Abstract
Myxococcus fulvus HW-1, a salt-tolerant bacterial strain, which was isolated from a coastal environment, changes its behavior with different salinities. To study the relationship between behavioral shifts and the adaption to oceanic conditions, the HW-1 strain was randomly mutagenized using transposon insertion, producing a dispersed-growing mutant, designated YLH0401. The mutant did not develop fruiting bodies and myxospores, was deficient in S-motility, produced less extracellular matrix and was less salt tolerant. The YLH0401 strain was determined to be mutated by a single insertion in a large gene of unknown function (7011 bp in size), which is located in a horizontally transferred DNA fragment. The gene is expressed during the vegetative growth stage, as well as highly and stably expressed during the development stage. This horizontally transferred gene may allow Myxococcus to adapt to oceanic conditions.
Similar content being viewed by others
Introduction
Myxobacteria exhibit remarkable complex social behaviors that are essential for their survival in natural environments (Shimkets, 1990). Myxobacterial cells crawl on solid surfaces in swarms and feed cooperatively on macromolecules and on other microbial cells. When food is exhausted, hundreds of thousands of myxobacterial cells accumulate on solid surfaces to form multicellular fruiting bodies, inside which myxospores that are resistant to adverse conditions develop. Using specific identification techniques on the basis of their social characteristics, such as the ability to form fruiting bodies and characteristic swarms, myxobacterial strains have been identified in various terrestrial habits (Reichenbach, 1999; Dawid, 2000). These cultured soil myxobacterial strains are normally unable to grow in a high-salt environment (Reichenbach, 1999; Li et al., 2002). Furthermore, when placed in liquid, soil myxobacterial cells typically grow in clumps or by adhering to the container walls. Continuously subculturing the soil myxobacterium Myxococcus xanthus in liquid causes cells to lose social behavior (Velicer et al., 1998). However, some halophilic and halotolerant myxobacterial strains have been isolated from marine environments (Iizuka et al., 1998, 2003a, 2003b; Fudou et al., 2002; Li et al., 2002). On the basis of phylogenetic analyses and living patterns, the halophilic myxobacteria are considered to be indigenous to the ocean, whereas the halotolerant myxobacteria are probably soil myxobacterial strains that have adapted to oceanic conditions (Zhang et al., 2005). There is much interest in understanding how salt-tolerant myxobacteria and their social behaviors have adapted to oceanic conditions.
Myxococcus fulvus HW-1 is a typical salt-tolerant marine strain, which was isolated from a coastal sample (Li et al., 2002). When cultured in a low concentration of saltwater, the strain exhibits a culture phenotype similar to soil myxobacterial strains, such as developing fruiting bodies and myxospores in response to starvation on solid surfaces. However, when the saltwater concentration is increased, the strain changes its living pattern by forming myxospores directly from vegetative cells (Zhang et al., 2005). These characteristics make HW-1 an ideal model for studying the adaptation mechanisms of salt-tolerant myxobacteria to marine environments.
Our previous studies have shown that salt-tolerant Myxococcus strains enhance their S-motility in high seawater concentrations, suggesting an adaptation to oceanic conditions (Wang et al., 2007). Further mechanistic investigations have demonstrated that the salt tolerance of these strains is often closely associated with the social behavior of Myxococcus. For example, mutation of the S-motility-related gene locus mts (homolog of which in the model strain M. xanthus DK1622 is located at MXAN_1333–1337) led to a simultaneous decrease in salt tolerance (Zhang et al., 2007). Two seawater-regulated genes—the two component system gene Tc105 (MXAN_4042 in DK1622) and outer-membrane protein gene Om031 (MXAN_3016 in DK1622)—are both involved in social behavior and salt tolerance (Pan et al., 2009). In this study, we report an association between the salt-tolerance adaptation and social behavior in a new mutant of M. fulvus HW-1. A single insertion mutation, which was introduced by random transposition of the pMiniHimar1-lacZ plasmid, enabled HW-1 to grow completely dispersed in liquid CTT medium. Phenotypic characteristics of the mutant showed that, concomitant with the loss of salt tolerance, its social behavior was also completely lost. Interestingly, the transposition insertion site was located in a horizontally transferred gene that encodes a hypothetical protein containing 2336 amino acids. The gene was expressed during both the vegetative growth and development stages. The gene was also identified in some salt-tolerant Myxococcus strains but not in the type strains of different Myxococcus species that were isolated from soil samples. We speculate that this horizontally transferred gene confers some advantages for the adaption of Myxococcus to oceanic conditions.
Materials and methods
Transposon mutagenesis
The plasmid pMiniHimar1-lacZ (H. B. Kaplan, University of Texas) was transformed into the wild-type M. fulvus strain (HW-1), as previously described (Kashefi and Hartzell, 1995), with minor modifications. The M. fulvus cells were grown to the early exponential phase in CTT medium. The cells were then collected by centrifugation at 8000 × g for 5 min, and washed three times in sterile deionized water. The cell pellet was re-suspended in sterile deionized water, and cooled on ice. A 40-μl aliquot (1 × 1010 cells per ml) was mixed with 50−100 ng DNA and transferred to a 0.2-cm electroporation cuvette (Bio-Rad, Hercules, CA, USA). The electroporation was performed with a Bio-Rad Gene Pulser at 1.25 kV, 400 Ω and 25 μF. After the electroporation, cells were immediately transferred to 2.5 ml CTT and incubated 4−6 h at 30 °C. After centrifugation, the cells were mixed with 2.5 ml CTT medium containing 0.3% agar and poured onto CTT plates containing 40 μg ml1 Km1. After incubation at 30 °C for 6−7 days, colonies that grew dispersed in liquid CTT broth were chosen for further study.
Determination of the insertion site
The genomic DNA of YLH0401 was extracted using the cetyltrimethyl ammonium bromide method described by Ward et al. (1998) and digested with SacII. After purification by alcohol precipitation, fragments were self-circularized using T4 DNA ligase. The ligation product was transformed into the Escherichia coli DH5α λpir strain, and the kanamycin-resistant (Kmr) transformants were selected. Plasmid DNA was extracted from the resistant transformants using the eZNA Plasmid Mini Kit I (Omega Bio-Tek, Norcross, GA, USA) according to the manufacturer's instructions and subcloned into the plasmid pGEM-7Zf (+) (Promega, Madison, WI, USA) for sequencing. Besides SacII, other restriction enzymes, such as BamHI, SphI, ScaI, ApaI and so on, were also used to digest the genomic DNA of YLH0401 to determine the insertion site as described above.
Isolation of the segments flanking the inserted locus
Thermal asymmetric interlaced PCR (Liu and Whittier, 1995) was performed to amplify the segments flanking the insertion site of the transposon. The nested specific primers and arbitrary degenerate primers used are listed in Supplementary Table S2. The 20-μl PCR mixture consisted of 0.25 mM of dNTPs, 0.3 μM of the specific primer, 6 μM of the arbitrary degenerate primer, and 1 U of Ex Taq DNA polymerase (TaKaRa Biotechnol, Dalian, China) in 1 × GC buffer. Chromosomal DNA (300 ng) was added to the PCR mixture for the primary reaction. After first-round PCR, the reaction mixture was diluted 100 × with sterile water, and 2 μl of the dilution was used as the template for the second PCR reaction, which was further diluted for the third PCR reaction. The thermal cycling conditions followed the protocol described previously (Liu and Whittier, 1995).
Social behavior analysis of YLH0401
The social behavior of YLH0401 was analyzed as previously described (Shi and Zusman, 1993; Kearns et al., 2000), with minor modifications. To assay cell swarming capacity, aliquots (2 μl, 5 × 109 cells per ml) were inoculated onto CTT medium containing 1.5 or 0.3% agar. After 5 days of incubation at 30 °C, the size of the swarming colonies was measured, and the edges of colonies were observed under a dissection microscope (Nikon-SMZ1500, Tokyo, Japan). To assay cell development, cells were grown to the mid-logarithmic growth phase, collected, and resuspended in TPM buffer at a density of 5 × 109 cells per ml. Aliquots of 10 μl were spotted onto TPM agar plates and incubated at 30 °C for 5 days. Cell development was examined under a dissection microscope. Five day-old TPM cultures were used for the sporulation assay as follows: cells were collected, resuspended in 100 μl TPM buffer and then slightly sonicated to a homogenized state. After 2-h incubation at 50 °C, the cells were serially diluted, mixed with CTT containing 0.3% agar and poured onto CTT plates. The number of colonies was counted after 5−7 days of incubation at 30 °C.
Agglutination assay
Cell agglutination was assayed using the method as described by Pan et al. (Pan et al., 2009).
Examination of extracellular polysaccharide production
To examine the extracellular polysaccharide production in YLH0401 and HW-1 cells, the fluorescent dye Calcofluor white M2R (Dana and Shimkets, 1993; Black et al., 2006) and liquid colorimetry with Congo red and Trypan blue (Black and Yang, 2004) were used.
RT–PCR
Total RNA was isolated using the Promega SV Total RNA Purification Kit as described by Pan et al. (2009). The isolated RNA was treated with the DNA Free Kit (Ambion, Austin, TX, USA) to remove residual genomic DNA. The first strand of DNA was synthesized from the RNA template using the primer 5′-GACAAGCCGCTACTGGAAGTGGATG-3′. Then the PCR fragment was amplified with the primer pair 5′-GACAAGCCGCTACTGGAAGTGGATG-3′ and 5′-CGAGCGACTTCTTGTGCCCCTGGAG-3′.
β-Galactosidase assays
The β-galactosidase activity was measured using the method described by Müller and Jakobsen (2008) with small modifications. The YLH0401 and HW-1 cells were scraped from TPM plates every 12 h. The scraped cells were suspended in TPM buffer. After centrifugation at 10 000 × g for 5 min at room temperature, the supernatant was discarded and the cell pellets were resuspended to a density of 5 × 109 cells per ml. The cells were then disrupted in a homogenization machine by beads, followed by centrifugation to remove cell debris. A total of 100 μl of the extract was mixed with 400 μl Z buffer containing 1 mg ml1 of o-nitrophenyl-β-D-galactoside as the substrate. The mixtures were incubated at 37 °C until the solution turned sufficiently yellow. The reaction was stopped by the addition of 0.5 ml of 1 M Na2CO3. The absorption of the solution was read at 420 nm. The protein concentration was determined using the Easy Quant Protein Quantitative Kit (TransGene Biotech, Beijing, China). β-Galactosidase activity was calculated as follows:

Amplification of hdsp from different Myxococcus strains
Five salt-tolerant Myxococcus strains (listed in Supplementary Table S1) isolated from oceanic conditions (Wang et al., 2007), as well as three typical soil strains, were selected for amplification of the hdsp gene. The primers used for the amplification are listed in Supplementary Table S2.
Sequences
The predicted horizontally transferred sequences were deposited in GenBank (accession number GU190823).
Results
Random transposon mutagenesis of HW-1 for dispersed growth in liquid culture
We hypothesized that the ability to grow in clumps is one of the key traits enabling salt-tolerant myxobacteria to live in the ocean. Thus, we wanted to mutate the salt-tolerant Myxococcus fulvus HW-1 strain to produce mutants that grow dispersed in liquid to study other associated phenotypic changes. The HW-1 strain was mutated by electroporation of the random-insertion plasmid pMiniHimar1-lacZ. After mutation, colonies that grew on kanamycin selection plates were individually transferred into liquid CTT medium to screen for dispersed growth. From more than 300 transformants, a single strain, designated YLH0401, was able to grow in a completely dispersed state in liquid CTT broth (Figure 1a).
Comparison of the phenotypic characteristics of the mutant YLH0401 and the wild type strain HW-1. (a) The growth form in liquid CTT for 48 h. (b and c) The swarming ability on solid CTT containing 1.5 and 0.3% agar for 120 h. (d) Fruiting body formation on TPM for 120 h. The bar is equal to 0.5 cm for (b and c) and 1.0 mm for (d).
Phenotypic characteristics of the mutant YLH0401
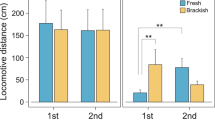
The YLH0401 mutant formed smaller colonies than wild-type HW-1 cells on CTT plates containing either 1.5% (hard) or 0.3% (soft) agar (Figures 1b and c). At the edge of colonies on hard agar, HW-1 cells showed a bigger swarming size than YLH0401 cells (Figure 2b). On TPM plates, the wild-type strain HW-1 formed visible fruiting bodies after 48-h incubation, and the sporulation capacity was approximately 6 × 105 per colony in the 5-day culture. However, under the same conditions, the mutant failed to develop fruiting bodies, and no spores were generated (sporulation was measured morphologically and by counting the colony forming unit number after heat treatment at 50 °C for 2 h). Figure 1d demonstrates the morphological characteristics of YLH0401 and HW-1 on TPM plates. Concurrently, the salt tolerance of YLH0401 also decreased; although HW-1 cells were able to grow in 130% HVY or 3% NaCl (Zhang et al., 2005), mutant cells were able to grow only in 30% seawater (30% HVY, VY-2 (Reichenbach and Dworkin, 1992) prepared in 30% seawater) or 1% NaCl. Therefore, the insertion mutation in YLH0401 produced a series of phenotypic changes related to both salt tolerance and social behavior.
To measure the production of extracellular matrix by the mutant, we performed binding assays with the dyes Congo red and Trypan blue, which are able to bind to extracellular polysaccharides (Dana and Shimkets, 1993; Yang et al., 2000). The ability of the mutant cells to bind to Congo red and Trypan blue was approximately 66.7 and 16.8% lower, respectively, than those observed in the wild-type strain. Decreased production of extracellular polysaccharide by the mutant was also visualized using the Calcofluor white binding assay on CTT plates containing 0.3% agar (Figure 2a). The clumping assay for intercellular cohesion and agglutination ability indicated that the mutant exhibited less cohesion than the wild-type cells (Figure 2c).
The mutagenized gene in YLH0401
A 3.0-kb segment flanking the insertion site was obtained using Thermal asymmetric interlaced PCR and self-circularized plasmid DNA containing the inserted transposon and flanking sequences, as described in the Materials and Methods section. In the plasmid-rescue experiment, several restriction enzymes were used to digest the genome of the mutant, and all transformants were found to be at the same site by sequencing. For further confirmation, a Southern blot experiment was conducted using a fragment of the Km-resistance gene from the plasmid as the probe (Supplementary Figure S1). The results indicated that there was only one insertion site of the transposon plasmid in the genome of the YLH0401 mutant (in addition, RT–PCR results also indicated no expression of the fragment in the mutant, see Figure 3b below). Therefore, the striking phenotypic changes in the mutant strain were caused by the insertional inactivation of a single gene. Bioinformatic analysis showed that the sequence had no homologs in the genome of Myxococcus xanthus DK1622. On the basis of our incomplete HW-1 genome (there are seven gaps, but cover 187 times the length of the genome, unreported), this 3.0-kb sequence is a part of a predicted open reading frame (ORF) that is 7011 bp in length. We have designated this ORF hdsp (HW-1 dispersed growth).
Expression of the hdsp gene in YLH0401 and HW-1. (a) Expression analysis of the lacZ protein in YLH0401 and HW-1 during development. The lacZ gene was transcriptionally fused under the hdsp promoter in YLH0401. (b) RT–PCR analysis of the expressions of hdsp during the vegetative growth stage of HW-1 and YLH0401. The PCR template in each lane: 1, negative; 2, genomic DNA from HW-1; 3, total RNA from HW-1; 4, complimentary DNA (cDNA) from HW-1; 5, total RNA from YLH0401; 6, cDNA from YLH0401. M: DL2000.
Flanking the hdsp ORF are seven predicted ORFs that also have no homologs in the genome of Myxococcus xanthus DK1622. It is interesting to note that six of these eight ORFs are in a typical horizontally transferred segment approximately 12.7 kb in length (the fragment locates in the largest contig that contains 6.9 million bases). Excluding hdsp, the five other ORFs in this region are all predicted to encode transposase (Table 1). The transposon MiniHimar1-lacZ was inserted between nucleotides 6760 and 6761 of the hdsp ORF (Supplementary Figure S2). A search of the GenBank database using basic local alignment search tool showed that the hypothetical Hdsp protein has 44% identity to the hypothetical sce0919 protein of Sorangium cellulosum So ce56. The predicted Hdsp protein has four conservative TPRs (Tetratricopeptide repeats) domains, as predicted by simple modular architecture research tools (Schultz et al., 1998; Letunic et al., 2009).
Transcription of hdsp
Because transposase-encoding genes on either side of hdsp transcribed in the opposite direction of hdsp, mutant phenotype was suggested to be not caused by polar effects of the insertion (Lee et al., 2005). The promoter-less lacZ gene in plasmid pMiniHimar1-lacZ was integrated in the same direction as the putative promoter of hdsp (Supplementary Figure S2); therefore, β-galactosidase activity could reflect the expression level of the mutated hdsp gene. The lacZ gene was determined to be expressed at a high and stable level during the development stage (Figure 3a). Because the β-galactosidase protein is rather stable during the growth stage of Myxococcus (Li et al., 1992), the expression of the hdsp gene during the growth stage was determined using RT–PCR. We found that the hdsp gene was also expressed during vegetative growth (Figure 3b).
Existence of hdsp in Myxococcus strains
To determine whether the hdsp gene occurs often in salt-tolerant Myxococcus, five salt-tolerant Myxococcus strains that were isolated from oceanic environments and the model strains of M. xanthus (listed in Supplementary Table S1) were analyzed for the presence of an hdsp homolog. These strains are highly similar of their 16S rRNA gene sequences with more than 98% identity. Because the DK1622 strain has been reported to lack a 222-kb segment compared with its parental strains FB and DZ2 (Chen et al., 1990), FB and DZ2 were also analyzed. Three pairs of primers were designed for PCR amplification (Figure 4). The first and second primer pairs were used to amplify the hdsp gene from the above mentioned salt-tolerant Myxococcus strains. Interestingly, hdsp PCR products were amplified from all five salt-tolerant Myxococcus strains, but not from the soil Myxococcus strains (Supplementary Figure S3). All of the PCR products were cut out for sequencing that revealed that the hdsp gene is highly conserved (99% identical) among the five salt-tolerant Myxococcus strains. The PCR products from the third pair of primers in the three strains of DK1622, FB and DZ2 were the same size, but did not include the inserted segment (Supplementary Figure S3). Furthermore, the existence of the hdsp gene in the type strains of five Myxococcus species (Supplementary Table S1) that were obtained from soil samples were also assayed. Because the PCR amplification protocol of the hdsp gene is not suitable in these five Myxococcus strains, dot blotting was used. The results indicated that the hdsp homolog was neither present in these five strains (Supplementary Figure S4).
Discussion
Among prokaryotes, myxobacteria have the largest known genomes and the most complicated gene-regulation systems that are probably necessary for their complex social lifestyle (Goldman et al., 2006; Zusman et al., 2007). The questions of how these large and complex genomes evolved and how they confer the capacity for sophisticated social behavior have generated much interest. Bioinformatic analyses (Goldman et al., 2007) have indicated that the genes involved in polysaccharide biosynthesis were probably acquired by lateral gene transfer, whereas the genes for sensory systems were more likely to be vertically inherited. This suggests that horizontally transferred genes may have important roles in genome expansion and the evolution of complex social behavior in myxobacteria. The functions of the hdsp gene in salt-tolerant Myxococcus provide the first experimental evidence supporting this hypothesis.
The well-known social behaviors of myxobacteria occur on solid surfaces. In liquid culture conditions, myxobacterial cells stick together to form loose or tight clumps. However, subculturing of myxobacteria strains in liquid medium can produce mutants that are capable of dispersed growth in liquid medium (Reichenbach and Dworkin, 1992; Velicer et al., 1998). Thus, the major obstacles to the survival of soil myxobacteria in marine conditions are the liquid environment and the high salinity. Salt-tolerant Myxococcus fulvus strain HW-1 exhibits remarkable changes in the social behavior when cultured in media of different saltwater concentrations. Our previous studies have indicated that maintenance and reinforcement of social behavior are important for HW-1 to adapt to oceanic conditions (Wang et al., 2007; Zhang et al., 2007; Pan et al., 2009). In this study, the phenotype of the mutant strain YLH0401 indicated that the insertional inactivation of a single horizontally transferred gene led to dramatic phenotypic changes, including dispersed growth, defective fruiting body formation and sporulation, decreased ECM production and S-motility, and seawater intolerance. Perhaps the dramatic decrease of ECM production led to these phenotypic changes in YLH0401. The hdsp gene is conserved in some salt-tolerant Myxococcus strains but does not exist in the type strains of Myxococcus that were obtained from soil samples. However, if the claim that the lateral transfer of the hdsp gene had a role in the genome expansion that gave rise to myxobacteria among the delta proteobacteria is true, then one would have expected to find hdsp in one or all of detected soil strains of Myxococcus species, whose genomes have been expanded. The fact that hdsp was not found in any of those strains suggests that salt tolerance was acquired by terrestrial strains that already were endowed with social behavior, when those strains were adapting to the high salt condition of a near-ocean habitat. The profound phenotypic changes by the mutation of this gene suggest it has important roles in maintaining the salt tolerance and social behavior in marine environments. The hdsp gene was probably acquired through lateral gene transfer and conserved in salt-tolerant Myxococcus strains for adaptation to oceanic habitats. Bioinformatic analyses revealed four TPR motifs in the hypothetical protein product of hdsp. The TPR motifs have been identified in wide variety of proteins in different organisms, and mediate protein–protein interactions and the assembly of multi-protein complexes (Blatch and Lässle, 1999). Proteins that contain TPRs are normally involved in a variety of biological processes, such as cell cycle regulation, transcriptional control, mitochondrial and peroxisomal protein transport, neurogenesis and protein folding (Blatch and Lässle, 1999). Proteins containing TPR were also reported to be involved in motility and fruiting body formation in DK1622 (Youderian et al., 2003; Nariya and Inouye, 2005; Yu and Kaiser, 2007; Scott et al., 2008). It is therefore possible that the putative TPR motifs in Hdsp also have a role in mediating interactions between Hdsp and other proteins. In addition to up- or downregulating the expression of multifunctional genes at transcriptional level to quickly and efficiently respond and acclimate to environmental changes (Pan et al., 2009), the marine salt-tolerant Myxococcus strains probably also horizontally exchange genes to maintain and reinforce social behavior to adapt to oceanic conditions.
Accession codes
References
Black WP, Xu Q, Yang ZM . (2006). Type IV pili function upstream of the Dif chemotaxis pathway in Myxococcus xanthus EPS regulation. Mol Microbiol 61: 447–456.
Black WP, Yang Z . (2004). Myxococcus xanthus chemotaxis homologs DifD and DifG negatively regulate fibril polysaccharide production. J Bacteriol 186: 1001–1008.
Blatch GL, Lässle M . (1999). The tetratricopeptide repeat: a structural motif mediating protein–protein interactions. BioEssays 21: 932–939.
Chen HW, Keseler IM, Shimkets LJ . (1990). Genome size of Myxococcus xanthus determined by pulsed-field gel electrophoresis. J Bacteriol 172: 4206–4213.
Dana JR, Shimkets LJ . (1993). Regulation of cohesion-dependent cell interactions in Myxococcus xanthus. J Bacteriol 175: 3636–3647.
Dawid W . (2000). Biology and global distribution of myxobacteria in soils. FEMS Microbiol Rev 24: 403–427.
Fudou R, Jojima Y, Iizuka T, Yamanaka S . (2002). Haliangium ochraceum gen. nov., sp. nov. and Haliangium tepidum sp. nov.: novel moderately halophilic myxobacteria isolated from coastal saline environments. J Gen Appl Microbiol 48: 109–116.
Goldman B, Bhat S, Shimkets LJ . (2007). Genome evolution and the emergence of fruiting body development in Myxococcus xanthus. PLoS ONE 2: e1329.
Goldman BS, Nierman WC, Kaiser D, Slater SC, Durkin AS, Eisen J et al. (2006). Evolution of sensory complexity recorded in a myxobacterial genome. Proc Natl Acad Sci USA 103: 15200–15205.
Iizuka T, Jojima Y, Fudou R, Hiraishi A, Ahn JW, Yamanaka S . (2003a). Plesiocystis pacifica gen. nov., sp. nov., a marine myxobacterium that contains dihydrogenated menaquinone, isolated from the Pacific coasts of Japan. Int J Syst Evol Microbiol 53: 189–195.
Iizuka T, Jojima Y, Fudou R, Tokura M, Hiraishi A, Yamanaka S . (2003b). Enhygromyxa salina gen. nov., sp. nov., a slightly halophilic myxobacterium isolated from the coastal areas of Japan. Syst Appl Microbiol 26: 189–196.
Iizuka T, Jojima Y, Fudou R, Yamanaka S . (1998). Isolation of myxobacteria from the marine environment. FEMS Microbiol Lett 169: 317–322.
Kashefi K, Hartzell PL . (1995). Genetic suppression and phenotypic masking of a Myxococcus xanthus frzF− defect. Mol Microbiol 15: 483–494.
Kearns DB, Campbell BD, Shimkets LJ . (2000). Myxococcus xanthus fibril appendages are essential for excitation by a phospholipid attractant. Proc Natl Acad Sci USA 97: 11505–11510.
Lee B, Higgs PI, Zusman DR, Cho K . (2005). EspC is involved in controlling the timing of development in Myxococcus xanthus. J Bacteriol 187: 5029–5031.
Letunic I, Doerks T, Bork P . (2009). SMART 6: recent updates and new developments. Nucleic Acids Res 37 (Database issue): D229–D232.
Li SF, Lee BU, Shimkets LJ . (1992). CsgA expression entrains Myxococcus xanthus development. Genes Dev 6: 401–410.
Li YZ, Hu W, Zhang YQ, Qiu ZJ, Zhang Y, Wu BH . (2002). A simple method to isolate salt-tolerant myxobacteria from marine samples. J Microbiol Methods 50: 205–209.
Liu YG, Whittier RF . (1995). Thermal asymmetric interlaced PCR: automatable amplification and sequencing of insert end fragments from P1 and YAC clones for chromosome walking. Genomics 25: 674–681.
Müller FD, Jakobsen JS . (2008). Myxobacteria: Multicellularity and Differentiation. ASM Press: Washington DC, pp 479–489.
Nariya H, Inouye S . (2005). Modulating factors for the Pkn4 kinase cascade in regulating 6-phosphofructokinase in Myxococcus xanthus. Mol Microbiol 56: 1314–1328.
Pan HW, Liu H, Liu T, Li CY, Li ZF, Cai K et al. (2009). Seawater-regulated genes for two-component systems and outer membrane proteins in Myxococcus. J Bacteriol 191: 2102–2111.
Reichenbach H . (1999). The ecology of myxobacteria. Environ Microbiol 1: 15–21.
Reichenbach H, Dworkin M . (1992). The myxobacteria. In: Balows A, Trüper HG, Dworkin M, Harder W, Schleifer KH (eds). The Prokaryotes, 2nd edn. Springer-Verlag: Berlin. pp 3416–3487.
Schultz J, Milpetz F, Bork P, Ponting CP . (1998). SMART, a simple modular architecture research tool: identification of signaling domains. Proc Natl Acad Sci USA 95: 5857–5864.
Scott AE, Simon E, Park SK, Andrews P, Zusman DR . (2008). Site-specific receptor methylation of FrzCD in Myxococcus xanthus is controlled by a tetra-trico peptide repeat (TPR) containing regulatory domain of the FrzF methyltransferase. Mol Microbiol 69: 724–735.
Shi W, Zusman DR . (1993). The two motility systems of Myxococcus xanthus show different selective advantages on various surfaces. Proc Natl Acad Sci USA 90: 3378–3382.
Shimkets LJ . (1990). Social and developmental biology of myxobacteria. Microbiol Rev 54: 473–501.
Velicer GJ, Kroos L, Lenski RE . (1998). Loss of social behaviors by Myxococcus xanthus during evolution in an unstructured habitat. Proc Natl Acad Sci USA 95: 12376–12380.
Wang B, Hu W, Zhang CY, Zhao JY, Jiang DM, Wu ZH et al. (2007). Adaptation of salt-tolerant Myxococcus strains and their motility systems to the ocean conditions. Microb Ecol 54: 43–51.
Ward MJ, Mok KC, Astling DP, Lew H, Zusman DR . (1998). An ABC transporter plays a developmental aggregation role in Myxococcus xanthus. J Bacteriol 180: 5697–5703.
Yang Z, Ma X, Tong L, Kaplan HB, Shimkets LJ, Shi W . (2000). Myxococcus xanthus dif genes are required for biogenesis of cell surface fibrils essential for social gliding motility. J Bacteriol 182: 5793–5798.
Youderian P, Burke N, White DJ, Hartzell PL . (2003). Identification of genes required for adventurous gliding motility in Myxococcus xanthus with the transposable element mariner. Mol Microbiol 49: 555–570.
Yu R, Kaiser D . (2007). Gliding motility and polarized slime secretion. Mol Microbiol 63: 454–467.
Zhang CY, Cai K, Liu H, Zhang Y, Pan HW, Wang B et al. (2007). A new locus important for Myxococcus social motility and development. J Bacteriol 189: 7937–7941.
Zhang YQ, Li YZ, Wang B, Wu ZH, Zhang CY, Gong X et al. (2005). Characteristics and living patterns of marine myxobacterial isolates. Appl Environ Microbiol 71: 3331–3336.
Zusman DR, Scott AE, Yang ZM, Kirby JR . (2007). Chemosensory pathways, motility and development in Myxococcus xanthus. Nat Rev Microbiol 5: 862–872.
Acknowledgements
We thank Drs D Kaiser (Stanford University), HB Kaplan (University of Texas Medical School), DR Zusman (University of California, Berkeley) and Z Yang (Virginia Polytechnic Institute and State University) for sharing strains and plasmids. We also thank Nature Publishing Group Language Editors for their help in editing this paper. This study was financially supported by National Science Foundation for Distinguished Young Scholars (No. 30825001), National Natural Science Foundation (No. 30600007) and National High Technology Research and Development program (No. 2006AA02Z171) of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Pan, Hw., Tan, Zg., Liu, H. et al. Hdsp, a horizontally transferred gene required for social behavior and halotolerance in salt-tolerant Myxococcus fulvus HW-1. ISME J 4, 1282–1289 (2010). https://doi.org/10.1038/ismej.2010.52
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2010.52
Keywords
This article is cited by
-
Phylogenetic perspectives of nitrogen-fixing actinobacteria
Archives of Microbiology (2012)