Abstract
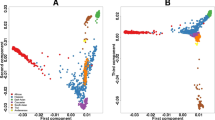
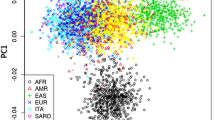
Pharmacogenomically relevant markers of drug response and adverse drug reactions are known to vary in frequency across populations. We examined minor allele frequencies (MAFs), genetic diversity (FST) and population structure of 1156 genetic variants (including 42 clinically actionable variants) in 212 genes involved in drug absorption, distribution, metabolism and excretion (ADME) in 19 populations (n=1478). There was wide population differentiation in these ADME variants, reflected in the range of mean MAF (ΔMAF) and FST. The largest mean ΔMAF was observed in African ancestry populations (0.10) and the smallest mean ΔMAF in East Asian ancestry populations (0.04). MAFs ranged widely, for example, from 0.93 for single-nucleotide polymorphism (SNP) rs9923231, which influences warfarin dosing to 0.01 for SNP rs3918290 associated with capecitabine metabolism. ADME genetic variants show marked variation between and within continental groupings of populations. Enlarging the scope of pharmacogenomics research to include multiple global populations can improve the evidence base for clinical translation to benefit all peoples.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Table of pharmacogenomic biomarkers in drug labels http://www.fda.gov/drugs/scienceresearch/researchareas/pharmacogenetics/ucm083378.htm.
Wang L, McLeod HL, Weinshilboum RM . Genomics and drug response. N Engl J Med 2011; 364: 1144–1153.
Schwab M, Schaeffeler E . Pharmacogenomics: a key component of personalized therapy. Genome Med 2012; 4: 93.
Deeken J . The Affymetrix DMET platform and pharmacogenetics in drug development. Curr Opin Mol Therapeut 2009; 11: 260–268.
VeraCode ADME core panel. http://res.illumina.com/documents/products/datasheets/datasheet_veracode_adme_core_panel.pdf.
Rotimi CN, Dunston GM, Berg K, Akinsete O, Amoah A, Owusu S et al. In search of susceptibility genes for type 2 diabetes in West Africa: the design and results of the first phase of the AADM study. Ann Epidemiol 2001; 11: 51–58.
Adeyemo A, Gerry N, Chen G, Herbert A, Doumatey A, Huang H et al. A genome-wide association study of hypertension and blood pressure in African Americans. PLoS Genet 2009; 5: e1000564.
Shriner D . Investigating population stratification and admixture using eigenanalysis of dense genotypes. Heredity (Edinb) 2011; 107: 413–420.
Li J, Zhang L, Zhou H, Stoneking M, Tang K . Global patterns of genetic diversity and signals of natural selection for human ADME genes. Hum Mol Genet 2011; 20: 528–540.
Wang C, Szpiech ZA, Degnan JH, Jakobsson M, Pemberton TJ, Hardy JA et al. Comparing spatial maps of human population-genetic variation using Procrustes analysis. Stat Appl Genet Mol Biol 2010; 9, Article 13.
Limdi NA, Veenstra DL . Warfarin pharmacogenetics. Pharmacotherapy 2008; 28: 1084–1097.
Schwab M, Zanger UM, Marx C, Schaeffeler E, Klein K, Dippon J et al. Role of genetic and nongenetic factors for fluorouracil treatment-related severe toxicity: a prospective clinical trial by the German 5-FU Toxicity Study Group. J Clin Oncol 2008; 26: 2131–2138.
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 2010; 467: 1061–1073.
Tishkoff SA, Reed FA, Friedlaender FR, Ehret C, Ranciaro A, Froment A et al. The genetic structure and history of Africans and African Americans. Science (New York, NY) 2009; 324: 1035–1044.
Shriner D . Improved eigenanalysis of discrete subpopulations and admixture using the minimum average partial test. Human Heredity 2012; 73: 73–83.
Clayton DG, Walker NM, Smyth DJ, Pask R, Cooper JD, Maier LM et al. Population structure, differential bias and genomic control in a large-scale, case-control association study. Nat Genet 2005; 37: 1243–1246.
Rosenberg NA, Huang L, Jewett EM, Szpiech ZA, Jankovic I, Boehnke M . Genome-wide association studies in diverse populations. Nat Rev Genet 2010; 11: 356–366.
Larussa T, Suraci E, Lentini M, Nazionale I, Gallo L, Abenavoli L et al. High prevalence of polymorphism and low activity of thiopurine methyltransferase in patients with inflammatory bowel disease. Eur J Intern Med 2012; 23: 273–277.
Dai Z, Papp AC, Wang D, Hampel H, Sadee W . Genotyping panel for assessing response to cancer chemotherapy. BMC Med Genomics 2008; 1: 24.
Santos PC, Soares RA, Santos DB, Nascimento RM, Coelho GL, Nicolau JC et al. CYP2C19 and ABCB1 gene polymorphisms are differently distributed according to ethnicity in the Brazilian general population. BMC Med Genet 2011; 12: 13.
Tegretol Drug Label http://www.accessdata.fda.gov/drugsatfda_docs/label/2012/016608s107,018281s055,018927s048,020234s040lbl.pdf.
Langley MR, Booker JK, Evans JP, McLeod HL, Weck KE . Validation of clinical testing for warfarin sensitivity: comparison of CYP2C9-VKORC1 genotyping assays and warfarin-dosing algorithms. J Mol Diagn 2009; 11: 216–225.
Ross KA, Bigham AW, Edwards M, Gozdzik A, Suarez-Kurtz G, Parra EJ . Worldwide allele frequency distribution of four polymorphisms associated with warfarin dose requirements. J Hum Genet 2010; 55: 582–589.
Sung C, Lee PL, Tan LL, Toh DS . Pharmacogenetic risk for adverse reactions to irinotecan in the major ethnic populations of Singapore: regulatory evaluation by the health sciences authority. Drug Saf 2011; 34: 1167–1175.
Bhatia G, Patterson N, Pasaniuc B, Zaitlen N, Genovese G, Pollack S et al. Genome-wide comparison of African-ancestry populations from CARe and other cohorts reveals signals of natural selection. Am J Human Genet 2011; 89: 368–381.
Johnson JA, Gong L, Whirl-Carrillo M, Gage BF, Scott SA, Stein CM et al. Clinical Pharmacogenetics Implementation Consortium Guidelines for CYP2C9 and VKORC1 genotypes and warfarin dosing. Clin Pharmacol Ther 2011; 90: 625–629.
Rotimi CN, Jorde LB . Ancestry and disease in the age of genomic medicine. N Engl J Med 2010; 363: 1551–1558.
Serrano D, Lazzeroni M, Zambon CF, Macis D, Maisonneuve P, Johansson H et al. Efficacy of tamoxifen based on cytochrome P450 2D6, CYP2C19 and SULT1A1 genotype in the Italian Tamoxifen Prevention Trial. Pharmacogenomics J 2011; 11: 100–107.
He YJ, Shapero MH, McLeod HL . Novel tagging SNP rs1495741 and 2-SNPs (rs1041983 and rs1801280) yield a high prediction of the NAT2 genotype in HapMap samples. Pharmacogenet Genomics 2012; 22: 322–324.
Ramos E, Callier SL, Rotimi CN . Why personalized medicine will fail if we stay the course. Personalized Med 2012; 9: 839–847.
Acknowledgements
The informatics expertise of Kevin Long, University of North Carolina, Chapel Hill is greatly appreciated. The study was supported by National Institutes of Health grants S06GM008016-320107 to CNR and S06GM008016-380111 to AA. HUFS participants were enrolled at the Howard University General Clinical Research Center, which is supported by grant 2M01RR010284 from the former National Center for Research Resources, National Institutes of Health. This research was supported in part by the Intramural Research Program of the Center for Research on Genomics and Global Health. The Center for Research on Genomics and Global Health is supported by the National Human Genome Research Institute, the National Institute of Diabetes and Digestive and Kidney Diseases, the Center for Information Technology and the Office of the Director at the National Institutes of Health (Z01HG200362).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Ramos, E., Doumatey, A., Elkahloun, A. et al. Pharmacogenomics, ancestry and clinical decision making for global populations. Pharmacogenomics J 14, 217–222 (2014). https://doi.org/10.1038/tpj.2013.24
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2013.24
Keywords
This article is cited by
-
Genetic polymorphisms associated with upper gastrointestinal bleeding: a systematic review
The Pharmacogenomics Journal (2021)
-
Population structure and pharmacogenomic risk stratification in the United States
BMC Biology (2020)
-
Phenotype prediction and characterization of 25 pharmacogenes in Thais from whole genome sequencing for clinical implementation
Scientific Reports (2020)
-
Evaluating the promise of inclusion of African ancestry populations in genomics
npj Genomic Medicine (2020)
-
Polymorphisms of ADME-related genes and their implications for drug safety and efficacy in Amazonian Amerindians
Scientific Reports (2019)