Abstract
Allantopyrone A is a fungal metabolite that uniquely possesses two α,β-unsaturated carbonyl moieties. We recently reported that allantopyrone A inhibited the nuclear factor-κB (NF-κB) signaling pathway induced by tumor necrosis factor (TNF)-α in human lung carcinoma A549 cells. In the present study, the mechanism by which allantopyrone A inhibits the TNF-α-induced signaling pathway was investigated in more detail. Allantopyrone A blocked extensive modifications to receptor-interacting protein 1 (RIP1) in the TNF receptor 1 (TNF-R1) complex. Allantopyrone A augmented the high-MW bands of TNF-R1, TNF receptor-associated factor 2, RIP1, the NF-κB subunit RelA and inhibitor of NF-κB kinase β in A549 cells, suggesting that it binds to and promotes the crosslinking of these proteins. The extracellular cysteine-rich domains of TNF-R1 were crosslinked by allantopyrone A more preferentially than its intracellular portion. The present results demonstrate that allantopyrone A interferes with multiple components of the TNF-R1 complex and blocks RIP1 modifications in the TNF-α-induced NF-κB signaling pathway.
Similar content being viewed by others
Introduction
Tumor necrosis factor (TNF)-α is a pro-inflammatory cytokine that is required to regulate the immune system.1 The deregulation of TNF-α is involved in chronic inflammation, and has been implicated in autoimmune diseases.2 Therapies that target TNF superfamily members, including TNF-α, have been developed for autoimmune and inflammatory diseases.3 TNF receptor 1 (TNF-R1) is a transmembrane protein that contains four cysteine-rich domains (CRDs) in the extracellular portion and a common death domain (DD) in the intracellular portion.4, 5 In addition, the pre-ligand-binding assembly domain overlapping CRD1 is necessary for the association of TNF-R1 in the absence of its ligands.6 The pre-ligand-binding assembly domain-mediated receptor preassociation is required for signaling by TNF-R1 and the TNF receptor family member Fas.6, 7
TNF-α binds to TNF-R1 in many types of cells, and mainly triggers the activation of the nuclear factor-κB (NF-κB) signaling pathway.8 TNF-R1 is trimerized in response to TNF-α, which induces the association of its intracellular DD with the TNF receptor-associated DD protein (TRADD).9 TRADD interacts with TNF receptor-associated factor 2 (TRAF2) and recruits receptor-interacting protein 1 (RIP1).9 This leads to the formation of the TNF-R1 complex (termed complex I), which is essential for the activation of NF-κB.10 The cellular inhibitor of apoptosis (cIAP) proteins cIAP1 and cIAP2 mediate the formation of polyubiquitin chains on RIP1, which is responsible for the recruitment of the inhibitor of NF-κB (IκB) kinase complex.9, 11 In the basal state, the NF-κB heterodimer RelA/p50 associates with IκBα.8, 9 Upon a TNF-α stimulation, IκBα is phosphorylated at two serine residues by IκB kinase, leading to its ubiquitination.12 The proteasome-dependent degradation of IκB promotes the translocation of the RelA/p50 heterodimer into the nucleus, in which it induces the expression of NF-κB target genes.8, 9
Allantopyrone A (Figure 1a) was originally discovered from the endophytic fungus Allantophomopsis lycopodina KS-97, and was found to induce apoptosis and DNA fragmentation in human premyelocytic leukemia HL-60 cells.13 We recently reported that allantopyrone A protected rat pheochromocytoma PC12 cells from oxidative stress-induced cell death by activating the Keap1–Nrf2 pathway.14 In addition to these biological activities, we previously showed that allantopyrone A inhibited the TNF-α-induced NF-κB signaling pathway upstream of IκBα degradation in human lung carcinoma A549 cells.15 In the present study, the molecular mechanisms by which allantopyrone A inhibits the TNF-α-induced NF-κB pathway were investigated in more detail.
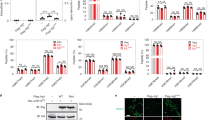
Allantopyrone A inhibited nuclear factor-κB (NF-κB) reporter activity induced by tumor necrosis factor-α (TNF-α). (a) Structure of allantopyrone A. (b) A549 cells were transfected with an NF-κB-responsive firefly luciferase reporter plasmid together with a cytomegalovirus (CMV) promoter-driven Renilla luciferase reporter plasmid for 24 h. A549 cells were pre-treated with or without allantopyrone A (10–100 μm) for 1 h and then stimulated with (+) or without (−) TNF-α (2.5 ng ml−1) for 2.5 h. Cell lysates were used to analyze luciferase reporter activity. Data are shown as the mean±s.e. of three independent experiments. **P<0.01 and ***P<0.001.
Materials and methods
Cell culture
Human lung carcinoma A549 cells (JCRB0076) were provided by the National Institutes of Biomedical Innovation, Health, and Nutrition JCRB Cell Bank (Osaka, Japan). A549 cells and human embryonic kidney (HEK) 293T cells were grown in RPMI 1640 medium (Thermo Fisher Scientific, Grand Island, NY, USA) and Dulbecco’s modified Eagle’s medium (Thermo Fisher Scientific), respectively, which were supplemented with heat-inactivated fetal calf serum (Nichirei Bioscience, Tokyo, Japan) and penicillin–streptomycin (Nacalai Tesque, Kyoto, Japan). All cell lines were cultured at 37 °C under 5% CO2.
Reagents
Allantopyrone A was prepared as described previously.13 Glutathione (reduced form) was commercially obtained from Wako Pure Chemical Industries (Osaka, Japan).
Expression vectors
The coding regions of human TNF-α and human TNF-R1 were amplified by PCR from cDNA libraries prepared from human acute monocytic leukemia THP-1 cells and human cervical adenocarcinoma HeLa cells, respectively. Human TNF-α (77–233) was inserted into pGEX-4T-1 to construct a fusion protein with glutathione S-transferase (GST). Human TNF-R1 was inserted into pCR3 expression vectors with a C-terminal FLAG tag. The coding region of human TRADD was amplified by PCR from the A549 cDNA library and was inserted into the pCR3 expression vector with an N-terminal FLAG tag.
Reporter assay
The luciferase reporter assay was performed as described previously.16
Pull-down assay
Human TNF-α (77–233) fused to GST (GST-TNF-α) in the pGEX-4T-1 expression vector was expressed in Escherichia coli. Bacterial cells were lyzed by 2% Triton X-100 in phosphate-buffered saline (PBS) and sonicated, followed by centrifugation (15 300 g, 10 min). Supernatants were treated with glutathione Sepharose 4B beads (GE Healthcare, Uppsala, Sweden). The beads were washed three times with PBS and eluted with 10 mm glutathione in 50 mm Tris-HCl (pH 8.0), followed by ultrafiltration. To analyze the formation of the TNF-R1 complex, A549 cells were stimulated with GST-TNF-α (1 μg ml−1) for 5 min. Cells were washed with PBS, lyzed with the DISC lysis buffer (0.5% Nonidet P-40, 150 mm NaCl, 10% glycerol, 20 mm Tris-HCl (pH 7.4), 2 mm sodium vanadate and Complete (Roche Diagnostics, Indianapolis, IN, USA)) and centrifuged (15 300 g, 5 min). In the TNF-α-binding assay, GST-TNF-α (1 μg ml−1) was directly added to cell lysates. Glutathione Sepharose 4B beads were used to pull down GST-TNF-α. The beads were washed three times with DISC lysis buffer and analyzed by western blotting.
Western blotting
The preparation of cell lysates and western blotting were performed as described previously.16, 17 Protein samples (30 μg) in sample buffer (62.5 mm Tris, 2% SDS, 10% glycerol, 0.003% bromophenol blue and 288 mm 2-mercaptoethanol) were boiled at 100 °C for 5 min, fractionated by SDS-polyacrylamide gel electrophoresis and transferred onto nitrocellulose membranes. The membranes were incubated with primary antibodies specific to γ1-actin (2F3; Wako Pure Chemical Industries), FLAG (1E6; Wako Pure Chemical Industries), IκBα (clone 25; BD Biosciences, San Jose, CA, USA), IκB kinase β (D30C6; Cell Signaling Technology, Danvers, MA, USA), RelA (C-20; Santa Cruz Biotechnology, Santa Cruz, CA, USA), RIP1 (38/RIP; BD Biosciences), TNF-R1 (H-5; Santa Cruz Biotechnology), TRADD (37/TRADD; BD Biosciences), TRAF2 (C-20; Santa Cruz Biotechnology) and horseradish peroxidase-linked secondary antibodies (Jackson ImmunoResearch Laboratories, West Grove, PA, USA). Blots were analyzed by ImageQuant LAS 4000 mini (GE Healthcare).
Transfection
Cells were dispersed in new culture media 1 day before transfection. A549 cells and HEK293T cells were transiently transfected with expression vectors by lipofection using HilyMax transfection reagent (Dojindo Laboratories, Kumamoto, Japan) and the calcium phosphate method, respectively. Culture media were changed before the treatment with allantopyrone A.
Statistical analysis
Statistical analyses were performed by a one-way analyis of variance followed by Tukey’s test.
Results
Allantopyrone A did not inhibit TNF-α binding to TNF-R1
A549 cells were transfected with luciferase reporter plasmids, pre-treated with allantopyrone A for 1 h and then stimulated with TNF-α. Consistent with our previous findings,15 allantopyrone A inhibited NF-κB luciferase reporter activity at concentrations >30 μm (Figure 1b). To clarify whether allantopyrone A prevents the binding of TNF-α to TNF-R1, GST-TNF-α was added directly to cell lysates prepared from allantopyrone A-treated A549 cells. Glutathione Sepharose 4B beads were used to pull down GST-TNF-α, which bound to TNF-R1. Allantopyrone A did not inhibit the binding of TNF-α to TNF-R1 at concentrations up to 50 μm (Figure 2). In contrast, allantopyrone A at 100 μm partially decreased the binding of TNF-α to TNF-R1 (Figure 2). At this concentration, allantopyrone A slightly decreased the amount of TNF-R1 and conversely increased high-MW bands reactive to the anti-TNF-R1 antibody (Figure 2).
Allantopyrone A did not inhibit tumor necrosis factor (TNF)-α binding to TNF receptor 1 (TNF-R1). A549 cells were pre-treated with or without allantopyrone A (10–100 μm) for 1 h. GST-TNF-α (1 μg) was directly added to cell lysates and then isolated by glutathione Sepharose 4B beads. The asterisk indicates the high-MW bands of TNF-R1. Pull-down samples and cell lysates were analyzed by western blotting. Blots are representative of three independent experiments. The amount of TNF-R1 in pull-down samples and the amount of TNF-R1 relative to actin in cell lysates are shown as the mean±s.e. of three independent experiments. *P<0.05. GST, glutathione S-transferase.
Allantopyrone A inhibited RIP1 modifications in the TNF-R1 complex
TNF-α rapidly induces the formation of the TNF-R1 complex (complex I) at the plasma membrane.10 Complex I contains the adaptor proteins, TRADD, TRAF2 and RIP1.10 RIP1 is known to be extensively modified by K63-linked polyubiquitination in the TNF-R1 complex.18 A549 cells were pre-treated with allantopyrone A for 1 h and stimulated with GST-TNF-α for 5 min. Consistent with previous findings,10, 18 RIP1 underwent extensive modifications when A549 cells were stimulated with TNF-α (Figure 3). TNF-α-induced RIP1 modifications were decreased by allantopyrone A at concentrations >30 μm (Figure 3). In addition, allantopyrone A partially decreased the association of TRADD, TRAF2 and RIP1 under conditions in which TNF-α bound to TNF-R1 (Figure 3). These results indicate that allantopyrone A inhibits RIP1 modifications in the TNF-R1 complex.
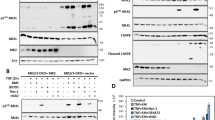
Allantopyrone A inhibited receptor-interacting protein 1 (RIP1) modifications in the tumor necrosis factor receptor 1 (TNF-R1) complex. A549 cells were pre-treated with or without allantopyrone A (10–100 μm) for 1 h and then stimulated with (+) or without (−) GST-TNF-α (1 μg ml−1) for 5 min. GST-TNF-α (0.1 μg) was added directly to the GST-TNF-α (−) cell lysate. The TNF-R1 complex was isolated by glutathione Sepharose 4B beads. Western blotting was used to analyze pull-down samples and cell lysates. The arrowhead indicates nonspecific bands. The asterisk indicates the high-MW bands of RIP1. GST, glutathione S-transferase.
Allantopyrone A augmented crosslinked bands of TNF-R1, TRAF2 and RIP1
We investigated the effects of allantopyrone A on components of the TNF-R1 complex in A549 cells. The longer exposure of western blots revealed that allantopyrone A augmented the high-MW bands of TNF-R1, TRAF2 and RIP1, but had a negligible effect on TRADD (Figure 4a). As these bands of TNF-R1 and TRAF2 were observed at sizes that were approximately twofold and threefold the MW of each monomer, respectively, under reducing conditions on SDS-polyacrylamide gel electrophoresis, they may have included homodimers or homotrimers, which were crosslinked by covalent bonds other than disulfide bonds. TRAF2 appeared to be crosslinked more preferentially by allantopyrone A than RIP1.
Allantopyrone A augmented crosslinked bands of tumor necrosis factor receptor 1 (TNF-R1), TNF receptor-associated factor 2 (TRAF2) and receptor-interacting protein 1 (RIP1). (a) A549 cells were treated with or without allantopyrone A (10–100 μm) for 1 h. Cell lysates were analyzed by western blotting. The asterisks indicate the high-MW bands of TNF-R1, TRAF2 and RIP1. (b) Structure of human TNF-R1. CRD, cysteine-rich domain; DD, death domain; SP, signal peptide; TM, transmembrane domain. The number of amino acids and number of cysteines are indicated. (c) A549 cells were transfected with (+) or without (−) an expression vector encoding TNF-R1 fused to C-terminal FLAG, and treated with or without allantopyrone A (10–100 μm) for 1 h. Western blotting was used to analyze cell lysates. The asterisk indicates the high-MW bands of TNF-R1. (d) A549 cells were transfected with (+) or without (−) an expression vector encoding TRADD fused to N-terminal FLAG, and treated with or without allantopyrone A (10–100 μm) for 1 h. Western blotting was used to analyze cell lysates. The asterisks indicate the high-MW bands of TRADD.
TNF-R1 possesses 24 cysteines in extracellular CRDs and 5 cysteines in the intracellular portion (Figure 4b). The 24 extracellular cysteine residues are bridged by disulfide bonds in a single TNF-R1 receptor.19 The anti-TNF-R1 monoclonal antibody (clone H5) was raised against the extracellular domain (amino acids 30–301) of human TNF-R1. To investigate whether allantopyrone A affects antibody reactivity, TNF-R1 fused to a C-terminal FLAG tag was expressed in A549 cells (Figure 4c). In contrast to endogenous TNF-R1 migrating at slightly lower than 59 kDa as the main band (Figure 4a), transfected TNF-R1 was expressed as two bands migrating at ~195 and 59 kDa in A549 cells (Figure 4c). The larger bands of transfected TNF-R1, which might correspond to homotrimers, were clearly increased by allantopyrone A at concentrations >20 μm (Figure 4c). These results indicate that allantopyrone A augmented the crosslinked bands of TNF-R1 and conversely reduced antibody reactivity.
The anti-TRADD monoclonal antibody (37/TRADD) was raised against human TRADD (163–312), which contains two cysteine residues. We also investigated whether the anti-TRADD antibody efficiently recognized the high-MW bands of TRADD. A549 cells were transiently transfected with an expression vector encoding TRADD fused to the N-terminal FLAG tag. In allantopyrone A-treated A549 cells, the high-MW bands of TRADD were detected by both the anti-FLAG antibody and the anti-TRADD antibody (Figure 4d). These results suggest that the reactivity of the anti-TRADD antibody still remains when TRADD is crosslinked by allantopyrone A, and that transfected TRADD is crosslinked by allantopyrone A more preferentially than endogenous TRADD. Further experiments are necessary to address the different effects of allantopyrone A on endogenous TRADD and transfected TRADD.
Allantopyrone A augmented crosslinked bands of TNF-R1 CRDs
HEK293T cells have the ability to overexpress transiently transfected genes encoded in the pCR3 expression vector. To establish whether allantopyrone A promotes the crosslinking of TNF-R1 on an extracellular portion or intracellular portion, TNF-R1 (1–232), TNF-R1 (233–455) and TNF-R1 (1–455) (Figure 5a) were overexpressed in HEK293T cells, followed by a 1 h treatment with allantopyrone A. Similar to A549 cells (Figure 4c), TNF-R1 (1–455) was expressed as two bands migrating at ~197 and 60 kDa in HEK293T cells (Figure 5b). The larger bands of TNF-R1 (1–455) were markedly increased by allantopyrone A at concentrations >10 μm (Figure 5b). In agreement with Figure 4c, the larger bands were less reactive to the anti-TNF-R1 antibody than to the anti-FLAG antibody. While TNF-R1 (1–232) was mainly expressed as a monomer, the high-MW bands of TNF-R1 (1–232) were strongly augmented by allantopyrone A at concentrations >10 μm (Figure 5c). In contrast to TNF-R1 (1–232), TNF-R1 (233–455) was expressed as two major bands, which may have corresponded to a monomer and dimer (Figure 5d). Allantopyrone A did not increase the larger major band of TNF-R1 (233–455) at concentrations up to 10 μm (Figure 5d). These results indicate that allantopyrone A promotes the crosslinking of TNF-R1 extracellular CRDs more preferentially than its intracellular portion.
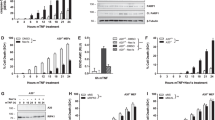
Allantopyrone A augmented crosslinked bands of the extracellular domain of tumor necrosis factor receptor 1 (TNF-R1). (a) Structures of TNF-R1 (1–455), TNF-R1 (1–232) and TNF-R1 (233–455). CRD, cysteine-rich domain; DD, death domain; SP, signal peptide; TM, transmembrane domain. The arrows indicate cleavage sites by signal peptidase. (b–d) Human embryonic kidney 293T cells were transfected with (+) or without (−) TNF-R1 (1–455) (b), TNF-R1 (1–232) (c) and TNF-R1 (233–455) (d), and then treated with or without allantopyrone A (0.1–30 μm) for 1 h. The amount of TNF-R1 was analyzed by western blotting. The asterisks indicate the high-MW bands of TNF-R1.
Glutathione suppressed the augmentation of crosslinked bands of TNF-R1 and TRAF2, and the inhibition of IκBα degradation in allantopyrone A-treated cells
Allantopyrone A has two α,β-unsaturated carbonyl moieties, which are highly reactive to cysteinyl thiol groups. As shown in Figure 5b, allantopyrone A augmented the high-MW bands of TNF-R1 (1–455) in HEK293T cells (Figure 6a). Glutathione suppressed this augmentation of the high-MW bands of TNF-R1 (1–455; Figure 6a). These results appear to support allantopyrone A binding to the cysteine residues of proteins and, thus, promoting crosslinking.
Glutathione suppressed the biological activities of allantopyrone A. (a) Human embryonic kidney 293T cells were transfected with or without an expression vector encoding tumor necrosis factor receptor 1 (TNF-R1; 1–455), pre-treated with (+) or without (−) glutathione (5 mm) for 1 h, then treated with (+) or without (−) allantopyrone A (30 μm) for 1 h. Western blotting was used to analyze cell lysates. The asterisk indicates the high-MW bands of TNF-R1. (b) A549 cells were pre-treated with (+) or without (−) glutathione (5 mm) for 1 h, treated with (+) or without (−) allantopyrone A (50 μm) for 1 h and then stimulated with (+) or without (−) TNF-α (2.5 ng ml−1) for 15 min. Western blotting was used to analyze cell lysates. The asterisks indicate the high-MW bands of inhibitor of nuclear factor-κBα (IκBα), TNF-R1 and TNF receptor-associated factor 2 (TRAF2).
We further investigated the effects of glutathione on the inhibitory activity of allantopyrone A against the TNF-α-induced NF-κB signaling pathway. Allantopyrone A was previously shown to inhibit TNF-α-induced IκBα degradation at concentrations of more than 32 μm.15 Consistent with this finding, allantopyrone A at 50 μm inhibited TNF-α-induced IκBα degradation (Figure 6b). Glutathione reversed the inhibition of TNF-α-induced IκBα degradation by allantopyrone A (Figure 6b). The high-MW bands of TNF-R1 and TRAF2 promoted by allantopyrone A were also diminished by glutathione (Figure 6b). These results demonstrate that crosslinking by allantopyrone A is responsible for the inhibition of the NF-κB signaling pathway.
Allantopyrone A augmented crosslinked bands of IκB kinase β and RelA
A large number of compounds possessing α,β-unsaturated carbonyl moieties have been shown to inhibit RelA and IκB kinase β by targeting their cysteine residues.20 We investigated whether allantopyrone A crosslinked RelA and IκB kinase β in A549 cells. The high-MW bands of RelA and IκB kinase β were increased in allantopyrone A-treated A549 cells (Figure 7).
Discussion
We previously reported that allantopyrone A inhibited TNF-α-induced IκBα phosphorylation, but did not directly block IκB kinase β activity.15 In the present study, we showed that allantopyrone A did not affect TNF-α binding to TNF-R1, but inhibited RIP1 modifications in the TNF-R1 complex in A549 cells. Moreover, allantopyrone A exhibited the ability to crosslink TNF-R1, TRAF2, RIP1, RelA and IκB kinase β. The present results revealed that allantopyrone A interferes with multiple components of the TNF-R1 complex and blocks RIP1 modifications in the TNF-α-induced NF-κB signaling pathway.
Upon a TNF-α stimulation, the TNF-R1 complex consisting of TRADD, TRAF2 and RIP1 (complex I) is initially formed at the plasma membrane, which mediates NF-κB activation.10 TNF-α is known to induce RIP1 polyubiquitination.9, 11 In Jurkat cells stimulated with GST-TNF-α, the high-MW bands of RIP1 in the TNF-R1 complex isolated by glutathione Sepharose beads were shown to be polyubiquitinated.18 The pull-down assay using GST-TNF-α showed that the high-MW bands of RIP1 were augmented in the TNF-R1 complex upon TNF-α stimulation in A549 cells. Thus, the extensively modified RIP1 observed in A549 cells are most likely to be polyubiquitinated. Although allantopyrone A did not inhibit the binding of TNF-α to TNF-R1 at concentrations up to 50 μm, it inhibited RIP1 modifications at concentrations >30 μm. The association of adaptor proteins to TNF-R1 was partially affected by allantopyrone A. These results indicate that allantopyrone A mainly targets components of the membrane-bound TNF-R1 complex. We assume that crosslinked TNF-R1, TRAF2 and RIP1 by allantopyrone A are unable to undergo the proper conformational changes required for recruitment to the TNF-R1 complex and/or RIP1 modifications.
Many compounds possessing an α,β-unsaturated carbonyl moiety inhibit the canonical NF-κB signaling pathway by targeting cysteine 179 of IκB kinase β or cysteine 38 of RelA.20 As a very unique property, allantopyrone A possesses two α,β-unsaturated carbonyl moieties in a single molecule, which may be responsible for the crosslinking of two cysteine residues. In addition to RelA and IκB kinase β, allantopyrone A augmented the high-MW bands of TNF-R1, TRAF2 and RIP1, suggesting that it binds to these proteins and crosslinks them at higher concentrations, possibly forming dimers, trimers or even oligomers. Moreover, comparisons between western blots using the anti-TNF-R1 antibody raised against its extracellular portion and the anti-FLAG antibody showed that allantopyrone A reduced the reactivity of the anti-TNF-R1 monoclonal antibody used in the present study, possibly due to the covalent modification of one or multiple cysteine residues scattered in the CRDs.
Allantopyrone A strongly augmented the crosslinked bands of full-length TNF-R1 and extracellular TNF-R1 in HEK293T cells, which exhibited greater sensitivity to allantopyrone A than A549 cells. The extracellular portions of TNF receptor family members are highly conserved and consist of CRDs.4, 5 TNF-R1 possesses four CRDs (CRD1–CRD4), in which six cysteine residues form three-disulfide bonds.2 While CRD2 and CRD3 form the TNF-α-binding surface, CRD1 acts as a pre-ligand-binding assembly domain, which enables TNF-R1 to form a homotypic trimer.21, 22 A small amount of endogenous TNF-R1 was crosslinked as dimers by allantopyrone A, while allantopyrone A promoted the formation of the crosslinked trimers of transfected TNF-R1. Thus, the pre-ligand-binding assembly domain-dependent TNF-R1 association may explain the efficient crosslinking of TNF-R1 by allantopyrone A as trimers and dimers. However, it is currently unclear the mechanism by which allantopyrone A promotes the formation of different crosslinked products between endogenous TNF-R1 and transfected TNF-R1. The intracellular portion of TNF-R1 contains a DD, which is critical for the activation of NF-κB or induction of apoptosis.5 The DD of TNF-R1 undergoes self-association.23 However, the intracellular portion of TNF-R1 was not crosslinked by allantopyrone A as efficiently as extracellular TNF-R1.
In addition to TNF-R1, TRAF2 and RIP1 were crosslinked by allantopyrone A in A549 cells. In the presence of allantopyrone A, TRAF2 may be largely crosslinked to become homotrimers, because TRAF2 forms a trimeric coiled-coil structure.24 RIP1 may also be crosslinked to become higher-MW products by allantopyrone A because RIP1 has the ability to interact by itself or with other proteins via a DD and RIP homotypic interaction motif.25 We previously reported that allantopyrone directly bound to Keap1 and this was accompanied by an increase in the high-MW band reactive to an anti-Keap1 antibody.14 These findings suggest that allantopyrone A targets multiple cellular proteins. Further studies are needed to address the selectivity of allantopyrone A to binding proteins.
In conclusion, allantopyrone A was shown to block RIP1 modifications in the TNF-R1 complex during the TNF-α-induced NF-κB signaling pathway. Many compounds possessing a single α,β-unsaturated carbonyl moiety have been reported to inhibit the NF-κB signaling pathway by blocking IκB kinase β and RelA.20 In addition to the TNF-R1 complex components, allantopyrone A augmented the crosslinked bands of RelA and IκB kinase β. These results indicate that allantopyrone A is not selective to the components of the TNF-R1 complex. Nevertheless, allantopyrone A differs from many known compounds, particularly because it possesses two α,β-unsaturated carbonyl moieties in a molecule. We propose that two α,β-unsaturated carbonyl moieties as the unique structure of allantopyrone A promote the crosslinking of adjacent proteins in close proximity and, unlike other single α,β-unsaturated carbonyl moiety-containing compounds, interfere with multiple components of the TNF-R1 complex at the initial step in the TNF-α-induced NF-κB signaling pathway.
Change history
17 December 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41429-021-00479-2
References
Aggarwal, B. B. Signalling pathways of the TNF superfamily: a double-edged sword. Nat. Rev. Immunol. 3, 745–756 (2003).
Fischer, R., Kontermann, R. E. & Maier, O. Targeting sTNF/TNFR1 signaling as a new therapeutic strategy. Antibodies 4, 48–70 (2015).
Croft, M., Benedict, C. A. & Ware, C. F. Clinical targeting of the TNF and TNFR superfamilies. Nat. Rev. Drug Discov. 12, 147–168 (2013).
Locksley, R. M., Killeen, N. & Lenardo, M. J. The TNF and TNF receptor superfamilies: integrating mammalian biology. Cell 104, 487–501 (2001).
Park, H. H. et al. The death domain superfamily in intracellular signaling of apoptosis and inflammation. Annu. Rev. Immunol. 25, 561–586 (2007).
Chan, F. K. M. et al. A domain in TNF receptors that mediates ligand-independent receptor assembly and signaling. Science 288, 2351–2354 (2000).
Siegel, R. M. et al. Fas preassociation required for apoptosis signaling and dominant inhibition by pathogenic mutations. Science 288, 2354–2357 (2000).
Wajant, H. & Scheurich, P. TNFR1-induced activation of the classical NF-κB pathway. FEBS J. 278, 862–876 (2011).
Hayden, M. S. & Ghosh, S. Regulation of NF-κB by TNF family cytokines. Semin. Immunol. 26, 253–266 (2014).
Micheau, O. & Tschopp, J. Induction of TNF receptor I-mediated apoptosis via two sequential signaling complexes. Cell 114, 181–190 (2003).
Sasaki, K. & Iwai, K. Roles of linear ubiquitinylation, a crucial regulator of NF-κB and cell death, in the immune system. Immunol. Rev. 266, 175–189 (2015).
Perkins, N. D. Post-translational modifications regulating the activity and function of the nuclear factor kappaB pathway. Oncogene 25, 6717–6730 (2006).
Shiono, Y. et al. Allantopyrone A, a new α-pyrone metabolite with potent cytotoxicity from an endophytic fungus, Allantophomopsis lycopodina KS-97. J. Antibiot. 63, 251–253 (2010).
Uesugi, S. et al. Allantopyrone A activates Keap1-Nrf2 pathway and protects PC12 cells from oxidative stress-induced cell death. J. Antibiot. 70, 429–434 (2017).
Yokoigawa, J. et al. Allantopyrone A, an α-pyrone metabolite from an endophytic fungus, inhibits the tumor necrosis factor α-induced nuclear factor κB signaling pathway. J. Antibiot. 68, 71–75 (2015).
Matsuda, I. et al. The C-terminal domain of the long form of cellular FLICE-inhibitory protein (c-FLIPL inhibits the interaction of the caspase 8 prodomain with the receptor-interacting protein (RIP1) death domain and regulates caspase 8-dependent nuclear factor κB (NF-κB) activation. J. Biol. Chem. 289, 3876–3887 (2014).
Hirano, S. & Kataoka, T. Deoxynivalenol induces ectodomain shedding of TNF receptor 1 and thereby inhibits the TNF-α-induced NF-κB signaling pathway. Eur. J. Pharmacol. 701, 144–151 (2013).
Ea, C. K., Deng, L., Xia, Z. P., Pineda, G. & Chen, Z. J. Activation of IKK by TNFα requires site-specific ubiquitination of RIP1 and polyubiquitin binding by NEMO. Mol. Cell 22, 245–257 (2006).
Banner, D. W. et al. Crystal structure of the soluble human 55 kd TNFβ receptor-human TNFβ complex: implication for TNF receptor activation. Cell 73, 431–445 (1996).
Kataoka, T. Chemical biology of inflammatory cytokine signaling. J. Antibiot. 62, 655–667 (2009).
Chan, F. K. M. Three is better than one: pre-ligand receptor assembly in the regulation of TNF receptor signaling. Cytokine 37, 101–107 (2007).
Wu, H. & Siegel, R. M. Progranulin resolves inflammation. Science 332, 427–428 (2011).
Boldin, M. P. et al. Self-association of the ‘death domain’ of the p55 tumor necrosis factor (TNF) receptor and Fas/APO1 prompts signaling for TNF and Fas/APO1 effects. J. Biol. Chem. 270, 387–391 (1995).
Zheng, C., Kabaleeswaran, V., Wang, Y., Cheng, G. & Wu, H. Crystal structures of the TRAF2: cIAP2 and the TRAF1: TRAF2: cIAP2 complexes: affinity, specificity, and regulation. Mol. Cell 38, 101–113 (2010).
Wu, X. N. et al. Distinct roles of RIP1-RIP3 hetero- and RIP3-RIP3 homo-interaction in mediating necroptosis. Cell Death Differ. 21, 1709–1720 (2014).
Acknowledgements
This work was partly supported by JSPS KAKENHI Grant numbers 25292061 (to TK) and 16H04910 (to TK and KK).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
The original online version of this article was revised: In the section “Western blotting” in “Materials and Methods”, sentence 3, “β-actin” has been replaced with “γ1 -actin”. All other mentions of “β-actin” have been replaced with “actin”. Figures 2, 3, 4, 5, 6 and 7 have also been replaced to change “β-actin” to “actin”.
Rights and permissions
About this article
Cite this article
Quach, H., Tanigaki, R., Yokoigawa, J. et al. Allantopyrone A interferes with multiple components of the TNF receptor 1 complex and blocks RIP1 modifications in the TNF-α-induced signaling pathway. J Antibiot 70, 929–936 (2017). https://doi.org/10.1038/ja.2017.74
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2017.74