Abstract
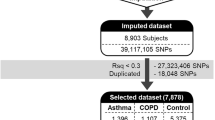
We developed a genetic marker set of single nucleotide polymorphisms (SNPs) by summing risk scores of 14 SNPs showing a significant association with aspirin-exacerbated respiratory disease (AERD) from our previous 660 W genome-wide association data. The summed scores were higher in the AERD than in the aspirin-tolerant asthma (ATA) group (P=8.58 × 10−37), and were correlated with the percent decrease in forced expiratory volume in 1 s after aspirin challenge (r2=0.150, P=5.84 × 10−30). The area under the curve of the scores for AERD in the receiver operating characteristic curve was 0.821. The best cutoff value of the summed risk scores was 1.01328 (P=1.38 × 10−32). The sensitivity and specificity of the best scores were 64.7% and 85.0%, respectively, with 42.1% positive and 93.4% negative predictive values. The summed risk score may be used as a genetic marker with good discriminative power for distinguishing AERD from ATA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Stevenson DD, Sanchez-Borges M, Szczeklik A . Classification of allergic and pseudoallergic reactions to drugs that inhibit cyclooxygenase enzymes. Ann Allergy Asthma Immunol 2001; 87: 177–180.
Samter M, Beers RF Jr . Concerning the nature of intolerance to aspirin. J Allergy 1967; 40: 281–293.
Mascia K, Haselkorn T, Deniz YM, Miller DP, Bleecker ER, Borish L et al. Aspirin sensitivity and severity of asthma: evidence for irreversible airway obstruction in patients with severe or difficult-to-treat asthma. J Allergy Clin Immunol 2005; 116: 970–975.
Szczeklik A, Nizankowska E, Duplaga M . Natural history of aspirin-induced asthma. AIANE Investigators. European Network on Aspirin-Induced Asthma. Eur Respir J 2000; 16: 432–436.
Chang HS, Park JS, Jang AS, Park SW, Uh ST, Kim YH et al. Diagnostic value of clinical parameters in the prediction of aspirin-exacerbated respiratory disease in asthma. Allergy Asthma Immunol Res 2011; 3: 256–264.
Hedman J, Kaprio J, Poussa T, Nieminen MM . Prevalence of asthma, aspirin intolerance, nasal polyposis and chronic obstructive pulmonary disease in a population-based study. Int J Epidemiol 1999; 28: 717–722.
Marquette CH, Saulnier F, Leroy O, Wallaert B, Chopin C, Demarcq JM et al. Long-term prognosis of near-fatal asthma. A 6-year follow-up study of 145 asthmatic patients who underwent mechanical ventilation for a near-fatal attack of asthma. Am Rev Respir Dis 1992; 146: 76–81.
Picado C . Aspirin-intolerant asthma: role of cyclo-oxygenase enzymes. Allergy 2002; 57: 58–60.
Schafer D, Maune S . Pathogenic mechanisms and in vitro diagnosis of AERD. J Allergy (Cairo) 2012; 2012: 789232.
Gollapudi RR, Teirstein PS, Stevenson DD, Simon RA . Aspirin sensitivity: implications for patients with coronary artery disease. JAMA 2004; 292: 3017–3023.
De Weck AL, Sanz ML, Gamboa PM, Jermann JM, Kowalski M, Medrala W et al. Nonsteroidal anti-inflammatory drug hypersensitivity syndrome: a multicenter study. II. Basophil activation by nonsteroidal anti-inflammatory drugs and its impact on pathogenesis. J Investig Allergol Clin Immunol 2010; 20: 39–57.
Schafer D, Schmid M, Gode UC, Baenkler HW . Dynamics of eicosanoids in peripheral blood cells during bronchial provocation in aspirin-intolerant asthmatics. Eur Respir J 1999; 13: 638–646.
Devouassoux G, Pachot A, Laforest L, Diasparra J, Freymond N, Van Ganse E et al. Galectin-10 mRNA is overexpressed in peripheral blood of aspirin-induced asthma. Allergy 2008; 63: 125–131.
Min JW, Jang AS, Park SM, Lee SH, Lee JH, Park SW et al. Comparison of plasma eotaxin family level in aspirin-induced and aspirin-tolerant asthma patients. Chest 2005; 128: 3127–3132.
Kim HJ, Park JS, Heo JS, Moon KY, Park CS . Plasma apolipoprotein H levels are different between aspirin induced respiratory diseases and aspirin tolerant asthma. Pulm Pharmacol Ther 2014; 27: 184–189.
Sanak M, Kielbasa B, Bochenek G, Szczeklik A . Exhaled eicosanoids following oral aspirin challenge in asthmatic patients. Clin Exp Allergy 2004; 34: 1899–1904.
Shin S, Park JS, Kim YJ, Oh T, An S, Park CS . Differential gene expression profile in PBMCs from subjects with AERD and ATA: a gene marker for AERD. Mol Genet Genomics 2012; 287: 361–371.
Park JS, Chang HS, Park CS, Lee JH, Lee YM, Choi JH et al. Association analysis of cysteinyl-leukotriene receptor 2 (CYSLTR2) polymorphisms with aspirin intolerance in asthmatics. Pharmacogenet Genomics 2005; 15: 483–492.
Jinnai N, Sakagami T, Sekigawa T, Kakihara M, Nakajima T, Yoshida K et al. Polymorphisms in the prostaglandin E2 receptor subtype 2 gene confer susceptibility to aspirin-intolerant asthma: a candidate gene approach. Hum Mol Genet 2004; 13: 3203–3217.
Sanak M, Szczeklik A . Leukotriene C4 synthase polymorphism and aspirin-induced asthma. J Allergy Clin Immunol 2001; 107: 561–562.
Choi JH, Park HS, Oh HB, Lee JH, Suh YJ, Park CS et al. Leukotriene-related gene polymorphisms in ASA-intolerant asthma: an association with a haplotype of 5-lipoxygenase. Hum Genet 2004; 114: 337–344.
Kim SH, Bae JS, Holloway JW, Lee JT, Suh CH, Nahm DH et al. A polymorphism of MS4A2 (- 109T>C) encoding the beta-chain of the high-affinity immunoglobulin E receptor (FcepsilonR1beta) is associated with a susceptibility to aspirin-intolerant asthma. Clin Exp Allergy 2006; 36: 877–883.
Dekker JW, Nizankowska E, Schmitz-Schumann M, Pile K, Bochenek G, Dyczek A et al. Aspirin-induced asthma and HLA-DRB1 and HLA-DPB1 genotypes. Clin Exp Allergy 1997; 27: 574–577.
Akahoshi M, Obara K, Hirota T, Matsuda A, Hasegawa K, Takahashi N et al. Functional promoter polymorphism in the TBX21 gene associated with aspirin-induced asthma. Hum Genet 2005; 117: 16–26.
Kim BS, Park SM, Uhm TG, Kang JH, Park JS, Jang AS et al. Effect of single nucleotide polymorphisms within the interleukin-4 promoter on aspirin intolerance in asthmatics and interleukin-4 promoter activity. Pharmacogenet Genomics 2010; 20: 748–758.
Oh SH, Park SM, Park JS, Jang AS, Lee YM, Uh ST et al. Association analysis of peroxisome proliferator-activated receptors gamma gene polymorphisms with asprin hypersensitivity in asthmatics. Allergy Asthma Immunol Res 2009; 1: 30–35.
Park SM, Park JS, Park HS, Park CS . Unraveling the genetic basis of aspirin hypersensitivity in asthma beyond arachidonate pathways. Allergy Asthma Immunol Res 2013; 5: 258–276.
Kim JH, Park BL, Cheong HS, Bae JS, Park JS, Jang AS et al. Genome-wide and follow-up studies identify CEP68 gene variants associated with risk of aspirin-intolerant asthma. PLoS One 2010; 5: e13818.
Park BL, Kim TH, Kim JH, Bae JS, Pasaje CF, Cheong HS et al. Genome-wide association study of aspirin-exacerbated respiratory disease in a Korean population. Hum Genet 2013; 132: 313–321.
Shin SW, Park J, Kim YJ, Uh ST, Choi BW, Kim MK et al. A highly sensitive and specific genetic marker to diagnose aspirin-exacerbated respiratory disease using a genome-wide association study. DNA Cell Biol 2012; 31: 1604–1609.
Bateman ED, Hurd SS, Barnes PJ, Bousquet J, Drazen JM, FitzGerald M et al. Global strategy for asthma management and prevention: GINA executive summary. Eur Respir J 2008; 31: 143–178.
Kraft P, Wacholder S, Cornelis MC, Hu FB, Hayes RB, Thomas G et al. Beyond odds ratios—communicating disease risk based on genetic profiles. Nat Rev Genet 2009; 10: 264–269.
Ritchie MD, Hahn LW, Roodi N, Bailey LR, Dupont WD, Parl FF et al. Multifactor-dimensionality reduction reveals high-order interactions among estrogen-metabolism genes in sporadic breast cancer. Am J Hum Genet 2001; 69: 138–147.
Shigemizu D, Abe T, Morizono T, Johnson TA, Boroevich KA, Hirakawa Y et al. The Construction of Risk Prediction Models Using GWAS Data and Its Application to a Type 2 Diabetes Prospective Cohort. PLoS One 2014; 9: e92549.
Klein TE, Altman RB, Eriksson N, Gage BF, Kimmel SE, Lee MT et al. Estimation of the warfarin dose with clinical and pharmacogenetic data. N Engl J Med 2009; 360: 753–764.
Horne BD, Lenzini PA, Wadelius M, Jorgensen AL, Kimmel SE, Ridker PM et al. Pharmacogenetic warfarin dose refinements remain significantly influenced by genetic factors after one week of therapy. Thromb Haemost 2012; 107: 232–240.
Verhoef TI, Ragia G, de Boer A, Barallon R, Kolovou G, Kolovou V et al. A randomized trial of genotype-guided dosing of acenocoumarol and phenprocoumon. N Engl J Med 2013; 369: 2304–2312.
Qiu FH, Devchand PR, Wada K, Serhan CN . Aspirin-triggered lipoxin A4 and lipoxin A4 up-regulate transcriptional corepressor NAB1 in human neutrophils. FASEB J 2001; 15: 2736–2738.
Serhan CN, Takano T, Chiang N, Gronert K, Clish CB . Formation of endogenous "antiinflammatory" lipid mediators by transcellular biosynthesis. Lipoxins and aspirin-triggered lipoxins inhibit neutrophil recruitment and vascular permeability. Am J Respir Crit Care Med 2000; 161: S95–s101.
Krause G, Winkler L, Mueller SL, Haseloff RF, Piontek J, Blasig IE . Structure and function of claudins. Biochim Biophys Acta 2008; 1778: 631–645.
Oshima T, Miwa H, Joh T . Aspirin induces gastric epithelial barrier dysfunction by activating p38 MAPK via claudin-7. Am J Physiol Cell Physiol 2008; 295: C800–C806.
Krymskaya VP . Targeting the phosphatidylinositol 3-kinase pathway in airway smooth muscle: rationale and promise. BioDrugs 2007; 21: 85–95.
Jandrot-Perrus M, Busfield S, Lagrue AH, Xiong X, Debili N, Chickering T et al. Cloning, characterization, and functional studies of human and mouse glycoprotein VI: a platelet-specific collagen receptor from the immunoglobulin superfamily. Blood 2000; 96: 1798–1807.
Acknowledgements
This study was supported by a grant of the Korea Healthcare Technology R&D Project, Ministry for Health, Welfare & Family Affairs, Republic of Korea (HI13C0319), by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MEST) (2012R1A1A2038396) and by a research grant of Soonchunhyang University. The DNA samples were generously provided by Soonchunhyang University, Bucheon Hospital Biobank, a member of the National Biobank of Korea, supported by the Ministry of Health, Welfare and Family Affairs, Republic of Korea.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Chang, H., Shin, S., Lee, T. et al. Development of a genetic marker set to diagnose aspirin-exacerbated respiratory disease in a genome-wide association study. Pharmacogenomics J 15, 316–321 (2015). https://doi.org/10.1038/tpj.2014.78
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2014.78
This article is cited by
-
Aspirin exacerbated respiratory disease (AERD): molecular and cellular diagnostic & prognostic approaches
Molecular Biology Reports (2021)
-
Olfaction and sinonasal symptoms in patients with CRSwNP and AERD and without AERD: a cross-sectional and longitudinal study
European Archives of Oto-Rhino-Laryngology (2017)