Abstract
Stress during pregnancy may impact subsequent generations, which is demonstrated by an increased susceptibility to childhood and adulthood health problems in the children and grandchildren. Although the importance of the prenatal environment is well reported with regards to future physical and emotional outcomes, little is known about the molecular mechanisms that mediate the long-term consequences of early stress across generations. Recent studies have identified DNA methylation as a possible mediator of the impact of prenatal stress in the offspring. Whether psychosocial stress during pregnancy also affects DNA methylation of the grandchildren is still not known. In the present study we examined the multigenerational hypothesis, that is, grandmaternal exposure to psychosocial stress during pregnancy affecting DNA methylation of the grandchildren. We determined the genome-wide DNA methylation profile in 121 children (65 females and 56 males) and tested for associations with exposure to grandmaternal interpersonal violence during pregnancy. We observed methylation variations of five CpG sites significantly (FDR<0.05) associated with the grandmother’s report of exposure to violence while pregnant with the mothers of the children. The results revealed differential methylation of genes previously shown to be involved in circulatory system processes (FDR<0.05). This study provides support for DNA methylation as a biological mechanism involved in the transmission of stress across generations and motivates further investigations to examine prenatal-dependent DNA methylation as a potential biomarker for health problems.
Similar content being viewed by others
Introduction
The impact of early-life adversity likely persists in subsequent generations and may influence the individual’s physiology and behavior. There is considerable evidence that exposure to prenatal stress can induce stable changes that influence neurodevelopment, mental health and the risk for psychiatric disorders.1, 2, 3, 4 Studies with animal models have provided strong support for the idea that the effects induced by early stress are not limited to the offspring. In guinea pigs, for example, the impact of maternal nutrition during pregnancy on hypothalamic pituitary adrenal (HPA) axis function and heart structure can be observed in both the second and third generation.5 Rats exposed to excess glucocorticoids in utero showed reduced birth weight and glucose intolerance into the third generation.6 Animal models also show an accumulation of the effects of prenatal stress over several generations on brain function7 and physiological response to stress.8 In humans, the available body of findings indicating multigenerational effects of early stress (that is, effects observed in the children and grandchildren) stem mainly from epidemiological studies supporting the role of grandparental adversities in shaping grandchildren health outcomes.9, 10 For instance, the paternal grandfather's nutrition in Överkalix Sweden, was linked to the mortality risk ratio of grandsons, while the paternal grandmother’s nutrition was linked to the mortality risk ratio of the granddaughters.10 Another example is the effect of famine during grandmaternal pregnancy in the time of the Dutch Hunger Winter (1944–1945) on adiposity in grandchildren.9
There is growing support for the role of epigenetic mechanisms, especially DNA methylation, in the persistent effects of prenatal stress on child’s development11, 12, 13, 14, 15, 16, 17, 18, 19 (for a review Serpeloni et al20). Studies in animal models suggest that DNA methylation modifications have the potential to be transmitted to further generations.21, 22, 23 Despite increasing evidence of the effect of prenatal stress on the offspring’s DNA methylation, the existing evidence in humans relies on investigations of mother-child dyads. Recently, a multigenerational study in humans showed changes in grandchild DNA methylation following environmental lead exposure of their pregnant grandmothers.24 In the present study, we were interested in examining the impact of psychological stress during pregnancy using a multigenerational design (Figure 1a). It must be emphasized that this is not a transgenerational study, since the first-generation (G1) interacts not only with the fetal offspring (first-generation offspring, G2) but also with the germ cells developing within those offspring, which mature into the sperm and eggs that give rise to the grandchildren (second-generation offspring, G3). Using a genome-wide DNA methylation approach, we aimed at investigating to what extent grandmaternal stress during pregnancy was associated with genome-wide methylation changes in the grandchildren (G3) (Figure 1b).
Illustration of the study design. (a) G1 represents the grandmother, G2 the developing fetus, and G3 the germ cells (future grandchild). (b) Flow diagram of the design of the study indicating the psychosocial data and the biological data. Retrospective interviews of 121 maternal grandmothers were conducted about interpersonal violence exposure during pregnancy. Saliva samples were collected from their grandsons/granddaughters. Epigenome-wide association analysis was performed between exposure to interpersonal violence during grandmaternal pregnancy and DNA methylation of the grandchild. CDV, community and domestic violence; EWAS, epigenome-wide association study; IPV, intimate partner violence.
Materials and methods
Participants
In a cross-sectional study design, a sample of 121 teenage youth (M age=13.63 years, s.d.=2.48, age range=8–18 years; 65 females) and their maternal grandmothers (N=121, M age=64.12 years, s.d.=8.19, age range=46–88 years) participated in the study. This investigation was conducted in the context of a larger study25 dealing with the impact of early exposure to violence across generations. The study was carried out with families living in São Gonçalo, a city located in the state of Rio de Janeiro, Brazil. São Gonçalo has a population of over 1 million, with a high proportion of low-income families and high levels of domestic and community violence.26, 27, 28 Local health community agents contacted the families registered in the Family Health Program of the city to participate in the study. The families were contacted if the maternal grandmother and the grandchild (8–18 years old) were both living in the city of São Gonçalo. The study received approval from the research ethics board of the University of Konstanz (DE) and the National Commission for Ethics in Research—CONEP (BR). The participation was voluntary and no monetary compensation was offered. We obtained written informed consent from the adult participants, as well as written informed consent from parents and written assent from the youth. A team of five trained interviewers carried out structured interviews with the participants. Grandmother and grandchild were interviewed individually in their home setting. Demographic information was collected, including the participant’s age and sex.
Interpersonal violence during grandmaternal pregnancy
The Composite Abuse Scale (CAS)29 was used to measure grandmaternal exposure to intimate partner violence (IPV). It consisted of 30 items, which are scaled from a frequency of zero (never) to five (daily). A score equal to or higher than three was used as the criterion for exposure to IPV, as recommended29, 30 (IPV+ and IPV−). A Cronbach’s alpha of 0.90 suggested excellent internal consistency. To assess grandmaternal exposure to community and domestic violence (CDV) we applied ‘Things I have seen and heard’.31 This instrument has 20 items and an additional 6 that measure exposure to CDV. Items are presented in a 5-point format ranking from zero (never) to four (many times). Higher scores reflect higher exposure to violence.31 Cronbach’s alpha was 0.83. Under the assumption that the greatest effect of stress during pregnancy on DNA methylation is observed in higher exposure to adversities32 we divided the participants in groups of high and low or none exposure to violence during grandmaternal pregnancy (CDV+ and CDV−). Grandmaternal exposure to violence during pregnancy was conducted three times, separately focusing on the periods before, during and after pregnancy.
Mental health problems
Post-traumatic stress disorder (PTSD) symptom severity was measured using the University of California at Los Angeles Post-traumatic Stress Disorder Reaction Index DSM-IV-UCLA33 (Cronbach’s α=0.88, possible range 0–88). Depression symptom severity was assessed with the Patient Health Questionnaire PHQ-934 (Cronbach’s α=0.80, possible range 0–27).
DNA methylation and data preprocessing
Saliva samples (2 ml) for DNA methylation analysis were collected in the end of the interview using a non-invasive tool, the OrageneDiscover (OGR-500) Collection Kit (DNA Genotek; Ottawa, ON, Canada). Genome-wide DNA methylation profiling was performed using the Infinium HumanMethylation450 BeadChip Kit and arrays by the Queen Mary University of London Genome Center, 500 ng genomic DNA was prepared and hybridized according to manufacturer’s specification (Illumina, San Diego, CA, USA, catalog # WG-914-1002, Part # 15019522 Rev. A, 2010). We verified the bisulfite conversion, hybridization and other quality measures for the remaining samples with qcReport in Minfi R package.35 Quantile normalization and probe filtering were also done using Minfi.35 To avoid problems with multiple testing, we choose to apply a conservative approach to filter out probes with any sign of low signal to background ratio. For this we included a total data set of 396 samples/arrays, which beside the children samples also contained samples from the grandmaternal and maternal generation (not part of the present study but quality controlled the same way as the children). A probe was excluded if the signal intensities were not distinguishable from the background intensity at a significance level of 0.01 in any of the 396 samples (N=270 296). After that we excluded cross-reactive probes (remaining probes N=247 548), SNP probes at interrogated CpG and within probes36 (remaining probes N=160 022) and sex chromosomes, resulting in a final data set of 155337 probes of highest quality. Approximately 40% of all the probes identified in the detection P-value process were within cross-reactive, SNPs (targeted and within), as well as sex chromosomes.
We validated the biological relationships of the family members and screened for contaminants/bad quality samples using the 65 control SNP probes included on the Infinium HumanMethylation450 BeadChip as follows. The variation in beta values of the 65 control probes provides an empirical and experiment-specific reference for quality control of preprocessing and data acquisition (for example, R packages wateRmelon or Leiden450K).37 In the present study, we extended the use of the 65 control SNPs to validate the parent-offspring relations using a simple exclusion criterion, the presence of mendelian incompatibilities. The beta values for the SNP probes present a trimodal distribution after appropriate preprocessing. This distribution allowed for straightforward genotyping of 99.8% of the SNP genotypes (total number of genotypes>25 000) using the R function cut through the categorization of beta values from each of the three peaks corresponding to the different genotypes. Very few beta values (0.2%, n=54) were intermediate between the otherwise non-overlapping peaks and were considered ambiguous (i.e. treated as missing data). A very large proportion of the ambiguous genotype calls (42/54) stemmed from only three samples (all other 12 ambiguous genotypes were unique), indicating poor data quality. These samples (one child and two adults) were excluded from further analysis. The obtained genotypic frequencies were tested for accordance to the Hardy-Weinberg proportions to confirm that our genotyping approach was accurate and unbiased. We then calculated a matrix of Mendelian incompatibilities (that is, CC individuals cannot be parents or offspring of GG individuals, and vice-versa) between all individuals in the data set. One case of incompatible parent-offspring pair involved samples that were completely genotypically identical to other individuals in the data set, indicating cross-contamination. This sample was also excluded from further analysis.
In summary, in the child subset, samples that were contaminated (n=1) or had insufficient genotyping quality (n=1) were excluded. As well as samples whose DNA (n=2) failed bisulfite conversion based on Nanodrop measurements following the conversion stage were removed from further analysis. Furthermore, three participants had to be excluded due missing data regarding grandmaternal stress during pregnancy. In total, 121 (out of 128) saliva samples from the children original data set were included in the final analysis.
In addition to the quality control, the match between reported sex and predicted epigenetic sex, was also performed using the Minfi R package. To control for genetic structure and cell-type heterogeneity bias between grandmaternal IPV+ and IPV−, as well as CDV+ and CVD− groups, we applied a combination of methods available within GLINT 1.0.338 (http://glint-epigenetics.readthedocs.io/en/latest/). EPISTRUCTURE was used for genetic ancestry analysis,39 while ReFACTor was used for reference free,40 and the Houseman method was used for reference based cell-type analysis.41 Since, saliva contains many white-blood cells we used the Houseman blood reference available in GLINT, as well as a custom-made reference (Supplementary Table S1) based on differences between saliva, buccal and a pool of the other tissues available in the GEO-data set GSE48472 published by Slieker et al.42 Our analysis showed strong similarities to the Slieker et al. saliva samples, with an expected small ratio of buccal cells, and no signs of differences in ancestry or cellular heterogeneity between groups (Supplementary Table S2).
Statistical analysis
All analyses were conducted using R 3.2.1. To investigate to what extent prenatal stress has an impact on DNA methylation in the grandchildren, logit-transformed beta values (M-values) were subjected to a robust linear regression model to identify significantly differentially methylated probes in association with grandmaternal exposure to violence during pregnancy. The analysis was adjusted for multiple testing (Benjamini–Hochberg method), using the Limma R package.43 We used robust linear modeling to minimize the effect of sporadic outliers (such as unusually high abundance of rare SNPs affecting methylation sites in this particular cohort). The false discovery rate (FDR) associated loci was later categorized into either the high-confidence group (5% or less FDR) or a medium-confidence group (25% or less FDR) as used earlier.44, 45 IPV and CDV were used as categorical variables in the model (probe_meth~ipv+cdv+age+sex). For IPV, according to the CAS manual, a sensitivity test suggest that a cutoff score of three would correctly detect 100% of abused woman.30 Regarding the CDV, the variable was split in two groups. A group of extreme exposure and a group with lower or none exposure based on the upper quartile of the sum scores (scores higher than four). We believe that the precision of the interviews may be not accurate enough to robustly detect a difference between, for example, 2 and 4 score. In such case exposure vs low exposure may be more accurate. To control for the possible confounding effects of sex and age, both were included as covariates in the model. Correlation analysis was performed to further study the relationship between epigenetic and psychological variables.
Technical validation by NGS sequencing
Targeted bisulfite sequencing was used to validate the methylation values of the CpGs. Primer design and validation was performed on 10 probes by Zymo Research (Irvine, CA, USA) on bisulfite-converted control DNA.46 The targeted relevant regions from the bisulfite-converted DNA were amplified and the PCR product were sequenced. The PCR amplicons were between 100 and 300 bp. Samples were bisulfite converted using the EZ DNA Methylation- LightningTM Kit (www.zymoresearch.com—catalog number D5030) according to the manufacturer’s instructions. Multiplex amplification of all samples using ROI-specific primer pairs and the Fluidigm Access Array System (Irvine, CA, USA) was performed according the to the manufacturer’s instructions. The resulting amplicons were pooled for harvesting and subsequent barcoding according to the Fluidigm instrument’s guidelines. After barcoding, samples were purified (ZR-96 DNA Clean & Concentrator—ZR, Cat#D4023, Irvine, CA, USA) and then prepared for massively parallel sequencing using a MiSeq V2 300 bp Reagent Kit and paired-end sequencing protocol according to the manufacturer’s guidelines. Sequence reads were identified using standard Illumina base-calling software. Low-quality nucleotides and adapter sequences were trimmed off during QC: Sequence reads were aligned back to the reference genome using Bismark (http://www.bioinformatics.babraham.ac.uk/projects/bismark/), an aligner optimized for bisulfite sequence data and methylation calling.47 The methylation level of each sampled cytosine was estimated as the number of reads reporting a C, divided by the total number of reads reporting a C or T.
Results
Demographics, grandmaternal stress and grandchild outcomes
In our study, 98% of the grandmothers reported being exposed to at least one event of CDV during their life. In total, 26% (n=31) of the grandmothers were exposed to high levels of CDV during their lifetime and 22% (n=27) during the pregnancy of interest. Regarding IPV, 20% (n=24) experienced IPV during pregnancy and 44% (n=53) during their lifetime. Only 7% (n=9) of the grandmothers reported having experienced both CDV and IPV during pregnancy. Demographic and mental health measures of participating grandchildren are shown in Table 1.
Association of grandmaternal exposure to violence during pregnancy and genome-wide methylation in the grandchildren
Genome-wide methylation analysis revealed five CpG sites significantly (FDR<0.05) associated with grandmaternal exposure to violence during pregnancy and an additional list of 22 CpG sites significantly associated (FDR<0.25), according to the criteria of medium-confidence as shown in Table 2.
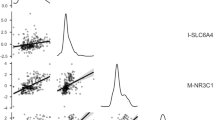
We observed that grandmaternal exposure to CDV during pregnancy was significantly associated at high confidence (FDR<0.05) with decreased methylation at the CpG sites within CORIN (logFC=−0.30, P=8.12E−08), SMYD3 (logFC=−0.88, P=3.29E−07), and BARX1 (logFC=−0.17, P=6.09E−07), as well as increased methylation at the CpG sites within CFTR (logFC=0.32, P=1.63E−07) and CpG cg26684363 (logFC=0.22, P=8.10E−07), which had no gene associated, according the UCSC genome browser (Figure 2). To validate the final model (probe_meth~ipv+cdv+age+sex), we used bootstrapping validation in the rms package.48 Using a resampling rate of 1000 showed that all significant CpGs (Table 2) increased their R2-values when we sequentially added each covariate to the model (R2 differences between final model and models only including Grandmaternal IPV or CDV: mean=0.08, min=0.02, max=0.19; the estimated optimism of the final model: mean relative =0.04, min relative=0.03, max relative=0.06; mean absolute=−0.005, min absolute=−0.045, max absolute=−0.001).
Volcano plot for genome-wide DNA methylation analysis of the grandchild (N=121) associated with grandmaternal exposure to CDV during pregnancy using a linear regression. Each point represents a CpG site. The red dots are significant CpG sites after correction for multiple comparisons in the whole genome with high confidence (FDR <0.05), and the orange dots are CpG sites with medium confidence (FDR<0.25). CDV, community and domestic violence during grandmaternal pregnancy; FDR, false discovery rate.
To further investigate the biological processes involved in the differentially methylated CpG sites associated with grandmaternal exposure to CDV, we performed enrichment analysis using the WEB-based Gene SeT AnaLysis Toolkit (WebGestalt; http://www.webgestalt.org/).49 As input data, we used the genes from the high-confidence group above (FDR<0.05) from the genome-wide methylation analysis (n=4, significant probes overlapping with genes, Table 2). After correction for multiple testing using the Benjamini-Hochberg method (FDR<0.05), the analysis revealed an over-representation for the genes CORIN and CFTR, involving diseases associated with circulatory system processes as well as congenital abnormalities. The results show that there was no significant correlation within the neighboring probes (P<0.05) of the top five CpGs (FDR 0.05; Supplementary Table S3). We screened the http://www.mqtldb.org/ database for mQTL50 that are known to influence the methylation of our 27 significant CpGs. Several thousand genetic variants have small effects on the methylation levels of 19/27 of our significant CpG’s, particularly for cg23275840 and cg24478129 (Supplementary Table S4).
To explore the association between mental health variables (depression and PTSD) and DNA methylation at the top four CpG probes affected by grandmaternal stress during pregnancy, Pearson’s correlations were performed (Table 3). The analysis reveals that CFTR methylation was significantly correlated with depression (r (121),=0.21, P<0.05) and PTSD (r (121),=0.20, P<0.05). None of the mental health variables were correlated with grandmaternal exposure to CDV during pregnancy.
Technical validation by NGS sequencing
We validated the arrays by technically replicating 10 CpGs, including the top significant CpG site cg21212505_CFTR, on the BeadChip in 33 chosen individuals using targeted bisulfite resequencing, showing strong inter-assay reliability within subjects (mean r2=0.94; max=0.96; min=0.90; each subject had P<0.001).
Discussion
The aim of this study was to examine the multigenerational epigenetic effects of psychosocial stress. Our results show that grandmaternal exposure to interpersonal violence during pregnancy is associated with 27 differentially methylated CpG sites in the children that map to 22 uniquely annotated genes. Within the CpG sites associated with grandmaternal prenatal stress, five had a high-confidence significance (FDR<0.05).
The most differentially methylated genes (high-confidence group) associated with grandmaternal stress during pregnancy were: CFTR, CORIN, BARX1 and SMYD3. CFTR (cystic fibrosis transmembrane conductance regulator) and CORIN (corin, serine peptidase) genes are both involved in circulatory system processes and congenital abnormalities. Furthermore, changes in methylation of CFTR was previously shown in adolescents from institutions for orphaned and abandoned children with a history of adversities.51 Prior studies have shown that dysregulation of CFTR has an impact on the release of vitamin D, which may affect depression symptoms.52 CORIN is associated with blood pressure regulation and preeclampsia and is essential for physiological changes at the maternal-fetal interface. Animal studies have shown that this gene plays an important role in preventing high blood pressure during pregnancy.53 CORIN was pointed to as a possible methylation-based marker gene involved in pregnancy-associated diseases.54 Dysregulation of CORIN expression is implicated in cardiovascular diseases such as heart failure and hypertension. BARX1 (BARX homeobox 1) is involved in embryonic development and plays a role in craniofacial development, odontogenesis, and stomach organogenesis.
Since CORIN and CFTR are associated with circulatory system processes, future studies integrating clinical characteristics might shed light on whether differential DNA methylation play a role in ill-health outcomes. Prenatal exposure to stress has been associated with cardiovascular disease risk in previous studies.55 Children that were exposed to the Dutch Hunger Winter during prenatal period experienced elevated rates of obesity, altered lipid profiles and cardiovascular disease.55 Blood pressure and increased stress responsiveness were also associated with prenatal exposure to the Dutch Hunger Winter.56 Studies in animal models have reported multigenerational impacts of stress during pregnancy on heart structure in the second and third generation.5
The possibility that environmental factors promote epigenetic susceptibility to disease in subsequent generations has medical and public health implications. In our data there was a significant association of methylation in the CFTR gene with depression and PTSD symptoms in the children. Little is known about the role of CFTR in mental health problems. As indicated above, dysregulation of CFTR has an impact on the release of vitamin D and might be implicated in depression.52 This is supported in patients with cystic fibrosis, a condition that may be caused by a genetic abnormality in the CFTR, where a link between vitamin D insufficiency and depression symptoms has been demonstrated.57 The comorbidity of cardiovascular disease and major depressive disorder represents the fourth leading cause of morbidity and mortality worldwide.58 Future studies investigating the relationship between mental health and epigenetic programming of CFTR are needed to better characterize the pathways involved in depression and PTSD. Our findings suggest that epigenetic modifications might mediate the impact of stress on health outcomes across generations.59, 60, 61, 62, 63, 64
Genome-wide methylation studies allow the discovery of new signatures of stress, providing a better understanding of the complex pathways involved in different regulatory mechanisms susceptible to stress during pregnancy. Whether those variations can be considered as a reliable biomarker of a specific trait needs to be further investigated. Future studies on DNA methylation alterations and it’s role in the development of health and disease might clarify the question of using DNA methylation as a biomarker of stress. Based on the LogFC and the P-values we observe that the effects on DNA methylation of the third generation are modest, which is expected based on the literature on the impact of environmental stress on DNA methylation17 specially considering the complexity of the path between grandmaternal experience and epigenetic regulation in the third generation. Larger LogFC represents stronger differential methylation, but current knowledge does not allow a prediction of the observed effects on gene expression59 and therefore potential functional effects are unclear. In the field of cancer research, studies tend to report larger effects60, 61, 62 but this setting is not straight comparable with variation associated to psychiatric traits because the path is more direct and the effects are likely to be more extreme. However, it has been argued that modest changes might have functional impact in certain conditions such as psychiatry disorders, if they are stable and produce meaningful changes59 and this could be tested in future work on model organisms. Our study does not allow us to establish causality between grandmaternal stress and mental health in the grandchildren. The phenotypic consequences of epigenetic modifications in the grandchild associated with grandmaternal stress, such as stress reactivity, brain function, and behavior, remains to be explored in future studies. Considering that brain tissue is not accessible in living humans, epigenetic studies are usually conducted in peripheral tissues, such as blood and saliva. Saliva has the advantage of being readily collectable63 and is a non-invasive method. While DNA methylation patterns are tissue specific, different studies have shown correlations between saliva and blood,32 as well as between peripheral tissues and brain.63, 64 Therefore, DNA methylation from saliva is a valuable opportunity for studies of DNA methylation signatures of psychosocial exposure. Nevertheless, extrapolating the results across tissues must be done with caution.
To the best of our knowledge, this is the first investigation in humans of the impact of grandmaternal psychosocial stress during pregnancy on grandchild DNA methylation. Our data suggest that not only exposure to toxin during pregnancy24 but also grandmaternal psychosocial stress can be translated into DNA methylation changes in the grandchild. Although the significance level is not considered high, it is comparable with other studies on the epigenetic impact of prenatal adversities. For instance, differential DNA methylation after early gestational famine exposure, prenatal psychosocial stress16, 17 and psychiatry disorders.46 Evidence of replication of DNA methylation findings,46 brings confidence in the validity of the results. However, lack of replication have been also reported51 and raise concerns about the reliability of the results. Our conclusions and results are somewhat limited by the difficulty of obtaining such data in large numbers, which likely leads to false-negatives given that the effect sizes tend to be rather small.65, 66, 67 Furthermore, the absence of published multigenerational studies on humans precludes strong conclusions, which will require future work on independent populations.
While it is possible that different external factors, such as smoking68, 69 and nutrition70, 71 have an impact on DNA methylation patterns, they were not examined in the present study. Future studies should covariate for other external factors such as nutrition and smoking. Another potential confounding effect is the existence of genetic differences at loci that influence methylation patterns (mQTL). Although it is possible that genetic variation at mQTLs might have had an influence in our results, it is not possible to test for differences in allele frequency between the relevant groups in the absence of high-throughput genotyping data.
The effect sizes are in fact expected to be rather small and a number of factors might have further reduced these effects in our analysis. For instance, we did not account for different timescales during pregnancy and stress exposure during different trimesters has indeed shown to account for offspring differences in methylation profiles.72 Furthermore, we relied on retrospective information of events happening during pregnancy reported by the grandmothers. However, both the lack of time resolution and reliance on retrospective information is likely to reduce effects rather than produce them. Nevertheless, we detected epigenome-wide significant effects in spite of these limitations.
The results highlight, on a molecular basis, the importance of women’s wellbeing during pregnancy and the epigenetic consequence of grandmaternal stress on the grandchild. Considering ancestral stress on lifetime health trajectories is critical to improve the strategies that support healthy development of children. While guidelines for proper nutrition or avoidance of toxin exposure, such as tobacco and alcohol, have been incorporated in the routine care of pregnant women, psychosocial risk factors have largely been neglected. Given that prenatal stress may affect the mother, child, and (as we have shown) the grandchild, pregnancy is a critical period to screen for psychosocial risk factors and apply early interventions to promote health in the general public.
References
Huizink AC, Mulder EJ, Buitelaar JK . Prenatal stress and risk for psychopathology: specific effects or induction of general susceptibility? Psychological bulletin 2004; 130: 115.
Lupien SJ et al. Effects of stress throughout the lifespan on the brain, behaviour and cognition. Nature Reviews Neuroscience 2009; 10: 434–445.
O'connor TG et al. Maternal antenatal anxiety and children's behavioural/emotional problems at 4 years. The British Journal of Psychiatry 2002; 180: 502–508.
DiPietro JA, Novak MFSX, Costigan KA, Atella LD, Reusing SP . Maternal psychological distress during pregnancy in relation to child development at age two. Child Dev 2006; 77: 573–587.
Bertram C, Khan O, Ohri S, Phillips DI, Matthews SG, Hanson MA . Transgenerational effects of prenatal nutrient restriction on cardiovascular and hypothalamic-pituitary-adrenal function. J Physiol 2008; 586: 2217–2229.
Drake AJ, Walker BR, Seckl JR . Intergenerational consequences of fetal programming by in utero exposure to glucocorticoids in rats. Am J Physiol 2004; 288: 34–38.
Skelin l, Needham MA, Molina LM, Metz GA, Gruber AJ . Multigenerational prenatal stress increases the coherence of brain signaling among cortico-striatal-limbic circuits in adult rats. Neuroscience 2015; 289: 270–278.
Ward ID, Zucchi FCR, Robbins JC, Falkenberg EA, Olson DM, Benzies K et al. Transgenerational programming of maternal behaviour by prenatal stress. BMC Pregnancy Childbirth 2013; 13: 1–8.
Painter RC, Osmond C, Gluckman PD, Hanson M, Phillips DIW, Roseboom TJ . Transgenerational effects of prenatal exposure to the Dutch famine on neonatal adiposity and health in later life. BJOG 2008; 115: 1243–1249.
Pembrey ME, Bygren LO, Kaati G, Edvinsson S, Northstone K, Sjostrom M et al. Sex-specific, male-line transgenerational responses in humans. Eur J Hum Genet 2005; 14: 159–166.
Babenko O, Kovalchuk I, Metz GAS . Stress-induced perinatal and transgenerational epigenetic programming of brain development and mental health. Neurosci Biobehav Rev 2015; 48: 70–91.
Oberlander TF, Weinberg J, Papsdorf M, Grunau R, Misri S, Devlin AM . Prenatal exposure to maternal depression, neonatal methylation of human glucocorticoid receptor gene (NR3C1) and infant cortisol stress responses. Epigenetics 2008; 3: 97–106.
Radtke KM, Ruf M, Gunter HM, Dohrmann K, Schauer M, Meyer A et al. Transgenerational impact of intimate partner violence on methylation in the promoter of the glucocorticoid receptor. Transl Psychiatry 2011; 1: e21.
Conradt E, Lester BM, Appleton AA, Armstrong DA, Marsit CJ . The roles of DNA methylation of NR3C1 and 11β-HSD2 and exposure to maternal mood disorder in utero on newborn neurobehavior. Epigenetics 2013; 8: 1321–1329.
Hompes T, Izzi B, Gellens E, Morreels M, Fieuws S, Pexsters A et al. Investigating the influence of maternal cortisol and emotional state during pregnancy on the DNA methylation status of the glucocorticoid receptor gene (NR3C1) promoter region in cord blood. J Psychiatr Res 2013; 47: 880–891.
Kertes DA, Kamin HS, Hughes DA, Rodney NC, Bhatt S, Mulligan CJ . Prenatal maternal stress predicts methylation of genes regulating the hypothalamic–pituitary–adrenocortical system in mothers and newborns in the Democratic Republic of Congo. Child Dev 2016; 87: 61–72.
Cao-Lei L, Elgbeili G, Massart R, Laplante DP, Szyf M, King S . Pregnant women/'s cognitive appraisal of a natural disaster affects DNA methylation in their children 13 years later: Project Ice Storm. Transl Psychiatry 2015; 5: e515.
Nemoda Z, Massart R, Suderman M, Hallett M, Li T, Coote M et al. Maternal depression is associated with DNA methylation changes in cord blood T lymphocytes and adult hippocampi. Transl Psychiatry 2015; 5: e545.
Mansell T, Novakovic B, Meyer B, Rzehak P, Vuillermin P, Ponsonby AL et al. The effects of maternal anxiety during pregnancy on IGF2/H19 methylation in cord blood. Transl Psychiatry 2016; 6: e765.
Serpeloni F, Radtke KM, Hecker T, Elbert T . Epigenetic biomarkers of prenatal maternal stress In: Spengler D, Binder E (eds) Epigenetics and Neuroendocrinology: Clinical Focus on Psychiatry vol. 2. Springer International Publishing: Switzerland, 2016, pp 177–196.
Roth TL, Lubin FD, Funk AJ, Sweatt JD . Lasting epigenetic influence of early-life adversity on the BDNF gene. Biological Psychiatry 2009; 65: 760–769.
Skinner MK, Haque CG-BM, Nilsson E, Bhandari R, McCarrey JR . Environmentally induced transgenerational epigenetic reprogramming of primordial germ cells and the subsequent germ line. PLoS One 2013; 8: e66318.
Weaver IC, Cervoni N, Champagne FA, D'Alessio AC, Sharma S, Seckl JR et al. Epigenetic programming by maternal behavior. Nature Neuroscience 2004; 7: 847–854.
Sen A, Heredia N, Senut M-C, Land S, Hollocher K, Lu X et al. Multigenerational epigenetic inheritance in humans: DNA methylation changes associated with maternal exposure to lead can be transmitted to the grandchildren. Sci Rep 2015; 5: 14466.
Serpeloni F Interpersonal violence and epigenetic modifications: the impact of prenatal and lifetime stress across three generations Doctoral Dissertation Thesis. University of Konstanz: Konstanz, Germany, 2016.
Assis SGd, Avanci JQ, Pesce RP, Ximenes LF . The situation of Brazilian children and adolescents with regard to mental health and violence. Ciência & Saúde Coletiva 2009; 14: 349–361.
Pinto LW, Assis SGd . Family and community violence of schoolchildren from the city of São Gonçalo, Rio de Janeiro, Brazil. Revista Brasileira de Epidemiologia 2013; 16: 288–300.
Avanci J, Assis SGd, Oliveira R . A cross-sectional analysis of women’s mental health problems: examining the association with different types of violence among a sample of Brazilian mothers. BMC Womens Health 2013; 13: 1.
Hegarty K, Bush R, Sheehan M . The composite abuse scale: further development and assessment of reliability and validity of a multidimensional partner abuse measure in clinical settings. Violence Victims 2005; 20: 529–547.
Hegarty K, Valpied J . Composite Abuse Scale Manual. Department of General Practice, University of Melbourne: Melbourne, Australia, 2007.
Richters JE, Martinez P . Things I have seen and heard: A Structured Interview for Assessing Young Children’s Violence Exposure. National Institute of Mental Health: Rockville, MD,USA, 1990.
Cao-Lei L, Massart R, Suderman MJ, Machnes Z, Elgbeili G, Laplante DP et al. DNA methylation signatures triggered by prenatal maternal stress exposure to a natural disaster: project ice storm. PLoS ONE 2014; 9: e107653.
Steinberg AM, Brymer MJ, Decker KB, Pynoos RS . The University of California at Los Angeles post-traumatic stress disorder reaction index. Curr Psychiatry Rep 2004; 6: 96–100.
Richardson LP, McCauley E, Grossman DC, McCarty CA, Richards J, Russo JE et al. Evaluation of the Patient Health Questionnaire-9 Item for detecting major depression among adolescents. Pediatrics 2010; 126: 1117–1123.
Aryee MJ, Jaffe AE, Corrada-Bravo H, Ladd-Acosta C, Feinberg AP, Hansen KD et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 2014; 30: 1363–1369.
Chen Y, Lemire M, Choufani S, Butcher D, Grafodatskaya D, Zanke B et al. Discovery of cross-reactive probes and polymorphic CpGs in the Illumina Infinium HumanMethylation450 microarray. Epigenetics 2013; 8: 203–209.
Pidsley R, Wong C, Volta M, Lunnon K, Mill J . A data-driven approach to preprocessing Illumina 450K methylation array data. BMC Genomics 2013; 14: 293.
Rahmani E, Yedidim R, Shenhav L, Schweiger R, Weissbrod O, Zaitlen N et al. GLINT: a user-friendly toolset for the analysis of high-throughput DNA-methylation array data. Bioinformatics 2017; 1: 3.
Rahmani E, Shenhav L, Schweiger R, Yousefi P, Huen K, Eskenazi B et al. Genome-wide methylation data mirror ancestry information. Epigenet Chromatin 2017; 10: 1.
Rahmani E, Zaitlen N, Baran Y, Eng C, Hu D, Galanter J et al. Sparse PCA corrects for cell type heterogeneity in epigenome-wide association studies. Nat Methods 2016; 13: 443–445.
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinformatics 2012; 13: 86.
Slieker RC, Bos SD, Goeman JJ, Bovée JV, Talens RP, van der Breggen R et al. Identification and systematic annotation of tissue-specific differentially methylated regions using the Illumina 450k array. Epigenet Chromatin 2013; 6: 26.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 2015; 43: 47–47.
Lam LL, Emberly E, Fraser HB, Neumann SM, Chen E, Miller GE et al. Factors underlying variable DNA methylation in a human community cohort. Proceedings of the National Academy of Sciences of the United States of America 2012; 109 (Suppl 2): 17253–17260.
Essex MJ, Boyce WT, Hertzman C, Lam LL, Armstrong JM . Epigenetic Vestiges of Early Developmental Adversity: Childhood Stress Exposure and DNA Methylation in Adolescence. Child Dev 2011; 84: 58–75.
Wilmot B, Fry R, Smeester L, Musser ED, Mill J, Nigg JT . Methylomic analysis of salivary DNA in childhood ADHD identifies altered DNA methylation in VIPR2. J Child Psychol Psychiatry 2016; 57: 152–160.
Krueger F, Andrews SR . Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 2011; 27: 1571–1572.
Harrell FE Jr, Harrell MFE Jr, Hmisc D . Package ‘rms’. Vanderbilt University, 2017; 229.
Wang J, Duncan D, Shi Z, Zhang B . WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): update 2013. Nucleic Acids Res 2013; 41: W77–W83.
Gaunt TR, Shihab HA, Hemani G, Min JL, Woodward G, Lyttleton O et al. Systematic identification of genetic influences on methylation across the human life course. Genome Biol 2016; 17: 61.
Esposito EA, Jones MJ, Doom JR, MacIsaac JL, Gunnar M, Kobor MS . Differential DNA methylation in peripheral blood mononuclear cells in adolescents exposed to significant early but not later childhood adversity. Dev Psychopathol 2016; 1–15.
Hoang MT, DeFina LF, Willis BL, Leonard DS, Weiner MF, Brown ES . Association between low serum 25-Hydroxyvitamin D and depression in a large sample of healthy adults: the Cooper Center longitudinal study. Mayo Clin Proc 2011; 86: 1050–1055.
Zhou Y, Wu Q . Corin in natriuretic peptide processing and hypertension. Curr Hypertens Rep 2014; 16: 415–415.
Oudejans C, Van Dijk M . (Epi) genetics of pregnancy-associated diseases. Front Genet 2013; 4: 180.
Roseboom T, de Rooij S, Painter R . The Dutch famine and its long-term consequences for adult health. Early Hum Dev 2006; 82: 485–491.
Painter RC, de Rooij SR, Bossuyt PM, Phillips DI, Osmond C, Barker DJ et al. Blood pressure response to psychological stressors in adults after prenatal exposure to the Dutch famine. J Hypertens 2006; 24: 1771–1778.
Smith BA, Cogswell A, Garcia G . Vitamin D and depressive symptoms in children with cystic fibrosis. Psychosomatics 2014; 55: 76–81.
Goldstein JM, Handa RJ, Tobet SA . Disruption of fetal hormonal programming (prenatal stress) implicates shared risk for sex differences in depression and cardiovascular disease. Front Neuroendocrinol 2014; 35: 140–158.
Daskalakis NP, Yehuda R . Site-specific methylation changes in the glucocorticoid receptor exon 1F promoter in relation to life adversity: systematic review of contributing factors. Front Neurosci 2014; 8: 369.
Stone A, Zotenko E, Locke WJ, Korbie D, Millar EKA, Pidsley R et al. DNA methylation of oestrogen-regulated enhancers defines endocrine sensitivity in breast cancer. Nat Commun 2015; 6: 7758.
Martino D, Maksimovic J, Joo JH, Prescott SL, Saffery R . Genome-scale profiling reveals a subset of genes regulated by DNA methylation that program somatic T-cell phenotypes in humans. Genes Immun 2012; 13: 388–398.
Bergman Y, Cedar H . DNA methylation dynamics in health and disease. Nat Struct Mol Biol 2013; 20: 274–281.
Smith AK, Kilaru V, Klengel T, Mercer KB, Bradley B, Conneely KN et al. DNA extracted from saliva for methylation studies of psychiatric traits: Evidence tissue specificity and relatedness to brain. Am J Med Genet B Neuropsychiatr Genet 2015; 168: 36–44.
Stenz L, Zewdie S, Laforge-Escarra T, Prados J, La Harpe R, Dayer A et al. BDNF promoter I methylation correlates between post-mortem human peripheral and brain tissues. Neurosci Res 2015; 91: 1–7.
Tobi EW et al. DNA methylation signatures link prenatal famine exposure to growth and metabolism. Nature Communications 2014; 5: doi:10.1038/ncomms6592.
Lee KW et al. Prenatal exposure to maternal cigarette smoking and DNA methylation: epigenome-wide association in a discovery sample of adolescents and replication in an independent cohort at birth through 17 years of age. Environmental Health Perspectives (Online) 2015; 123: 193.
Rijlaarsdam J et al. An epigenome-wide association meta-analysis of prenatal maternal stress in neonates: A model approach for replication. Epigenetics 2016; 11: 140–149.
Joubert BR . 450K epigenome-wide scan identifies differential DNA methylation in newborns related to maternal smoking during pregnancy. Environ Health Perspect 2012; vol 120: 1425–1431, 2012; 120(12): A455-A455.
Zeilinger S, Kühnel B, Klopp N, Baurecht H, Kleinschmidt A, Gieger C et al. Tobacco smoking leads to extensive genome-wide changes in DNA methylation. PLoS ONE 2013; 8: e63812.
Heijmans BT, Tobi EW, Stein AD, Putter H, Blauw GJ, Susser ES et al. Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci USA 2008; 105: 17046–17049.
Ehrlich S, Weiss D, Burghardt R, Infante-Duarte C, Brockhaus S, Muschler MA et al. Promoter specific DNA methylation and gene expression of POMC in acutely underweight and recovered patients with anorexia nervosa. J Psychiatr Res 2010; 44: 827–833.
Tobi EW, Lumey L, Talens RP, Kremer D, Putter H, Stein AD et al. DNA methylation differences after exposure to prenatal famine are common and timing- and sex-specific. Hum Mol Genet 2009; 18: 4046–4053.
Acknowledgements
We are thankful to the families who participated and contributed to this work, as well as to the NGO ‘Mulheres em Movimento’, the ‘Estratégia de Saúde da Família’ and the health community agents from São Gonçalo-RJ for their support facilitating the fieldwork. We are also gratefully to Prof. Dr. Elizabeth Wieling, Jeanine Narrog, Dr. Maggie Schauer, Dr. Susanne Schaal and Dr. Joviana Avanci for their contributions in this research. This work was funded by a grant from the European Research Council (ERC) through ERC advanced grant 323977 (to T.E), and a grant from the CAPES/DAAD 18354/12-2 (to F.S).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Translational Psychiatry website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Serpeloni, F., Radtke, K., de Assis, S. et al. Grandmaternal stress during pregnancy and DNA methylation of the third generation: an epigenome-wide association study. Transl Psychiatry 7, e1202 (2017). https://doi.org/10.1038/tp.2017.153
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tp.2017.153
This article is cited by
-
Advances in pathogenesis of preeclampsia
Archives of Gynecology and Obstetrics (2024)
-
A novel approach to risk exposure and epigenetics—the use of multidimensional context to gain insights into the early origins of cardiometabolic and neurocognitive health
BMC Medicine (2023)
-
Dissecting early life stress-induced adolescent depression through epigenomic approach
Molecular Psychiatry (2023)
-
Inheritance of paternal lifestyles and exposures through sperm DNA methylation
Nature Reviews Urology (2023)
-
Grandmaternal smoking during pregnancy is associated with differential DNA methylation in peripheral blood of their grandchildren
European Journal of Human Genetics (2022)