Abstract
Reproduction and aging evolved to be intimately associated. Experimental selection for early-life reproduction drives the evolution of decreased longevity in Drosophila whereas experimental selection for increased longevity leads to changes in reproduction. Although life history theory offers hypotheses to explain these relationships, the genetic architecture and molecular mechanisms underlying reproduction–longevity associations remain a matter of debate. Here we show that mating triggers accelerated mortality in males and identify hundreds of genes that are modulated upon mating in the fruit fly Drosophila melanogaster. Interrogation of genome-wide gene expression in virgin and recently mated males revealed coherent responses, with biological processes that are upregulated (testis-specific gene expression) or downregulated (metabolism and mitochondria-related functions) upon mating. Furthermore, using a panel of genotypes from the Drosophila Synthetic Population Resource (DSPR) as a source of naturally occurring genetic perturbation, we uncover abundant variation in longevity and reproduction-induced mortality among genotypes. Genotypes displayed more than fourfold variation in longevity and reproduction-induced mortality that can be traced to variation in specific segments of the genome. The data reveal individual variation in sensitivity to reproduction and physiological processes that are enhanced and suppressed upon mating. These results raise the prospect that variation in longevity and age-related traits could be traced to processes that coordinate germline and somatic function.
Similar content being viewed by others
Introduction
Reproduction and longevity evolved to be intimately associated, with life history theory providing hypotheses to explain why a tight relationship between reproduction and rate of aging is expected (Stearns, 1992; Falconer and Mackay, 1996; Stearns et al., 2000). Studies of laboratory evolution have shown that selection on the age of reproduction can lead to a correlated response in longevity, an observation that is in accord with predictions of the evolutionary theory of aging (Rose and Charlesworth, 1981; Stearns et al., 2000). Similarly, direct selection for increased longevity can result in correlated changes in reproduction (Rose and Charlesworth, 1981; Stearns et al., 2000). Furthermore, experimental studies have shown that removal of the germline cells in the nematode Caenorhabditis elegans substantially extends lifespan, and identified candidate genes modulating the relationship (Hsin and Kenyon, 1999; Arantes-Oliveira et al., 2002; Kenyon, 2010). However, longevity outcomes emerging from perturbing specific genes or tissues can be surprising. For instance, in C. elegans, the daf-2 gene encodes for the insulin-like growth factor-1 and the severity of its knockdown can either weaken or strengthen the effect of reproduction on aging (Kenyon, 2010). Furthermore, in worms, removal of the somatic gonad compensates for removal of the germline (Kenyon, 2010). In mammals, placing the reproductive tissues of young female rodents into older females results in increased longevity (Cargill et al., 2003; Mason et al., 2011).
Evidently, the coordinated modulation of reproduction and lifespan is remarkably complex (Flatt et al., 2008; Ghazi et al., 2009; Tatar, 2010; Flatt, 2011; McCormick et al., 2011). Mechanisms through which male–female interactions affect mortality and longevity are varied, and include: (1) a direct role of the germline in somatic aging (Hsin and Kenyon, 1999; Kirkwood and Austad, 2000; Arantes-Oliveira et al., 2002; Cargill et al., 2003; Ghazi et al., 2009; Austad, 2010; Kenyon, 2010; Tatar, 2010; Mason et al., 2011; McCormick et al., 2011; Gendron et al., 2014; Maures et al., 2014; Shi and Murphy, 2014, 2) male and female harm to each other during copulation (Chapman et al., 1995; Wolfner, 2009, 3) costs to gamete production (Sirot et al., 2011; Dowling and Simmons, 2012) and (4) observations of diffusible sex-harming factors transmitted between the sexes and between individuals of the same sex (Gendron et al., 2014; Maures et al., 2014; Shi and Murphy, 2014). These costs can themselves be modulated with age. For instance, female attributes and male age have been implicated in the modulation of sperm production costs (Sirot et al., 2011; Dowling and Simmons, 2012). Nevertheless, naturally occurring genetic variation in males and in females might limit the generality of studies that address molecular mechanisms of reproduction and longevity within a single genotype. Thus, although much progress has been made in understanding the feedbacks between germline and somatic tissues during aging, it is not surprising that identifying the underlying genetic factors and molecular mechanisms that mediate the relationship has remained a challenge (Kirkwood and Austad, 2000; Austad, 2010; Kenyon, 2010; Tatar, 2010).
Natural genetic variation is a rich source of perturbation that modifies the expressivity and penetrance of traits. For instance, natural variation can even affect canonical pathways and mutations that have been abundantly studied, as is the case of the scalloped mutation in Drosophila melanogaster (Dworkin et al., 2009; Chari and Dworkin, 2013). Diversity panels such as the Drosophila Genetic Reference Panel (DGRP) and the Drosophila Synthetic Population Resource (DSPR) harnessed naturally occurring variation to identify candidate segments contributing to phenotypes as varied as starvation stress, body weight and resistance to cold stress (Mackay et al., 2012) as well as modifiers of alcohol dehydrogenase activity (King et al., 2012a, 2012b) and genome-wide gene expression (King et al., 2014). Furthermore, within- and between-population variation in male-induced costs of mating to females have been well documented in Drosophila (Fiumera et al., 2006; Reinhart et al., 2015; Filice and Long, 2016). Hence, natural variation in reproduction-induced male mortality and aging might similarly be expected to be abundant in genetically diverse panels.
Here we show that mating triggers genome-wide gene expression responses in Drosophila males and induces increased post-reproduction mortality. Investigation of genome-wide patterns of gene expression in virgin and recently mated males revealed genes that are modulated upon mating and provide insights into biological processes that are enhanced or suppressed by mating. Furthermore, using a panel of genotypes (strains) from the DSPR as a source of naturally occurring genetic perturbation (King et al., 2012a, 2012b), we uncover abundant variation in reproduction-induced aging across genotypes and identify candidate genomic segments harboring loci that are likely to mediate the impact of reproduction on male aging. We conclude that genotypes from the DSPR panel harbor substantial variation for longevity and reproduction-induced male aging that can be harnessed in quantitative trait locus (QTL) analyses.
Materials and methods
Gene expression assays
Global gene expression levels were compared before and after reproduction for males of a selected genotype (BL4361). Virgin males (35 individuals per bottle) were collected within 7 h of eclosion and aged without females. At 33 days of age, males were transferred to bottles with 45 virgin females for 2 days. On day 35, males were isolated, transferred to fresh food and set to rest for 4 h before being flash frozen. Dead males were excluded from the control and treatment groups before flash freezing. Virgin control males were subjected to the same procedure but were placed in bottles without females. We contrasted global gene expression between virgin and recently mated males at 35 days of age. Microarrays were ~18 000 feature arrays spotted with D. melanogaster complementary DNA PCR products. Total RNA was extracted from whole flies using TRIzol (Life Technologies, Carlsbad, CA, USA). Synthesis and labeling of complementary DNA with fluorescent dyes (Cy3 and Cy5) and hybridization were carried out using 3DNA (Genisphere, Hatfield, PA, USA) reagents and protocols. Four replicates of each sample were hybridized and scanned using an Axon 400B scanner (Axon Instruments, Union City, CA, USA) and processed with GenePix Pro 6.0 software (Union City, CA, USA). Gene expression data can be obtained from GSE86345.
Drosophila strains and lifespan assays
DSPR strains were kindly obtained from Stuart Macdonald (University of Kansas, Kansas City, MO, USA). From the DSPR population B (pB) set, we randomly selected 71 strains for analyses. The strains were previously genotyped (King et al., 2012a, 2012b). We scored male mortality daily in duplicate within each genotype. Accordingly, males were collected as virgins (within 7 h of eclosion) under light CO2 anesthesia. Flies were aged in cohorts of 45 males per bottle (~177 ml), and each group was transferred to a new bottle containing fresh food every 2 days. On day 21, males of the treatment set were subject to CO2 anesthesia and transferred to bottles with ~90 young virgin females of a common genotype for 48 h. The control set was similarly subjected to the CO2 anesthesia and transferred to new bottles that did not contain females. After this 48 h exposure to females, treated males were moved (day 23) to a new bottle with fresh food and scored daily for survival; all males are expected to mate at least once but probably several times during the 48 h interval. Control males were similarly transferred to a fresh bottle on day 23 and scored daily for survival. This was a labor-intensive assay requiring collection and manipulation of >15 000 virgin male and female flies. Hence, we conducted it sequentially in four batches that were run with ∼18 genotypes in each. We did not observe significant differences in pre- and post-exposure mortality between batches (for example, P=0.40 for batch effect at day 21; P=0.41 at day 23; P=0.87 at day 25; P=0.44 at day 27; analysis of variance (ANOVA)). Flies were maintained at 23 °C, with standard diet (Bloomington recipe), ~60% humidity and a 16 h/8 h light/dark cycle throughout the experiment. These choices were arbitrary. Our design does not separate costs because of physical harm from mating, courting, post-mating events or male–female interactions during mating through diffusible factors.
Statistical analyses
Analyses of survival curves for treatment and control populations were conducted with the survival package (R Development Core Team, 2011); the Cox proportional hazards model was fit with treatment as fixed effect. Kaplan–Meier survival curves were estimated for the average survival across all strains; P-values were also calculated using a log-rank test. Test of proportional hazards was conducted with Schoenfeld’s residuals using the cox.zph function. There is no evidence that the proportional hazard assumption is violated (ρ=0.13, χ2=1.72, P-value=0.19). In addition, differences in survivorship between treatment and control groups at specific ages as well as batch and genotype effects were assessed with mixed linear models using percent mortality as the response variable.
For the QTL analyses, we used the R/DSPRqtl analytical software developed by the DSPR team (King et al., 2012a, 2012b). Briefly, the DSPR team genotyped the recombinant inbred lines, and derived eight probabilities for the presence of one of the eight founder genotypes along the chromosomes of each recombinant inbred line. Multiple regressions of the eight additive probabilities on the mean mortality metric were calculated with no covariates for the pB population. The resulting F-statistic was converted to a LOD (logarithm (base 10) of odds) score. Significance was estimated with 1000 permutations.
For the analyses of gene expression, we used stringent quality control criteria to ensure reliable foreground intensity estimates for both Cy3 and Cy5 channels (Branco et al., 2013). Foreground fluorescence of dye intensities was normalized by the Loess method in Bioconductor/Limma (Smyth and Speed, 2003; Smyth, 2005). Statistical significance of gene expression variation was assessed with both Bayesian models in BAGEL and with linear models in Limma (Townsend and Hartl, 2002; Smyth, 2004). Results are nearly identical with both methods, as we observed previously (Branco et al., 2013). False discovery rates were empirically estimated by permutation of the data set. Tissue specificity of differentially expressed genes was assessed as previously described (Branco et al., 2013) using the FlyAtlas resource (Chintapalli et al., 2007). Discovery of gene ontology enrichments were done with the FlyMine platform and enrichment P-values corrected with Holm–Bonferroni (Lyne et al., 2007).
Results
Mating induces upregulation of testis-specific gene expression
To gain insights into functional responses to mating and assist in identifying processes relevant for accelerated mortality after reproductive activity, we profiled genome-wide gene expression immediately after the pulse of reproduction. To that end, we identified a genotype that displays good longevity as virgins (~74% survivorship at age 33) and acute response to the 48 h mating treatment with a threefold higher mortality in the treatment group relative to control. Virgin 33-day-old males were exposed to females for 48 h, and flash frozen at day 35; dead males were excluded from the control and treatment groups before flash freezing. We observed acute gene expression responses to the treatment with 681 genes induced after mating (Bayesian posterior probability >0.95, false discovery rate <10%). Genes upregulated are significantly enriched for targets displaying testis-specific expression (Figures 1 and 2a; P=0.001, Fisher’s exact test; odds ratio=1.48; 95% confidence interval=1.16–1.86), whereas the set of genes downregulated after mating displays a deficit of testis-specific genes (P=2.2e−16, Fisher’s exact test; odds ratio=0.16; 95% confidence interval=0.08–0.28). The set of upregulated genes also includes a strong enrichment for Chaperonin-containing T-complex (GO:0005832; 7 genes, P=3.40e−7), a complex that is known to be relevant for spermatogenesis (Soues et al., 2003; Dun et al., 2011). For instance, this category includes the genes T-cp1zeta, T-complex chaperonin 5, sarah and CCT-γ that are involved in meiosis and unfolded protein response. Analysis of the KEGG (Kyoto Encyclopedia of Genes and Genomes)/Reactome databases for the set of upregulated genes further reinforces the pattern with significant enrichment in the pathway ‘formation of tubulin folding intermediates by CCT/TriC’ (12 genes; P=2.10e−12). The upregulated set also displays significant enrichments in the classes of ‘response to stimulus’ (175 genes, P=0.8.10e−4), ‘response to biotic stimulus’ (35 genes, P=0.001) and ‘response to stress’ (GO:0006850, 78 genes, P=0.002).
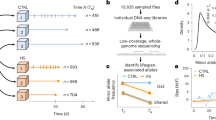
Mating triggers upregulation of testis-specific genes and downregulation of broadly expressed metabolic genes. Number of differentially expressed genes that are upregulated (yellow) and downregulated (blue) in each category. The excess of downregulated or upregulated genes is significant for all categories (P<0.001, χ2 test in each category). A full color version of this figure is available at the Heredity journal online.
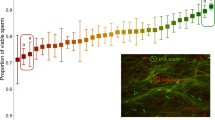
Heat maps with differentially expressed genes from selected categories modulated in recently mated males. (a) Testis-specific genes. (b) Genes whose protein products localize to the mitochondria. Contrast between two treatments: virgin males and recently mated males. Scale reflects values of fold change relative to 1. For each gene, the expression in the treatment with the lower expression level was set to 1. A full color version of this figure is available at the Heredity journal online.
Mating induces downregulation of metabolic processes
On the other hand, we observed 688 genes repressed after mating (Bayesian posterior probability >0.95, false discovery rate <10%). The genes downregulated with mating are broadly expressed and display highly significant enrichment in candidates belonging to ‘metabolic processes’ (364 genes, P=4.85e−10; GO:0008152), including ‘enrichment in cellular amino acid metabolic process’ (32 genes, P=4.07e−5; GO:0006520), ‘organonitrogen compound metabolic process’ (65 genes, P=0.02; GO:1901564) and ‘oxidation-reduction process’ (90 genes, P=2.18e−22; GO:0055114). The gene candidates display, moreover, significant localization to the mitochondrion (56 genes, P=4.62e−5; GO:0005739) and lipid particles (32 genes, P=1.32e−6; GO:0005811). Furthermore, in accordance with these observations, the proteins whose mRNA precursors are downregulated with mating contain NAD(P)-binding domain (29 genes, P=5.43e−5, IPR016040). Analysis of the KEGG and Reactome databases further reinforce the pattern, with significant enrichments in downregulated genes belonging to ‘metabolic pathways’ (112 genes, P=1.50e−4) and in pathways of ‘amino acid and derivative metabolism’ (P=9.48e−4), ‘fatty acid metabolism’ (13 genes, P=2.49e−5) and ‘glycolysis’ (15 genes, P=2.87e−4). Finally, we observed a greater number of downregulated than upregulated genes among differentially expressed candidates involved in oxidation-reduction processes, amino acid metabolism and localized to the mitochondrion and lipid particles (Figures 1 and 2b). The shifts to repression are statistically significant within each functional category (P<0.01, Fisher’s exact test).
Reproduction accelerates male mortality in Drosophila
The DSPR pB panel consists of recombinant inbred lines generated with a round-robin design of 8 wild-type founder lines. The 8 donor lines were collected from distinct localities to capture a broad range of natural allelic diversity. Using this resource, we first addressed the extent of natural genetic variation for the rate of aging in 71 genotypes (strains) of the DSPR panel. The data revealed large variation in longevity across genotypes, ranging from 18 to 97% mortality at 21 days of age (P=8.22e−08 for genotype effect; ANOVA; Figure 3a). These results are congruent with a recent study uncovering similarly high variability in longevity among 40 strains of the DGRP (Mackay et al., 2012). Thus, in order to investigate the genetic determinants underpinning the association between reproduction and mortality, we tested whether a 48 h pulse of reproduction in early-to-mid life (at days 21–23) could trigger accelerated mortality across genotypes of the DSPR panel (Figure 3b). As expected, there were no differences between control and treatment populations immediately before exposure to females at day 21 (P=0.78 for treatment effect at day 21, ANOVA). Similarly, there were no differences in male mortality between the control and treatment populations of each genotype immediately following the mating treatment (P=0.13, for treatment effect at day 23, ANOVA). This suggests that the experimental manipulation did not disproportionally affect treatment or control males and that exposure to females did not immediately cause a difference in male survival. Nevertheless, exposure to females triggered faster mortality rates in males resulting in significant differences in survival curves (Figures 3b and 4; Cox hazard ratio=2.15, P=0.001; Log rank test score=11.16, P=0.0008), with statistically significant survival differences between treatment and control populations starting to emerge at age 25 days (P=0.015, ANOVA for the treatment effect on day 25; P=0.0568, Mann–Whitney); the difference remains statistically significant throughout the experiment and peaks at age 49 days (P<0.0001, ANOVA for the treatment effect on day 49; 19% difference in mortality between treatment and control populations across all genotypes). Collectively, these observations point to substantial variation in longevity across the DSPR and indicate that reproduction triggers accelerated mortality across diverse genotypes.
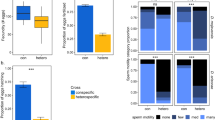
Longevity variation in virgin and mated males across a genetically diverse panel. (a) Extensive variation in the longevity of virgin males across genotypes of the DSPR. Each line represents a genotype. (b) Extensive variation in reproduction-induced male mortality across genotypes of the DSPR. Each line represents a genotype. Virgin males (21 days old) were exposed to females for 48 h. Both control (a) and treatment (b) males were handled identically, except that bottles from the control group did not contain females during the 48 h exposure interval. A full color version of this figure is available at the Heredity journal online.
Mating induces accelerated mortality in Drosophila male genotypes. Shown are survival curves and 95% confidence bands. Black line represents longevity profiles of virgin controls. Red line represents longevity profiles of mated treatment. Confidence bands estimated with variation across genotypes. The treatment consists of a 48 h female exposure to virgin males (days 21–23). A full color version of this figure is available at the Heredity journal online.
Identifying determinants of longevity and reproduction-induced mortality
QTLs modulating aging and reproduction-induced mortality in males could be detectable in the DSPR panel. To address the possibility we first performed QTL analyses for longevity in the DSPR (King et al., 2012a, 2012b) using survivorship at 21 days of age as the phenotype and R scripts with default parameters. The analysis identified a single peak with LOD score above 6. This 60 kb X-linked segment with higher probability of containing genetic factors that contribute to variation in aging among genotypes (Figure 5) harbors 7 protein-coding genes. One of these genes is the protein-coding gene hemipterous (hep) that encodes a protein kinase that has long been known to affect aging in Drosophila (Seong et al., 2001; Wang et al., 2003, 2005; Paaby and Schmidt, 2009; Omelyanchuk et al., 2015). In view of this observation we proceeded to investigate the association between reproduction-induced male mortality and genetic variation in the DSPR. We used the difference in mortality between control virgin males and matched treatment males from the same genotype as a phenotype. In agreement with our expectations, we observed substantial variation in postreproductive mortality across genotypes. To illustrate, we measured the difference in survivorship between treatments and controls for each of the 71 genotypes at day 49. The data showed 50 strains with lower survivorship in the treatment relative to controls, 16 strains with identical survivorship in both groups and 5 strains with lower survivorship in the control group. This represents a significant bias toward genotypes with greater mortality rate post reproduction relative to the mortality in virgin controls (P<0.001, Fisher’s exact test). In addition, all five strains with higher mortality in the control were within 10% of the mortality observed in the treatment populations with reproduction, suggesting this might represent stochastic variation. On the other hand, 15 strains displayed mortality that was >40% higher in the reproduction regime relative to virgin controls. Similar patterns were replicated at 33 days and 35 days of age. Collectively, these observations point to variation in reproduction-induced mortality across genotypes that could be traceable to individual loci in the DSPR. Hence, QTL analyses for reproduction-induced male mortality in the DSPR suggest candidate genomic segments; the candidate segment with the highest LOD score contains six protein-coding genes (Figure 6), namely Approximated (app), CG32100, Porphobilinogen synthase (Pbgs), CG4300, CG10426 and CG4328; only one gene (app) is differentially expressed in recently mated males. Another segment emerging with higher likelihood of harboring relevant variation is at chromosome 2R: this is a broad 280 kb segment (from gene CG3045 to gene CG10384) containing >20 protein-coding genes and lacking a sharply defined peak. The highest peak in the interval defined a 20 kb intergenic region between genes pickpocket 12 (ppk12) and CG10384. The second highest peak in the interval defines a 20 kb segment overlapping the 3′ end of the gene defective proventriculus (dve).
Genome scan for genetic determinants of longevity. The trait analyzed is variation in survivorship at 21 days of age in a set of 71 strains from DSPR pB. Chromosomes and chromosome location are represented along the x axis. LOD scores across 10 000 intervals along the Drosophila genome. The LOD line is set at 6. The segment with the highest LOD is displayed above the profile. The analyses identified a peak containing the hep gene, a candidate that is known to modulate aging in Drosophila.
Genome scan for genetic determinants of reproduction-induced male mortality. The trait analyzed is variation in the difference in survivorship between the control and treatment groups at 33 days of age in a set of 71 strains from DSPR pB. Chromosomes and chromosome location are represented along the x axis. LOD scores across 10 000 intervals along the Drosophila genome. The LOD line is set at 6. The segment with the highest LOD is displayed above the profile.
Discussion
The consequences of reproduction and germline tissues on organismal longevity are evolutionarily conserved (Rose and Charlesworth, 1981; Stearns, 1992; Hsin and Kenyon, 1999; Stearns et al., 2000; Flatt et al., 2008; Kenyon, 2010). Here we observed that reproductive activity triggers gene expression changes in Drosophila males and induces decreased post-reproductive longevity. Specifically, our observations indicate that mating induces the expression of testis-specific genes, whereas it represses broadly expressed genes involved in metabolic processes and whose protein products are localized to the mitochondria; changes in gene expression are accompanied by accelerated male mortality post reproduction. Furthermore, analyses of longevity and reproduction-induced male mortality across genotypes indicate that the relationship between reproduction and longevity is not equally manifested across all genotypes. The observations point to genomic segments displaying higher likelihood of harboring genetic factors contributing to variation in male sensitivity to reproduction. The segments contain protein coding genes and microRNAs that are best candidates for further analyses aiming at pinpointing the identity of loci mediating the phenotype. Although analysis of variation in longevity among lines pointed to a segment containing the gene hep, a known factor associated with aging in Drosophila (Seong et al., 2001; Wang et al., 2003, 2005; Paaby and Schmidt, 2009; Omelyanchuk et al., 2015), the genes residing in the segments associated with accelerated mortality post-reproduction are less amenable to functional interpretations.
Male–female interactions can be experimentally modeled through a number of designs that themselves make the contribution of copulation, courting, diffusible compounds, ejaculate composition and/or other attributes more or less salient. Accordingly, in Mediterranean flies, Ceratitis capitata, costs of male courtship were higher than reproduction-induced costs because of sperm production or mating (Papadopoulos et al., 2010). On the other hand, data from Drosophila and crickets point to their ability to modulate the composition of the ejaculate (Sirot et al., 2011; Dowling and Simmons, 2012), and potentially alter the allocation of reproductive costs. Nevertheless, observations of gamete production costs would need to be reconciled with data from worms indicating that removal of the somatic gonad compensates for removal of the germline (Kenyon, 2010). The molecular pathways underlying how the compensation occurs have emerged in the context of insulin and steroid hormone signaling (Hsin and Kenyon, 1999; Flatt et al., 2008; Yamawaki et al., 2010); the observations highlight physiological complexity beyond simple costs of gamete production. Indeed, several studies have raised the expectation that interactions between age, reproduction and nutritional status might operate in Drosophila (Sgro and Partridge, 1999; Tu and Tatar, 2003; Toivonen and Partridge, 2009), with results indicating a key role of the insulin pathway in females. Nevertheless, alternative pathways have also been suggested to play a role in modulating reproduction–longevity relationships. For instance, evidence for tradeoffs between protein quality control in somatic tissues and eggs has recently emerged in Drosophila (Fredriksson et al., 2012). It is not far-fetched to imagine that similar tradeoffs might also operate in Drosophila males, in which protein quality control could be essential for the progression of the carefully orchestrated events that occur during spermatogenesis long after most gene transcription had been shut off. Hence, detailed molecular characterization of a variety of genomic attributes coupled with larger analyses of reproduction-induced male mortality that simultaneously consider the role of diet, age, reproduction and genotype could yield essential baseline data.
Furthermore, the data presented here as well as previous observations linking reproduction and longevity point to events in reproductive and somatic tissues, with downregulation of metabolic genes and genes whose protein products localize to the mitochondria. This observation is concordant with functional hypotheses about the role of metabolism (McCormick et al., 2011), insulin signaling (Flatt et al., 2008) and steroid hormones (Yamawaki et al., 2010) on reproduction–longevity associations, and could aid in narrowing the candidate genes or processes for further analyses. Indeed, our observations are in agreement with suggestions that germline removal promotes longevity in part by modulating insulin signaling and lipid metabolism (Hansen et al., 2013), and with expectations that naturally occurring variation in mitochondrial function and metabolism specifically affects male aging (Zeh and Zeh, 2005; Camus et al., 2012). Moreover, mitochondrial–nuclear genetic interactions tied to aging are likely to include nuclear-encoded genes associated with reproduction-induced mortality (Tanaka et al., 1998; De Benedictis et al., 1999; Rand et al., 2006; Clancy, 2008; Zhu et al., 2014). A recent study found that interactions between genes spanning the mitochondrial and nuclear genomes were associated with components of male reproductive aging via direct effects on ejaculate weight and indirect effects on the size of eggs produced by females (Immonen et al., 2016). Interestingly, mitochondrial DNA variation in Drosophila preferentially affects genes that are exclusively expressed in the testis (Innocenti et al., 2011). When probed in engineered genotypes that differ only in their mitochondrial DNA haplotype, males display significantly higher genetic variance for longevity than females (Camus et al., 2012; Dowling, 2014; Wolff et al., 2014). Variation in Y-linked heterochromatin is also known to modulate the expression of genes exclusively transcribed in reproductive tissues as well as genes associated with mitochondrial function in Drosophila (Lemos et al., 2008, 2010). The Y-linked variation is, moreover, conditional on the genetic background (Chippindale and Rice, 2001), differentially manifested in somatic and reproductive tissues (Branco et al., 2013) and recently implicated in aging (Griffin et al., 2015). In view of these, an interesting possibility is that genetic elements displaying sex-biased occurrence or transmission, such as the Y chromosome or the mitochondrial DNA, might evolve interactions that contribute to the modulation of reproduction-induced mortality.
In conclusion, genomic analyses and natural genetic variation can serve as platforms to identify molecular pathways in reproduction-induced male mortality. Although the effect size of naturally occurring alleles is typically lesser than the effect size of alleles used in mechanistic studies (for example, loss-of-function mutations), the data highlight the relevance of genetic background in studies of reproduction-induced male mortality and longevity. The analyses reveal strains in which specific mechanisms of reproduction-induced male mortality might be manifested with greater or lesser salience. Although the phenotypic characterization of a greater number of strains is labor intensive (the DSPR panel contains >1000 genotypes), efforts have the benefit of being agnostic and would be worthwhile in view of the scarcity of expectations about the likely functions of loci mediating the association between reproduction and longevity. On the other hand, analyses using targeted loss-of-function mutations or sequential deletions in the candidate segments could be efficient to zero in on specific genetic elements within these regions. Whether gene expression responses to mating and reproduction-induced male mortality differ during aging or display genotype by age interaction remains to be determined. Collectively, our results further raise the prospect that determinants of longevity and age-related phenotypes might be traced to genetic variation and molecular processes affecting the germline, and potentially intersecting with pathways modulated by the mitochondria and the Y chromosome. Measuring the various layers of regulatory control that are expected to have evolved to coordinate functional processes in reproductive and somatic tissues during aging is likely to remain an important challenge for many years to come.
Data archiving
Data available from the Dryad Digital Repository: http://dx.doi.org/10.5061/dryad.74940.
References
Arantes-Oliveira N, Apfeld J, Dillin A, Kenyon C . (2002). Regulation of life-span by germ-line stem cells in Caenorhabditis elegans. Science 295: 502–505.
Austad SN . (2010). Animal models of reproductive aging: what can they tell us? Ann NY Acad Sci 1204: 123–126.
Branco AT, Hartl DL, Lemos B . (2013). Chromatin-associated proteins HP1 and Mod(mdg4) modify Y-linked regulatory variation in the drosophila testis. Genetics 194: 609–618.
Camus MF, Clancy DJ, Dowling DK . (2012). Mitochondria, maternal inheritance, and male aging. Curr Biol 22: 1717–1721.
Cargill SL, Carey JR, Muller HG, Anderson G . (2003). Age of ovary determines remaining life expectancy in old ovariectomized mice. Aging Cell 2: 185–190.
Chapman T, Liddle LF, Kalb JM, Wolfner MF, Partridge L . (1995). Cost of mating in Drosophila melanogaster females is mediated by male accessory gland products. Nature 373: 241–244.
Chari S, Dworkin I . (2013). The conditional nature of genetic interactions: the consequences of wild-type backgrounds on mutational interactions in a genome-wide modifier screen. PLoS Genet 9: e1003661.
Chintapalli VR, Wang J, Dow JA . (2007). Using FlyAtlas to identify better Drosophila melanogaster models of human disease. Nat Genet 39: 715–720.
Chippindale AK, Rice WR . (2001). Y chromosome polymorphism is a strong determinant of male fitness in Drosophila melanogaster. Proc Natl Acad Sci USA 98: 5677–5682.
Clancy DJ . (2008). Variation in mitochondrial genotype has substantial lifespan effects which may be modulated by nuclear background. Aging Cell 7: 795–804.
De Benedictis G, Rose G, Carrieri G, De Luca M, Falcone E, Passarino G et al. (1999). Mitochondrial DNA inherited variants are associated with successful aging and longevity in humans. FASEB J 13: 1532–1536.
Dowling DK . (2014). Evolutionary perspectives on the links between mitochondrial genotype and disease phenotype. Biochim Biophys Acta 1840: 1393–1403.
Dowling DK, Simmons LW . (2012). Ejaculate economics: testing the effects of male sexual history on the trade-off between sperm and immune function in Australian crickets. PLoS One 7: e30172.
Dun MD, Smith ND, Baker MA, Lin M, Aitken RJ, Nixon B . (2011). The chaperonin containing TCP1 complex (CCT/TRiC) is involved in mediating sperm-oocyte interaction. J Biol Chem 286: 36875–36887.
Dworkin I, Kennerly E, Tack D, Hutchinson J, Brown J, Mahaffey J et al. (2009). Genomic consequences of background effects on scalloped mutant expressivity in the wing of Drosophila melanogaster. Genetics 181: 1065–1076.
Falconer DS, Mackay TFC . (1996) Introduction to Quantitative Genetics. Benjamin Cummings: England.
Filice DC, Long TA . (2016). Genetic variation in male-induced harm in Drosophila melanogaster. Biol Lett 12 (pii): 20160105.
Fiumera AC, Dumont BL, Clark AG . (2006). Natural variation in male-induced 'cost-of-mating' and allele-specific association with male reproductive genes in Drosophila melanogaster. Philos Trans R Soc Lond B Biol Sci 361: 355–361.
Flatt T . (2011). Survival costs of reproduction in Drosophila. Exp Gerontol 46: 369–375.
Flatt T, Min KJ, D'Alterio C, Villa-Cuesta E, Cumbers J, Lehmann R et al. (2008). Drosophila germ-line modulation of insulin signaling and lifespan. Proc Natl Acad Sci USA 105: 6368–6373.
Fredriksson A, Johansson Krogh E, Hernebring M, Pettersson E, Javadi A, Almstedt A et al. (2012). Effects of aging and reproduction on protein quality control in soma and gametes of Drosophila melanogaster. Aging Cell 11: 634–643.
Gendron CM, Kuo TH, Harvanek ZM, Chung BY, Yew JY, Dierick HA et al. (2014). Drosophila life span and physiology are modulated by sexual perception and reward. Science 343: 544–548.
Ghazi A, Henis-Korenblit S, Kenyon C . (2009). A transcription elongation factor that links signals from the reproductive system to lifespan extension in Caenorhabditis elegans. PLoS Genet 5: e1000639.
Griffin RM, Le Gall D, Schielzeth H, Friberg U . (2015). Within-population Y-linked genetic variation for lifespan in Drosophila melanogaster. J Evol Biol 28: 1940–1947.
Hansen M, Flatt T, Aguilaniu H . (2013). Reproduction, fat metabolism, and life span: what is the connection? Cell Metab 17: 10–19.
Hsin H, Kenyon C . (1999). Signals from the reproductive system regulate the lifespan of C. elegans. Nature 399: 362–366.
Immonen E, Collet M, Goenaga J, Arnqvist G . (2016). Direct and indirect genetic effects of sex-specific mitonuclear epistasis on reproductive ageing. Heredity (Edinb) 116: 338–347.
Innocenti P, Morrow EH, Dowling DK . (2011). Experimental evidence supports a sex-specific selective sieve in mitochondrial genome evolution. Science 332: 845–848.
Kenyon C . (2010). A pathway that links reproductive status to lifespan in Caenorhabditis elegans. Ann NY Acad Sci 1204: 156–162.
King EG, Macdonald SJ, Long AD . (2012a). Properties and power of the Drosophila Synthetic Population Resource for the routine dissection of complex traits. Genetics 191: 935–949.
King EG, Merkes CM, McNeil CL, Hoofer SR, Sen S, Broman KW et al. (2012b). Genetic dissection of a model complex trait using the Drosophila Synthetic Population Resource. Genome Res 22: 1558–1566.
King EG, Sanderson BJ, McNeil CL, Long AD, Macdonald SJ . (2014). Genetic dissection of the Drosophila melanogaster female head transcriptome reveals widespread allelic heterogeneity. PLoS Genet 10: e1004322.
Kirkwood TB, Austad SN . (2000). Why do we age? Nature 408: 233–238.
Lemos B, Araripe LO, Hartl DL . (2008). Polymorphic Y chromosomes harbor cryptic variation with manifold functional consequences. Science 319: 91–93.
Lemos B, Branco AT, Hartl DL . (2010). Epigenetic effects of polymorphic Y chromosomes modulate chromatin components, immune response, and sexual conflict. Proc Natl Acad Sci USA 107: 15826–15831.
Lyne R, Smith R, Rutherford K, Wakeling M, Varley A, Guillier F et al. (2007). FlyMine: an integrated database for Drosophila and Anopheles genomics. Genome Biol 8: R129.
Mackay TF, Richards S, Stone EA, Barbadilla A, Ayroles JF, Zhu D et al. (2012). The Drosophila melanogaster Genetic Reference Panel. Nature 482: 173–178.
Mason JB, Cargill SL, Griffey SM, Reader JR, Anderson GB, Carey JR . (2011). Transplantation of young ovaries restored cardioprotective influence in postreproductive-aged mice. Aging Cell 10: 448–456.
Maures TJ, Booth LN, Benayoun BA, Izrayelit Y, Schroeder FC, Brunet A . (2014). Males shorten the life span of C. elegans hermaphrodites via secreted compounds. Science 343: 541–544.
McCormick M, Chen K, Ramaswamy P, Kenyon C . (2011). New genes that extend Caenorhabditis elegans' lifespan in response to reproductive signals. Aging Cell 11: 192–202.
Omelyanchuk LV, Shaposhnikov MV, Moskalev AA . (2015). Drosophila nervous system as a target of aging and anti-aging interventions. Front Genet 6: 89.
Paaby AB, Schmidt PS . (2009). Dissecting the genetics of longevity in Drosophila melanogaster. Fly (Austin) 3: 29–38.
Papadopoulos NT, Liedo P, Muller HG, Wang JL, Molleman F, Carey JR . (2010). Cost of reproduction in male medflies: the primacy of sexual courting in extreme longevity reduction. J Insect Physiol 56: 283–287.
Rand DM, Fry A, Sheldahl L . (2006). Nuclear-mitochondrial epistasis and drosophila aging: introgression of Drosophila simulans mtDNA modifies longevity in D. melanogaster nuclear backgrounds. Genetics 172: 329–341.
Reinhart M, Carney T, Clark AG, Fiumera AC . (2015). Characterizing male-female interactions using natural genetic variation in Drosophila melanogaster. J Hered 106: 67–79.
R Development Core Team (2011). R: A Language and Environment for Statistical Computing. The R Foundation for Statistical Computing: Vienna, Austria..
Rose MR, Charlesworth B . (1981). Genetics of life history in Drosophila melanogaster. II. Exploratory selection experiments. Genetics 97: 187–196.
Seong KH, Matsuo T, Fuyama Y, Aigaki T . (2001). Neural-specific overexpression of drosophila plenty of SH3s (DPOSH) extends the longevity of adult flies. Biogerontology 2: 271–281.
Sgro CM, Partridge L . (1999). A delayed wave of death from reproduction in Drosophila. Science 286: 2521–2524.
Shi C, Murphy CT . (2014). Mating induces shrinking and death in Caenorhabditis mothers. Science 343: 536–540.
Sirot LK, Wolfner MF, Wigby S . (2011). Protein-specific manipulation of ejaculate composition in response to female mating status in Drosophila melanogaster. Proc Natl Acad Sci USA 108: 9922–9926.
Smyth GK . (2004). Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3: Article 3.
Smyth GK . (2005) Bioinformatics and Computational Biology Solutions using R and Bioconductor. Springer: New York.
Smyth GK, Speed T . (2003). Normalization of cDNA microarray data. Methods 31: 265–273.
Soues S, Kann ML, Fouquet JP, Melki R . (2003). The cytosolic chaperonin CCT associates to cytoplasmic microtubular structures during mammalian spermiogenesis and to heterochromatin in germline and somatic cells. Exp Cell Res 288: 363–373.
Stearns SC . (1992) The Evolution of Life Histories. Oxford University Press: London, UK. p 249.
Stearns SC, Ackermann M, Doebeli M, Kaiser M . (2000). Experimental evolution of aging, growth, and reproduction in fruitflies. Proc Natl Acad Sci USA 97: 3309–3313.
Tanaka M, Gong JS, Zhang J, Yoneda M, Yagi K . (1998). Mitochondrial genotype associated with longevity. Lancet 351: 185–186.
Tatar M . (2010). Reproductive aging in invertebrate genetic models. Ann NY Acad Sci 1204: 149–155.
Toivonen JM, Partridge L . (2009). Endocrine regulation of aging and reproduction in Drosophila. Mol Cell Endocrinol 299: 39–50.
Townsend JP, Hartl DL . (2002). Bayesian analysis of gene expression levels: statistical quantification of relative mRNA level across multiple strains or treatments. Genome Biol 3: RESEARCH0071.
Tu MP, Tatar M . (2003). Juvenile diet restriction and the aging and reproduction of adult Drosophila melanogaster. Aging Cell 2: 327–333.
Wang MC, Bohmann D, Jasper H . (2003). JNK signaling confers tolerance to oxidative stress and extends lifespan in Drosophila. Dev Cell 5: 811–816.
Wang MC, Bohmann D, Jasper H . (2005). JNK extends life span and limits growth by antagonizing cellular and organism-wide responses to insulin signaling. Cell 121: 115–125.
Wolff JN, Ladoukakis ED, Enriquez JA, Dowling DK . (2014). Mitonuclear interactions: evolutionary consequences over multiple biological scales. Philos Trans R Soc Lond B Biol Sci 369: 20130443.
Wolfner MF . (2009). Battle and ballet: molecular interactions between the sexes in Drosophila. J Hered 100: 399–410.
Yamawaki TM, Berman JR, Suchanek-Kavipurapu M, McCormick M, Gaglia MM, Lee SJ et al. (2010). The somatic reproductive tissues of C. elegans promote longevity through steroid hormone signaling. PLoS Biol 8: e1000468.
Zeh JA, Zeh DW . (2005). Maternal inheritance, sexual conflict and the maladapted male. Trends Genet 21: 281–286.
Zhu CT, Ingelmo P, Rand DM . (2014). GxGxE for lifespan in Drosophila: mitochondrial, nuclear, and dietary interactions that modify longevity. PLoS Genet 10: e1004354.
Acknowledgements
We thank members of the Lemos laboratory for critical reading of the manuscript, Stuart Macdonald for kindly sharing strains, and four anonymous reviewers for their valuable comments and helpful insights. This project was partly supported by an Ellison Foundation New Scholar in Aging Award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Branco, A., Schilling, L., Silkaitis, K. et al. Reproductive activity triggers accelerated male mortality and decreases lifespan: genetic and gene expression determinants in Drosophila. Heredity 118, 221–228 (2017). https://doi.org/10.1038/hdy.2016.89
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2016.89
This article is cited by
-
Selecting on age of female reproduction affects lifespan in both sexes and age-dependent reproductive effort in female (but not male) Ceratitis cosyra
Behavioral Ecology and Sociobiology (2021)
-
Photoresponsiveness affects life history traits but not oxidative status in a seasonal rodent
Frontiers in Zoology (2019)