to distinguish trans- from cis-activation of Notch signalling in ECs. Tg(kdrl:Hsa.HRAS-mcherry)s916 marks donor EC membranes in red, while Tg(TP1:GFP)um14 marks donor cells with Notch signalling pathway activation in green. (f–h) Schematic detailing different outcomes of cell transplantations. (i–o) Embryo 2. (p–v) Embryo 3 (i,p) NCA cells stained with a Fli1b antibody detecting EC nuclei (white). (j,q) Transplanted dll4 knockdown ECs. Arrows mark tip cells. (k,r) Cells with activated Notch signalling identified via GFP expression (green arrows). (l,s) Overlay of transplanted NCA cells with host NCA cells, showing that only cell 1 is donor-derived, while cells 2, 3, 4 and 5 are host-derived. (m,t) Overlay of transplanted cells showing Notch pathway activation (green arrows) with host cells. Only the tip cell (number 1) shows GFP expression. (n,u) Overlay of all three channels, identifying transplanted dll4 knockdown tip cell being GFP positive. (o) Zoom in of boxed area in n. Note tip cell (number 1) being next to a wt host cell (number 2). (v) Zoom in of boxed area in u. Note tip cell (number 1) being next to a wt host cell (number 2). Green arrows denote Notch pathway activation (n = 3 mosaic embryos with single donor cell at tip position, 3 independent experiments). Scale bars, 50 μm. Ab-antibody; EC-endothelial cell; hpf-hours post fertilization; CrDI-Cranial division of the internal carotid artery; PMBC-primordial midbrain channel; NCA-nasal ciliary artery.
Supplementary Figure 6 Notch signalling does not mediate competition between endothelial cells during intersegmental blood vessel sprouting.
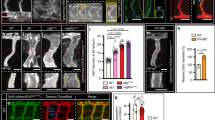
(a) Transplantation scheme. Tg(kdrl:EGFP)s843 marks donor cells that were either injected with a ctr or dll4 MO in green. Tg(kdrl:Hsa.HRAS-mcherry)s916 marks host vasculature in red. (b) Representative image of mosaic embryos at 32 hpf. (c) Quantification of contribution of ctr MO (n = 19 mosaic embryos) or dll4 MO (n = 23 mosaic embryos) injected ECs to different positions within the trunk vasculature (d) Transplantation of dll4 mutant cells reveals no preference for cell position within trunk blood vessels. Number of embryos with indicated distribution of transplanted cells indicated (n = 25 dll4+/− to WT mosaic embryos; n = 10 dll4−/− to WT mosaic embryos in 3 independent experiments). (e) Transplantation scheme. Tg(fli1a:EGFP)y1 marks donor cells that were either wt, hetero- or homozygous for mib in green. Tg(kdrl:Hsa.HRAS-mcherry)s916 marks host vasculature in red. (f) Representative image of mosaic embryos at 32 hpf. Arrowheads mark donor cells at tip positions. (g) Quantification of contribution of mib sibling (n = 17 mosaic embryos in 3 independent experiments) or homozygous mib mutant (n = 12 mosaic embryos in 3 independent experiments) ECs to different positions within the trunk vasculature. Images are representative of three independent experiments. (h) Still images of video 1 at indicated time points of ISV sprouting. Tg(fli1a:nEGFP)y7 marks EC nuclei in green. Numbers delineate individual ECs or their progeny. Out of 10 ISVs observed, we only detected one shuffling event (white arrowhead in g). (i) Still images of video 2 at indicated time points of ISV sprouting. Tg(fli1a:nEGFP)y7 marks EC nuclei in green. Numbers delineate individual ECs or their progeny. Out of 10 ISVs observed, we did not detect EC shuffling. (n = 2 embryos, 2 independent experiments). Scale bars, 50 μm. aISV-arterial intersegmental vessel; ctr-control; DA-dorsal aorta; DLAV-dorsal longitudinal ananstomotic vessel; hpf-hours post fertilization; MO-morpholino; PCV-posterior cardinal vein.
Supplementary Figure 7 The chemokine receptor cxcr4a affects tip cell migration, while dll4 knockdown causes aberrant blow flow patterns.
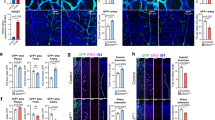
(a) Wt ctr video 1 showing sprouting of the NCA from the PMBC at indicated time points. Note migratory behaviours of cells 1 and 2 towards the CrDI during the 3:30 h time course. Tg(kdrl:EGFP)s843 marks ECs in green. (b) Wt ctr video 2 showing sprouting of the NCA from the PMBC at indicated time points. Note migratory behaviours of cells 1 and 2 towards the CrDI during the 3:30 h time course. (c) NCA sprouting in cxcr4a mutant embryo 1. Note normal initiation of NCA sprouting, but subsequent stalling of EC 1. Even at the 8:14 h time point, no connection between CrDI and NCA can be observed. At 16:47 h time point, a small sprout extends from the NCA towards the CrDI (arrowhead). (d) NCA sprouting in cxcr4a mutant embryo 2. Note stalling of ECs 1 and 2. At the 6:46 h time point, a sprout emanates from the CrDI (arrow) and subsequently connects to the NCA, but without generating a proper lumen. (n = 3 embryos, 3 independent experiments). (e) Quantification of blood flow patterns at 54 hpf and 72 hpf in ctr MO injected embryos (n = 19 embryos, 2 independent experiments). Note regular flow patterns and absence of ectopic sprouts. (f) Quantification of blood flow patterns at 54 hpf and 72 hpf in dll4 MO injected embryos (n = 19 embryos, 2 independent experiments). Note lack of blood flow, increase in venous ISVs and ectopic sprouts (green bars). (g) Aberrant blood flow patterns correlate with increased ectopic blood vessel sprouting in dll4 knockdown embryos (n = 19, 2 independent experiments). White arrows indicate ectopic blood vessel sprouts. These are more frequently found in areas containing ISVs without blood flow. (h) Blocking blood flow from 48 to 52 hpf leads to upregulation of cxcr4a expression in the DA and DLAV (arrows). ‘n/N’ reports the ‘number of embryos with staining pattern in image’/‘total embryos from three experiments’ (N = 20 embryos for DMSO and N = 15 embryos for Nifedipine treatment). ∗∗∗P < 0.0001, Fischer’s exact test. Scale bars, 50 μm. CrDI-Cranial division of the internal carotid artery; EC-endothelial cell; PMBC-primordial midbrain channel; NCA-nasal ciliary artery; h-hour; DA-dorsal aorta; PCV-posterior cardinal vein.
Supplementary Figure 8 dll4 and cxcr4a are not expressed in endothelial cells during venous plexus blood vessel sprouting.
(a) Expression of the dll4 reporter Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a(green) cannot be detected in sprouting venous plexus ECs (arrowheads) marked with Tg(kdrl:Hsa.HRAS-mcherry)s916 (red) at 28 hpf, 32 hpf or 38 hpf (n = 10 for each time point). (b) Expression of cxcr4a (red) cannot be detected in sprouting venous plexus ECs marked with an anti-GFP antibody staining in Tg(kdrl:EGFP)s843 (green) embryos (n = 10 for each time point). Visible red staining comes from overlying fin fold tissue. Images are representative of three independent experiments. (c) Current model of tip/stalk cell specification through lateral inhibition. VEGF signalling induces the Notch ligand dll4 in tip cells. High dll4 expression in tip cells activates Notch signalling in neighbouring stalk cells thereby preventing them from acquiring a tip cell phenotype. (d) Model integrating artery formation and Notch pathway activation during angiogenesis. Initially, Notch negative tip cells sprout from veins. These tip cells subsequently express dll4. Stalk cells also start expressing dll4. This leads to Notch activation via trans-activation first in tip and then also in stalk cells. Notch activation induces expression of the chemokine receptor cxcr4a. This allows proper tip cell migration towards the existing artery, ultimately allowing blood flow from arteries to veins. Scale bars, 50 μm. CVP-caudal vein plexus; DA-dorsal aorta; EC-endothelial cell; hpf-hours post fertilization.
Supplementary information
Supplementary Information
Supplementary Information (PDF 33516 kb)
Supplementary Information
Supplementary Information (PDF 69 kb)
Time lapse imaging of dll4 reporter line during eye vessel development- Embryo 1.
Time lapse confocal imaging of NCA development in Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a; Tg(kdrl:NLS-mcherry)is4 triple transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes:seconds. (MP4 671 kb)
Time lapse imaging of dll4 reporter line during eye vessel development- Embryo 2.
Time lapse confocal imaging of NCA development in Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a; Tg(kdrl:NLS-mcherry)is4 triple transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes:seconds. (MP4 726 kb)
Time lapse imaging of dll4 reporter line during eye vessel development- Embryo 3.
Time lapse confocal imaging of NCA development in Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a; Tg(kdrl:NLS-mcherry)is4 triple transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes:seconds. (MP4 698 kb)
Time lapse imaging of Notch reporter line during eye vessel development- Embryo 1.
Time lapse confocal imaging of NCA development in Tg(TP1:GFP)um14; Tg(kdrl:NLS-mcherry)is4 double transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes:seconds. (MP4 902 kb)
Time lapse imaging of Notch reporter line during eye vessel development- Embryo 2.
Time lapse confocal imaging of NCA development in Tg(TP1:GFP)um14; Tg(kdrl:NLS-mcherry)is4 double transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes:seconds. (MP4 682 kb)
Time lapse imaging of Notch reporter line during eye vessel development- Embryo 3.
Time lapse confocal imaging of NCA development in Tg(TP1:GFP)um14; Tg(kdrl:NLS-mcherry)is4 double transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes:seconds. (MP4 601 kb)
Time lapse imaging of destabilized Notch reporter line during eye vessel development- Embryo 1.
Time lapse confocal imaging of NCA development in Tg(TP1:Venus-PEST)s940; Tg(kdrl:NLS-mcherry)is4 double transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes. (MP4 742 kb)
Time lapse imaging of destabilized Notch reporter line during eye vessel development- Embryo 2.
Time lapse confocal imaging of NCA development in Tg(TP1:Venus-PEST)s940; Tg(kdrl:NLS-mcherry)is4 double transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes:seconds. (MP4 613 kb)
Time lapse imaging of destabilized Notch reporter line during eye vessel development- Embryo 3.
Time lapse confocal imaging of NCA development in Tg(TP1:Venus-PEST)s940; Tg(kdrl:NLS-mcherry)is4 double transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes:seconds. (MP4 783 kb)
Simultaneous imaging of dll4 expression and Notch activation during eye vessel development- Embryo 1.
Time lapse confocal imaging of NCA development in Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a; Tg(TP1:H2B-mcherry)s939 triple transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes. (MP4 486 kb)
Simultaneous imaging of dll4 expression and Notch activation during eye vessel development- Embryo 2.
Time lapse confocal imaging of NCA development in Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a; Tg(TP1:H2B-mcherry)s939 triple transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes. (MP4 578 kb)
Simultaneous imaging of dll4 expression and Notch activation during eye vessel development- Embryo 3.
Time lapse confocal imaging of NCA development in Tg(dll4:gal4)mu106; Tg(UAS:GFP)nkuasgfp1a; Tg(TP1:H2B-mcherry)s939 triple transgenic embryo. Image acquisition started at 26 hpf. Elapsed time in hours:minutes. (MP4 792 kb)
Time lapse imaging of Notch reporter line during eye vessel development with dll4 knockdown- Embryo 1.
Time lapse confocal imaging of NCA development in Tg(TP1:GFP)um14; Tg(kdrl:NLS-mcherry)is4 double transgenic embryo injected with dll4 morpholino. Image acquisition started at 26 hpf. Elapsed time in hours:minutes. (MP4 518 kb)
Time lapse imaging of Notch reporter line during eye vessel development with dll4 knockdown- Embryo 2.
Time lapse confocal imaging of NCA development in Tg(TP1:GFP)um14; Tg(kdrl:NLS-mcherry)is4 double transgenic embryo injected with dll4 morpholino. Image acquisition started at 26 hpf. Elapsed time in hours:minutes. (MP4 221 kb)
Time lapse imaging of Notch reporter line during eye vessel development with dll4 knockdown- Embryo 3.
Time lapse confocal imaging of NCA development in Tg(TP1:GFP)um14; Tg(kdrl:NLS-mcherry)is4 double transgenic embryo injected with dll4 morpholino. Image acquisition started at 26 hpf. Elapsed time in hours:minutes. (MP4 312 kb)
Time lapse imaging of Tg(kdrl:EGFP)s843 during eye vessel development- Embryo 1.
Time lapse confocal imaging of NCA development in Tg(kdrl:EGFP)s843 wt embryo starting at 30 hpf. Elapsed time in hours:minutes. (MOV 4014 kb)
Time lapse imaging of Tg(kdrl:EGFP)s843 during eye vessel development- Embryo 2.
Time lapse confocal imaging of NCA development in Tg(kdrl:EGFP)s843 wt embryo starting at 30 hpf. Elapsed time in hours:minutes. (MOV 4051 kb)
Time lapse imaging of cxcr4aum20 mutant during eye vessel development- Embryo 1.
Time lapse confocal imaging of NCA development in cxcr4aum20 Tg(kdrl:EGFP)s843 embryo starting at 26 hpf. Elapsed time in hours:minutes. (MOV 311 kb)
Time lapse imaging of cxcr4aum20 mutant during eye vessel development- Embryo 2.
Time lapse confocal imaging of NCA development in cxcr4aum20 Tg(kdrl:EGFP)s843 embryo starting at 26 hpf. Elapsed time in hours:minutes. (MOV 237 kb)
Time lapse imaging of caudal vein plexus development in control morpholino injected embryo.
Time lapse confocal imaging of caudal vein plexus development in Tg(fli1a:nEGFP)y7; Tg(kdrl:Hsa.HRAS-mcherry)s916 double transgenic embryo injected with control morpholino. Image acquisition started at 24 hpf. Elapsed time in hours:minutes:seconds. (MP4 1170 kb)
Time-lapse imaging of caudal vein plexus development in dll4 morpholino injected embryo.
Time lapse confocal imaging of caudal vein plexus development in Tg(fli1a:nEGFP)y7; Tg(kdrl:Hsa.HRAS-mcherry)s916 double transgenic embryo injected with dll4 morpholino. Image acquisition started at 24 hpf. Elapsed time in hours:minutes:seconds. (MP4 1847 kb)
Rights and permissions
About this article
Cite this article
Hasan, S., Tsaryk, R., Lange, M. et al. Endothelial Notch signalling limits angiogenesis via control of artery formation. Nat Cell Biol 19, 928–940 (2017). https://doi.org/10.1038/ncb3574
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3574
This article is cited by
-
Role of cell rearrangement and related signaling pathways in the dynamic process of tip cell selection
Cell Communication and Signaling (2024)
-
Eph-ephrin signaling couples endothelial cell sorting and arterial specification
Nature Communications (2024)
-
Parenchymal cues define Vegfa-driven venous angiogenesis by activating a sprouting competent venous endothelial subtype
Nature Communications (2024)
-
The Role of Resveratrol on Spinal Cord Injury: from Bench to Bedside
Molecular Neurobiology (2024)
-
Vascular endothelial cell development and diversity
Nature Reviews Cardiology (2023)