Abstract
Adiponectin is an adipocyte-secreted protein that modulates insulin sensitivity and whose low circulating concentration is associated with insulin resistance. In the present study, we analysed the association between two single-nucleotide polymorphisms (SNPs) in the adiponectin gene and insulin resistance in 253 nondiabetic subjects. In addition, we investigated whether this association is modulated by body mass index (BMI) levels. The SNPs +45T>G and +276G>T in the human adiponectin gene were detected in real-time PCR with LightCycler. No association was found with the +45T>G SNP. The +276G>T SNP was associated with higher BMI (P<0.01), plasma insulin (P<0.02) and HOMAIR (P<0.02). To analyse the possible interaction between BMI and the adiponectin gene on insulin resistance, the study group was divided into two subgroups, according to the BMI below or above the median of 26.2 kg/m2. In both subgroups, subjects carrying the +276G>T SNP had higher HOMAIR; however, the difference was highly significant among leaner (P<0.001), but not among heavier individuals, indicating that BMI status and the adiponectin gene interact in modulating insulin resistance. Among individuals with BMI <26.2 kg/m2, the relative risk of insulin resistance was 9.7 (CI: 1.32–87.7, P<0.035). In a subgroup of 67 subjects, carriers of the +276G>T SNP had significantly (P<0.05) lower mean serum adiponectin levels (25.7 ng/ml) compared to noncarriers (37.0 ng/ml), suggesting a possible influence of the +276G>T SNP on adiponectin levels. In summary, we observed an association between the +276G>T SNP in the adiponectin gene and insulin resistance. In particular, among leaner individuals, the adiponectin gene appears to determine an increased risk to develop insulin resistance.
Similar content being viewed by others
Introduction
Insulin resistance is considered the core factor in the pathogenesis of common disorders such as type II diabetes, atherosclerosis and the metabolic syndrome, and is often associated with obesity. Genetic and epidemiological studies strongly suggest that insulin resistance is, at least in part, genetically determined. The involved genes are, however, mostly unknown.
Adiponectin, an adipocyte-derived protein, is likely to modulate insulin sensitivity, and to play a role in both human and animal models of insulin resistance. In patients with diabetes, obesity and coronary artery disease (CAD), adiponectin levels have been found consistently to be lower than in control subjects.1,2,3 Recently, Tschritter et al 4 have shown that high adiponectin predicts increased insulin sensitivity, and this effect is independent of body fat mass. In recent longitudinal studies, individuals with higher plasma adiponectin concentrations were less likely to develop type II diabetes.5,6 Furthermore, administration of thiazolidinediones (TZD), an insulin-sensitising class of drugs, to insulin-resistant subjects significantly increased the plasma adiponectin levels, and this effect was correlated with the amelioration of insulin resistance in these subjects.7 Finally, a genome scan for insulin resistance loci has detected a quantitative trait locus on 3q27, where the adiponectin gene, also known as APM1 or ACRP30 gene, has been located,8 and this result was replicated in a French study.9
The adiponectin gene, therefore, has been suggested as a potential candidate gene for insulin resistance. A few studies have, in fact, reported the association between single-nucleotide polymorphisms (SNPs) of the adiponectin gene and type II diabetes10,11 or insulin resistance.12,13 While, in the study from Stumvoll and et al, a single SNP (ie +45T>G) was associated with obesity and related derangement of insulin sensitivity, in the study of Menzaghi et al,12 a haplotype of the adiponectin gene (ie the combination of the two SNPs +45T>G and +276G>T), was associated with several features of insulin resistance in nondiabetic individuals, including low serum adiponectin levels. Recently, Vasseur et al14 have reported on the association of an additional adiponectin gene haplotype (including SNPs −11 391 and −11 377, both located in the 5′ sequences) with plasma adiponectin levels, although no association with insulin-resistance indexes and type II diabetes was observed. The reasons for partially discrepant results are not known, and may reside in the different genetic background of the studied populations.
In the present study, we have studied the previously reported association between two SNPs (+45T>G and +276G>T) and insulin resistance in a population from the Lazio region (ie Centre-West Coast of Italy).11,12,13 This population was selected without subjects with diabetes and/or the metabolic syndrome, in order to avoid the possible modifying effects of these disorders on insulin sensitivity. In addition, since adiponectin and obesity are likely to modulate insulin sensitivity by different mechanisms, we also investigated whether adiponectin gene variations interact with obesity in determining the risk of insulin resistance.
Research design and methods
Subjects
A total number of 253 Caucasian subjects were studied. All subjects were recruited in the Lazio region of Italy, most from Rome and its surrounding towns. All the subjects were unrelated individuals randomly selected from a population of free-living individuals screened for CAD risk factors. The exclusion criteria were: (1) the presence of type II diabetes or of first-grade relatives with type II diabetes, (2) the presence of hypertension, (3) the presence of CAD and (4) the presence of the metabolic syndrome. CAD was excluded by use of the Rose questionnaire 15 and ECG (Minnesota coding). In all subjects, a complete medical history was obtained by the questionnaire. History taking included questions about smoking habits, history of hypertension and type II diabetes and current medication used. Diagnosis of type II diabetes was based on the history of hypoglycaemic treatment and/or confirmed fasting blood glucose >126 mg/dl (7.0 mmol/l).16 The metabolic syndrome was diagnosed according the WHO criteria.17 Diagnosis of hypertension was based on the presence of elevated systolic (>140 mmHg) and/or diastolic (>90 mmHg) blood pressure, and/or the current use of antihypertensive medications.
Homeostasis model assessment for insulin resistance (HOMAIR) was calculated as described by Matthews et al.18
Laboratory measurements
Cholesterol and triglyceride concentrations in total plasma and lipoprotein fractions were measured with a Technicon RA-1000 Autoanalyzer. High-density lipoprotein (HDL) cholesterol was determined in the whole plasma after precipitation of apoB-containing lipoproteins with phosphotungstic acid/MgCl2. Plasma glucose was determined by the glucose oxidase method (Autoanalyzer, Beckman Coulter, USA; coefficient of variation (CV), 1.9±0.2%). Plasma insulin concentration was measured on frozen samples, using a radio immunoassay (Biodata Insulin Kit, Milan, Italy) with an interassay CV of 7.5%.
Serum adiponectin levels were measured by commercial radioimmunoassay (LINCO Research Inc., USA), according to the manufacturer's instructions.
Mutation detection using fluorescence resonance energy transfer
The SNPs +45T>G and the SNP +276G>T of the human adiponectin gene were detected in real-time PCR with LightCycler hybridisation probes,19 using fluorescent-labelled nucleotides.20 Fluorescence resonance energy transfer occurs as two probes hybridise to adjacent sequences on the same unlabelled complementary strand.21 When a probe hybridises over a sequence variant, a mismatch is formed and the duplex is destabilised. This is reflected by a shift in melting temperature (Tm) from the completely complementary duplex. Hybridisation probe Tm is the temperature at which 50% of the probe has strand-separated from a template, and can be estimated from the inflection point of the melting curve or the centre of derivative melting curves.21
Design of primers and fluorogenic probes
The primers used to amplify the 250 bp fragment for the SNP +45T>G and the 196 bp fragment for the SNP +276G>T of the human adiponectin gene (GeneBank ref. AF304467) are shown in Table 1.
The detection probe (Table 1) was a 19-mer oligonucleotide for the SNP +45, labelled at the 3′-end with fluorescein, complementary to the leading strand of the G allele. The anchor probe was a 22-mer, labelled at the 5′-end with LightCycler Red 640, and modified at the 3′-end by phosphorylation to block extension. This probe binds at a distance one base 3′ from the detection probe.
For the SNP +276, the detection probe was a 22-mer oligonucleotide complementary to the leading strand of the G allele. The anchor was a 24-mer labelled at the 5′-end with LightCycler Red 640, and it binds at a distance one base 3′ from the detection probe.
Fluorescence protocol
The analysis was carried out on a LigthCycler apparatus (Roche Molecular Biochemical). For SNP +45T>G and SNP +276G>T, fragments were amplified by rapid-cycling in a reaction volume of 20 μl with 0.5 μmol/l each primer, 0.15 μmol/L anchor and detection probes, 200 ng of genomic DNA, 1 × DNA Master Hybridisation Probes (Roche Molecular Biochemical) containing nucleotides, Taq DNA polymerase and 10 mM Mg2+. The final Mg2+ concentration in the reaction mixture was adjusted to 4 mM. The samples were loaded into glass capillary cuvettes (Roche Molecular Biochemicals) and centrifuged. After an initial denaturation step at 95°C for 5 min, amplification was performed using 50 cycles of denaturation (95°C for 3 s), annealing (57°C for 12 s for SNP +45T>G and 55°C for 12 s for SNP +276G>T) and extension (72°C for 10 s for SNP +45T>G and 72°C for 8 s for SNP +276G>T) on the LightCycler. The temperature-transition rates were programmed at 20°C/s from denaturation to annealing, 20°C/s from annealing to extension and 20°C/s from extension to denaturation. Fluorescence was measured at the end of the annealing period of each cycle to monitor amplification. After amplification was completed, a final melting curve was recorded by heating to 95°C for 30 s with 20°C/s, holding at 40°C for 30 s with 20°C/s, and then heating slowly at 0.3°C/s until 80°C. Fluorescence was measured continuously during the low-temperature ramp to monitor the dissociation of the fluorescein-labelled detection probe. The fluorescence signal (F) was plotted in real time against temperature (T) to produce melting curves for each sample (F vs T). Melting curves were then converted to derivative melting curves by plotting the negative derivative of the fluorescence with respect to temperature against temperature (−(dF/dT) vs T). The entire process took approximately 30 min, with no separate manipulation of the product necessary.
Statistical analysis
Categorical variables were compared by χ2 or Fisher's exact test. Differences between continuous variables were evaluated by two-tailed Student's t-test and by ANOVA with age correction.22 Logarithmic transformation was used to normalise the distributions of BMI, HOMAIR, plasma insulin, total and HDL cholesterol, and triglycerides. Genotype distributions and allele frequencies between the study groups were compared by construction of 2 × 2 and 2 × 3 contingency tables and χ2 analysis. The study was powered to allow detection with 80% and an error rate of 5% for differences in allele frequencies of 13%. Haplotype frequencies were determined by gene counting, as previously described.23 Linkage disequilibrium between the +45T>G and +276G>T SNPs was assessed by calculating the disequilibrium statistics Δ 24 and D′.25 The sign of D′ (positive or negative) depends on the arbitrary choice of the alleles paired at the two loci, and indicates whether the same or opposite allelic association is present.
Analysis of covariance (ANCOVA) and multiple regression analysis were applied in all associations between the genetic variants and clinical or biochemical parameters that were significant in the univariate analysis, controlling each significant association for the other variables. Furthermore, regression analysis was performed to test the interaction between genotypes and BMI on insulin resistance (measured by HOMAIR), dividing the subjects into two groups, according to the median BMI of the sample. To estimate the risk of insulin resistance associated with the adiponectin gene SNPs, odds ratios (ie odds of insulin resistance, given the presence of the variant genotype) were calculated by multiple regression analysis, after adjustment for other modulators known to affect these conditions (including sex, age and BMI). For each odds ratio, we estimated two-tailed P-values and 95% confidence intervals (CI). P-values <0.05 or less were taken as statistically significant. All statistical analyses were performed with SPSS statistical package.
Results
Detection of the adiponectin gene +45T>G and +276G>T SNPs
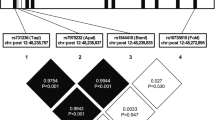
The +45T>G and +276G>T SNPs were detected in real-time PCR with LightCycler hybridisation probes. In each amplification, two controls (one homozygous and one heterozygous subject, kindly provided by Dr Claudia Menzaghi (Scientific Institute Casa Sollievo della Sofferenza, S Giovanni Rotondo, Italy) were added to ensure correct genotyping. In the whole study group, allele frequencies for the +45T>G SNP were 0.86 for the T allele and 0.14 for the G allele, and allele frequencies for the +276G>T SNP were 0.70 for the G allele and 0.30 for the T allele, comparable to previous findings.12 Genotype frequencies for both SNPs were in Hardy–Weinberg equilibrium (data not shown). Estimation of linkage disequilibrium between the two SNPs by the disequilibrium statistics Δ and D′ showed that the two SNPs are in strong linkage disequilibrium (Δ =12.05; D′=−1, both P<0.01). The +45/+276 haplotype frequencies were 0.59 for the ‘TG’, 0.29 for the ‘TT’ and 0.12 for the ‘GG’ haplotypes.
Association between the adiponectin gene SNPs and clinical and metabolic parameters
The clinical and metabolic characteristics of the study subjects were analysed according to genotypes at the +45T>G and +276G>T SNPs (Table 2). No association was found with the +45T>G SNP. The T/T genotype of the +276G>T SNP was associated in univariate analysis with a higher BMI (P<0.01), a higher plasma insulin (P<0.02) and a higher HOMAIR (P<0.02), suggesting a possible effect of the adiponectin gene on parameters of insulin resistance and on body weight. The +45/+276 haplotypes showed identical association as the +276G>T SNP (data not shown); thus results are presented for the single SNPs.
Interaction between adiponectin gene and BMI
In ANCOVA and in regression analysis, the +276G>T SNP did not show an independent role on HOMAIR, with age and BMI being the strongest predictors of it (P<0.01), thus suggesting that a significant proportion of the gene effect on insulin sensitivity is dependent upon changes in body weight. The possible interaction between the adiponectin gene and the BMI status on insulin resistance was therefore tested by subdividing the study group according to individual BMI (ie below or above 26.2 kg/m2, the median value of the entire cohort). As expected, the two subgroups (below or above the median BMI) differed significantly for mean (±SD) blood glucose (75.5±20.1 and 81.4±14.9 mg/dl, respectively, P<001), plasma insulin (7.2±4.5 and 13.2±9.6 μU/ml, P<0.0001), HOMAIR (1.32±1.4 and 2.88±2.3, P<0.0001), HDL (60.8±15.5 and 54.0±13.8 mg/dl, P<0.001) and LDL cholesterol (130.4±30.6 and 137.6±38.4 mg/dl, P<0.03). Genotype and allele frequencies for the +276G>T SNP were significantly different between subjects with BMI above or below the median (lean vs obese subjects: genotypes χ2=6.1, P<0.05; allele frequencies χ2=5.10, P<0.02). The comparison of genotype and allele frequencies for the +45T>G SNP between lean and obese subjects was not significant (data not shown), possibly because of the low prevalence of this SNP in our population.
In both subgroups, subjects carrying the +276 G>T SNP had higher HOMAIR; however, the difference was highly significant among leaner (P<0.001), but not among heavier individuals (Table 3), thus indicating that the BMI status and the adiponectin gene interact in modulating insulin resistance (P for interaction being <0.01). The relative risk of insulin resistance, identified as having an HOMAIR in the top quintile of our sample (>2.99), was 9.7 (95% CI: 1.32–87.7, P<0.035) in the subgroup with a BMI <26.2, after adjusting for age, sex and BMI.
Serum adiponectin level and +276G>T SNP
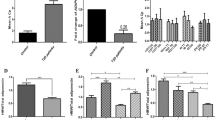
Circulating adiponectin concentration was measured in 67 (out of the 253 recruited) individuals, whose serum was available and who were representative, in terms of age, gender distribution and BMI, of the entire cohort studied (data not shown). The mean serum adiponectin levels were significantly different between the two genotype groups (25.7±13.1 vs 37.0±21.3 ng/ml in +276G>T SNP carriers (n=15) and noncarriers (n=52), respectively, P<0.05), thus suggesting a possible influence of the +276G>T SNP on adiponectin levels.
Discussion
The present study provides evidences of association between a variant in the adiponectin gene and insulin resistance. We found that the +276G>T SNP associates with higher BMI and the insulin-resistance index HOMAIR in our Italian population. In addition, the greater insulin resistance observed among subjects with the at-risk TT genotype was more pronounced in lean compared to obese individuals, thus suggesting that, when obesity is present, the effect of the gene variant is too small to be detected. To our knowledge, this is the first report of such a relationship between adiponectin and BMI in the modulation of insulin sensitivity. One possible limitation in interpreting this finding is that the interaction between the adiponectin gene SNP and BMI could be tested in a relatively small-size population. It should be, therefore, considered provisional, until validated by an independent study in a larger cohort.
How does the +276G>T SNP affect the adiponectin gene function is still an open question. Although a possible effect on gene expression of SNPs with no apparently biological significance cannot be ruled out and, in fact, has been recently reported also for the adiponectin gene,26 it appears more likely that this SNP is in linkage disequilibrium with another mutation either within, or in other genes close to, the adiponectin gene that determines its negative effects. Also in favour of this latter hypothesis is the fact that, in our study, it is the T allele at the +276 SNP that is associated with insulin resistance, whereas, in other reports, the G allele has been associated with features of insulin resistance.12,13 Menzaghi et al12 have previously shown that the +276G>T SNP is in almost complete linkage disequilibrium with an ‘A’ insertion in the 3′ UTR of the adiponectin gene (SNP +2019). 3′ UTR regions are generally recognised to play a central role in the regulation of gene expression,27,28 and the +2019 insertion may disrupt one of the regulatory elements of the adiponectin 3′ UTR region, affecting mRNA processing, translation or degradation. Interestingly, polymorphisms in the 3′ UTR of other genes affecting mRNA stability have recently been associated to insulin resistance.29,30,31 This possibility is also suggested by the observation that two independent studies have shown decreased adiponectin levels associated with genetic variants of the adiponectin gene,12,14 including the +276G>T SNP. Furthermore, the mean serum adiponectin levels measured in a sample of our subjects were lower in carriers of the T allele compared to noncarriers, confirming previous observations12,14 that hypo-adiponectinaemia may be genetically determined.
Adiponectin serum levels have been demonstrated to be reduced in patients with diabetes, obesity and CAD,1,2,3 suggesting a central role in the mechanisms leading to the metabolic abnormalities present in these disorders. A very recent study in Japanese subjects has demonstrated a strong relationship between serum adiponectin and insulin resistance index, lipids and blood pressure,32 showing, as previously described, lower levels of adiponectin in insulin-resistant subjects. Tschritter et al4 have also demonstrated that adiponectin levels predict insulin sensitivity, independently from body mass, and that inter-individual variability is probably genetically determined. In view of our results, it could be speculated that the adiponectin gene variant, or a mutation in linkage disequilibrium with it, determines lower adiponectin gene expression, causing in turn an increased risk to develop insulin resistance. When obesity ensues, other metabolic factors with much stronger effects are at play, decreasing the influence of the genetic variant.
In summary, we have observed an association between the +276G>T SNP in the adiponectin gene and insulin resistance, particularly among leaner individuals. Also, carriers of the T allele had lower serum adiponectin levels. Further studies aimed at screening for polymorphisms the entire adiponectin gene and other candidate genes located in the 3q27 region and subsequent functional studies of the identified SNPs are warranted to understand the molecular mechanisms of the observed association.
References
Weyer C, Funahashi T, Tanaka S et al: Hypoadiponectinemia in obesity and type 2 diabetes: close association with insulin resistance and hyperinsulinemia. J Clin Endocrinol Metab 2001; 86: 1930–1935.
Hotta K, Funahashi T, Arita Y et al: Plasma concentrations of a novel, adipose-specific protein, adiponectin, in type 2 diabetic patients. Arterioscler Thromb Vasc Biol 2000; 20: 1595–1599.
Kumada M, Kihara S, Sumitsuji S et al: Association of hypoadiponectinemia with coronary artery disease in men. Arterioscler Thromb Vasc Biol 2003; 23: 85–89.
Tschritter O, Fritsche A, Thamer C et al: Plasma adiponectin concentrations predict insulin sensitivity of both glucose and lipid metabolism. Diabetes 2003; 52: 239–243.
Lindsay RS, Funahashi T, Hanson RL et al: Adiponectin and development of type 2 diabetes in the Pima Indian population. Lancet 2002; 360: 57–58.
Spranger J, Kroke A, Mohlig M et al: Adiponectin and protection against type 2 diabetes mellitus. Lancet 2003; 361: 226–228.
Maeda N, Takahashi M, Funahashi T et al: PPARgamma ligands increase expression and plasma concentrations of adiponectin, an adipose-derived protein. Diabetes 2001; 50: 2094–2099.
Kissebah AH, Sonnenberg GE, Myklebust J et al: Quantitative trait loci on chromosomes 3 and 17 influence phenotypes of the metabolic syndrome. Proc Natl Acad Sci USA 2000; 97: 14478–14483.
Vionnet N, Hani El-H, Dupont S et al: Genome wide search for type 2 diabetes-susceptibility genes in French whites: evidence for a novel susceptibility locus for early-onset diabetes on chromosome 3q27-qter and independent replication of a type 2-diabetes locus on chromosome 1q21–q24. Am J Hum Genet 2000; 67: 1470–1480.
Kondo H, Shimomura I, Matsukawa Y et al: Association of adiponectin mutation with type 2 diabetes: a candidate gene for the insulin resistance syndrome. Diabetes 2002; 51: 2325–2328.
Hara K, Boutin P, Mori Y et al: Genetic variation in the gene encoding adiponectin is associated with an increased risk of type 2 diabetes in the Japanese population. Diabetes 2002; 51: 536–540.
Menzaghi C, Ercolino T, Di Paola R et al: A haplotype in the adiponectin locus is associated with obesity and other features of the insulin resistance syndrome. Diabetes 2002; 51: 2306–2312.
Stumvoll M, Tschritter O, Fritsche A et al: Association of the T-G polymorphism in adiponectin (exon 2) with obesity and insulin sensitivity: interaction with family history of type 2 diabetes. Diabetes 2002; 51: 37–41.
Vasseur F, Helbecque N, Dina C et al: Single-nucleotide polymorphism haplotypes in the both proximal promoter and exon 3 of the APM1 gene modulate adipocyte-secreted adiponectin hormone levels and contribute to the genetic risk for type 2 diabetes in French Caucasians. Hum Mol Genet 2002; 11: 2607–2614.
Rose GA, Blackburn H : Cardiovascular Survey Methods, 1st edn., No. 56. Geneva (Switzerland): World Health Organization; 1968.
Report of the Expert Committee on the Diagnosis and Classification of Diabetes Mellitus: Diabetes Care 2001; 24 (Suppl 1): S5.
Definition, Diagnosis and Classification of Diabetes Mellitus and its Complications: Report of a WHO Consultation 1999.
Matthews DR, Hosker JP, Rudenski AS, Naylor BA, Treacher DF, Turner RC : Homeostasis model assessment: insulin resistance and beta cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia 1985; 28: 412–419.
Wittwer CT, Ririe KM, Andrew RV, David DA, Gundry RA, Balis UJ : The LightCycler: a microvolume multisample fluorimeter with rapid temperature control. Biotechniques 1997; 22: 176–181.
Ririe KM, Rasmussen RP, Wittwer CT : Product differentiation by analysis of DNA melting curves during the polymerase chain reaction. Anal Biochem 1997; 245: 154–160.
Bernard PS, Pitham GH, Wittwer CT : Color multiplexing hybridization probes using the aploprotein E locus as a model system for genotyping. Anal Biochem 1999; 273: 221–228.
Daly LE, Bourke GJ, McGilvray J : Interpretation and Uses of Medical Statistics, Fifth edition. Oxford: Blackwell Scientific Publications; 2000.
Baroni MG, Alcolado JC, Pozzilli P, Cavallo MG, Li SR, Galton DJ : Polymorphisms at the GLUT2 (beta-cell/liver) glucose transporter gene and non-insulin-dependent diabetes mellitus (NIDDM): analysis in affected pedigree members. Clin Genet 1992; 41: 229–234.
Hill WG, Robertson A : Linkage disequilibrium of finite populations. Theor Appl Genet 1968; 38: 226–231.
Lewontin RC : The interaction of selection and linkage 1. General considerations: heterotic models. Genetics 1984; 49: 49–67.
Yang WS, Tsou PL, Lee WJ et al: Allele-specific differential expression of a common adiponectin gene polymorphism related to obesity. J Mol Med 2003, May 16 (Epub ahead of print).
Jupe ER, Badgett AA, Neas BR et al: Single nucleotide polymorphism in prohibitin 39 untranslated region and breast-cancer susceptibility. Lancet 2001; 357: 1588–1589.
Wong PM, Yuan Q, Chen H, Sultzer BM, Chung SW : A single point mutation at the 3′-UTR of Ran mRNA leads to profound changes in lipopolysaccharide endotoxin-mediated responses. J Biol Chem 2001; 276: 33129–33138.
Xia JB, Sherer SW, Cohen PT et al: A common variant in PPP1R3 associated with insulin resistance and type 2 diabetes. Diabetes 1998; 47: 1519–1524.
Frittitta L, Erccolino T, Bozzali M et al: A cluster of 3 single nucleotide polymorphisms in the 3′-untranslated region of human glycoprotein PC-1 gene stabilizes PC-1 mRNA and associates with increased PC-1 protein content and insulin resistance related abnormalities. Diabetes 2001; 50: 1952–1955.
Di Paola R, Frittitta L, Miscio G et al: A variation in 3′ UTR of hPTP1B increases specific gene expression and associates with insulin resistance. Am J Hum Genet 2002; 70: 806–812.
Yamamoto Y, Hirose H, Saito I et al: Correlation of the adipocyte-derived protein adiponectin with insulin resistance index and serum high-density lipoprotein-cholesterol, independent of body mass index, in the Japanese population. Clin Sci (Lond) 2002; 103: 137–142.
Acknowledgements
Financial support to this work was provided by the Faculty of Medicine (grant no. C26F022871) of the University of Rome ‘La Sapienza’, by the Ministry of the University and Scientific Research (MURST grant no. 2002065949–007) and by the Ministry of Health (grant RF2002), all to MG Baroni. We thank Drs Claudia Menzaghi (Osp. Casa Sollievo della Sofferenza, S. Giovanni Rotondo, Italy) and Alessandro Doria (Joslin Diabetes Center, Boston, USA) for comments and suggestions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Filippi, E., Sentinelli, F., Trischitta, V. et al. Association of the human adiponectin gene and insulin resistance. Eur J Hum Genet 12, 199–205 (2004). https://doi.org/10.1038/sj.ejhg.5201120
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201120
Keywords
This article is cited by
-
Associations between adipokines gene polymorphisms and knee osteoarthritis: a meta-analysis
BMC Musculoskeletal Disorders (2022)
-
The Association of SNP276G>T at Adiponectin Gene with Insulin Resistance and Circulating Adiponectin in Morbid Obese Patients After a Biliopancreatic Diversion Surgery
Obesity Surgery (2017)
-
Polymorphisms of the adiponectin gene in gestational hypertension and pre-eclampsia
Journal of Human Hypertension (2014)
-
Meta-analysis of the association of ADIPOQ G276T polymorphism with insulin resistance and blood glucose
Endocrine (2014)
-
Adiponectin: an adipocyte-derived hormone, and its gene encoding in children with chronic kidney disease
BMC Research Notes (2012)