Abstract
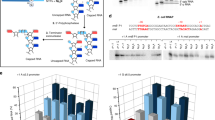
The long-standing assumption that messenger RNA (mRNA) degradation in Escherichia coli begins with endonucleolytic cleavage has been challenged by the recent discovery that RNA decay can be triggered by a prior non-nucleolytic event that marks transcripts for rapid turnover: the rate-determining conversion of the 5′ terminus from a triphosphate to a monophosphate1. This modification creates better substrates for the endonuclease RNase E, whose cleavage activity at internal sites is greatly enhanced when the RNA 5′ end is monophosphorylated2,3. Moreover, it suggests an explanation for the influence of 5′ termini on the endonucleolytic cleavage of primary transcripts, which are triphosphorylated4,5,6,7,8. However, no enzyme capable of removing pyrophosphate from RNA 5′ ends has been identified in any bacterial species. Here we show that the E. coli protein RppH (formerly NudH/YgdP) is the RNA pyrophosphohydrolase that initiates mRNA decay by this 5′-end-dependent pathway. In vitro, RppH efficiently removes pyrophosphate from the 5′ end of triphosphorylated RNA, irrespective of the identity of the 5′-terminal nucleotide. In vivo, it accelerates the degradation of hundreds of E. coli transcripts by converting their triphosphorylated 5′ ends to a more labile monophosphorylated state that can stimulate subsequent ribonuclease cleavage. That the action of the pyrophosphohydrolase is impeded when the 5′ end is structurally sequestered by a stem-loop helps to explain the stabilizing influence of 5′-terminal base pairing on mRNA lifetimes. Together, these findings suggest a possible basis for the effect of RppH and its orthologues on the invasiveness of bacterial pathogens. Interestingly, this master regulator of 5′-end-dependent mRNA degradation in E. coli not only catalyses a process functionally reminiscent of eukaryotic mRNA decapping but also bears an evolutionary relationship to the eukaryotic decapping enzyme Dcp2.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Celesnik, H., Deana, A. & Belasco, J. G. Initiation of RNA decay in Escherichia coli by 5′ pyrophosphate removal. Mol. Cell 27, 79–90 (2007)
Mackie, G. A. Ribonuclease E is a 5′-end-dependent endonuclease. Nature 395, 720–723 (1998)
Jiang, X. & Belasco, J. G. Catalytic activation of multimeric RNase E and RNase G by 5′-monophosphorylated RNA. Proc. Natl Acad. Sci. USA 101, 9211–9216 (2004)
Emory, S. A., Bouvet, P. & Belasco, J. G. A 5′-terminal stem-loop structure can stabilize mRNA in Escherichia coli . Genes Dev. 6, 135–148 (1992)
Bouvet, P. & Belasco, J. G. Control of RNase E-mediated RNA degradation by 5′-terminal base pairing in E. coli . Nature 360, 488–491 (1992)
Bricker, A. L. & Belasco, J. G. Importance of a 5′ stem-loop for longevity of papA mRNA in Escherichia coli . J. Bacteriol. 181, 3587–3590 (1999)
Mackie, G. A. Stabilization of circular rpsT mRNA demonstrates the 5′-end dependence of RNase E action in vivo. J. Biol. Chem. 275, 25069–25072 (2000)
Baker, K. E. & Mackie, G. A. Ectopic RNase E sites promote bypass of 5′-end-dependent mRNA decay in Escherichia coli . Mol. Microbiol. 47, 75–88 (2003)
McLennan, A. G. The Nudix hydrolase superfamily. Cell. Mol. Life Sci. 63, 123–143 (2006)
Mildvan, A. S. et al. Structures and mechanisms of Nudix hydrolases. Arch. Biochem. Biophys. 433, 129–143 (2005)
Mackie, G. A. & Parsons, G. D. Tandem promoters in the gene for ribosomal protein S20. J. Biol. Chem. 258, 7840–7846 (1983)
Mackie, G. Specific endonucleolytic cleavage of the mRNA for ribosomal protein S20 of Escherichia coli requires the product of the ams gene in vivo and in vitro. J. Bacteriol. 173, 2488–2497 (1991)
Mackie, G. A. Secondary structure of the mRNA for ribosomal protein S20. Implications for cleavage by ribonuclease E. J. Biol. Chem. 267, 1054–1061 (1992)
Ow, M. C., Perwez, T. & Kushner, S. R. RNase G of Escherichia coli exhibits only limited functional overlap with its essential homologue, RNase E. Mol. Microbiol. 49, 607–622 (2003)
Tock, M. R., Walsh, A. P., Carroll, G. & McDowall, K. J. The CafA protein required for the 5′-maturation of 16 S rRNA is a 5′-end-dependent ribonuclease that has context-dependent broad sequence specificity. J. Biol. Chem. 275, 8726–8732 (2000)
Jiang, X., Diwa, A. & Belasco, J. G. Regions of RNase E important for 5′-end-dependent RNA cleavage and autoregulated synthesis. J. Bacteriol. 182, 2468–2475 (2000)
Lee, K., Bernstein, J. A. & Cohen, S. N. RNase G complementation of rne null mutation identifies functional interrelationships with RNase E in Escherichia coli . Mol. Microbiol. 43, 1445–1456 (2002)
Feng, Y. & Cohen, S. N. Unpaired terminal nucleotides and 5′ monophosphorylation govern 3′ polyadenylation by Escherichia coli poly(A) polymerase I. Proc. Natl Acad. Sci. USA 97, 6415–6420 (2000)
Mitchell, S. J. & Minnick, M. F. Characterization of a two-gene locus from Bartonella bacilliformis associated with the ability to invade human erythrocytes. Infect. Immun. 63, 1552–1562 (1995)
Badger, J. L., Wass, C. A. & Kim, K. S. Identification of Escherichia coli K1 genes contributing to human brain microvascular endothelial cell invasion by differential fluorescence induction. Mol. Microbiol. 36, 174–182 (2000)
Ismail, T. M., Hart, C. A. & McLennan, A. G. Regulation of dinucleoside polyphosphate pools by the YgdP and ApaH hydrolases is essential for the ability of Salmonella enterica serovar typhimurium to invade cultured mammalian cells. J. Biol. Chem. 278, 32602–32607 (2003)
Edelstein, P. H. et al. Legionella pneumophila NudA Is a Nudix hydrolase and virulence factor. Infect. Immun. 73, 6567–6576 (2005)
Bessman, M. J. et al. The gene ygdP, associated with the invasiveness of Escherichia coli K1, designates a Nudix hydrolase, Orf176, active on adenosine (5′)-pentaphospho-(5′)-adenosine (Ap5A). J. Biol. Chem. 276, 37834–37838 (2001)
Muhlrad, D., Decker, C. J. & Parker, R. Deadenylation of the unstable mRNA encoded by the yeast MFA2 gene leads to decapping followed by 5′→3′ digestion of the transcript. Genes Dev. 8, 855–866 (1994)
Dunckley, T. & Parker, R. The DCP2 protein is required for mRNA decapping in Saccharomyces cerevisiae and contains a functional MutT motif. EMBO J. 18, 5411–5422 (1999)
Wang, Z., Jiao, X., Carr-Schmid, A. & Kiledjian, M. The hDcp2 protein is a mammalian mRNA decapping enzyme. Proc. Natl Acad. Sci. USA 99, 12663–12668 (2002)
Coleman, T. M., Wang, G. & Huang, F. Superior 5′ homogeneity of RNA from ATP-initiated transcription under the T7 φ2.5 promoter. Nucleic Acids Res. 32, e14 (2004)
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006)
Arnold, T. E., Yu, J. & Belasco, J. G. mRNA stabilization by the ompA 5′ untranslated region: two protective elements hinder distinct pathways for mRNA degradation. RNA 4, 319–330 (1998)
Emory, S. A. & Belasco, J. G. The ompA 5′ untranslated RNA segment functions in Escherichia coli as a growth-rate-regulated mRNA stabilizer whose activity is unrelated to translational efficiency. J. Bacteriol. 172, 4472–4481 (1990)
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000)
Acknowledgements
We are grateful to D. Guttman for his assistance in the discovery that purified RppH has RNA pyrophosphohydrolase activity. This research was supported by a grant to J.G.B. from the National Institutes of Health.
Author Contributions A.D., H.C. and J.G.B. planned the studies, interpreted the data and wrote the manuscript. A.D. and H.C. performed the experiments.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
The file contains Supplementary Figures S1-S5 with Legends and Supplementary Tables S1-S2 (PDF 2542 kb)
Rights and permissions
About this article
Cite this article
Deana, A., Celesnik, H. & Belasco, J. The bacterial enzyme RppH triggers messenger RNA degradation by 5′ pyrophosphate removal. Nature 451, 355–358 (2008). https://doi.org/10.1038/nature06475
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature06475
This article is cited by
-
N-terminal truncation of PhaCBP-M-CPF4 and its effect on PHA production
Microbial Cell Factories (2024)
-
Bacteriophages suppress CRISPR–Cas immunity using RNA-based anti-CRISPRs
Nature (2023)
-
Is mRNA decapping by ApaH like phosphatases present in eukaryotes beyond the Kinetoplastida?
BMC Ecology and Evolution (2021)
-
NUDT2 initiates viral RNA degradation by removal of 5′-phosphates
Nature Communications (2021)
-
Complete genome sequence and annotation of the laboratory reference strain Shigella flexneri serotype 5a M90T and genome-wide transcriptional start site determination
BMC Genomics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.