ENCODE 3
A collection of research articles and related content describing the Encyclopedia of DNA Elements, its datasets and tools.
How cells, tissues and organisms interpret the information encoded in the genome has vital implications for our understanding of development, health and disease. Launched in 2003, the ENCyclopedia Of DNA Elements (ENCODE) project has the aim of mapping the functional elements in the human genome (later expanded to include model organisms).
During the first phase of ENCODE, published in 2007, microarray-based technologies were used to detect regions associated with transcription factors, certain histone modifications and open chromatin within a pre-specified 1% of the human genome.
ENCODE’s second phase saw a switch to sequencing-based technologies, the addition of new assay types and the analysis of functional elements genome-wide, described in a collection of research articles in 2012.

2020 marks the year of the publication of the third phase of the ENCODE project, which has been further expanded in terms of assays used, and cells and tissues analysed.
The Encyclopedia paper of ENCODE 3, published in Nature, gives an overview of the various assays that were performed in human and mouse cell lines and tissues and describes a Registry of human and mouse candidate cis-regulatory elements (cCREs).
A suite of companion papers gives additional in-depth insights into the new datasets that were generated, their application to questions about how the activity of human and mouse functional elements is regulated, the organization of elements in the genome and nucleus, and the computational tools that were developed.
ENCODE in numbers
In this section, we want to highlight the growth of the ENCODE project since its inception, in size, in scope and in breadth, and how its data have been used for a better understanding of diseases in a multitude of publications. A Perspective article accompanying the research articles of ENCODE 3 gives further insights into the history and future of ENCODE.
Nature
Perspectives on ENCODE.

Number of Assays
In the early phase of ENCODE, assays were mostly performed in a selection of human model cell lines.
Over the years, additional assay types and experiments in a larger range of cell and tissue types, including primary cells and tissues from model organisms, were added.
By the end of phase 3 of ENCODE, more than 9,000 experiments had been carried out.
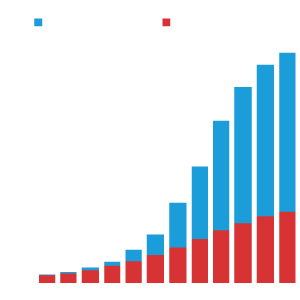
Number of Publications
With the growth of the data resource, the number of publications that make use of the ENCODE data has increased over the years.
This includes publications from the ENCODE consortium and ‘Community publications’ (which make use of ENCODE data but do not indicate grant support from ENCODE).
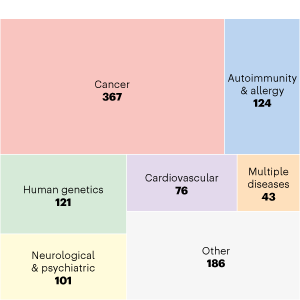
Publications by Disease
ENCODE data have been used in many studies of human diseases.
Shown here is a subset of community publications labelled as ‘Human disease’ studies, broken further down by type of disease studied, showcasing the breadth of research questions in which ENCODE data are of use.
ENCODE data and tools in use
The datasets produced and computational tools developed for ENCODE have facilitated insights into the regulation of the genome from many angles. In this section we showcase a non-exhaustive selection of the discoveries made in the ENCODE 3 papers.

Transcriptomes and Gene Regulation
The activity of genes must be regulated in a time-, cell-type- and tissue-specific manner, and the ENCODE project has generated datasets that help us to understand how this works in a large number of human and mouse cells and tissues.
Genome-wide maps of DNase I hypersensitive sites in 243 human cell and tissue types have made it possible to infer footprints of specific transcription factors and their activity at regulatory elements in the human genome.
Bulk RNA-sequencing data from up to 12 mouse embryonic tissues collected between embryonic days E10.5 and E16.5 and 17 tissues at birth (P0) allowed the identification of temporal driver genes during embryonic development in the mouse.
Matching DNA methylome maps, generated by whole-genome bisulfite sequencing of the same biosamples, highlighted the dynamics of changes in DNA methylation at fetal enhancers.
In addition, by integrating genomic and functional data, a mouse pseudogene catalogue was curated and the biogenesis of pseudogenes further explored.
Nature Communications
Transcriptional activity and strain-specific history of mouse pseudogenes.

3D Organization of the Genome
The primary sequence of DNA may be linear, but within a cellular context DNA is wrapped around histone proteins as part of the nucleosome, and chromatin is organized into further higher-order structures.
The three-dimensional organization of the genome has important implications for the activity of genes and other DNA-associated processes such as replication and repair.
By integrating chromatin immunoprecipitation with sequencing (ChIP–seq) assays for histone modifications and assay for transposase-accessible chromatin using sequencing (ATAC–seq) data from mouse embryonic tissues, the authors traced the changes in chromatin accessibility during embryonic development in mouse.
In addition to the extensive mapping of chromatin-associated proteins (CAPs; transcription factors and chromatin modifiers) described in the Encyclopedia paper, the binding profiles of almost a quarter of all CAPs that are active in a human cell line were generated for detailed insights into CAP-regulated transcriptional networks.
In 24 human cell lines, the three-dimensional physical interactions of distant genomic regions were mapped and integrated with epigenomic and transcriptome datasets to infer regulatory relationships.

RNA Binding and Regulation
RNA-binding proteins (RBPs) regulate gene expression by affecting the processing, stability, localization and translation of RNA.
For the third phase of ENCODE, RBPs were systematically studied in two human cell lines using a range of different biological assays which allowed, among other results, the identification of consensus RNA-binding motifs and binding affinities of 150 RBPs in vivo.

Human Variation and Disease
ENCODE data have found many uses in studies that aim to connect the functions of genomic regions with human health and disease.
The ENCODEC resource contains data-rich biosample pairings of cancer cell lines and normal cell types that enable cancer genomic annotations and inference of cancer-specific distal regulatory networks.
In a paper reporting the generation of high-resolution maps of DNase I hypersensitive sites (DHSs) in 733 human biosamples and their integration into an index of human regulatory elements, it was also found that heritability for human traits and diseases mediated by genetic variants identified from genome-wide association studies was enriched at DHSs.
Nature Communications
An integrative ENCODE resource for cancer genomics.

Using ENCODE
Beside the generation of comprehensive genomics datasets, another major focus of the consortium is the development of methods and tools for the scientific community to be able to further access, analyse and integrate genomic datasets for their own research questions.
The Encyclopedia paper introduces SCREEN, a Registry of 926,535 human and 339,815 mouse candidate cis-regulatory regions (cREs), which can be accessed through a browser.
A framework that integrates mammalian epigenomic and fly STARR-seq datasets into a matched filter score allows human and mouse enhancers to be predicted.
It is possible to detect donor mislabelling of next-generation sequencing datasets using the method CrosscheckFingerprints.
RADAR (RNA Binding Protein Regulome Annotation and Prioritization) is a framework that uses the ENCODE RNA-binding protein (RBP) datasets to annotate and prioritize genetic variants that may exert their functional effects through RBP-related processes.
Browse the ENCODE Portal to access these and many more ENCODE datasets, methods and information about associated projects.
Nature Communications
Detecting sample swaps in diverse NGS data types using linkage disequilibrium.
Browse the collection

Browse the Collection of ENCODE 3 publications, additional research papers, audio content, and more.