Abstract
Although host-plant selection is a central topic in ecology, its general underpinnings are poorly understood. Here, we performed a case study focusing on the publicly available data on Japanese butterflies. A combined statistical analysis of plant–herbivore relationships and taxonomy revealed that some butterfly subfamilies in different families feed on the same plant families, and the occurrence of this phenomenon more than just by chance, thus indicating the independent acquisition of adaptive phenotypes to the same hosts. We consequently integrated plant–herbivore and plant–compound relationship data and conducted a statistical analysis to identify compounds unique to host plants of specific butterfly families. Some of the identified plant compounds are known to attract certain butterfly groups while repelling others. The additional incorporation of insect–compound relationship data revealed potential metabolic processes that are related to host plant selection. Our results demonstrate that data integration enables the computational detection of compounds putatively involved in particular interspecies interactions and that further data enrichment and integration of genomic and transcriptomic data facilitates the unveiling of the molecular mechanisms involved in host plant selection.
Similar content being viewed by others
Introduction
Herbivorous insects and their host plants have been engaged in a chemical arms race for more than 420 million years1. Whilst host plants have developed various chemicals for defence against herbivorous insect damage, herbivorous insects have evolved various countermeasures2,3. Some insects even use plant chemicals for their own benefit4. Most chemical ecology studies have focused on a small number of specific species rather than systematically exploring a wide range of species.
Many herbivorous butterflies feed on specific host plants, thus forming plant–herbivore networks. In recent years, large-scale data on ecological networks, including plant–herbivore networks, have become available with the development of new observation techniques and with improvements in databases and other infrastructure. Network analysis techniques resulting from the development of network science5 have been used in ecology to actively investigate ecological networks, with respect to both basic scientific research (e.g., structure–stability relationships) and applied ecology (e.g., biodiversity maintenance and environmental assessment)6,7,8.
Taxonomic families of herbivorous butterflies generally correspond to those of host plants, thus reflecting the close co-evolutionary relationship between butterflies and plants. A correlation has been observed between changes in host plants and species diversification in the butterfly superfamily Papilionidea9. Numerous observations of host–plant selection have established that this selection is controlled by chemical constraints, i.e., several insect species recognise plants by detecting their chemical components10,11,12,13,14,15,16. Nevertheless, the molecular mechanisms that determine host–plant shifts are poorly understood16. A general understanding of the molecular factors involved in host–plant selection requires detailed chemical and genomic studies on a wide range of insects and plants.
To address this problem, we integrated biological data to gain a comprehensive picture of the contribution of plant chemical compounds to host plant selection and to identify candidate plant compounds involved in specific interactions between butterfly and plant families. This required us to first comprehensively collect data on plant–herbivore relationships involving Japanese butterflies, which were not previously available in the HOSTS lepidopteran host plant database [ http://www.nhm.ac.uk/our-science/data/hostplants/].
The Japanese Islands lie at the convergence of three terrestrial ecozones, namely, the Palearctic (including Europe, Asia north of the Himalayan foothills, northern Africa, and the northern and central Arabian Peninsula), the Oriental (extending across most of South and Southeast Asia and into southern East Asia), and Oceania. Although the flora and fauna of the Japanese Islands are influenced by all three ecozones, they retain their unique status in these islands, which extend from the subarctic to subtropical zones and include many highlands. These geographic and ecological features, as well as an abundance of accumulated knowledge from observations of area-specific populations of butterflies, are attractive targets for research on species diversification. We have generated a complete plant–herbivore network for the Japanese Islands. This network, designated the InsectInDB database (http://insect-plant.org/), comprises 545 insects, 1,922 plants and 3,435 insect–plant relationships.
In this study, we used network analysis combined with taxonomic information to clearly demonstrate that butterfly subfamilies in different families feed on the same plant groups, thereby indicating the independent acquisition of the same adaptive phenotypes (i.e., to the same hosts). The incorporation of chemical compound data allowed us to consider the contributions of plant compounds to butterfly host–plant selection by comparing two hypotheses: a “feed-on-family” hypothesis in which butterflies preferentially feed on host plants in the same family, and a “feed-on-compound” hypothesis in which butterflies select plants that produce similar compounds. Statistical analysis enabled the identification of compounds specific to host plants of particular butterfly families. We determined that some of these compounds are known to attract limited groups of butterflies and repel others. An integration of the insect–compound relationship data revealed potential metabolic processes related to host-plant selection. Our results successfully demonstrated the value of data integration for the computational detection of candidate host plant selection compounds. Finally, the further integration of genomic and transcriptomic data allowed us to unveil the putative molecular mechanism of host plant selection.
Results
Study strategy overview
Before describing the approach used in this study, we first will define the terms used in this paper to describe different plant–insect relationships. Several sets of words, including generalist/specialist and monophagous/oligophagous/polyphagous, have been used to represent the range of insect host preferences17. When discussing plant–herbivore relationships, a specialist is an herbivore that has evolved a specific mechanism to feed on a particular host plant, whereas a generalist is one that has not. Monophagous, oligophagous and polyphagous refer to the number of host plants fed on, i.e., only one, a few, or many host plants, respectively. Alternatively, oligophagous is defined as the use of multiple host plant species from the same family18. Generalist/specialist can also refer to the breadth of plant selection by pollinator insects19. In many cases, however, these words are ambiguous. For example, “specialist” and “monophagous” typically indicate that an herbivore feeds on a single host plant species, but they are sometimes applied to an herbivore feeding on an entire family. To avoid confusion, we define the number of host plant species and host plant families as “mono-species-phagous/oligo-species-phagous/poly-species-phagous” and “mono-family-phagous/oligo-family-phagous/poly-family-phagous”, respectively (Fig. 1a). Similar to oligophagous/polyphagous, the distinction between “oligo-” (several) and “poly-” (many) is somewhat subjective. In this paper, we use these words to avoid ambiguity when discussing plant–herbivore relationships at the species and family levels.
(a) Proposed word definitions to describe the range of host-plant preferences. (b) Plant–herbivore matrix. Rows and columns represent plants and herbivorous butterflies, respectively. A dot at an intersection indicates that the butterfly feeds on the corresponding plant. Butterfly taxonomy is indicated by the dendrogram. (c) Plant–compound matrix. Rows and columns represent plants and compounds, respectively. Dots are used to indicate that a plant possesses a compound. Plant taxonomy is indicated by the dendrogram. (d) Plant–herbivore network consisting of butterfly species with their host plant species as the nodes. (e) Plant–herbivore family network consisting of butterfly families with their host plant families as the nodes. (f) Compound effect on host plant selection. Arrows and dotted lines indicate that a plant compound attracts or repels herbivorous butterflies, respectively. (g) Plant–herbivore metabolism. Some compounds (such as C1) found in herbivorous butterflies are believed to be directly obtained from the host plant, whereas other compounds (such as C2′) are derived from plant compounds via enzymatic reactions or from other sources (such as C6). (h) Feed-on-family hypothesis.
To achieve a general understanding of host plant selection by Japanese butterflies, we considered plant–herbivore relationships (Fig. 1b) and the presence of compounds in various plant species (Fig. 1c) as follows. First, plant–herbivore relationships between butterflies and their host plants were represented as a bipartite network (Fig. 1d) in which a node represents a butterfly or plant species and an edge indicates that the butterfly feeds on the host plant. Second, the network was divided into sub-networks according to taxonomic family (Fig. 1e). The combined use of plant–herbivore relationship (Fig. 1b) and plant–compound relationship (Fig. 1c) data allowed us to identify compounds common to a group of host plants that are specific to a group of butterflies (in order to distinguish from just “common” compounds, we refer to this type of compounds as “common specific” compounds in this paper). For example, suppose that butterflies B4 and B5 feed specifically on plants P3 and P4 (Fig. 1b). Among the compounds shown in Fig. 1c, C1 is “common” to all plants, which indicates that C1 may be important for plant survival in general but may not be involved in interactions with specific butterflies. Compound C2 is common specific to plants P3 and P4 but not P1 and P2, indicating the possibility that C2 functions to attract butterflies B4 and B5 and repel B1, B2 and B3 (Fig. 1f). When we examined the contributions of these common specific compounds to host plant selection, we found that some were, in fact, important in the plant–herbivore relationship. Finally, a comparison of compounds identified as participants in plant–herbivore relationships unveiled insect metabolic processes that may be involved in the interaction (Fig. 1g).
General architecture of the Japanese butterfly plant–herbivore network
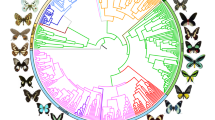
Several seminal studies have revealed that taxonomically related host plants are used by particular butterfly families20,21. In the present study, we adopted the most recently reported phylogenies of angiosperm plants (APGIII)22 and butterflies23 to examine the relationships between butterfly–host plant associations in the Japanese Islands. To analyse the network architecture as a whole, we conducted a comprehensive literature survey in which 3,435 plant–herbivore relationships encompassing 1,922 plant species and all 545 butterfly species living in Japan were examined. All members of the superfamily Rhopalocera except for Riodinidae are found in the Japanese Islands; these butterflies are classified into the families Hesperiidae, Papilionidae, Pieridae, Lycaenidae and Nymphalidae. Figure 2a shows the distribution of butterfly families in the Japanese Islands and global ecozones. We conducted a similarity analysis based on Pearson’s correlation coefficients (r) of the relative proportions of butterfly families in different ecozones, which indicated that the butterfly composition of the Japanese Islands is similar to those of the Holarctic (r = 0.92) and Oriental (r = 0.91) ecozones. This result may imply that butterfly distributions in the Japanese Islands are more influenced by distributions in the Holarctic and Oriental ecozones than by those in Oceania or other ecozones, however, it still lacks solid evidence and analysis. The number of butterfly species in Japan (302) is much smaller than the numbers in other ecozones (2,224, 2,411, 1,274, 3,964 and 7,784 in Holarctic, Oriental, Oceania, Afrotropical and Neotropical ecozones, respectively). Another factor that may need to be considered is that Papilionidae butterflies are especially popular in Japan and have been extensively catalogued.
(a) Distribution of butterfly families across the Japanese Islands and other global ecozones. (b) Distribution of host plants among butterfly families. Horizontal and vertical axes represent the number of host plant families per butterfly species and the number of butterfly species, respectively. (c) Plant–herbivore matrix of some Japanese butterfly species. Mono-family-phagous butterflies are Daimio tethys, Erynnis montanus, Pelopidas mathias, Polytremis pellucida, Thoressa varia, Isoteinon lamprospilus, Potanthus flavum, Parnara guttata, Parnassius citrinarius, Luehdorfia japonica, Byasa alcinous, Papilio macilentus, Papilio bianor, Papilio protenor, Papilio memnon Linnaeus, Papilio xuthus, Papilio maackii, Papilio helenus, Anthocharis scolymus, Pieris melete, Colias erate, Gonepteryx aspasia, Eurema laeta Boisduval, Curetis acuta paracuta, Everes argiades, Lycaena phlaeas, Japonica lutea, Iratsume orsedice orsedice, Artopoetes pryeri, Antigius attilia, Narathura japonica, Libythea celtis, Dichorragia nesimachus, Argyronome laodice, Argyronome ruslana, Fabriciana adippe, Nephargynnis anadyomene, Argyreus hyperbius, Argynnis paphia, Damora sagana, Vanessa cardui, Araschnia burejana, Limenitis glorifica, Limenitis camilla, Neptis pryeri, Hestina persimilis japonica, Sasakia charonda, Apatura metis, Ypthima argus, Mycalesis gotama, Minois dryas, Neope goschkevitschii, Lethe sicelis, Zophoessa callipteris, Lethe diana, Parantica sita. Oligo-family-phagous butterflies are Choaspes benjaminii, Graphium sarpedon, Pieris rapae, Eurema hecabe, Lampides boeticus, Pseudozizeeria maha, Neozephyrus japonicus, Vanessa indica, Polygonia c-aureum, Kaniska canace, Mycalesis francisca, Melanitis phedima. Poly-family-phagous butterflies are Papilio machaon, Celastrina argiolus, Callophrys ferrea, Rapala arata, Nymphalis xanthomelas, Neptis sappho. Taraka hamada is not herbivore but carnivore.
Figure 2b, which shows the relationship between the number of host-plant families and butterfly species, clearly demonstrates that host plant preferences differ among butterfly families. For example, Hesperiidae and Lycaenidae both feed on a wide range of host plant species; however, the butterflies in the first family feed on members of one or two specific host plant families, whereas butterflies in the latter family feed on a wider variety of host plant families. To distinguish between these behaviours, we therefore prefer to use mono-species-phagous/oligo-species-phagous/poly-species-phagous and mono-family-phagous/oligo-family-phagous/poly-family-phagous, as proposed above, rather than the existing terms, specialist/generalist and monophagous/oligophagous/polyphagous. The matrix representation in Fig. 2c clearly reveals some characteristics of the host plant selectivity of Japanese butterflies. For example, the genus Papilio in Papilionidae and the subfamilies Pierinae in Pieridae and Heliconiinae in Nymphalidae are generally mono-family-phagous and have specific relationships with host plants in Rutaceae, Brassicaceae and Violaceae, respectively. In contrast, some butterfly subfamilies in different families exhibit shared host plant selectivity: for example, the subfamily Coliadinae in Pieridae and the subfamily Polyommatinae in Lycaenidae both feed on plants in Fabaceae, whereas the subfamily Hesperiinae in Hesperiidae and the subfamily Satyrinae in Nymphalidae both interact with host plants in Poaceae. Poly-family-phagous butterflies can also be identified from the matrix. For example, butterflies in the subfamily Theclinae in Lycaenidae generally feed on more than three host-plant families.
An examination of the entire network of plant–herbivore relationships (Fig. 3) also reveals that butterfly species in the same family tend to share the same host-plant species and families. The Papilionidae, Nymphalidae and Pieridae families have their own predominant sub-networks (upper left, upper middle and upper right in Fig. 3, respectively). Rutaceae and Brassicaceae are examples of plant families predominantly fed on by single butterfly families (Papilionidae and Pieridae, respectively). In contrast, two or more butterfly families (Hesperiidae, Lycaenidae and Nymphalidae) share for the plants of the families Poaceae and Fabaceae.
Circles and rectangles represent butterfly and plant species, respectively, with the edges between them representing their relationship. Nodes (circles and rectangles) are coloured to indicate families of butterflies and plants, respectively. Node sizes are proportional to degree (i.e., number of edges per node).
To further analyse plant–herbivore relationships, we simplified the network to describe significant interactions between plant and butterfly families (Fig. 4a) with significance evaluated based on real vs. randomised network Z-scores (see Methods). Positive Z-scores indicated that plants in a particular family are significantly more likely to be eaten by insects from a certain family than by those in the randomised networks. As shown in Fig. 4a, the five butterfly families clearly form selective plant–herbivore relationships. For example, Papilionidae butterflies interact specifically with the families Rutaceae, Apiaceae, Aristolochiaceae and Lauraceae, thus forming an isolated network. The remaining four butterfly families also have their own specific host plant families, with two exceptions: Coliadinae subfamily in Pieridae and Polyommatinae subfamily in Lycaenidae butterflies share Fabaceae plants, and Hesperiinae subfamily in Hesperiidae and Satyrinae subfamily in Nymphalidae butterflies share plants in Poaceae. These butterfly subfamilies are in different families. Nonetheless, the families have independently acquired the ability to use the same groups of plants as hosts.
(a) Family-level plant–herbivore network. Circles and rectangles represent butterfly and plant families, respectively, with the edges between them representing their relationships. Node sizes are proportional to degree (i.e., number of edges per node). Edge widths represent the Z-scores. Node colours (circles and triangles) indicate that the butterfly and plant families are the same as in Fig. 3. (b) Plant–herbivore matrix of plant families and butterfly subfamilies. Closed and open circles indicate plant–herbivore relationships with positive and negative interaction frequency Z-scores, respectively. Areas of circles are proportional to Z-score absolute values.
Figure 4b shows significant plant–herbivore relationships between plant families and butterfly subfamilies. For example, the two subfamilies of Papilionidae (Parnassiinae and Papilioninae) have significantly different host plant families, and the same is true for the two Pieridae families (Pierinae and Coliadinae). Among the butterfly subfamilies in Lycaenidae, Curetinae and Miletinae have no specific host plant families, Polyommatinae and Theclinae are poly-family-phagous, and Lycaeninae is mono-family-phagous. The subfamily Satyrinae in Nymphalidae and the subfamily Hesperiinae in Hesperiidae might be regarded as generalist taxa because they feed on two or more host plant families. Because their host-plants are mostly restricted to a single order, we instead consider these latter butterflies to be mono-order-phagous.
With respect to butterfly families (Fig. 4a), the plant families Poaceae and Fabaceae are significantly shared by two butterfly families. At the butterfly subfamily level (Fig. 4b), plants in Fabaceae are consumed by the subfamily Coliadinae in Pieridae and by the subfamily Polyommatinae in Lycaenidae. Members of Poaceae serve as host plants for both the subfamily Satyrinae in Nymphalidae and the subfamily Hesperiinae in Hesperiidae. We found that these two plant families differ with respect to the types of butterflies that consume them. In particular, Poaceae is significantly eaten by subfamilies that are mono-order-phagous (Hesperiinae and Satyrinae), whilst Fabaceae is eaten at a significant level by poly-family-phagous subfamilies (Coliadinae, Polyommatinae, Cyrestinae and Limenitidinae).
A negative Z-score indicates that plants belonging to a particular family are significantly less subject to predation by a given insect family compared with those in randomised networks. Thus far, we have shown that Poaceae and Fabaceae are eaten by a wide range of butterflies (Fig. 4a). However, having analyzed the relationships with butterfly subfamilies, we also found that Poaceae and Fabaceae are consumed differently (Fig. 4b). Poaceae members have no significant relationships exhibiting negative Z-scores, indicating that there are no butterfly subfamilies that significantly avoid Poaceae plants. On the other hand, Fabaceae members have some relationships exhibiting negative Z-scores, i.e., significantly less numbers of host-plant relationships with Papilioninae, Theclinae, Nymphalinae and Satyrinae. To explain this difference, we hypothesised the presence/absence of some factors that work as repellent for some butterflies but as attractant for some other butterflies. In other words, Fabaceae plants are eaten by a wide range of butterflies because they possess some factors that exert both significantly positive (i.e., attractive) and negative (i.e., repellent) effects, whereas Poaceae plants are devoured by a wide range of butterflies because they have attraction factors but not repellent factors.
Influence of plants on host-plant selection
In the previous section, we demonstrated that host-plant selection is highly associated with butterfly taxonomic classifications and implied the existence of factors influencing host-plant selection. To provide more conclusive evidence for this idea, we explored two possible hypotheses: the “feed-on-family” hypothesis or the “feed-on-compound” hypothesis (Fig. 1h). To test the likelihood of the feed-on-family hypothesis, we calculated Z-score statistics based on family-based evenness for butterfly category (family or subfamily) i, Ereal(i), and the evenness of randomised networks, Erand(i) (see Methods). We found that the average Ereal(i) (0.03) was significantly lower than the average Erand(i) (0.16) (Z = –61.9; p < 2.2 × 10–16 using the Z-test).
We likewise examined the possibility of the feed-on-compound hypothesis. To determine whether a set of host plants of a butterfly species tended to possess a higher number of common specific compounds compared with those from randomly selected sets, we applied the food-pairing hypothesis24 to our datasets. For the statistical test, we used the average number of compounds shared among host plants by butterfly species k, Ns(k)24. From the original plant–herbivore network, we extracted all plant–herbivore relationships in which at least one compound was present in the host plant according to the species-metabolite relationship database KNApSAcK [25]. This extracted network, which consists of 216 butterfly and 405 plant species, is subsequently referred to as the real-world network. A comparison of real-world and randomised network Ns(k) values (Nsreal(k) and Nsrand(k), respectively) revealed that the average Nsreal (k) (0.37) was significantly larger than the average Nsrand (k) (0.09) (Z = 8.9; p < 2.2 × 10–16 using the Z-test).
We focused on the statistics for each butterfly family for further evaluation. In particular, we used the average family-based evenness for butterfly family i, E(i), and the average number of compounds shared among host plants by butterfly species k, Ns(k). Figure 5a shows the relationship between Z-scores of average plant family-based evenness for butterfly families, ZE(i) = (Ereal(i) – Erand(i))/SDE(i)rand, and the Z-scores of the average number of common specific compounds in host plants, ZNs(k) = (Nsreal(k) – Nsrand(k))/SDNs(k)rand. Ereal(i) [Nsreal(k)] and Erand(i) [Nsrand(k)] correspond to E(i) [Ns(k)] obtained from the real-world network and randomized networks, respectively. SDE(i)rand, and SDNs(k)rand are the standard deviations for Erand(i) and Nsrand(k), respectively (Data and Methods). Figure 5b is the same scatterplot for the butterfly subfamilies. For butterfly families and subfamilies, the Z-scores of evenness were mostly negative, which indicated that the actual host-plant relationship was more selective than random. The Z-scores of Ns were positive for the butterfly families Pieridae, Lycaenidae, and Papilionidae as well as for many subfamilies, which indicated that the host plants possessed more common specific compounds than would be expected by random chance. As clearly shown in Fig. 5a and b, the Z-scores of evenness and those of Ns were negatively correlated, indicating that the common specific compounds positively contributed to host-plant selection. For example, Pieridae and its subfamilies (Pierinae and Coliadinae) are significantly lower in evenness and higher in Ns compared with other families and subfamilies. Together with the previously known phenomenon that many phytophagous insects are highly adapted to defense chemicals produced by plants and use them as the specific host-finding cues14,15, the results obtained in this study provide quantitative support for our hypothesis that insects tend to selectively feed on closely-related plants sharing common specific chemical compounds.
(a) Relationships among Z-scores of average plant family-based evenness for butterfly families and Z-scores of the average number of common specific compounds in host plants. Circles represent butterfly families. Circle areas are proportional to degrees (i.e., numbers of host-plant species) of butterfly families. (b) The same plot for butterfly subfamilies. Circles represent butterfly subfamilies, with their colours corresponding to those given for butterfly families in (a). (c) Relationship between the compound contributions of plant species and degree (i.e., number of butterflies consuming the respective plant species). Circles represent plant species. Circle areas are proportional to the number of common specific compounds registered in the KNApSAcK database. Plant species in the Poaceae, Fabaceae, Brassicaceae and Rutaceae families are indicated by light green, dark green, yellow, and orange, respectively. (d) Relationship between the compound contributions (χi) of different plant families and average neighbour degree (i.e., average number of host-plant families used by herbivorous butterflies on the respective plant species). Circle areas are proportional to the average number of common specific compounds. Colours are the same as in (c).
We assumed that the global trend of host-plant selectivity was a mixture of different levels of feed-on-compound effects, i.e., some plants express a strong feed-on-compound effect, whereas others do not. Based on this idea, we determined the compound contribution of each plant species i, denoted as χi (see Methods). According to our calculations, most plant species had a positive value of χi, thus contributing positively to the feed-on-compound effect (Fig. 5c and d).
The relationship between χi and degree (i.e., the number of butterflies that predate on the respective plant species) is shown in Fig. 5c. Two Rutaceae species (Zanthoxylum ailanthoides and Phellodendron amurense) exerted particularly high feed-on-compound effects, whilst a species in Poaceae (Oryza sativa) had a negative effect. We found that χi was moderately positively correlated with degree (Spearman’s rank correlation coefficient rs = 0.53, p < 2.2 × 10–16). This correlation indicated that the presence of common specific plant compounds generally attracts phytophagous butterflies, an observation that supports the feed-on-compound hypothesis.
Figure 5d shows the relationship between plant family χi and average neighbour degree (i.e., the average number of host-plant families consumed by the butterfly herbivores of the respective plant species). Although the figure indicates that most plant families are eaten by mono-family-phagous butterflies (lower neighbour degree), we also observed that poly-family-phagous butterflies tend to feed on host plants with a lower χi. This result reveals the relationship between χi and host selectivity.
Common specific compounds in host plants
To identify specific compounds shared by host plants of a particular butterfly family, we used Fisher’s exact test, which, analogous to its application in genome-wide association studies, is the simplest method for inferring associations or correlations. We extracted all plant–herbivore relationships involving at least one plant compound registered in KNApSAcK25 from the original plant–herbivore network mentioned in the previous section, thereby obtaining 890 plant–herbivore relationships encompassing 217 butterfly species, 406 plant species and 3,507 plant compounds. Statistical significance based on Fisher’s exact test was evaluated using Cramer’s V; for example, given a sample size of 890 and a Cramer’s V value of ~0.6, the threshold P-value was set to 1.0 × 10−6.
Common specific compounds in host plants of Pierinae (family Pieridae), Lycaenidae, and Papilionae (family Papilionidae) butterflies are shown in Fig. 6a,b and c, respectively. Host plants of the Pierinae specifically possess a number of glucosinolates and flavonoids (Fig. 6a), whereas Papilionae (family Papilionidae) and Lycaenidae host plants are characterised by skimmianine and genistein, respectively (Fig. 6b,c). Among these identified compounds, glucosinolates are attractive to certain members of Pierinae but repel other butterflies10,11,26. Some plants, such as members of Brassicaceae, have a glucosinolate-myrosinase chemical defence system (Fig. 7a); to avoid this system, some Pieridae butterflies have a nitrile-specifier protein (NSP) (Fig. 7b)14. In addition, skimmianine (Fig. 6c) has been reported to attract certain Papilionidae butterflies, whereas a study on the interactions between Papilionidae and Rutaceae families27 has revealed that skimmianine has a prohibitive effect on these other butterflies. As clearly shown in the plant-compound matrix in Fig. 6d, plants belonging to the Brassicaceae and Rutaceae families typically possess glucosinolates and skimmianine, respectively. We found that genistein is specific to host plants of Lycaenidae butterflies; however, those plants belong to the family Fabaceae, which hosts a wide variety of Lycaenidae, Nymphalidae and Pieridae butterflies (Fig. 6e). Taking all of these results into consideration, we concluded that the identified common specific compounds contribute to the host-plant selectivity of mono-family-phagous butterflies.
(a–c) Compounds specific to host plants of Pieridae (a) Lycaenidae (b) and Papilionidae (c) families. (d) Matrix representation of plant species possessing the compounds shown in (a), (b) and (c). Among Pieridae-specific plant compounds, glucosinolates are represented by green dots; all others are represented by black dots. Lycaenidae-specific and Papilionidae-specific compounds are indicated by blue and red dots, respectively. (e) Plant–herbivore matrix of the plant species in (d) and the butterfly species that consume them.
Numbers below arrows indicate predictive values generated by E-zyme and E-zyme2 webservers. (a) Arabidopsis thaliana (plant family Brassicaceae) produces glucosinalbate. When plants are consumed by herbivores, the plant enzyme myrosinase acts on this compound to produce a toxic isothiocyanate compound. (b) Pieris rapae, a Pierinae butterfly, can prevent the production of the toxic isothiocyanate compound and instead produces a nitrile compound by altering the myrosinase activity with the help of nitrile-specifier protein (NSP). (c) Arabidopsis thaliana plants can also produce nitrile compounds from glucosinolates. (d–f) Putative chemical transformations of plant compounds (from KNApSAcK and Pherobase) to insect compounds (from Pherobase) identified by chemical structural comparison. The predicted transformations included a deduced reaction producing a compound possessed by a butterfly species (e).
Enzyme catalysis in plant–herbivore relationships
Because herbivorous butterflies incorporate plant chemicals whilst feeding, an herbivorous butterfly and its host plants can be generally assumed to share more compounds (C1 in Fig. 1g) than other butterfly–plant pairs. This assumption is consistent with the results of previous studies. For example, some insects have been found to use plant-derived chemicals as signalling materials or as precursors of chemical messengers (e.g., pheromones)15. In another study, volatile organic compounds produced by insects and plants had an overlap of 87%. Lepidoptera (butterflies and moths) members share significantly more aromatics than do Hymenoptera (bees and ants), whilst no significant difference in shared monoterpenes has been observed between the two orders28.
In this study, we expanded the above assumption to compounds not shared between butterflies and their host plants (C2, C2′, C5 and C6 in Fig. 1g). Even when compounds in a butterfly are not exactly the same as those in host plants, they may be derived from the latter (C2 and C2′ in Fig. 1g) via the insect’s metabolism. To overcome this problem, we used the Pherobase29 and KNApSAcK25 databases to find potential substrate–product pairs for all possible combinations of compound pairs (see Methods for details)30,31.
Our database search revealed four insect compounds that could possibly be such products (right-hand side of Fig. 7d–g). The corresponding substrates of all four compounds (Fig. 7d–g, left) are present in Brassicaceae plants; of these four compounds, three (Fig. 7d,e and g, left) were found to be Pieridae host-plant specific (Fig. 6a). We used the E-zyme webserver (see Methods) to search for known enzymatic reactions having the same chemical transformation patterns as the query reaction32,33. Only one of the four reactions (Fig. 7g) was shown to have the same chemical transformation pattern as the known glucosinolate-myrosinase reaction (Fig. 7a). The remaining three reactions (Fig. 7d–f) had the same chemical transformation patterns as those of an avoidance reaction (Fig. 7b,c)11,26. Among the possible products (Fig. 7d–g, right), only allylnitrile (Fig. 7e, right) was found to be possessed by a butterfly species, Pieris brassicae. No butterfly species possessed any other products, according to Pherobase. Although there is no direct evidence, we showed that the integration of plant–insect relationships, plant compounds and insect compounds provides indirect evidences to infer potential enzyme catalysis. If we integrate additional data, i.e., gene or protein sequence information, further analysis would be available.
To cope with glucosinolates, Pieridae family butterflies have developed the NSP protein34. In the presence of NSP, insect myrosinase acts on glucosinolates to produce nitrile compounds (Fig. 7b,c) that are less toxic than the isothiocyanate compounds produced when NSP is absent (Fig. 7a). These two alternative reactions correspond to the reactions predicted in Fig. 7 (i.e., the reactions in Fig. 7d–f and Fig. 7g correspond to the myrosinase reaction in the presence or absence of NSP, respectively). This NSP protein is not found in any other organisms and is thought to be specific to Pieridae family butterflies.
We therefore searched genome and transcriptome sequences to determine whether myrosinase and NSP genes are present in a wider range of insects. At the time of our survey, the National Center for Biotechnology Information (NCBI) database contained 17 complete and 36 draft genome sequences of insect species, whilst Sequence Read Archive (SRA)35 had genome and transcriptome sequences for 48 and 131 insect species, respectively. Although some protein sequences of Pieridae butterflies have been registered at NCBI, none of their genomes have yet been sequenced. We conducted a comprehensive search for myrosinase and NSP against all published insect genomic and transcriptomic data (see Methods). Myrosinase-like sequences were detected in all sequenced butterflies and moths (Bombyx mori, Chilo suppressalis, Danaus plexippus, Graphium sarpedon, Heliconius melpomene, Manduca sexta, Melitaea cinxia, Papilio glaucus, Papilio machaon, Papilio memnon, Papilio polytes, Papilio xuthus and Plutella xylostella), but no NSP-like sequences were detected. This result is consistent with butterfly host selection of glucosinolate-containing plants11.
Discussion
We have demonstrated that insects selectively feed on closely-related plants containing common specific compounds. Nevertheless, distinguishing between taxonomic and compound effects on plant–herbivore interactions (or host-plant selection) is difficult because plants in the same family may share many compounds. To overcome this complication, more careful examinations may be required. In particular, phylogenetic signals must be removed when evaluating the association between chemical compounds and plant–herbivore interactions. In this context, though comparative phylogenetic analysis36 is useful, this approach generally considers a simple evolutionary model in which species traits change in a Brownian fashion on a phylogenetic tree with accurate branch lengths. Phylogenetic comparative analysis was not applied in this study for two reasons. First, the phylogenetic tree was not accurate; thus, a phylogenetic comparative analysis may have led to misleading conclusions. Second, our dataset included only a few samples of specific plant–herbivore interactions and plant–compound relationships. In comparative phylogenetic analysis, a loss of statistical power is known to occur in small datasets37. Although definite conclusions were not obtained, we expect that the effect of compounds on plant–herbivore interactions is dominant14,15. In particular, only weak (although significant) phylogenetic signals have been observed in studies of plant–pollinator38 and seed-dispersal39 networks.
Our study has shown that the common specific compounds are mostly toxic compounds (Fig. 6), however, such common specific compounds were not found in Poaceae or Fabaceae, either of which is shared by some butterfly subfamilies in different families (Figs 3 and 4). This result may be due to the limited information in the examined databases but may also be explained by another hypothesis: Poaceae and Fabaceae do not possess particularly toxic compounds, and these less toxic plant species may serve as stepping stones during a transition to new host plants, because the butterflies were able to use their existing detoxification mechanisms to adapt to these plants. At least in Papilionidae, the diversification of butterfly species is thought to be correlated with changes in host plants9. Papilio machaon, one of many Papilionidae butterfly species that strongly depend on Rutaceae plants (Fig. 4), is thought to have evolved recently and feeds on a wide range of host plants, including Apiaceae. Rather than expanding to Apiaceae from Rutaceae plants in a single bound, a reasonable hypothesis is that Papilio machaon temporarily adopted less toxic hosts, such as Fabaceae plants, during its evolution.
When investigating host plant selection in this study, we only considered plant compounds; however, many other factors (such as hardness, leaf shape, and especially trichomes) are known to affect host plant edibility. Although these factors are described in field guides, encyclopaedias, and databases, their comprehensive and quantitative acquisition for use in informatics analysis is difficult. Moreover, the effects of phylogenetic signals were not considered in this study despite the importance of phylogenetic comparative analysis40. This is because an accurate phylogenetic tree is still unavailable. However, the absence of this analysis poses little problem because several studies have reported that phylogenetic signals are weak in ecological networks41,42. Despite these limitations, we have successfully shown that plant compounds generally contribute to the host-plant selection of phytophagous butterflies. Although molecular phylogeny is a powerful tool for understanding evolution, we strongly maintain that the integration of various types of data currently scattered across published articles and databases would enhance the study of natural evolutionary diversity.
Concluding remarks
Diversity is one of the most appealing characteristics of insects. Because of this diversity, however, unstructured knowledge is scattered throughout the literature and various databases, hindering comprehensive and systematic analyses. The strength of the current study is the integration of various types of data, including those pertaining to plant–herbivore relationships and plant–compound relationships, which enabled us to obtain a list of compounds that putatively contributed to host plant selection. By itself, this approach is insufficient to enable conclusions, but it can provide insights about molecular mechanisms drawn from the relationship data. Further enrichment of these relationship data would enhance and widen the scope of possible analyses, such as the prediction of host plant relationships and the inference of responsible genes or proteins using various machine-learning techniques.
Data and Methods
Plant–herbivore relationships
Although the HOSTS database (http://www.nhm.ac.uk/research-curation/research/projects/hostplants/) maintained by the Natural History Museum in the UK contains information on plant–herbivore relationships involving butterflies and their host plants worldwide, it lacks data on some relationships, especially those in the Japanese Islands. We therefore conducted a literature survey to collect plant–herbivore relationship information relevant to the Japanese Islands. The data collected and used in this study are summarised in the InsectInDB database (http://insect-plant.org/). The current InsectInDB release (as of January 2016) contains approximately 545 insects, 1,922 edible plants and 3,435 plant–herbivore relationships. InsectInDB provides links to plant–herbivore relationships and metabolomic and genomic data via the scientific names of insects or plants.
Plant-compound and insect-compound relationships
We collected plant compound data (Fig. 1c) using the KNApSAcK database25. We used Pherobase (http://www.pherobase.com), a major online resource cataloguing pheromone compounds that influence the behaviour of insects and other animals29, for the analysis of compounds common to insects and their host plants (Fig. 1g).
Interaction frequency between insect and plant families
To evaluate the strength of the interaction between insect family i and plant family j, we computed Wij, the number of interactions between the insects in family i and the plants in family j appearing in the plant–herbivore network. We also evaluated the statistical significance of Wij in the observed network using values from randomised networks (see below for details).
Randomised networks and significance
A comparison with randomised model networks was performed to evaluate the statistical significance of the observed network measures24,43,44,45. We used the null model 2 46, which is similar to the fixed row-fixed column null model used by Bascompte et al.47. This null model generates random bipartite networks with degree sequences identical to those of real networks. In the null model, the probability that a plant connects to an animal is proportional to the product of node degrees for a given plant or animal.
The significance of the network measure X was evaluated based on the Z-score: ZX = (Xreal – Xrand)/SDrand, where Xreal is the network measure of a real-world network, Xrand is the average value of the network measure, and SDrand is the standard deviation obtained from 1,000 randomised model networks.
Family-based evenness index
To measure the diversity of host plants of a specific insect category, we used a family-based evenness index based on Pielou’s evenness index48, which is a Shannon diversity index used to evaluate species evenness. In particular, the evenness index of insect category (species, subfamily or family) i was defined as follows:

where mf indicates the set of plant families appearing in the plant–herbivore network and |mf| is the number of plant families. Si and Nij are the number of plants interacting with insect category i (i.e., node degree of insect category i) and the number of plants in family j connecting to insect category i, respectively.
To characterise the overall trend of the evenness, we also considered the average family-based evenness: , where A and |A| denote the set and number of insect species, respectively.
, where A and |A| denote the set and number of insect species, respectively.
Compound contribution to plant–herbivore networks
Drawing on terminology established for flavour networks24, we defined a coefficient of compound contribution, χi, to evaluate the contribution of plant i to the feed-on-compound effect in plant–herbivore networks as follows:

where A and P denote sets of insect and plant species, respectively, A(i) and P(j) are the set of insects feeding on plant i and the set of plant species consumed by insect j, respectively, |X| is the size (number of elements) of set X, Ci is the number of compounds in plant i, fi represents the number of insects feeding on plant i, and S (i) is the average number of plant species that serve as prey in A(i). The first term indicates the average degree of the overlap of compounds between plant i and other plants in A(i) for the observed data. The second term is the expected value of the average degree obtained when assuming no host plant selectivity (i.e., insects feed on randomly selected plants). Thus, a positive or negative χi indicates the positive or negative contribution, respectively, of plant i to the feed-on-compound effect.
Chemical compound-based reaction prediction
The reaction prediction based on chemical compounds consists of two steps. The first step is the problem of enzymatic reaction-likeness, i.e., whether a pair of metabolic compounds can be the substrate and product of a single enzymatic reaction30. The second step is the prediction of the actual enzymes responsible for the putative enzymatic reactions33. To accomplish the first step, we used known chemical transformation patterns encoded as SMIRKS strings49 to search for pairs of metabolic compounds potentially possessing the same chemical transformation pattern. For the second step, we used the E-zyme32 and E-zyme233 webservers. E-zyme predicts putative enzyme classifications (referred to as Enzyme Commission numbers) with generated values reflecting the similarity between the given transformation patterns and known patterns. E-zyme2 predicts putative enzyme orthologues (i.e., homologous proteins/genes thought to have the same function). The values output by this program reflect the similarities of the transformation patterns as well as those of the surrounding conserved substructures.
Massive-scale sequencing data
DNA and RNA sequences were obtained from NCBI (http://www.ncbi.nlm.nih.gov/) and DDBJ SRA (http://trace.ddbj.nig.ac.jp/dra/index_e.html) databases35. Raw RNA-sequencing data for each insect were downloaded from SRA and then assembled with Trinity50, one of the most effective RNA-sequencing assemblers. The obtained Trinity contigs were used as training datasets in the AUGUSTUS program51 to predict protein-coding sequences in the corresponding insect genome downloaded from NCBI. This analysis generated a list of putative protein sequences encoded in the corresponding insect genomes.
Data Availability
The data collected in this study are summarised in a publicly accessible database, InsectInDB (http://insect-plant.org).
Additional Information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Jensen, N. B. et al. Convergent evolution in biosynthesis of cyanogenic defence compounds in plants and insects. Nat. Commun. 2, 273 (2011).
Gatehouse, J. A. Plant resistance towards insect herbivores: a dynamic interaction. New Phytol. 156, 145–169 (2002).
Konno, K. Plant latex and other exudates as plant defense systems: roles of various defense chemicals and proteins contained therein. Phytochemistry. 72 (13), 1510–1530 (2011).
Fürstenberg-Hägg, J., Zagrobelny, M. & Bak, S. Plant Defense against Insect Herbivores. Int. J. Mol. Sci. 14, 10242–10297 (2013).
Barabási A.-L. Network science. Philos. Trans. R. Soc. A. 371, 20120375 (2013).
Bascompte, J. Structure and dynamics of ecological networks. Science. 329, 765–766 (2010).
Allesina, S. & Tang, S. Stability criteria for complex ecosystems. Nature. 483, 205–208 (2012).
Mougi, A. & Kondoh, M. Diversity of interaction types and ecological community stability. Science. 337, 349–351 (2012).
Thompson, J. N. Evolutionary genetics of oviposition preference in swallowtail butterflies. Evolution. 42, 1223–1234 (1988).
Renwick, J. A. A., Radke, C. D., Sachdev-Gupta, K. & Städler, E. Leaf surface chemicals stimulating oviposition by Pieris rapae (Lepidoptera: Pieridae) on cabbage. Chemoecology. 3, 33–38 (1992).
Agerbirk, N., Olsen, C. E., Topbjerg, H. B. & Sørensen, J. C. Host plant-dependent metabolism of 4-hydroxybenzylglucosinolate in Pieris rapae: substrate specificity and effects of genetic modification and plant nitrile hydratase. Insect Biochem. Mol. Biol. 37, 1119–1130 (2007).
Nakayama, T. & Honda, K. Chemical basis for differential acceptance of two sympatric rutaceous plants by ovipositing females of a swallowtail butterfly, Papilio polytes (Lepidoptera, Papilionidae), Chemoecology. 14, 199–205 (2004).
Ozaki, K. et al. A gustatory receptor involved in host plant recognition for oviposition of a swallowtail butterfly. Nat. Commun. 2, 542 (2011).
Nishida, R. Chemical ecology of insect-plant interactions: ecological significance of plant secondary metabolites. Biosci. Biotechnol. Biochem. 78, 1–13 (2014).
Heidel-Fischer, H.-M. & Vogel, H. Molecular mechanisms of insect adaptation to plant secondary compounds. Curr. Opin. Insect Sci. 8, 8–14 (2015).
Bass, C. et al. Gene amplification and microsatellite polymorphism underlie a recent insect host shift. Proc. Natl. Acad. Sci. USA 110, 19460–19465 (2013).
Bidart-Bouzat, M. G. & Kliebenstein . An ecological genomic approach challenging the paradigm of differential plant responses to specialist versus generalist insect herbivores. Oecologia. 167, 677–689 (2011).
Wiklund, C. Generalist vs. Specialist Oviposition Behaviour in Papilio Machaon (Lepidoptera) and Functional Aspects on the Hierarchy of Oviposition Preferences. Oikos, 36, 163–170 (1981).
Tudor, O., Dennis, R. L. H., Greatorex-Davies, J. N. & Sparks, T. H. Flower preference of woodland butterflies in the UK: nectaring specialists are species of conservation concern. Biol. Conserv. 119, 397–403 (2004).
Ehrlich, P. R. & Raven, P. H. Butterflies and plants: A study in coevolution. Evolution 18, 586–608 (1964).
Ferrer-Paris, J. R., Sánchez-Mercado, A., Viloria, A. L. & Donaldson, J. Congruence and Diversity of Butterfly-Host Plant Associations at Higher Taxonomic Levels. PLoS One. 8, e63570 (2013).
The Angiosperm Phylogeny group. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG III. Bot. J. Linn. Soc. 161, 105–121 (2009).
De Jong, R., Vane-Wright, R. I. & Ackery, P. R. The higher classification of butterflies (Lepidoptera): problems and prospects. Insect Systematics & Evolution, 27, 1, 65–101 (1996).
Ahn, Y. Y., Ahnert, S. E., Bagrow, J. P. & Barabási, A. L. Flavor network and the principles of food pairing. Sci. Rep. 1, 196 (2011).
Afendi, F. M. et al. KNApSAcK family databases: integrated metabolite-plant species databases for multifaceted plant research. Plant Cell Physiol. 53, e1 (2012).
Nastruzzi, C. et al. In vitro cytotoxic activity of some glucosinolate-derived products generated by myrosinase hydrolysis. J. Agric. Food Chem. 44, 1014–1021 (1996).
Heininger, E. E. Effects of furocoumarin and furoquinoline allelochemicals on host plant utilization by Papilionidae. Graduate Dissertations and Theses at Illinois. (1989).
Schiestl, F. P. The evolution of floral scent and insect chemical communication. Ecol. Lett. 13, 643–656 (2010).
Symonds, M. R. & Elgar, M. A. The evolution of pheromone diversity. Trends Ecol. Evol. 23, 220–228 (2008).
Kotera, M. et al. Metabolome-scale prediction of intermediate compounds in multistep metabolic pathways with a recursive supervised approach. Bioinformatics. 30, i165–i174 (2014).
Yamanishi, Y., Tabei, Y. & Kotera, M. Metabolome-scale de novo pathway reconstruction using regioisomer-sensitive graph alignments. Bioinformatics. 31, i161–i170 (2015).
Yamanishi, Y., Hattori, M., Kotera, M., Goto, S. & Kanehisa, M. E-zyme: predicting potential EC numbers from the chemical transformation pattern of substrate-product pairs. Bioinformatics. 25, i179–86 (2009).
Moriya, Y. et al. Identification of Enzyme Genes Using Chemical Structure Alignments of Substrate-Product Pairs. J. Chem. Inf. Model. 56, 510–516 (2016).
Wheat, C. W., Vogel, H., Wittstock, U., Braby, M. F., Underwood, D. & Mitchell-Olds, T. The genetic basis of a plant-insect coevolutionary key innovation. Proc. Natl. Acad. Sci. USA 104, 20427–204231 (2007).
Nakazato, T., Ohta, T. & Bono, H. Experimental design-based functional mining and characterization of high-throughput sequencing data in the sequence read archive. PLoS One. 8, e77910 (2013).
Garland, T., Bennett, A. F. & Rezende, E.-L. Phylogenetic approaches in comparative physiology. J. Exp. Biol. 208, 3015–3035 (2005).
Griffith, O. L., Moodie, G. E. E. & Civetta, A. Genome size and longevity in fish. Exp. Gerontol. 38, 333–337 (2003).
Rezende, E. L., Jordano, P. & Bascompte, J. Effects of phenotypic comple- mentarity and phylogeny on the nested structure of mutualistic networks. Oikos. 116, 1919–1929 (2007).
Schleuning, M. et al. Ecological, historical and evolutionary determinants of modularity in weighted seed-dispersal networks. Ecol. Lett. 17, 454–463 (2014).
Garland, T., Bennett, A. F. & Rezende, E. L. Phylogenetic approaches in comparative physiology. J. Exp. Biol. 208, 3015–3035 (2005).
Rezende, E. L., Jordano, P. & Bascompte, J. Effects of phenotypic complementarity and phylogeny on the nested structure of mutualistic networks. Oikos. 116, 1919–1929 (2007).
Schleuning, M., Ingmann, L., Strau, R., Fritz, S. A., Dalsgaard, B. & Matthias Dehling, D. et al. Ecological, historical and evolutionary determinants of modularity in weighted seed-dispersal networks. Ecol Lett. 17, 454–463 (2014).
Thébault, E. & Fontaine, C. Stability of ecological communities and the architecture of mutualistic and trophic networks. Science. 329, 853–856 (2010).
Toju, H., Guimarães, P. R., Olesen, J. M. & Thompson, J. N. Assembly of complex plant-fungus networks. Nat. Commun. 5, 5273 (2014).
Feng, W. & Takemoto, K. Heterogeneity in ecological mutualistic networks dominantly determines community stability. Sci. Rep. 4, 5912 (2014).
Bascompte, J., Jordano, P., Melián, C. J. & Olesen, J. M. The nested assembly of plant-animal mutualistic networks. Proc. Natl. Acad. Sci. USA 100, 9383–9387 (2003).
Almeida-Neto, M., Guimaraes, P., Guimaraes P. R., Loyola, R. D. & Ulrich, W. A consistent metric for nestedness analysis in ecological systems: reconciling concept and measurement. Oikos 117, 1227–1239 (2008).
Hill, M. O. Diversity and Evenness: A Unifying Notation and Its Consequences. Ecology. 54, 427–432 (1973).
Daylight Theory Manual, Chapter 5. http://www.daylight.com/dayhtml/doc/theory/theory.smirks.html (accessed August 1, 2016).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–439 (2006).
Acknowledgements
This work was financially supported by the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT), the Japan Society for the Promotion of Science (JSPS) (MEXT/JSPS Kakenhi grant number 25870379), and the Japan Science and Technology Agency. This work was also supported by the JST/MEXT Programme to Promote the Tenure Track System in Tokyo Institute of Technology.
Author information
Authors and Affiliations
Contributions
A.M., K.T., K.O. and M. Kotera designed the study. K.T. and M. Kotera performed the network analysis. A.M. and M. Kotera performed the metabolomic and genomic analyses and interpretation. K.O., T.N. and T.T. provided advice on the analyses. S.K. and N.M. helped with data integration and management. Y.C. and M. Kono collected and organised information on phytophagous relationships. A.M., K.T., and M. Kotera drafted the manuscript. All authors completed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Muto-Fujita, A., Takemoto, K., Kanaya, S. et al. Data integration aids understanding of butterfly–host plant networks. Sci Rep 7, 43368 (2017). https://doi.org/10.1038/srep43368
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep43368
This article is cited by
-
Polyhydroxy Acids as Fabaceous Plant Components Induce Oviposition of the Common Grass Yellow Butterfly, Eurema Mandarina
Journal of Chemical Ecology (2023)
-
Oviposition stimulants underlying different preferences between host races in the leaf-mining moth Acrocercops transecta (Lepidoptera: Gracillariidae)
Scientific Reports (2022)
-
LepTraits 1.0 A globally comprehensive dataset of butterfly traits
Scientific Data (2022)
-
Machine learning for the meta-analyses of microbial pathogens’ volatile signatures
Scientific Reports (2018)
-
Oviposition inhibitor in umbelliferous medicinal plants for the common yellow swallowtail (Papilio machaon)
Journal of Natural Medicines (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.