Abstract
Virulence plasmid (VP) acquisition was a key step in the evolution of Shigella from a non-pathogenic Escherichia coli ancestor to a pathogenic genus. In addition, the co-evolution and co-ordination of chromosomes and VPs was also a very important step in the evolutionary process. To investigate the cross-talk between VPs and bacterial chromosomes, we analyzed the expression profiles of protein complexes and protein monomers in three wild-type Shigella flexneri strains and their corresponding VP deletion mutants. A non-pathogenic wild-type E. coli strain and mutant E. coli strains harboring three Shigella VPs were also analyzed. Comparisons showed that the expression of chromosome-encoded proteins GadA/B and AtpA/D, which are associated with intracellular proton flow and pH tuning of bacterial cells, was significantly altered following acquisition or deletion of the VP. The acid tolerance of the above strains was also compared, and the results confirmed that the presence of the VP reduced the bacterial survival rate in extremely acidic environments, such as that in the host stomach. These results further our understanding of the evolution from non-pathogenic E. coli to Shigella, and highlight the importance of co-ordination between heterologous genes and the host chromosome in the evolution of bacterial species.
Similar content being viewed by others
Introduction
Bacillary dysentery, or shigellosis, is caused by bacteria belonging to the genus Shigella, and is the most common cause of bacterial diarrhea. Shigella species and Escherichia coli have a high degree of homology at the genomic level, with the main difference lying in the presence (or absence) of a 230-kb virulence plasmid (VP). It is generally believed that Shigella evolved from E. coli after acquiring the VP. Bacterial evolution is a dynamic process, which can be advanced by gene loss and horizontal gene transfer among distant bacterial strains. To date, whole-genome sequencing, including both VPs and chromosomes, has been completed for many strains of all four Shigella species1,2. This large amount of genetic information allows us to identify the continuous genetic events that led to the evolution of pathogenic Shigella from non-pathogenic E. coli. In addition, this information provides insight into changes in virulence traits among different Shigella strains.
Acquisition of the VP is a key step in the evolution of Shigella from a non-pathogenic ancestor to a pathogenic genus3. Studies have shown that the VPs carried by Shigella species are closely related to their pathogenicity. Deletion of the VP results in loss of invasion capability of the pathogen. However, E. coli K-12 transformed with the VP does not have the same pathogenicity profile as Shigella species. In addition, transformation of the E. coli K-12 chromosome into S. flexneri can lower the virulence of the pathogen4,5. As well as acquisition of the VP, the evolution of non-pathogenic E. coli into pathogenic Shigella also involved the acquisition of chromosomal virulence genes and the loss of anti-virulence genes (“black hole” genes). There may also be cross-talk between the VP and the chromosome of S. flexneri. For example, some functional proteins encoded by the host cell chromosome might be regulated by the VP. However, such regulatory hierarchy is relatively unstudied, with only one previous report in which a single VP-cured S. flexneri strain was analyzed6. Regardless, this initial information suggests that the co-evolution and co-ordination of chromosomes and VPs are very important steps in the evolutionary process7,8, and should be thoroughly analyzed for a more reliable and extensive understanding of the evolution of this pathogen.
Therefore, to study the influence of VPs on bacterial chromosomes at a proteomic level, we constructed VP-cured Shigella strains and VP-imported E. coli strains. The expression profiles of protein complexes in the wild-type, deletion mutant, and transformed mutant strains were determined by blue native/sodium dodecyl sulfate-polyacrylamide gel electrophoresis (BN/SDS-PAGE), and the protein monomers were analyzed by isoelectric focusing (IEF)/SDS-PAGE two-dimensional (2-D) electrophoresis. Importantly, three different S. flexneri strains were included in this study to gain a broader and more reliable understanding of the interaction between the VPs and the bacterial chromosomes.
Results
The VP regulates the expression of GadA/B
To examine the interactions between the VP and chromosomal genes, we deleted the VPs from several S. flexneri strains and introduced them into E. coli. The interaction between the VP and the chromosome is likely to be mediated by many different proteins, which form complexes with other proteins according to their structure, dynamics, and complex physical and chemical principles to carry out their biological functions9,10. We therefore performed BN/SDS-PAGE and IEF/SDS-PAGE 2-D electrophoresis to investigate the expression profiles of protein complexes and protein monomers in the wild-type, deletion mutant, and transformed mutant strains. Figure 1A shows a representative 2-D electropherogram obtained for analysis of wild-type strain 301, its deletion mutants, and transformed mutant strains. Corresponding results for the remaining strains are shown in Supplementary Fig. S2. MALDI-TOF mass spectrometry identified the most significant differentially expressed proteins as GadA/B, which play an important role in the acid resistance of Shigella species. As shown in Fig. 1, the expression patterns of GadA/B protein complexes and the GadA and GadB protein monomers had similar shifting tendencies. Compared with the wild-type strain, GadA/B expression levels were significantly reduced in E. coli strains harboring any of the three Shigella VPs (pCP, pSF, and pWR). In contrast, no changes in expression were observed for the plasmid-cured Shigella mutants when compared with their corresponding wild-type strains (Fig. 1B,C). Interestingly, abundance of GadA/B in wild-type S. flexneri strain 2457 T, deletion mutant ΔpSF, and transformed mutant MG1655/pSF was extremely low (Fig. 1A,B).
(A) Representative two-dimensional electrophoresis map. (B) Enlarged images of GadA/B protein complex. (C) Enlarged images of the GadA/GadB protein subunits. (D) Transcript levels of gadA/B were determined by quantitative reverse-transcriptase polymerase gel electrophoresis analysis normalized to the levels of the 16S rRNA gene in each sample.
To verify the proteomic data, the effects of the VP on gadA and gadB mRNA levels were quantified using quantitative real-time polymerase chain reaction (qRT-PCR) analysis. Consistent with the protein expression data, the presence of the VP was associated with reduced transcriptional levels of gadA and gadB mRNA in transformed mutant strains, but did not significantly alter transcription in VP-deficient Shigella strains. In addition, the expression of gadA and gadB was relatively low in 2457 T strains (Fig. 1D).
The VP regulates acid tolerance of the host bacterium
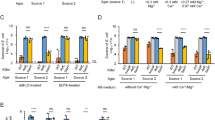
As the VP can regulate the expression of glutamic acid decarboxylase, which is the most effective acid-tolerance pathway, we examined whether the VP was associated with bacterial acid resistance. Cell survival rates of various strains were determined following growth at pH 2.5, and then compared with the survival rates at pH 5.0 using flow cytometry (injured cells were counted as surviving cells). Figure 2A is representative of results obtained for S. flexneri wild-type strain 301, its deletion mutants, and transformed mutant strains, while the results for the remaining strains are shown in Supplementary Fig. S3. As expected, the acid tolerance of the various strains was positively correlated with the GadA/B expression level. Strains with a higher abundance of GadA/B showed strong acid resistance, while the acid resistance of strains with lower abundance of GadA/B tended to be weaker (Fig. 2B).
(A) Flow cytometry was used to provide counts of living cells before and after acid treatment, calculate the viability of bacterial cells, and then infer the strength of the acid tolerance. R1 represented the whole cells; R2–R4 were set around the dead, injured, and live bacterial populations, respectively. (B) Survival rates of bacterial cells were calculated following culture in acid medium. Injured cells also counted as surviving cells.
The VP regulates the expression of the AtpA/D complex but not that of protein monomers
The AtpA/D protein complex is also associated with bacterial intracellular pH regulation. The γ, δ, and ε subunits of ATP synthase are difficult to detect by 2D electrophoresis as they have low molecular weights. Thus, the relative abundance of the α (AtpA) and β (AtpD) subunits is used to represent the expression of the ATP synthase complex. As shown in Fig. 3A and B, deletion of the VP greatly increased the abundance of the AtpA/D complex, while the abundance of the protein monomers remained virtually unchanged. This trend was consistent in all three of the strains analyzed. We also examined the expression of AtpD using western blot analysis, and the results mainly coincided with those generated by IEF/SDS-PAGE analysis (Fig. 3C). The expression of the AtpD protein monomer appeared to be identical in wild-type and deletion mutant strains. Therefore, the VP altered the expression of the ATP synthase complex, but had no effect on the monomers.
(A) and (B) Analysis of the AtpA/D protein complex (A) and AtpA (AtpD) protein subunit (B) from wild-type and deletion mutant Shigella strains using blue native-polyacrylamide gel electrophoresis or isoelectric focusing/sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). (C) Detection of AtpD in protein samples from the wild-type and deletion mutant Shigella strains using western blot analysis. The samples were separated by 12.5% SDS-PAGE, and immunoblotted with anti-AtpD.
An ATP Assay System Bioluminescence Detection Kit was then used to analyze the intracellular ATP levels of wild-type and VP-cured Shigella strains. According to the luminescence detection results (Fig. 4), the ATP concentrations were greatly increased in the three deletion mutant strains. Combined with the previous results of BN/SDS-PAGE 2D electrophoresis (Fig. 3A,B), we can infer that the VP reduced intracellular ATP synthesis by decreasing the expression of the ATP synthase complex.
The VP suppressed the assembly of ATP synthase
As the levels of the ATP synthase protein complex did not correlate with the abundance of each of the protein monomers, we speculated that the VP could block the assembly of ATP synthase. To test this hypothesis, the 385-kDa ATP synthase complex and the ~50-kDa AtpD monomer were isolated simultaneously using a non-denaturing 6–11% acrylamide gradient gel. AtpD antibody analysis detected two bands (Fig. 5). The upper bands on the western blot should be protein complexes, while the lower bands may be free AtpD monomers. The abundance of the two sets of bands showed a counter-balance relationship with each other. Based on our hypothesis, removal of the VP should allow more AtpD monomers to be assembled into ATP synthase complexes because the inhibitory effect on ATP synthase assembly would be abolished. Accordingly, in deletion mutant strains, the ATP synthase complex was highly expressed, and the abundance of the AtpD monomer was lower than in wild-type strains. As a membrane protein, ATP synthase is distributed on the cellular membrane of prokaryotes11. Notably, the type III secretion system (T3SS), which plays a key role during the invasion of pathogens, is also located on the bacterial cellular membrane12. Thus, there may be spatial competition between ATP synthase and the T3SS.
Together, these findings further our understanding of the complex evolution from non-pathogenic E. coli to pathogenic Shigella, and identified a novel role for the VP in regulating acid resistance-related genes present on the chromosome. We also confirmed that glutamate decarboxylase is the primary acid-resistance mechanism in Shigella, and that its expression is closely correlated with the acid tolerance of these pathogens.
Discussion
Shigella species are highly pathogenic, with reports that as few as 10 organisms can cause infection. With a long incubation period, the diagnosis and treatment of Shigella infection is even more difficult. All enteric pathogens must pass through the extremely acidic environment of the stomach before they reach the neutral environment of the intestinal lumen. Shigella strains are often more resistant to gastric acid than other intestinal bacteria such as Salmonella and E. coli, which may allow the survival of a small number of cells, and thus provide the opportunity for Shigella to invade the intestinal mucosa13. E. coli have acquired five acid resistance pathways to counteract the extreme acidity of the stomach, but there are only two main acid-tolerance pathways in Shigella species: the AR1 (decarboxylase independent pathway) and AR2 (glutamate decarboxylase system) pathways14,15. The glutamate decarboxylase system is particularly important for survival of extremely acidic environments16. Our analyses showed that GadA/B expression was significantly reduced in E. coli harboring the VP, but was not significantly altered in VP-cured Shigella strains.
The glutamate decarboxylase system encompasses three genes: gadA, gadB, and gadC. GadA and GadB have highly homologous sequences; thus, with similar structures, they can form a 330-kDa hexamer14. X-ray crystallography has been used to analyze the structures of GadA and GadB17,18, and each monomer comprises three domains: an N-terminal domain, a large pyridoxal-5′-phosphate-binding domain, and a C-terminal domain. Among these, the N-terminal domain is the most important as it ensures that GadA/B is bound preferentially to the intracellular membranes in a low pH environment. The pyridoxal 5′-phosphate-dependent enzyme activity converts the α-decarboxylation product of L-glutamate to γ-aminobutyric acid and carbon dioxide19, consuming a cytoplasmic proton in the process20. This pathway prevents the internal pH from decreasing to lethal levels, as protons leaking into the cell during acid stress are consumed and excreted21. Because GadA/B was closely related to acid resistance, we used flow cytometry to determine the survival rates of the strains under acidic stress. We confirmed that the acid tolerance of the various strains was positively correlated with GadA/B expression level.
In addition, analysis of protein expression profiles showed that deletion of the VP greatly increased levels of the AtpA/D complex, while the expression of the protein monomers remained unchanged. ATP synthesis is one of the main chemical reactions in most organisms. However, a hydrogen ion (H+) can be released during the synthesis of ATP from inorganic phosphate and ADP by the ATP synthase, which makes the intracellular environment even more acidic and unfavorable for bacterial survival. ATP synthase is distributed on the cell membrane in prokaryotes, and plays a key role in cellular energy exchange. ATP synthase consists of two parts. The F1 hydrophilic head consists of five different subunits (α, β, γ, δ, and ε), with a stoichiometry of α3β3γδε, which catalyze the synthesis of ATP from ADP. In contrast, the F0 hydrophobic tail consists of three subunits (a, b, and c), in a stoichiometry of ab2c12, forming an ion channel that allows the flow of protons22. Because ATP synthase is composed of a variety of subunits and the assembly process is relatively complex, many factors can affect its expression. BN-PAGE and western blotting analysis suggested that the VP suppressed the assembly of AtpA/D. Because the VP contains a large number of T3SS-related genes, the synthesis of these proteins and assembly of the T3SS, which is an ultra-structure across two layers of membrane, may hinder the fluidity of the bacterial outer membrane. This would be harmful for the translocation of ATP synthase subunit proteins, thereby interfering with the formation of ATP synthase complexes.
Combining all of the above results, we speculate that during the initial stages of Shigella evolution, the expression of glutamate decarboxylase rapidly decreased after E. coli acquired the VP, which decreased the acid resistance of the bacterium. However, as acid tolerance is required for intestinal bacteria to survive in the gut and colonize host cells, the decrease in acid resistance was unfavorable. In particular, it has been reported that lysine decarboxylase is absent in Shigella, suggesting that the glutamate decarboxylase system is the most important acid-resistance mechanism in these species14. Perhaps, during the long period of co-evolution of the VP and the chromosomal genes, GadA/B expression gradually increased again via some unknown mechanism, restoring acid resistance (Fig. 6). Thus, even if the VP was deleted (as in the current study), GadA/B expression levels would not be dramatically altered as a large part of this expression is no longer controlled by the VP. More simply, the acid resistance pathways of E. coli and S. flexneri developed via different evolutionary processes. In this hypothesis, S. flexneri strain 2457 T, which showed lower levels of GadA/B expression, may represent a kind of intermediate strain in the evolution of Shigella. Despite this decreased GadA/B expression, S. flexneri strain 2457 T is a clinical isolate that was responsible for a dysentery epidemic. The survival advantage of 2457 T might come from the R27 plasmid, which contains multiple antibiotic resistance genes that would aid in the survival of this pathogen in a clinical setting.
In the case of ATP synthase, the release of H+ during the synthesis of ATP from inorganic phosphate and ADP, resulting in an increase in the acidity of the intracellular environment (Fig. 6), may be a compensatory mechanism that has evolved during the long evolutionary period as an attempt to maintain the acquired virulence after the partial loss of acid tolerance23. Moreover, after a bacterium enters the host, its metabolism will be slowed (except for the synthesis of invasion- and virulence-related proteins) in an attempt to resist the extremely acidic environment of the stomach and evade the host immune response6. Its ATP requirements also decrease accordingly, meaning that high levels of ATP synthase are not needed. Taken together, these findings suggest that the VP decreases the production of ATP synthase, which could aid in the invasion of host cells by the pathogen.
Methods
Bacterial strains and growth conditions
E. coli strain DH5α was used for plasmid construction, and was maintained on Luria-Bertani (LB) agar or broth (Difco) at 37 °C. Wild-type S. flexneri serotype 2a strains 301 and 2457 T, and serotype 5a strain M90T, were grown on tryptic soy agar (Difco) containing 0.01% (w/v) Congo red or in LB broth at 30 °C and 37 °C. When necessary, 50 μg ml−1 nalidixic acid, 50 μg ml−1 streptomycin, or 30 μg ml−1 chloramphenicol were added to the growth media.
Construction of transconjugant and mutant S. flexneri strains
As shown in Supplemental Fig. S1, the VP gene fragment virG, amplified from S. flexneri strain 301 using primers virGp1/virGp2 (listed in Supplemental Table S1), was ligated into chloramphenicol resistance pir-dependent suicide vector pXL275, generating recombinant plasmid pXL275-virG. pXL275-virG was transformed into E. coli S17-λpir, and then conjugated into S. flexneri strain 301 according to the method of Klümper et al.24. Following homologous recombination into the VP at the virG site, the resulting VP contained the chloramphenicol resistance marker. Using helper plasmid pRK2013, the recombinant plasmid was conjugated into E. coli strain MG1655, as described previously25. Resulting transconjugants were named MG1655/pCP, and were isolated as pure cultures for use in further analyses. Two further transconjugant strains, named MG1655/pSF and MG1655/pWR and harboring the VPs from S. flexneri strains 2457 T and M90T, respectively, were prepared using the same method. VP-cured strains ΔpCP, ΔpSF, and ΔpWR were constructed using plasmid incompatibility methods6.
Two-dimensional polyacrylamide gel electrophoresis and data analysis
Preparation of whole-cell protein extracts (complex and monomer samples) and 2-D polyacrylamide gel electrophoresis analysis (BN-PAGE and IEF/SDS-PAGE) were performed as previously described26.
Protein identification by matrix-assisted laser desorption/ionization-time of flight tandem mass spectrometry (MALDI-TOF/TOF)
All of the protein spots were analyzed by MALDI-TOF/TOF mass spectrometry. The protein spots were carefully excised from the gel, destained using destaining solution (50% methyl cyanide, 25 mM acid ammonium carbonate), and then digested for 13 h with sequencing grade modified trypsin (Roche, USA). Peptides from digested proteins were used for MALDI-TOF/TOF analysis. The MALDI-TOF mass spectrometry measurement was performed on a Bruker UltraflexIII MALDI-TOF-MS instrument (Bruker Daltonics, Germany), operating in reflectron mode with 20 kV accelerating voltage and 23 kV reflecting voltage. A saturated solution of α-cyano-4-hydroxycinnamic acid in 50% acetonitrile and 0.1% trifluoroacetic acid was used as the matrix. A 1-μl volume of the matrix solution and the sample solution at a ratio of 1:1 was applied to the Score384 target well. The SNAP algorithm (S/N threshold: 5; quality factor threshold: 30) in FlexAnalysis 2.4 was used to select the 150 most prominent peaks in the mass range m/z 700–4000. The subsequent tandem mass spectrometry (MS/MS) analysis was performed in a data-dependent manner, and the 10 most abundant ions were subjected to high energy collision-induced dissociation analysis. The collision energy was set to 1 keV, and nitrogen was used as the collision gas.
Data interpretation and database searching
To deal with one peptide mass fingerprinting and multiple TOF/TOF spectra from one sample as a single combined dataset, the raw data were first merged into one Mascot generic format file using Biotools v3.0 software, and then searched using Mascot 2.1 (Matrix Science Ltd.) against the S. flexneri 2a 301 database. The search included all predicted open reading frames on both the chromosome (GenBank GI: 344915202) and virulence plasmid pCP (GenBank GI: 18462515) of S. flexneri 2a 301 to eliminate redundancy resulting from multiple members of the same protein family, and the results were checked against the NCBInr database (version 20061021, 4,072,503 sequences) to eliminate known contaminants. The search parameters were: trypsin digestion with one missed cleavage; carbamidomethyl modification of cysteine as a fixed modification and oxidation of methionine as a variable modification; peptide tolerance maximum, ±100 ppm; MS/MS tolerance maximum, ±0.6 Da; peptide charge, +1; monoisotopic mass. Scores greater than 21 were considered significant (P < 0.05) for a local MS/MS search. For unambiguous identification of proteins, more than five peptides must be matched.
RNA extraction and qRT-PCR analysis
Total RNA extraction and qRT-PCR analysis were performed as described previously26. Gene-specific primers (Supplemental Table S1) were designed using Primer Premier 5.0 software (Premier Biosoft). Relative amounts of cDNA were normalized to the amounts of 16 S rRNA cDNA in each sample.
Acid tolerance assay
Bacteria were grown to stationary phase at 37 °C in LB medium adjusted to pH 5.0. A 1-ml aliquot of culture was then centrifuged for 5 min at 2000 × g, and the resulting pellet resuspended in 1 ml of LB medium adjusted to either pH 2.5 or pH 5.027,28. The suspension was then incubated for 30 min at 37 °C with shaking at 220 rpm. Bacteria were collected by centrifugation as described above, and washed three times with PBS. Cells were then stained for 15 min using a BD cell viability kit, and measured using a BD FACS flow cytometer. The data were analyzed using CellQuest software.
Western blot analysis
The BN- and SDS-PAGE gels were transferred to polyvinylidene difluoride membranes at 15 V for 1.5 h. The membranes were blocked with 10% (w/v) skim milk powder in TBS (100 mM Tris-HCl, pH 7.5, 0.9% (w/v) NaCl) containing 0.1% (v/v) Tween 20 (TBST) for 1 h. Membranes were then incubated in anti-AtpD antibody (Abmart Corp.) diluted in TBST for 1–2 h at room temperature, or at 4 °C overnight, at the recommended concentration, followed by detection using ECL reagents (Thermo) and manual film development.
ATP assay system
The intracellular ATP levels of the wild-type and mutant Shigella strains were measured using an ATP bioluminescence assay kit (Promega, USA) as per the manufacturer’s protocol. Strains were grown in LB broth to stationary phase, diluted in dilution buffer, and lysed using the provided cell lysis reagent. After addition of the luciferase reagent, the light signal was detected. All measurements were within the linear range as determined for an ATP standard curve.
Additional Information
How to cite this article: Niu, C. et al. Role of the virulence plasmid in acid resistance of Shigella flexneri. Sci. Rep. 7, 46465; doi: 10.1038/srep46465 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Jin, Q. et al. Genome sequence of Shigella flexneri 2a: insights into pathogenicity through comparison with genomes of Escherichia coli K12 and O157. Nucleic acids research 30, 4432–4441 (2002).
Yang, F. et al. Genome dynamics and diversity of Shigella species, the etiologic agents of bacillary dysentery. Nucleic acids research 33, 6445–6458 (2005).
Ud-Din, A. & Wahid, S. Relationship among Shigella spp. and enteroinvasive Escherichia coli (EIEC) and their differentiation. Brazilian journal of microbiology:[publication of the Brazilian Society for Microbiology] 45, 1131–1138 (2014).
Formal, S. B., Gemski, P., Baron, L. S. & Labrec, E. H. A Chromosomal Locus Which Controls the Ability of Shigella flexneri to Evoke Keratoconjunctivitis. Infection and immunity 3, 73–79 (1971).
Formal, S. B., Labrec, E. H., Kent, T. H. & Falkow, S. Abortive Intestinal Infection with an Escherichia Coli-Shigella Flexneri Hybrid Strain. Journal of bacteriology 89, 1374–1382 (1965).
Zhu, L. et al. Global analysis of a plasmid-cured Shigella flexneri strain: new insights into the interaction between the chromosome and a virulence plasmid. Journal of proteome research 9, 843–854 (2010).
Nakata, N. et al. The absence of a surface protease, OmpT, determines the intercellular spreading ability of Shigella: the relationship between the ompT and kcpA loci. Molecular microbiology 9, 459–468 (1993).
Sokurenko, E. V., Hasty, D. L. & Dykhuizen, D. E. Pathoadaptive mutations: gene loss and variation in bacterial pathogens. Trends in microbiology 7, 191–195 (1999).
Wodak, S. J., Pu, S., Vlasblom, J. & Seraphin, B. Challenges and rewards of interaction proteomics. Molecular & cellular proteomics: MCP 8, 3–18 (2009).
von Mering, C. et al. Comparative assessment of large-scale data sets of protein-protein interactions. Nature 417, 399–403 (2002).
Nakanishi-Matsui, M., Sekiya, M. & Futai, M. ATP synthase from Escherichia coli: Mechanism of rotational catalysis, and inhibition with the epsilon subunit and phytopolyphenols. Biochimica et biophysica acta 1857, 129–140 (2016).
Hu, B. et al. Visualization of the type III secretion sorting platform of Shigella flexneri . Proceedings of the National Academy of Sciences of the United States of America 112, 1047–1052 (2015).
Hale, T. L. Genetic basis of virulence in Shigella species. Microbiological reviews 55, 206–224 (1991).
Zhao, B. & Houry, W. A. Acid stress response in enteropathogenic gammaproteobacteria: an aptitude for survival. Biochemistry and cell biology = Biochimie et biologie cellulaire 88, 301–314 (2010).
Bhagwat, A. A. & Bhagwat, M. Comparative analysis of transcriptional regulatory elements of glutamate-dependent acid-resistance systems of Shigella flexneri and Escherichia coli O157:H7. FEMS microbiology letters 234, 139–147 (2004).
Sherburne, C. K. et al. The complete DNA sequence and analysis of R27, a large IncHI plasmid from Salmonella typhi that is temperature sensitive for transfer. Nucleic acids research 28, 2177–2186 (2000).
Capitani, G. et al. Crystal structure and functional analysis of Escherichia coli glutamate decarboxylase. The EMBO journal 22, 4027–4037 (2003).
Dutyshev, D. I. et al. Structure of Escherichia coli glutamate decarboxylase (GADalpha) in complex with glutarate at 2.05 angstroms resolution. Acta crystallographica. Section D, Biological crystallography 61, 230–235 (2005).
Foster, J. W. Escherichia coli acid resistance: tales of an amateur acidophile. Nature reviews. Microbiology 2, 898–907 (2004).
Bearson, S., Bearson, B. & Foster, J. W. Acid stress responses in enterobacteria. FEMS microbiology letters 147, 173–180 (1997).
Wei, J. et al. Complete genome sequence and comparative genomics of Shigella flexneri serotype 2a strain 2457T. Infection and immunity 71, 2775–2786 (2003).
Peng, Y. et al. A blue native-PAGE analysis of membrane protein complexes in Clostridium thermocellum. BMC microbiology 11, 22 (2011).
Jennison, A. V. & Verma, N. K. The acid-resistance pathways of Shigella flexneri 2457T. Microbiology 153, 2593–2602 (2007).
Klumper, U., Droumpali, A., Dechesne, A. & Smets, B. F. Novel assay to measure the plasmid mobilizing potential of mixed microbial communities. Frontiers in microbiology 5, 730 (2014).
Zhang, P. Y. et al. Combined treatment with the antibiotics kanamycin and streptomycin promotes the conjugation of Escherichia coli . FEMS microbiology letters 348, 149–156 (2013).
Niu, C. et al. Analysis of Soluble protein complexes in Shigella flexneri reveals the influence of temperature on the amount of lipopolysaccharide. Molecular & cellular proteomics: MCP 12, 1250–1258 (2013).
Lin, J., Lee, I. S., Frey, J., Slonczewski, J. L. & Foster, J. W. Comparative analysis of extreme acid survival in Salmonella typhimurium, Shigella flexneri, and Escherichia coli . Journal of bacteriology 177, 4097–4104 (1995).
Yang, G. et al. hfq regulates acid tolerance and virulence by responding to acid stress in Shigella flexneri . Research in microbiology 166, 476–485 (2015).
Acknowledgements
This work was supported by the National Key Basic Research Program of China (973 Program, No. 2013CB910804 and 2011CB504901), and the National Natural Science Foundation of China (No. 81501719, 81125012, and 81171531).
Author information
Authors and Affiliations
Contributions
H.-L.W. and L.Z. conceived, organized, and interpreted experiments. C.N. generated the bulk of the results and wrote the manuscript. J.Y., H.-S.L., Y.C., H.-J.X., and R.-F.W. performed the experiments. D.-S.W. and C.P. assisted in performing and interpreting experiments. X.-K.L. and E.-L.F. analyzed the results and datasets. W.X. and X.-Q.L. provided advice and expertise.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Niu, C., Yang, J., Liu, H. et al. Role of the virulence plasmid in acid resistance of Shigella flexneri. Sci Rep 7, 46465 (2017). https://doi.org/10.1038/srep46465
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep46465
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.