Abstract
Various interspersed repeated sequences and elements (IRSs) can be utilized to generate PCR-based multilocus fingerprint profiles by amplifying the interelement segments, using primers matching the elements themselves. We assessed the utility of inter-IRS fingerprinting in phylogenetic comparisons among six artiodactyl species using several primers derived from two abundant genomic components: the Bov-tA short interspersed nuclear elements (SINEs) and simple sequence repeats or microsatellites (SSRs). Character- and distance-based analyses of the fingerprint data produced trees conforming to the established phylogenetic relationships of species. The strength of phylogenetic signal from different primers varied; combining data from different experiments resulted in robust trees. Within the Cervidae, the hierarchical relationship [(Odocoileus, Rangifer) Alces] was strongly supported. Both methods appear useful tools for systematic studies at time scales <30 Myr. To elucidate the material basis of inter-SINE fingerprints, we obtained the first sequences of the ‘bovid’ Bov-tA element also from two cervids (reindeer and white-tailed deer) and analysed their relationship to a number of paralogous bovid elements. The differences among sequences, both intra- and interspecific, were relatively high (mean 18.5%); the sequences showed no clear clustering with the species from which they had been isolated. Most individual elements probably date back to the cervid–bovid ancestor >25 Myr ago, which is in line with the observed fingerprint distributions.
Similar content being viewed by others
Introduction
Eukaryote genomes contain several classes of abundant interspersed repetitive sequences and elements (IRSs), which can be used as generalized anchor sites for PCR-based genomic fingerprinting. With primers designed to match the elements themselves, several anonymous inter-IRS fragments can be amplified simultaneously, provided primer sites in opposite orientations are occasionally found close enough for conventional PCR.
The most trivial and general class of IRSs are the simple sequence repeats (SSRs) or microsatellites, widespread in all eukaryote genomes. Individual SSR loci probably originate and evolve largely independently of one another, through errors intrinsic to the replication process of noncoding DNA (Levinson & Gutman, 1987; Stallings, 1995). Another common class of IRSs is the short interspersed nuclear elements (SINEs), which vary from tens to hundreds of basepairs in length. SINEs are thought to have multiplied and spread across the genome by retroposition, the reintegration of reverse transcribed copies of certain master elements during evolution. A particular SINE family may comprise hundreds of thousands of paralogous loci. SINEs have phylogenetically restricted distributions, often limited to a certain taxonomic order or family. The precise mechanisms and dynamics of their spread and their biological significance are largely unknown (e.g. Deininger & Batzer, 1993).
The applications of inter-IRS fingerprinting have so far mostly concerned intraspecific polymorphisms (e.g. Sinnet et al., 1990; Kaukinen & Varvio, 1992; Miller, 1994). As with any evolving feature of the genome, IRS fingerprints should also carry information on the evolutionary relationships of lineages, and thus provide useful characters for systematic studies (Zietkiewicz et al., 1994).
Here we assess the practical utility of two IRS–PCR primer systems in a study of intergeneric phylogenetic relationships among artiodactyl mammals. Our model data set involves six species from the Bovidae, Cervidae and Suidae, representing divergence times from ≈5 to 60 Myr. One set of experiments used SSR-anchored primers (CA)8XY, with a choice of the anchoring 3′ nucleotides X and Y. The other approach used primers derived from an artiodactyl SINE, the Bov-tA element. Bov-tA itself is a construct of two subunits: an 85-bp unit of tRNA ancestry, and an adjacent 117-bp A-unit (see Fig. 1). Besides making a part of the Bov-tA SINE, the A-subunit also occurs as a monomer, as a dimer, and as part of the Bov-B LINE and the 1.709 satellite (see Lenstra et al., 1993). The Bov-tA family has so far only been described in Bovidae, although Jobse et al. (1995) suggested that the A-subunit is also present in other true ruminants.
Alignment of paralogous Bov-tA element sequences from five artiodactyl species. The four short interspersed nuclear element (SINE) primers used for inter-SINE fingerprinting and the bovid tA consensus from Kaukinen & Varvio (1992) are also shown. The elements apparently truncated at 3′ have been sequenced from clones produced by inter-SINE PCR using the SINE2 primer. The ambiguities (?) at the 5′ end represent evidently nonhomologous bases omitted from the phylogenetic analyses. The GenBank accession numbers are (+ indicates a new sequence generated in this study): cattle1–3: Z33884+, Z33886+, AF018147+; sheep 1–3: X12817, X01610, X07973; wtdeer1–3: Z33896+, Z33897+, AF018148+; reindeer 1–3: AF018149+, AF018150+, Z33895+; muntjac1–3: X82880, X82883, X82886.
Because Bov-tA fingerprinting proved generally almost as efficient in Cervidae as in Bovidae, we went on to elucidate the basis of the observed priming site conservation at the deeper phylogenetic levels. We therefore analysed the relationships of a sample of (paralogous) SINE elements from bovids and cervids, including the first Bov-tA sequences from reindeer and white-tailed deer. This gives an initial insight into the depth and dynamics of Bov-tA evolution in the Artiodactyla.
Materials and methods
Inter-IRS fingerprinting
The data set involved five pecoran ruminants, two from the Bovidae (cattle: Bos taurus; sheep: Ovis aries) and three from Cervidae (moose: Alces alces; white-tailed deer: Odocoileus virginianus; reindeer: Rangifer tarandus). The pig [Sus scrofa (Suiformes)] was included as an outgroup. Generally, each species was represented by one to four unrelated individuals; however, the inter-SINE experiments included a total of 22 cattle and 10 sheep, of which 10 and four were unrelated individuals, respectively.
Three primers, SINE2, SINE11 and SINE10, derived from bovid Bov-tA sequences (Kaukinen & Varvio, 1992) were used for inter-SINE fingerprinting, each in a separate experiment. Note that because of a palindromic feature of the A-subunit, SINE2 can prime in two orientations (Fig. 1). Inter-SINE PCRs were carried out in 20 μL reaction volume using 20 ng of genomic DNA essentially as described in Kaukinen & Varvio (1992). Annealing temperatures were 63°C for SINE11 and 68°C for SINE2 and -10. Inter-SINE PCR products were resolved on polyacrylamide sequencing gels (6%, 7 M urea), and detected with autoradiography.
For inter-SSR fingerprinting, three different SSR-anchored primers (CA)8AT (CA)8AG and (CA)8GG, were used in three separate experiments, each with a different combination of two of the primers. The combinations will be denoted according to the 3′-terminal anchoring dinucleotides, e.g. AT/GG for the experiment involving (CA)8AT and (CA)8GG. Note that because of overlapping sets of primers, some overlap of fragments between the experiments (ideally 25%) is expected. The reaction conditions were essentially as described in Kaukinen & Varvio (1992), except that 10 pmol of each primer was used in 20 μL reaction volume. The cycling parameters were 94°C (2 min), 27 cycles with 94°C (30 s), 52°C (45 s), 72°C (2 min), and a 10-min final extension. The products were detected as above.
Fingerprint analysis
The fragments scored from the fingerprint profiles were treated as binary characters (presence/absence), and species relationships were assessed both by character- and distance-based methods. For the character-based approach, a fragment was scored as present if found in any individual of the given species. Character states were alternatively assumed as unordered, or as ordered Dollo characters (i.e. only one 0 → 1 change allowed). PAUP 3.1 software (Swofford, 1993) was employed in parsimony analysis and tree evaluation.
We assessed the performance of the data in recovering the reference tree topology in Fig. 3(b), representing the current consensus of the phylogeny of the taxa studied (see Discussion for references). For five ingroup species (treating the pig as an outgroup) this is one of 105 alternative possible rooted topologies. The trees recovered were evaluated with consistency and retention indices (CI, RI), bootstrap percentage support in parsimony analysis for the three critical internodes in Fig. 3(b) (BP, 500 replicates), and decay indices (DI). Consistency indices are a measure of relative homoplasy in the data (here, the number of informative characters/number of steps implied by the tree). Decay indices are the number of additional steps required to collapse a particular internode, or corresponding clade (Bremer, 1994); a negative value indicates the difference between lengths of the reference tree and the most parsimonious (MP) tree where the clade is not present. The DI value for the whole tree is here the number of steps required to collapse any internode of the reference tree (equal to the smallest individual DI), or, if negative, the difference between the MP and reference tree lengths.
(a) Neighbour-joining (NJ) clustering based on interspecies distance matrices, from each inter-SINE (short interspersed nuclear element)and inter-SSR (simple sequence repeat) fingerprinting experiment. Distances to pig for SINE10 (undefined as no bands were scored) were set to unity for clustering. (b) The reference phylogeny, based on previous molecular and fossil data (see Discussion for references). The circled numbers denote three internodes (and the three clades defined by them), for which phylogenetic support was evaluated in Table 2.
For the distance approach, a band-sharing similarity s was calculated for each pair of individuals as the number of shared fragments divided by the average total number in the two individuals; the corresponding divergence measure is d = 1 − s. Using the averages of d in all pairwise individual comparisons between species i and j (Dij), and averages of d within each species (Di and Dj), interspecific distance estimates corrected for intraspecific variation were obtained as D′ij = Dij − [(ni − 1)Di+(nj − 1)Dj]/(ni+nj − 2). The distances were illustrated by neighbour-joining (NJ) clustering (Saitou & Nei, 1987).
Analysis of SINE sequences
To elucidate intra- and interspecific variation and evolution among the Bov-tA elements, three (partial) Bov-tA sequences from each of a set of five artiodactyl species were aligned (Fig. 1), and sequence relationships were assessed using the maximum parsimony criterion using PHYLIP software (Felsenstein, 1993), and also illustrated as a NJ-tree calculated from Kimura’s 2-parameter sequence distances (DK). New Bov-tA element sequences from reindeer, white-tailed deer and cattle were obtained in connection with construction of genomic libraries targeted to inter-SINE segments: fragment pools produced by (nonradioactive) inter-SINE PCR using the SINE2 primer were subcloned, and random positive clones were sequenced, essentially as in Kaukinen & Varvio (1992) and Kostia et al. (1997). Database sequences from cattle, sheep and another cervid, the muntjac (Muntiacus muntjak) were used in addition (accession numbers in Fig. 1).
Results
Inter-IRS fingerprints
Electrophoresis of radioactively labelled IRS PCR products revealed primer and species-specific fingerprint profiles, with a low level of intraspecific polymorphism (Fig. 2).
In the five pecoran species, an average of 54, 35 and 13 different fragments were scored using the primers SINE2, -11 and -10, respectively; few, if any, fragments were seen in pig (0–4 per primer; Table 1). Primer-specific differences in the fragment numbers were also found between bovids and cervids: SINE11 produced twice the number of fragments in cervids than in bovids, whereas SINE2 and -10 produced fewer fragments in moose and white-tailed deer than in other taxa. About 6% of the fragments were scored as polymorphic within a species (Table 1). The polymorphisms appeared as dominant traits (presence/absence of a band).
The SSR-anchored primer combinations AT/GG, AT/AG and GG/AG produced on average 26, 28 and 36 different fragments per species. Generally, fragment numbers were slightly higher in bovids than in the other taxa. Seven percent of the fragments were polymorphic.
Phylogenetic information
The NJ-trees derived from each SINE- and SSR-primer experiment were similar to each other, and generally agreed with the reference topology (Fig. 3). The congruence of results is seen in the high correlations of D′-value matrices between different experiments: correlations among the different SINE experiments were in the range r = 0.95–0.99, those among the different SSR primer combinations r = 0.90–0.93, and those between individual SINE and SSR experiments r = 0.87–0.94 (see also Fig. 4).
Pairwise distance estimates from individual fingerprinting experiments plotted against the approximate divergence time (Fig. 3b).
The number of phylogenetically informative fragments in the character-based approach varied from 14 to 69 in the different experiments (Table 1). Treating character states as unordered, the reference tree was identical with the MP topology in five of the six experiments; in four, this was the unique MP solution. When the MP topology was different (AT/GG), it was only a single step shorter than the reference tree (Table 2).
The strength of the phylogenetic signal varied among the primers. SINE2 and AT/AG provided solid support for all three nodes (minimum DI 11 and four characters, respectively). In other experiments, individual nodes remained ambiguous; combining data over experiments yielded robust trees (Table 2). The alternative treatment with Dollo parsimony did not increase the overall tree robustness.
Divergence and relationships of Bov-tA element sequences
The comparison of Bov-tA sequences from five ruminant species did not suggest any taxon-related pattern of differentiation. The most prominent structural distinction was a 9-bp insertion/duplication in the tRNA related-part of the element, which was present occasionally both in the cervid and bovid elements. Because of internal sequence redundancy, the position of this insertion cannot be unequivocally decided (cf. Lenstra et al., 1993). Sequence relationships were analysed excluding the insertion and two flanking nucleotides at its either end in Fig. 1. The observed nucleotide distances (p) for the remaining 125 positions common to all sequences ranged from 6 to 29% (DK = 0.07–0.36) with an average of 18.5% (DK = 0.21) for both intra- and interspecific comparisons. This is similar to the average for 48 bovid sequences in Kaukinen & Varvio (1992). A NJ-tree did not indicate any consistent clustering with the taxonomic origin of the sequences, or with the presence/absence of the insertion (Fig. 5). Neither did parsimony analysis on the entire alignment (excluding the 9 + 4 bp; data not shown). Bootstrap support estimates for individual nodes of these NJ and MP trees were low, not exceeding 65%.
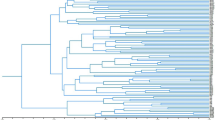
Relationships of the Bov-tA sequences in Fig. 1 as recovered by neighbour-joining (NJ)-clustering. Kimura’s two-parameter distances were calculated for a 125-bp portion of the alignment common to all sequences; this excluded the 9-bp insertion and two bases on its either side. Asterisks mark sequences that contain the insertion.
Discussion
Phylogenetic signal in inter-IRS fingerprints
SINEs and SSRs represent unstable components in the genome, each with their own complex dynamics, different from that of single-copy DNA. Changes in inter-IRS fingerprint patterns may be caused by length variations in the interelement segments, and by processes of priming site evolution. The latter may involve generation of new sites by SINE retroposition or by in situ origin of new microsatellites, and, on balance, degeneration through mutation or collapse of SSR length.
The presence of individual SINE copies at specific chromosomal locations has recently been noted as a new class of strong phylogenetic clade markers (Cook & Tristem, 1997). Inter-IRS fingerprints should carry information from those and similar phylogenetic events. However, as other more complex phenomena are also involved, an approach based on the strict assumptions of phylogenetic analysis (e.g. character homology and independence) could be questioned. No modelling of fingerprint divergence is possible, either. Yet, a phylogenetic signal is expected to be retained, and for an empirical test of that signal in a simple phylogeny we employed the standard character- and distance-based approaches, noting that they here represent pragmatic operational tools rather than theoretically grounded analyses. In a related study, Borowsky et al. (1995) assessed the utility of the AP-PCR fingerprinting method.
With these straightforward parsimony and clustering analyses, the fingerprint data did recover the correct phylogeny in most of our six experiments (Table 2, Fig. 3). There was a largely congruent relationship between the average phenetic distance and evolutionary time. At the deeper nodes, the divergence estimates evidently approach the saturation level, and resolution is lost (Figs 3 and 4). Yet, as a general conclusion, on time scales <30 Myr both fingerprinting techniques seem to reveal a considerable amount of true information on mammalian relationships. Particulary inter-SSR fingerprinting should then provide a universal and efficient, quick-and-dirty systematic tool, as SSR priming sites are abundant in all eukaryotes. By contrast, the use of specific inter-SINE primers is restricted by the phylogenetic distribution of the targeted SINEs, and requires prior sequence information from some member(s) of the group.
There was much variation in the strength of the phylogenetic signal between the experiments (Table 2). This was largely a function of the total fragment number, which, in turn, could sometimes be plausibly related to the number of potential priming sites. The larger number of fragments produced by SINE2 and -11 than SINE10 (Table 1) is in line with the fact that SINE2 and -11 match the A-subunit of Bov-tA, which, unlike the tRNA-subunit, is also involved in various other IRS families (see Introduction). For instance, Bov-tA and dimer Bov-A2 SINE loci are approximately equally numerous in the bovine genome (Lenstra et al., 1993). Furthermore, SINE2 produced more fragments than SINE11: this is expected from the capacity of SINE2 to prime in both orientations within the A-subunit. The distinctly smallest number of fragments found with SINE10 may also be related to the coincidence of the priming site with the occasional 9-bp insert, which would prevent priming in a large fraction of Bov-tA loci (Fig. 1). Furthermore, the difference in SINE10 fragment numbers between species could reflect different frequencies of the insert and noninsert elements in the lineages. Similar differences in hitherto unrecognized SINE-subfamilies could cause other differences in priming efficiency, such as the bovid–cervid contrast in SINE11 (Table 1).
Systematic resolution
Although the reference phylogeny of the six species (Fig. 3b) is fairly well accepted and age estimates are available both from fossils and molecules, there is still some uncertainty of the timing and even details of topology. In this context, the relative support values implied by our data to the various clades are of interest, as strong signals were obtained for the a priori weakest clades.
From fossil data, it has been difficult to discern the initial radiation of the pecoran families (Bovidae, Cervidae) and the subsequent split into the cattle and sheep lineages within the Bovidae; both have been referred to ≈20 Myr (review in Miyamoto et al., 1993). From molecular data the hierarchy is clearer, with estimates 23–28 Myr (Kraus & Miyamoto, 1991) and 14–24 Myr (Georgiadis et al., 1990; Allard et al., 1992), respectively. The radiation of the three cervids, all referred to subfamily Odocoileinae, was distinctly more recent, <10 Myr (Miyamoto et al., 1990). Therefore the relatively strong signal retained in our data for the bovid clade (internode 3) as compared to that for the cervids (2) is notable (Table 2). Distance data from both marker types illustrate almost similar divergence within the odocoileine and bovid clades, although the latter is considered twice as old (Fig. 3).
Previous information on the relationships among the three cervids has been contradictory. The [(reindeer, white-tailed deer) moose] hierarchy in Fig. 3(b) was suggested from morphology (Groves & Grubb, 1987); but alternatively, the three lineages have been assigned equal rank (Gustafson, 1985), and allozyme and mitochondrial RFLP data even pointed to a [(moose, white-tailed deer) reindeer] topology (Baccus et al., 1983; Cronin, 1991). The combined data from each fingerprinting approach, and from four of six individual experiments, now strongly support the [(reindeer, white-tailed deer) moose] topology. The relative power and information content of these approaches (Table 2, node 1) is illustrated by comparison to recent data from a total of 1517 bp of nuclear and mitochondrial sequence, which supported this topology (node) with DI = 10 (Randi et al., 1998).
Distribution and evolution of Bov-tA SINE in the Artiodactyla
The Bov-tA SINE family is represented by 285 000 copies in the bovine genome (Lenstra et al., 1993). It was described as a bovid-specific element (Rogers, 1985), but there have been scattered indications of a taxonomically wider distribution in the Artiodactyla (Kaukinen & Varvio, 1992; Jobse et al., 1995). Our results now include multiple lines of evidence that the Bov-tA family was already widespread in the common ancestor of the bovids and cervids. First, all the bovid consensus primers amplified comparable numbers of fragments from cervids and bovids, and retention of common fragments was implicated by the fingerprint phylogeny analyses. Secondly, sequencing of the inter-SINE PCR products of reindeer and white-tailed deer, and the database muntjac sequences (Fig. 5) directly verified the presence and identical structure of the Bov-tA SINE in the Cervidae.
Thirdly, the relationships among the paralogous Bov-tA sequences from cervids and bovids show no evidence of clustering with taxonomic origin, suggesting that most of them predate the radiation of the pecoran families, 20–25 Myr ago. This is further supported by the amount of sequence divergence, with an average corrected nucleotide distance DK = 0.21 (observed average P = 18.5%). This is clearly higher than a recent estimate of (orthologous) pseudogene divergence between cervids and bovids, DK = 0.12 (S. Kostia, S.-L. Varvio and R. Väinölä, unpubl. data); it might imply ≈40-Myr average age for the Bov-tA copies. This is also in line with the concept of SINE divergence rate based on orthologous comparisons of human and rodent mammalian-wide interspersed repeats (MIR) elements (DJC = 0.30 at 65–100 Myr; Jurka et al., 1995). Even with the more general pseudogene rate calibration of 0.4%/Myr (Li, 1997), the Bov-tA average would correspond to >25 Myr divergence time. The depth of the Bov-tA element diversity appears similar to that for a number of other mammalian SINE families, notably the porcine PRE-1 (Yasue & Wada, 1996). Although a few fragments were scored from pig in this study, it may be premature to argue for the presence of the Bov-tA in that lineage before a direct verification at sequence level, given the possibility of occasional random priming. Yet, with the divergence rates discussed above, the most divergent Bov-tA lineages in Fig. 5 (DK = 0.36) might well trace back to the ruminant–suiform ancestor, 50–60 Myr ago.
References
Allard, M. W., Miyamoto, M. M., Jarecki, L., Kraus, F. and Tennant, M. R. (1992) DNA systematics and evolution of the artiodactyl family Bovidae. Proc Natl Acad Sci USA. 89: 3972–3976.
Baccus, R., Ryman, N., Smith, M. H., Reuterwall, C. and Cameron, D. (1983) Genetic variability and differentiation of large grazing mammals. J Mammal. 64: 109–120.
Borowsky, R. L., Mcclelland, M., Cheng, R. and Welsh, J. (1995) Arbitrarily primed DNA fingerprinting for phylogenetic reconstruction in vertebrates: the Xiphophorus model. Mol Biol Evol. 12: 1022–1032.
Bremer, K. (1994) Branch support and tree stability. Cladistics. 10: 295–304.
Cook, J. M. and Tristem, M. (1997) ‘SINEs of the times’ — transposable elements as clade markers for their hosts. Trends Ecol Evol. 12: 295–297.
Cronin, M. A. (1991) Mitochondrial-DNA phylogeny of deer (Cervidae). J Mammal. 72: 533–566.
Deininger, P. L. and Batzer, M. A. (1993) Evolution of retroposons. Evol Biol. 27: 157–196.
Felsenstein, J. (1993) Phylip: phylogeny inference package, v. 3.5. Department of Genetics, University of Washington, Seattle, WA.
Georgiadis, N. J., Kat, P. W., Oketch, H. and Patton, J. (1990) Allozyme divergence within the Bovidae. Evolution. 44: 2135–2149.
Groves, C. P. and Grubb, P. (1987) Relationships of living deer. In: Wemmer, C. M. (ed.) Biology and Management of the Cervidae pp. 21–59. Smithsonian Institute Press, Washington, DC.
Gustafson, E. P. (1985) Antlers of Bretzia and Odocoileus (Mammalia, Cervidae) and the evolution of New World deer. Trans Nebraska Acad Sci. 13: 83–92.
Jobse, C., Buntjer, J. B., Haagsma, N., Breukelman, H. J., Beintema, J. J. and Lenstra, J. A. (1995) Evolution and recombination of bovine DNA repeats. J Mol Evol. 41: 277–283.
Jurka, J., Zietkiewicz, E. and Labuda, D. (1995) Ubiquitous mammalian-wide interspersed repeats (MIRs) are molecular fossils from the mesozoic era. Nucl Acids Res. 23: 170–175.
Kaukinen, J. and Varvio, S.-L. (1992) Artiodactyl retroposons: association with microsatellites and use in SINEmorph detection by PCR. Nucl Acids Res. 20: 2955–2958.
Kostia, S., Vilkki, J., Pirinen, M., Womack, J. E., Barendse, W. and Varvio, S. -L. (1997) SINE targeting of bovine microsatellites from bovine/rodent hybrid cell lines. Mamm Genome. 8: 365–367.
Kraus, F. and Miyamoto, M. M. (1991) Rapid cladogenesis among the pecoran ruminants: evidence from mitochondrial DNA sequences. Syst Zool. 40: 117–130.
Lenstra, J. A., Vanboxtel, J. A. F., Zwaagstra, K. A. and Schwerin, M. (1993) Short interspersed nuclear element (SINE) sequences of the Bovidae. Anim Genet. 24: 33–39.
Levinson, G. and Gutman, G. A. (1987) Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mol Biol Evol. 4: 203–221.
Li, W.-H. (1997) Molecular Evolution Sinauer Associates, Sunderland, MA.
Miller, J. R. (1994) Use of porcine interspersed repeat sequences in PCR-mediated genotyping. Mamm Genome. 5: 629–632.
Miyamoto, M. M., Kraus, F. and Ryder, O. A. (1990) Phylogeny and evolution of antlered deer determined from mitochondrial DNA sequences. Proc Natl Acad Sci USA. 87: 6127–6131.
Miyamoto, M. M., Kraus, F., Laipis, P. J., Tanhauser, S. M. and Webb, S. D. (1993) Mitochondrial DNA phylogenies within Artiodactyla. In: Szalay, F. S., Novacek, M. J. and McKenna, M. C. (eds) Mammal Phylogeny: Placentals pp. 268–281. Springer, New York.
Randi, E., Mucci, N., Pierpaoli, M. and Douzery, E. (1998) New phylogenetic perspectives on the Cervidae (Artiodactyla) are provided by the mitochondrial cytochrome B gene. Proc R Soc B. 265: 793–801.
Rogers, J. H. (1985) The origin and evolution of retroposons. Int Rev Cytol. 93: 187–279.
Saitou, N. and Nei, M. (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 4: 406–425.
Sinnett, D., Deragon, J.-M., Simard, L. R. and Labuda, D. (1990) Alumorphs—Human DNA polymorphisms detected by polymerase chain reaction using Alu-specific primers. Genomics. 7: 331–334.
Stallings, R. L. (1995) Conservation and evolution of (CT)n/(GA)n microsatellite sequences at orthologous positions in diverse mammalian genomes. Genomics. 25: 107–113.
Swofford, D. L. (1993) Paup: phylogenetic analysis using parsimony v. 3.1.1. Illinois Natural History Survey, Champaign, IL.
Yasue, H. and Wada, Y. (1996) A swine SINE (PRE-1 sequence) distribution in swine-related animal species and its phylogenetic analysis in swine genome. Anim Genet. 27: 95–98.
Zietkiewicz, E., Rafalski, A. and Labuda, D. (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics. 20: 176–183.
Acknowledgements
This work has been supported by the Academy of Finland. We thank Mikko Kolkkala for comments on the previous version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kostia, S., Ruohonen-Lehto, M., Väinölä, R. et al. Phylogenetic information in inter-SINE and inter-SSR fingerprints of the Artiodactyla and evolution of the Bov-tA SINE. Heredity 84, 37–45 (2000). https://doi.org/10.1046/j.1365-2540.2000.00629.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2000.00629.x
Keywords
This article is cited by
-
Analysis of genetic variability among bivoltine and multivoltine silkworm genotypes using inter simple sequence repeat and simple sequence repeat markers
The Nucleus (2023)
-
Development and application of SINE-based markers for genotyping of potato varieties
Theoretical and Applied Genetics (2012)
-
Genetic Divergence Detected by ISSR Markers and Characterization of Microsatellite Regions in Mytilus Mussels
Biochemical Genetics (2007)
-
Intraspecific Variation in isolates of Bursaphelenchus xylophilus (Nematoda: Aphelenchoididae) revealed by ISSR and RAPD fingerprints
Journal of Plant Diseases and Protection (2006)
-
Genomic sequence analysis of the 238-kb swine segment with a cluster of TRIM and olfactory receptor genes located, but with no class I genes, at the distal end of the SLA class I region
Immunogenetics (2005)