Abstract
Mechanisms of metal resistance have been reported in many plants but knowledge in woody species is scarce. The TonB-dependent receptors family (TBDTs) is a large group of proteins that facilitate the transport of molecules across the membrane of Gram-negative bacteria. Some evidence exists that TBDTs are involved in metal stress. The existence of a TonB-like mechanism in non-prokaryotes has not been established. The recent development of the Betula papyrifera (white birch) transcriptome has allowed the discovery of genes involved in plant adaptation to stress. The main objective of the present study was to identify novel genes associated with nickel resistance in B. papyrifera. Our results from next generation sequencing and RT-qPCR analyses show that genes involved in transport activities are upregulated in nickel-resistant genotypes compared with susceptible forms. Detailed analysis of gene expression and genome analysis shows for the first time the existence of a TonB-dependent receptor and TonB-like family protein in non-prokaryotes. In addition, we have found that these proteins are associated with nickel resistance in B. papyrifera. Our experiments suggest that the TonB-dependent receptor may be exclusive to the Betula genus, suggesting that Betula species may have acquired the gene via horizontal gene transfer from prokaryotes or fungi.
Similar content being viewed by others
Introduction
Metals are essential for proper homeostasis of all living organisms. They are involved in the stability of the tertiary structure of enzymes and are used as cofactors for enzymatic activities (Andreini et al., 2008). If metal balance is disrupted, it may lead to deficiency or toxicity, often associated with oxidative stress. The production of reactive oxygen species usually results in deregulation of protein activity, organelle damage, membrane deterioration and nucleic acid damage (Yadav, 2010). Plants are often exposed to metals due to expansion of industries and their pollutants. For example, the Greater Sudbury Region in Northern Ontario, Canada is home to one of the largest nickel/copper deposits and extractive operations in the world. A decade of mining and smelting operations has left the surrounding land acidified and contaminated with high levels of metals (Hutchinson and Whitby, 1977; Freedman and Hutchinson, 1980; Gratton et al., 2000). Years of exposure to toxic levels have led to development of metal-resistant plant population (Kirkey et al., 2012).
Mechanisms of metal resistance have been reported in many plant species, but knowledge of how woody plants deal with metals is scarce (Erãnen et al., 2009; Maron et al. 2013; Visioli et al. 2014). Resistance mechanisms differ from species to species and sometimes within the same species. Root exudation is one of the most commonly studied mechanisms of metal resistance. Plants using this strategy will lower the bioavailability of metals around the rhizosphere by secreting organic acids such as citrate and malate from their roots (Qin et al., 2007; Ryan et al., 2009; Maron et al., 2013). Other plants species increase production of chelating molecules such as phytochelatins, metallothioneins and small molecules from metabolic processes to cope with metal contamination (Freeman et al., 2005; Mari et al., 2006; Picault et al., 2006; Callahan et al., 2008). These compounds chelate excess metals in cells to reduce toxicity. Some plant groups accumulate metals in their tissues. Most notable are hyperaccumulators that compartmentalize metals mostly in cell vacuoles to decrease their toxic effects elsewhere in the cells (Schaaf et al., 2006; Merlot et al., 2014).
De Silva et al. (2012) results showed that even a small amount of nickel at the bioavailable levels in the Greater Sudbury Region cause a decrease of stomatal density and chlorophyll content in Acer rubrum. They further showed that these effects are amplified when metal contaminations are combined with drought leading to reductions in hydraulic conductance, xylem-specific conductivity and leaf-specific conductivity.
Several studies have demonstrated that transporters play an important role in metal resistance in plants. In the hyperaccumulator Psychotria gabriellae, the overexpression of the PgIREG1 transporter leads to accumulation of nickel in the tonoplast (Merlot et al., 2014). A similar mechanism has been reported in Arabidopsis thaliana where nickel is transported into the tonoplast via a IREG2/FPN transporter (Schaaf et al., 2006; Nishida et al., 2011). Transporters from the NRAMP family have been thought to play a role in nickel resistance (Wei et al., 2009). This has been confirmed in a recent study showing a strong association between NRAMP transporters with nickel accumulation and resistance in Betula papyrifera (Theriault et al., 2016c). TonB-dependent transports (TBDTs) have been reported to play a role in metal transport in bacteria. They are outer membrane proteins that bind and transport ferric chelates, nickel complexes and other compounds such as carbohydrates and vitamin B12 (Noinaj et al., 2010). In some cases, genes associated with microbial proteins have been transferred to eukaryotes during their evolution. Such horizontal transfers with significant functional implications have been reported in higher plants and animals (Keeling and Palmer, 2008).
The main objective of the present study was to identify novel genes associated with nickel resistance in B. papyrifera.
Material and methods
This study is a comprehensive extension of the baseline transcriptome data described in Theriault et al. (2016c).
Nickel treatment
White birch (B. papyrifera) seeds were collected from a Laurentian University research field site located in the Greater Sudbury Region in Northern Ontario (Canada). The site has been contaminated with metals for >100 years. Details of seed germination and seedlings treatment with nickel nitrate are presented in Theriault et al. (2016c). Likewise, assessment of nickel toxicity is described in Theriault et al. (2016a, 2016b). Gene expression in genotypes resistant and susceptible to a soil nickel concentration of 1600 mg kg−1 is analyzed in details in the present study.
De Novo Transcripts Assembly
Methods for extraction, RNA-seq libraries, new generation sequencing and De Novo Transcripts Assembly are described in Theriault et al. (2016c). The raw reads were mapped to Trinity assembled transcripts using bowtie (http://bowtie-bio.sourceforge.net/index.shtml), and RSEM (http://deweylab.biostat.wisc.edu/rsem) was used to quantify transcript and gene expression levels. Gene expression was calculated and expressed as reads per kilobase per million (RPKM) reads mapped. A differentially expressed gene analysis was performed between resistant and susceptible genotypes. The heatmap data describing the top 50 most expressed genes have be reported in Theriault et al. (2016c). The top 25 most upregulated genes based on Log2FC from the pairwise comparison were ranked in this study (Tables 1 and 2). Similar analysis was performed for the top 25 most downregulated genes. Baseline filtering of genes that likely are the effect of nitrate was conducted in order to make sure the selected candidate genes responded to nickel and not to nitrate (Theriault et al., 2016c).
Validation of the expression of the TonB-dependent receptor using RT-qPCR
RT-qPCR was used to verify the transcriptome data. The RNA was treated with DNase1 (#EN0521) from Life Technologies (Carlsbad, CA, USA). PCR primers were designed using the transcriptome sequence. The cDNA was synthesized using the High-Capacity cDNA Reverse Transcription Kit by Life Technologies. RT-qPCR was performed using the Dynamo HS SYBR Green qPCR Kit by Life Technologies according to the manufacturer’s protocol. Each sample was amplified with the MJ Research PTC-200 Thermal Cycler in triplicates. The process included (1) initial denaturing at 95 °C for 15 min; (2) denaturing at 94 °C for 30 s; (3) 30 s annealing; (4) elongation at 72 °C for 30 s; (5) read; (6) repeat step 2–6 for 41 cycles; (7) final elongation at 72 °C for 7 min; (8) melting curve 72–95 °C, every 1 °C, hold for 10 s; and (9) final elongation at 72 °C for 3 min. The qPCR was run three separate times with each sample in triplicate, resulting in a total of nine data point for each sample. The data were analyzed using the MJ Opticon Monitor 3.1 by BioRad and delta C(t) values were exported to Microsoft Excel. RNA concentrations were calculated using Delta C(t) values and standard curves. Expression was first normalized against α-tubulin (housekeeping gene) then the relative expression was calculated by dividing the expression of resistant/susceptible genotypes over water controls. We compared the mean expression of the TonB-dependent receptor between resistant/susceptible genotypes and water controls. Data were analyzed using SPSS 20 for Windows, with all data being log10 transformed to achieve a normal distribution. Variance ratio test was performed with an assumption of data normality in the underlying population distributions of the data. ANOVA, followed by Tukey’s HSD multiple comparison analysis, was performed to determine significant differences among means for qPCR (P<0.05).
TonB-dependent receptor in other species
PCR was performed using the cDNA or DNA template from B. papyrifera and other species (Table 3). Each PCR reaction included a forward primer (0.1 mmol), reverse primer (0.1 mmol), MgCl (2 mmol), dNTPs (0.2 mmol), 10 × buffer and Taq Polymerase. The PCR was performed using the MJ Research PTC-200 Thermal Cycler: (1) initial denaturing at 95 °C for 15 min; (2) denaturing at 94 °C for 30 s; (3) 30 s annealing; (4) elongation at 72 °C or 30 s; (5) read; (6) repeat steps 2–6 for 41 cycles; and (7) final elongation at 72 °C for 7 min. Amplification products were run on agarose gels and the fragment sizes verified using the BioRad Doc system (Hercules, CA, USA).
A basic local alignment search (BLAST) using the dwarf birch (Betula nana) genome (http://birchgenome.org/) was performed to determine sequence similarity (Wang et al., 2013).
Results
Transcriptome
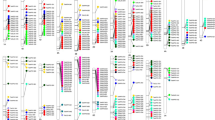
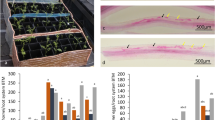
Differentially expressed genes between nickel-resistant, susceptible and water controls were identified. Broad data of the heatmap has been presented in Theriault et al. (2016c). The top upregulated and downregulated genes were ranked based on log 2FC. The study shows that nickel treatment triggers different regulation of several genes. In-depth analysis in the present study of molecular functions of the 25 most upregulated in resistant genotypes reveals that 32% were associated with catalytic activities, 12% with transport and 8% with binding (Figure 1). Among the top 25 most downregulated genes, 20, 0 and 8% were associated with catalytic, transport and binding activities, respectively (Figure 2). Detailed description of the top 10 most upregulated and downregulated is presented in Tables 1 and 2. Three of the 10 most upregulated genes in resistant genotypes were associated with transport (TR99219|c0_g1, TR91600|c2_g5, TR109068|c1_g1 and TR78530|c1_g1), three with binding (TR55738|c2_g6, TR117076|c0_g1 and TR96264|c0_g1), and four have unknown molecular function (TR86018|c0_g2, TR67414|c0_g5, and TR100502|c2_g1; Table 1). For downregulated genes, six were involved in binding (TR94889|c2_g1, TR53148|c0_g1, TR94980|c2_g4, TR56209|c7_g8, TR35375|c2_g19 and TR79073|c0_g1) and the other four (TR16661|c0_g1, TR55969|c0_g1, TR61028|c0_g1 and TR35437|c1_g1) had unknown molecular functions (Table 2).
The transcriptome assembly of B. papyrifera revealed that a TonB-dependent receptor and a TonB-like family protein transcript were the two most upregulated genes in resistant genotypes (RG) compared with susceptible (SG; Table 1). Their gene IDs in the B. papyrifera transcriptome are TR99219|c0_g1 and TR91600|c2_g5, respectively. This transcriptome has been deposited at DDBJ/EMBL/GenBank under the accession GEIC00000000 (Theriault et al., 2016c). The sequence length of the TonB-dependent receptor and the TonB-like family protein transcript were 1604 and 258 bp, respectively (Figures 3 and 4).
The mean expression values for water controls, resistant and susceptible genotypes were significantly different based on ANOVA followed by the Tukey’s HSD tests (P<0.05). In fact, the expressions of TonB-dependent receptor and TonB-like family protein transcripts were expressed 2978 × and 1448 × higher in RG compared with SG. We found also that the expression of TonB-dependent receptor transcript was 1082 × higher in the RG compared with controls. There was no significant difference in the expression of the TonB-dependent receptor transcript when the water control was compared with SG. But the expression of TonB-like family protein transcript was 2048 × higher in the water controls compared with Ni-treated susceptible genotypes. No significant difference was found in the expression of the TonB-like family protein transcript when RG were compared with water controls.
RT-qPCR analysis
Overall, qPCR data showed significant differences among genotypes based on Tukey’s HSD multiple comparison analysis (P<0.05). The expression of the TonB-dependent receptor was significantly higher in RG compared with SG or water controls (Figure 5).
Quantitative qPCR measurement of the expression of TonB-dependent family receptor (TR99219|c0_g1) in nickel resistant and susceptible white birch (Betula papyrifera) treated with 1600 mg kg−1 of nickel. Expression was standardized using the housekeeping gene ef1a. Normalization was then performed against water controls (all values divided by water). *Significant differences were found using ANOVA (Tukey’s HSD multiple comparison analysis, P<0.05).
TonB-dependent receptor/-like protein in other species
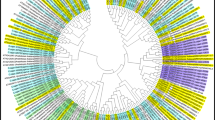
Betula nana database revealed the presence of TonB-dependent receptor and TonB-like family protein. We performed a BLAST alignment of the B. papyrifera transcripts with the Betula nana (dwarf birch) genome (Supplementary Figures S1 and S2). The sequence similarity between both birch species for the TonB-dependent receptor and TonB-like family protein was 93 and 83%, respectively. No match was found in plant species when using the NCBI BLAST tool (Supplementary Figures S3 and S4). BLAST was also performed in additional databases (DDBJ/EMBL/GenBank, Poplar genome database, dendrome, Plantgdb, etc.) and no match in plants species was found outside the Betula genus. We performed a search in the UniProt database and it revealed the presence of TonB-dependent receptors/proteins in a number of bacteria and fungi species. We found that almost 100% of the 183 372 TonB protein matches were from bacterial and fungal species (Figure 6).
Primers were designed to confirm the presence of the gene in the B. papyrifera genome and transcriptome. PCR amplification using root and leaf cDNA revealed a single band of around 300 bp for the TonB-dependent family receptor (Figure 7a). Several B. papyrifera populations were screened and the same size band was observed. The presence of the gene was also confirmed in yellow birch (B. alleghaniensis) and other Betula species (Figure 7a). Figure 7b shows the absence of the band in other tree species. A complete list of all the species screened for the TonB-dependent receptor is presented in Table 3.
(a) PCR amplification products on 1% agarose gel. A 300 bp band corresponds to the presence of the TonB-dependent receptor. Lanes 1, 11 and 16 are loaded with 1 kb+ ladder; 2–10 with white birch (Betula papyrifera) from different populations (leaf DNA); 12 Betula alleghaniensis (seedling DNA); 13 Betula minor (seedling DNA); 14 and 15 white birch (root cDNA and leaf cDNA). (b) PCR amplification products on 1% agarose gel. A 300 bp band corresponds to the presence of the TonB-dependent receptor (red arrow). Lane 1: white birch (Betula papyrifera), 2 and 23 are loaded with 1 kb+ ladder, 3–22 are the following species in order: Quercus rubra, Populus tremuloides, Acer rubrum, Pinus strobus, Pinus monticola, Pinus nigra, Pinus sylvestris, Pinus contorta, Pinus rigida, Pinus banksiana, Pinus resinosa, Pinus montana, Picea glauca, Picea sitchensis, Picea wilsonii, Picea jezoensis, Picea orientalis, Picea engelmannii, and Picea pungens (pop. 1) Picea pungens (pop. 2). A full color version of this figure is available online at the Heredity website.
Discussion
Recent general analysis of B. papyrifera transcriptome generated through shotgun methods revealed that the main mechanism involved in B. papyrifera resistance to nickel is downregulation of genes during translation (Theriault et al., 2016a, 2016b). In this study, we show that actually genes involved in transport activities are upregulated in resistant genotypes compared with susceptible forms. Those few genes are of particular importance in nickel resistance in many organisms.
TonB-dependent receptors and TonB protein family in prokaryotes
The TonB-dependent transporter (TBDT) family is a large group of proteins that facilitate the transport of molecules across the membrane of Gram-negative bacteria. To date, they have never been reported in plants. Their tertiary structure is a 22-stranded beta-barrel located in the outer membrane with a plug domain folded inside the barrel (Noinaj et al., 2010). TBDTs are more commonly known for their role in iron regulation but have been associated with transport of vitamins, nickel and carbohydrates (Schauer et al., 2008). In order to pass through the pore, two requirements must be met: the metal must be chelated to a siderophore and the inner membrane must provide energy. Transport through TBDT is dependent on the inner membrane protein complex TonB, ExbB and ExbD (Postle, 2007). The plug domain of TBDTs interacts with TonB to transfer energy via a proton motive force leading to the opening of the pore and passage of the sidosphore complex (Postle, 2007).
Some evidence exists that TBDTs are dealing with metal stress. In some species, TBDTs are repressed during heavy metal stress (Yoneyama and Nakae, 1996; Park and Ely, 2008) while in others, there is an induction (Hu et al., 2005; Brown et al., 2006). Hu et al. (2005) found that TBDTs were upregulated during heavy metal stress. However, their role in response to metal was unclear since TonB was downregulated (Hu et al., 2005). Increase in TBDT expression has been linked to metal resistance. High expression of opdT and OmpC porins have been associated with copper resistance in Pseudomonas aeruginosa and Escherichia coli (Egler et al., 2005; Teitzel et al., 2006).
TonB-like mechanism in plants
The existence of a TonB-like mechanism other than in prokaryotes has not been established. Only the TonB box, a consensus sequence in bacterial TBDTs, has been found in plant transporters and G proteins (Assmann, 2002; Duy et al., 2011). The role of these elements in eukaryotes is unknown. Plants, like bacteria and fungi, produce siderophores. In iron-deficient plants, siderophores with a high affinity for iron (III) are secreted from the roots via a transporter (Walker and Connolly, 2008). The siderophores will chelate non-soluble iron (III) and so form an iron-siderophore complex (Walker and Connolly, 2008). The complex is then carried back into the plant’s root system via a secondary transporter (Walker and Connolly, 2008). The presence of TonB-binding boxes and siderophore production in plants suggests that a mechanism involving a TonB receptor in plants is possible. Here we present the first data showing the presence of a TBDT in plants and its association with nickel tolerance in B. papyrifera.
Validating the presence of TonB-dependent receptor and TonB-like family protein
Expression of the TonB-dependent receptor transcript in resistant plants was much higher than susceptible genotypes and water control. Similar results were obtained by the confirmatory qPCR. This suggests that the expression of the TonB-dependent receptor is induced by nickel. The expression of the TonB-like family protein transcript was also significantly higher in resistant genotypes and water controls compared with susceptible. However, no significant difference was found between resistant and water controls, suggesting that the transcript is constitutively expressed.
To exclude the possibility of any bacterial contamination of the roots analyzed in this study, we tested for the presence of the transcript/gene in the areal parts of the plant. Both analyses of B. papyrifera leaf cDNA and genomic DNA confirmed the presence of the TonB-dependent receptor gene. While the contamination of foreign RNA is greatly diminished when using leaf tissue, bacteria can still colonize leaves. Phylobacteria are bacteria that survive on or inside plant leaves (Beattie and Lindow, 1999). Some live on the surface while others will enter the plant via lesions in the leaves or hydathodes (Beattie and Lindow, 1999). However, the fact that expression of this gene was found in both RG and SG exclude the possibility of contamination happening only in resistant plants.
Further analyses found that the TonB-dependent receptor and TonB-like family protein are also present in the B. nana (dwarf birch) transcriptome. The sequence alignment revealed a high degree of similarity between the B. papyrifera and B. nana genomes. This is expected since the two species (B. papyrifera and B. nana) are genetically closely related (Järvinen et al., 2004; Li et al., 2005). In addition, using PCR, we were able to confirm the presence of the TonB-dependent receptor in the genome of Betula species. We also screened several populations of B. payrifera from different locations in Northern Ontario and New Brunswick (Canada) and elsewhere and we confirmed the presence of the TonB-dependent receptor in all of them. When screening other species from different genera via BLAST or PCR, we found that the gene was exclusive to the Betula genus and was not found in any other tree species.
It should be noted that the lack of amplification with primer pairs targeting this gene in species outside the Betula genus could have been due to the absence of primer-binding sites or weak primer bindings. But such possibility is unlikely since the BLAST search of existing depositories did not report any match to gene for the TonB-dependent receptor or TonB-like family protein. In fact, NCBI (National Center for Biotechnology Information—USA), EMBL (European Molecular Biology Laboratory), DDP (DNA databank of Japan), Plantgdb (Resources for Plant Comparative Genomics), Dendrome (forest trees genome database), Populus and B. nana databases contain millions of sequences from different organisms but the BLAST search for the TonB-dependent receptor gene generated hits only for bacteria, fungi and Betula sequences.
It is possible that the gene for the TonB-dependent receptor or TonB-like family protein was transferred from bacteria to Betula through horizontal transfer during the evolution of the genus Betula. The notions of horizontal transfer between microorganisms and plants is a topic of great interest and has been discussed by many authors (Hanekamp et al., 1997; Kim et al., 2002; Nkongolo et al., 2004; Richardson and Palmer, 2007; Yue et al., 2012; Gao et al., 2014).
Yue et al. (2012) identified 57 family of genes transferred from prokaryotes, fungi or viruses to moss Physcomitrella patens. Most of these genes were directly or indirectly related to plant defence and stress tolerance. Notably the glutamate–cysteine ligase gene was acquired from bacteria by P. patens. Glutamate–cysteine ligase is one of the two genes that catalyze the formation of glutamine. This compound is involved in plant disease resistance, photo-oxidative stress defense; and heavy metal detoxification. Interestingly, our initial analysis of B. papyrifera transcriptome revealed that glutathione is also upregulated in nickel-resistant genotype compared with susceptible (Theriault et al., 2016c). The mechanism of possible transfer of TonB genes to Betula spp. is not established, but horizontal gene transfer is a widespread process involved in the evolution of multicellular eukaryotes.
Conclusion
This is the first study that shows and documents the existence of a TonB-dependent receptor and TonB-like family protein in plants. We found that they are associated with nickel resistance in white birch (B. papyrifera). A series of experiments showed that the TonB-dependent receptor could be exclusive to the Betula genus. This suggests that Betula species might have acquired the gene via a recent horizontal gene transfer from prokaryotes or fungi. Thanks to advances in next generation genome sequencing, gene depositories represent the main databases useful for identifying the presence of genes of interest in a species. Hence, it is possible that other plant genera or species whose sequences have not been deposited in a public gene bank might carry the TonB-dependent receptor and/or TonB protein family. Further studies including physical mapping of TonB-dependent receptor in Betula will be conducted using in situ hybridization. Transcriptional regulation by Ni on other plant species showing different resistance mechanisms to Ni toxicity such as red maple (A. rubrum), red oak (Quercus rubra) and trembling aspen (Populus tremuloides) will be also investigated.
Data archiving
We have deposited data in the following depository: Repository/DataBank Accession: EmbL Accession ID: GEIC00000000 Databank URL: http://www.ebi.ac.uk/ena Repository/DataBank Accession: NCBI genbank Accession ID: GEIC00000000 Databank URL: http://www.ncbi.nlm.nih.gov/genbank
Accession codes
References
Andreini C, Bertini I, Cavallaro G, Holliday GL, Thornton JM . (2008). Metal ions in biological catalysis: from enzyme databases to general principles. J Biol Inorg Chem 13: 1205–1218.
Assmann SM . (2002). Heterotrimeric and unconventional GTP binding proteins in plant cell signaling. Plant Cell 14: S355–S373.
Beattie GA, Lindow SE . (1999). Bacterial colonization of leaves: a spectrum of strategies. Phytopathology 89: 353–359.
Brown SD, Thompson MR, VerBerkmoes NC, Chourey K, Shah M, Zhou J et al. (2006). Molecular dynamics of the Shewanella oneidensis response to chromate stress. Mol Cell Proteomics 5: 1054–1071.
Callahan DL, Roessner U, Dumontet V, Perrier N, Wedd AG, Richard AJ et al. (2008). LC–MS and GC–MS metabolite profiling of nickel (II) complexes in the latex of the nickel-hyperaccumulating tree Sebertia acuminata and identification of methylated aldaric acid as a new nickel (II) ligand. Phytochemistry 69: 240–251.
de Silva ND, Cholewa E, Ryser P . (2012). Effects of combined drought and heavy metal stresses on xylem structure and hydraulic conductivity in red maple (Acer rubrum L.). J Exp Bot 63: 5957–5966.
Duy D, Stübe R, Wanner G, Philippar K . (2011). The chloroplast permease PIC1 regulates plant growth and development by directing homeostasis and transport of iron. Plant Physiol 155: 1709–1722.
Egler M, Grosse C, Grass G, Nies DH . (2005). Role of the extracytoplasmic function protein family sigma factor RpoE in metal resistance of Escherichia coli. J Bacteriol 187: 2297–2307.
Erãnen JK, Nilsen J, Zverev VE, Kozlov MV . (2009). Mountain birch under multiple stressors—heavy metal-resistant populations co-resistant to biotic stress but maladapted to abiotic stress. J Evol Biol 22: 840–851.
Freedman B, Hutchinson TC . (1980). Pollutant inputs from the atmosphere and accumulations in soils and vegetation near a nickel-copper smelter at Sudbury, Ontario, Canada. Can J Bot 58: 108–132.
Freeman JL, Garcia D, Kim D, Hopf A, Salt DE . (2005). Constitutively elevated salicylic acid signals glutathione-mediated nickel tolerance in Thlaspi nickel hyperaccumulators. Plant Physiol 137: 1082–1091.
Gao C, Ren X, Mason AS, Liu H, Xiao M, Li J et al. (2014). Horizontal gene transfer in plants. Funct Integr Genomics 14: 23–29.
Gratton WS, Nkongolo KK, Spiers GA . (2000). Heavy metal accumulation in soil and Jack Pine (Pinus banksiana Needles in Sudbury, Ontario, Canada. Bull Environ Contam Toxicol 64: 550–557.
Hanekamp T, Kobayashi D, Hayes S, Stayton MM . (1997). Avirulence gene D of Pseudomonas syringae pv. tomato may have undergone horizontal gene transfer. FEBS Lett 415: 40–44.
Hu P, Brodie EL, Suzuki Y, McAdams HH, Andersen GL . (2005). Whole-genome transcriptional analysis of heavy metal stresses in Caulobacter crescentus. J Bacteriol 187: 8437–8449.
Hutchinson TC, Whitby LM . (1977). The effects of acid rainfall and heavy metal particulates on a boreal Forest ecosystem near the sudbury smelting region of Canada. Water Air Soil Pollut 7: 421–438.
Järvinen P, Palmé A, Orlando Morales L, Lännenpää M, Keinänen M, Sopanen T et al. (2004). Phylogenetic relationships of Betula species (Betulaceae based on nuclear ADH and chloroplast matK sequences. Am J Bot 91: 1834–1845.
Keeling PJ, Palmer JD . (2008). Horizontal gene transfer in eukaryotic evolution. Nat Rev Genet 9: 605–618.
Kim N-S, Armstrong KC, Fedak G, Ho K, Park N-I . (2002). A microsatellite sequence from the rice blast fungus (Magnaporthe grisea distinguishes between the centromeres of Hordeum vulgare and H. bulbosum in hybrid plants. Genome 45: 165–174.
Kirkey FM, Matthews J, Ryser P . (2012). Metal resistance in populations of red maple (Acer rubrum L.) and white birch (Betula papyrifera Marsh.) from a metal-contaminated region and neighbouring non-contaminated regions. Environ Pollut 164: 53–58.
Li J, Shoup S, Chen Z . (2005). Phylogenetics of Betula Betulaceae inferred from sequences of nuclear ribosomal DNA. Rhodora 107: 69–86.
Mari S, Gendre D, Pianelli K, Ouerdane L, Lobinski R, Briat JF et al. (2006). Root-to-shoot long-distance circulation of nicotianamine and nicotianamine-nickel chelates in the metal hyperaccumulator Thlaspi caerulescens. J Exp Bot 57: 4111–4122.
Maron LG, Guimaraes CT, Kirst M, Albert PS, Birchler JA, Bradbury PJ et al. (2013). Aluminum tolerance in maize is associated with higher MATE1 gene copy number. Proc Natl Acad Sci USA 110: 5241–5246.
Merlot S, Hannibal L, Martins S, Martinelli L, Amir H, Lebrun M et al. (2014). The metal transporter PgIREG1 from the hyperaccumulator Psychotria gabriellae is a candidate gene for nickel tolerance and accumulation. J Exp Bot 65: 1551–1564.
Nishida S, Tsuzuki C, Kato A, Aisu A, Yoshida J, Mizuno T . (2011). AtIRT1, the primary iron uptake transporter in the root, mediates excess nickel accumulation in Arabidopsis thaliana. Plant Cell Physiol 52: 1433–1442.
Nkongolo KK, Kim NS, Michael P . (2004). Detection and physical mapping of the 18 S-5.8 S-26 S rDNA and the pKFJ660 probe with microsatellite sequences derived from the rice blast fungus (Magnaporthe grisea in conifer species. Hereditas 140: 70–78.
Noinaj N, Guillier M, Barnard TJ, Buchanan SK . (2010). TonB-dependent transporters: regulation, structure, and function. Annu Rev Microbiol 64: 43–60.
Park S, Ely RL . (2008). Candidate stress genes of Nitrosomonas europaea for monitoring inhibition of nitrification by heavy metals. Appl Environ Microbiol 74: 5475–5482.
Picault N, Cazalé AC, Beyly A, Cuiné S, Carrier P, Luu DT et al. (2006). Chloroplast targeting of phytochelatin synthase in Arabidopsis: effects on heavy metal tolerance and accumulation. Biochimie 88: 1743–1750.
Postle K . (2007). TonB system, in vivo assays and characterization. Methods Enzymol 422: 245–269.
Qin R, Hirano Y, Brunner I . (2007). Exudation of organic acid anions from poplar roots after exposure to Al, Cu and Zn. Tree Physiol 27: 313–320.
Richardson AO, Palmer JD . (2007). Horizontal gene transfer in plants. J Exp Bot 58: 1–9.
Ryan PR, Raman H, Gupta S, Horst WJ, Delhaize E . (2009). A second mechanism for aluminum resistance in wheat relies on the constitutive efflux of citrate from roots. Plant Physiol 149: 340–351.
Schaaf G, Honsbein A, Meda AR, Kirchner S, Wipf D, von Wirén N . (2006). AtIREG2 encodes a tonoplast transport protein involved in iron-dependent nickel detoxification in Arabidopsis thaliana roots. J Biol Chem 281: 25532–25540.
Schauer K, Rodionov DA, de Reuse H . (2008). New substrates for TonB-dependent transport: do we only see the ‘tip of the iceberg’? Trends Biochem Sci 33: 330–338.
Teitzel GM, Geddie A, Susan K, Kirisits MJ, Whiteley M, Parsek MR . (2006). Survival and growth in the presence of elevated copper: transcriptional profiling of copper-stressed Pseudomonas aeruginosa. J Bacteriol 188: 7242–7256.
Theriault G, Michael P, Nkongolo K . (2016a). Nickel and copper toxicity and plant response mechanisms in white birch (Betula papyrifera. Bull Envir Contam Toxicol 97: 171–176.
Theriault G, Michael P, Nkongolo K . (2016c). Comprehensive transcriptome analysis of response to nickel stress in White Birch (Betula papyrifera. PLoS One 11: e0153762.
Theriault G, Michael P, Nkongolo KK . (2016b). Decrypting the regulation and mechanism of nickel resistance in white birch (Betula papyrifera using cross-species metal-resistance genes. Genes Genom 38: 341–350.
Visioli G, Gullì M, Marmiroli N . (2014). Noccaea caerulescens populations adapted to grow in metalliferous and non-metalliferous soils: Ni tolerance, accumulation and expression analysis of genes involved in metal homeostasis. Environ Exp Bot 105: 10–17.
Walker EL, Connolly EL . (2008). Time to pump iron: iron-deficiency-signaling mechanisms of higher plants. Curr Opin Plant Biol 11: 530–535.
Wang N, Thomson M, Bodles WJA, Crawford RMM, Hunt HV, Featherstone AW et al. (2013). Genome sequence of dwarf birch (Betula nana and cross species RAD markers. Mol Ecol 22: 3098–3111.
Wei W, Chai T, Zhang Y, Han L, Xu J, Guan Z . (2009). The Thlaspi caerulescens NRAMP homologue TcNRAMP3 is capable of divalent cation transport. Mol Biotechnol 41: 15–21.
Yadav SK . (2010). Heavy metals toxicity in plants: an overview on the role of glutathione and phytochelatins in heavy metal stress tolerance of plants. South African J Bot 76: 167–179.
Yoneyama H, Nakae T . (1996). Protein C (OprC) of the outer membrane of Pseudomonas aeruginosa is a copper-regulated channel protein. Microbiology 142: 2137–2144.
Yue J, Hu X, Sun H, Yang Y, Huang J . (2012). Widespread impact of horizontal gene transfer on plant colonization of land. Nat Commun 3: 1152.
Acknowledgements
We thank Natural Sciences and Engineering Research Council of Canada (NSERC) for financial support. We are grateful to Dr Paul Michael for assistance with data analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Theriault, G., Nkongolo, K. Evidence of prokaryote like protein associated with nickel resistance in higher plants: horizontal transfer of TonB-dependent receptor/protein in Betula genus or de novo mechanisms?. Heredity 118, 358–365 (2017). https://doi.org/10.1038/hdy.2016.106
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2016.106