Abstract
Determinants of species diversity in microbial ecosystems remain poorly understood. Bacteriophages are believed to increase the diversity by the virtue of Kill-the-Winner infection bias preventing the fastest growing organism from taking over the community. Phage-bacterial ecosystems are traditionally described in terms of the static equilibrium state of Lotka-Volterra equations in which bacterial growth is exactly balanced by losses due to phage predation. Here we consider a more dynamic scenario in which phage infections give rise to abrupt and severe collapses of bacterial populations whenever they become sufficiently large. As a consequence, each bacterial population in our model follows cyclic dynamics of exponential growth interrupted by sudden declines. The total population of all species fluctuates around the carrying capacity of the environment, making these cycles cryptic. While a subset of the slowest growing species in our model is always driven towards extinction, in general the overall ecosystem diversity remains high. The number of surviving species is inversely proportional to the variation in their growth rates but increases with the frequency and severity of phage-induced collapses. Thus counter-intuitively we predict that microbial communities exposed to more violent perturbations should have higher diversity.
Similar content being viewed by others
Introduction
An important and largely unsolved question in microbial ecology is what determines the diversity of microbial ecosystems. Indeed, unbridled competition between microbes sharing common resources would eventually limit species diversity not to exceed the number of different nutrient types1. Predation by bacteriophages introduces the negative frequency-dependent selection2,3,4,5 which offers the possibility for a dramatically larger species diversity5. In the classical Kill-the-Winner (KtW) model of Thingstad5 virulent phages reduce populations of their susceptible hosts to a low steady state level, which is independent of hosts’ growth rate thus allowing multiple species per nutrient type. The number of co-existing bacterial species in the resulting ecosystem is determined exclusively by the parameters of phage predation5, the topology of the phage-bacterial infection network6,7,8, and the carrying capacity of the environment4,7,8,9.
Microbial populations are routinely exposed to more dynamics than assumed in the traditional steady state KtW model and its variants. Extended Lotka-Volterra equations for two layer ecosystems of phages and bacteria could predict persistently varying populations10, even without considering mutations. In addition, microbial systems are typically exposed to changes in interaction rules and new invading species. For example, in the lab experiments11 E. coli population suffered a dramatic collapse by a factor ~104–105 caused by a T7 phage infection. Collapse-driven dynamics is common in both natural12 and man-made13,14,15,16 ecosystems in which bacteria are engaged in the continuous arms race with phages17,18,19,20,21.
Here we propose and explore a particularly simplified dynamical interpretation of Kill-the Winner principle, in which bacterial populations are characterized by periods of competitive exponential growth punctuated by rapid and severe collapses. Larger bacterial populations in our model are proportionally more likely to be infected by phages. Furthermore, in larger and thus denser populations such infections once started are likely to eliminate a sizable fraction of susceptible hosts resulting in a severe collapse in the populations of individual bacterial strains. The proposed collapse scenario should should be understood as a very simplified version of a dynamics of an open system that is exposed to a variety of externally stresses. Stresses associated to new mutations of already present phages, or “epidemics” caused by new invading virulent phages.
When viewed over a long period of time any given species would repeatedly cycle between low and high population numbers. Such cyclic dynamics of populations of individual species masked by an approximately constant total population saturated at the carrying capacity of the environment is discussed in the ecological literature as “cryptic cycles”22,23,24.
Model
Consider a number of bacterial species/strains sharing the same environment and competing for the same rate-limiting nutrient defining its carrying capacity. Their populations sizes at time t are denoted as Pi(t), where i = 1, 2, …, N. Each of these individual species is exposed to rare but severe collapse events in which its population is suddenly and drastically reduced by a constant factor γ ≪ 1. We assume that these collapses happen relatively rarely so that the total population of all bacterial species has sufficient time to reach the steady state value given by the overall carrying capacity of the environment. Without loss of generality carrying capacity can be set to 1, so that in between collapses one has ∑jPj(t) = 1. In our model we assume that while the total population of all stays constant, relative population sizes of individual species continue to change exponentially in-between collapse due to differences in their fitness in the saturated environment.
In the spirit of Kill-the-Winner principle we assume that the rate of collapse of the species i is proportional to its population size Pi. Due to a broad distribution of population sizes this rule strongly biases collapses towards one or few largest populations. We assume that collapse events are independent of each other, so that the time interval between consecutive collapses is exponentially distributed with mean τ.
One update cycle in our model consist of three steps:
(1) Draw a time interval Δt until the next collapse event from the exponential distribution P(Δt) = exp(−Δt/τ)/τ.
(2) Calculate population sizes at the time of collapse. In between collapse events relative population sizes are assumed to change exponentially while the total population stays saturated at 1 (the carrying capacity of the environment):

(3) Select one species to collapse with the probability equal to its relative population size Pi(t + Δt) and multiply its population by γ.
In our simulations each of N species is assigned its individual growth rate drawn from the Gaussian distribution with zero mean and standard deviation σ. The value of the mean is not important since normalization of the overall population to 1 ensures that only relative growth rates matter. Furthermore, the exact form of the distribution of growth rates is not particularly important. In our mathematical analysis we will use a more convenient exponential distribution of growth rates: P(gi) = exp(−gi/σ)/σ, while delegating more cumbersome derivations for the Gaussian P(gi) to supplementary materials.
Results
Collapses supports Diversity
Figure 1A shows a typical outcome of a simulation of our model with γ = 10−3 and σ = 4 over around 100 population collapses after which only D = 3 out of N = 8 species survive. The relative growth rate gi of the species is the main predictor on whether it will survive or not. Indeed, as shown by the rainbow coloring of curves in Fig. 1A ranging from dark red (the slowest growing) to purple (the fastest growing) the 3 surviving species have the largest values of gi.
(A) Time courses of populations of N = 8 species with fixed growth rates assigned from the Gaussian distribution with standard deviation σ = 4. Rainbow colors correspond to growth rates ranging from the slowest (dark red) to the fastest (purple). For each of the species, the likelihood of collapse is proportional to its population sizes (“Kill-the-Winner” rule) and the collapse ratio γ = 10−3 is the same for all species. Only 3 fastest growing species survive in the long term. (B) The final diversity D counted as number of surviving species as function of σΔt - the spread of growth rates integrated over the average time between collapses. Each black dot represents the outcome of one simulation started with N = 500 species exposed to collapse ratio γ = 10−6. The dashed line is the analytical fit similar to Eq. 4 but here done for the Gaussian distribution of growth rates used in the simulation (see Supplementary Materials for details).
A natural question to ask is what determines the number of surviving species/strains in the steady state of the model? In the limit of very rare collapses the fastest growing species would diverge from the rest of the population so much that it will be the only one to survive, as indeed expected from the competitive exclusion principle1.
The situation is more complex for intermediate rate of collapses where more than one of the fastest growing species can coexist with each other but some of the slowest growers become extinct. In the steady state each of these surviving species repeatedly cycles between low and high populations. Faster growing species reach large population sizes more often which makes them to collapse more frequently thus eliminating their growth advantage. As we show below this balance can be sustained within a finite range of growth rates.
For each of the species its individual growth rate gi is reduced by the same negative number −gcc due to the overall resource competition quantified by the denominator in Eq. 1. In the steady state the excess growth rate of each of the surviving species (gi − gcc) must be exactly compensated by the logarithmic population losses |log γ| due to collapses happening at the species-specific probability ci:

Note that as the probability of collapse (per update) ci = (gi − gcc)Δt|log γ| needs to be positive and normalized. Positive values of ci means that only the fastest growing species with gi > gcc would survive in the long run. The collapse rates of these D surviving species are further constrained by normalization  , reflecting the requirement of one collapse per update. Using eq. 2 the threshold gcc is then determined by:
, reflecting the requirement of one collapse per update. Using eq. 2 the threshold gcc is then determined by:

For a given set of species, this allows us to self-consistently calculate gcc and D.
For gi selected from the exponential distribution with standard deviation σ the diversity D is given by (see Supplementary Information for derivation)

This expression holds for average diversity provided that it is larger than 1. This is because a single fastest growing species would always survive. Clearly D is also capped from above by N. Similar relation holds for the Gaussian distribution of growth rates and is in agreement with our numerical simulations of the model shown in Fig. 1B.
For the exponential distribution the growth rate threshold above which a species survives is given by gcc = σ log(N/D) = σ log(NσΔt/|log γ|). Note that while threshold explicitly depends on the starting number of species, the final diversity given by Eq. 4 is independent of N. This particular property of the exponential distribution would be modified for other distributions resulting in a mild dependence of D on N.
A Gaussian distribution of growth rates would slightly increase the diversity compared to the exponential distribution with the same spread, while a more fat-tailed *(say, power law) distribution would decrease it.
Our basic model can be generalized to the case where different species have different collapse ratios γi. This may for example reflect their different degrees of vulnerability to phages, or different ways to partition their population in physical space. The only consequence of this modification is that log γ in the equations above needs to be replaced by its average value across species (see supplementary materials for simulation results).
In our model the collapse probability of a given species is proportional to its population size. Thus time-averaged relative population size of each of the species species is equal to its overall collapse frequency 〈Pi(t)〉t = ci. This is consistent with “Kill-the-Winner” principles according to which species with larger populations collapse more often.
Figure 2B illustrates this cyclic dynamics in a system containing a mixture of slow and fast growing species. Surviving populations mostly grow, but do so at different rates. Their coexistence is possible only because of the negative feedback via “Kill-the-Winner” rule where populations of an individual species get severely reduced once it starts to dominate the overall biomass. The population of each of the species goes through approximately periodic cycles of growth and collapses with the period Ti = 1/ci = |log γi|/(gi − gcc)Δt (in units of collapse events). Thus the slowest surviving species (marked blue in Fig. 2B) nearly never collapse, whereas the fastest growing species (marked red in Fig. 2B) obtain dominance and expose themselves to a collapse on a much shorter timescale. Individual collapse events of these species are marked in Fig. 2B with red and blue arrows correspondingly. Note that the population of the slowest growing species often decreases not due to a phage-mediated collapse but simply because it gets temporarily outgrown by other species with a faster growth rate.
(A) D = 10 species with identical growth rates and collapse ratios γi = 10−6. The highlighted purple curve illustrates the characteristic growth and collapse cycle for a particular population. (B) Simulation with D = 10 surviving species (down from N = 60) each with growth rates selected from the Gaussian distribution with σ = 3, and identical collapse ratios γi = 10−6. The blue arrow and the red arrows mark times for collapse events of these two species. Note how the fastest growing species (red line) collapses much more often than the slowest growing species (blue line) which only collapsed once during the time shown. The growth rate difference between these two species is gmax − gmin = 2.44.
For comparison in Fig. 2A we show a system of the same size (D = 10) but where all species have exactly the same growth rate. In that case the system has a very long memory of the initially imposed order of species populations, because even after a long time each of the species would collapse the same number of times. That is, if one species have experienced one more collapse than the others, it would be smaller by a factor γ and thus be much less exposed to subsequent collapses until it would regrow to the size where it again may collapse with a non-negligible probability. Indeed, populations shown in Fig. 2A follow much more regular oscillatory dynamics than those with unequal growth rates shown in Fig. 2B.
Model with collapses to a fixed threshold
In our standard version of the KtW model the collapsing population is reduced by a constant factor: Pi → Pi ⋅ γ. An alternative possibility is that following a collapse the population starts at a fixed small threshold value γ irrespective of its earlier population size. This would be the case e.g. when following a collapse the local population is completely eliminated and is reintroduced by one individual from a neighboring region. It can also happen when a collapse drives one species extinct only to be quickly replaced by a single bacterium of a new species. In the thus defined fixed threshold kill-the winner model (KtWT) the diversity remains close to what was reported in Fig. 1B (data now shown). The dynamics is also characterized by individual population undergoing cycles of duration defined by their relative growth rates much similar to what is shown in Fig. 2 for our original model. However the long term memory of cycle order is reduced compared to the constant factor model discussed above, simply because every collapse completely erases the population history of the collapsed species. In what follows we explore the dynamical properties of the fixed threshold model and its variants.
“Kill-the-King” Model
To better understand the cyclic dynamics in the KtWT model we first consider its extreme and deterministic version in which the next collapse always happens at the largest population. We will refer to this version as Kill-the-King (KtK) model. Furthermore, we assume that the growth rates gi of all species are equal to each other. Thereby the asymptotic dynamics becomes periodic with period N when time is measured by collapse events.
To concentrate on slow trends in population size dynamics we only measure them between intervals where each population collapsed exactly once, which in KtK secure that the order of populations is exactly preserved. We relabel species in the order of decreasing population sizes and calculate the ratios δi = Pi+1/Pi < 1 between successive population sizes (The ratio δn for the currently smallest population Pn is defined by its value after the next collapse when it becomes the second smallest). As shown in the supplementary materials, in KtK model these ratios evolve according to the following discrete equation describing changes acquired after a full round of N collapses so that each member of the population collapsed exactly once:

The steady state of the equation is reached when all ratios are equal to each other, i.e. δi+1−δi = 0. In this case the logarithms of population sizes are equidistantly distributed in the interval of length |log γ| so that δi(∞) = δ* = γ1/N. Figure 3 shows a simulation of KtK model with N = 10 and γ = 10−6 One can see how it asymptotically approaches this steady state.
For clarity we show the state of the model only every 10 collapses, that is to say, after each species collapsed exactly once so that the population order is maintained. The steady state of KtK model where all ratios δi between rank-ordered populations are equal to each other and to γ1/N is approximately reached already after 300 collapses. The relaxation to this steady state is described by the discrete anisotropic Burgers equation 5 or its continuous counterpart Eq. 6.
The asymptotic dynamics of KtK is described by the discrete Eq. 5 which for large N can be approximated by a continuous PDE (see SI for more details) in which the continuous coordinate x replaces the species rank i:

Here δ(x) has periodic boundary conditions over x-interval [0, N]. As its discrete counterpart this equation describes the state of our system every N’th time-step. This equation is closely related to the Burgers equation25,26, although it differs in terms of the diffusion coefficient that instead of being constant as in refs 25,26 and is proportional to δ(t).
Having finished with Kill-the-King model we return to Kill-the-Winner fixed Threshold (KtWT) model. In the KtWT model population collapses do not always happen in the order dictated by their relative sizes. This results in a somewhat chaotic dynamics illustrated in Fig. 4A. When a smaller population collapses out of turn it causes only a very small rescaling of other population sizes. The (very likely) subsequent collapse of the largest population leads to a situation where these two just collapsed populations become nearly equal in size ( ). This dramatically increases the likelihood for further re-orderings between these two species, resulting in an extended period where these two species fight for dominance. This intermittent dynamics switching the order of populations is clearly visible in Fig. 4B with D = 3. The nearly vertical lines clustered around collapse events 5100, 5200, and 5400 correspond to frequent shifts in the population order of three species within the cycle.
). This dramatically increases the likelihood for further re-orderings between these two species, resulting in an extended period where these two species fight for dominance. This intermittent dynamics switching the order of populations is clearly visible in Fig. 4B with D = 3. The nearly vertical lines clustered around collapse events 5100, 5200, and 5400 correspond to frequent shifts in the population order of three species within the cycle.
(A) Dynamics of all 3 species, emphasizing that the cyclic order occasionally changes, caused by an “out of order” collapse of a population that is not the largest. (B) Same time series as in the above panel, but only showing every third time-point. This panel highlights the interplay between occasional intermittent alternations in the cyclic order (clustered vertical lines) and longer “quiet” periods during which ratios of rank-ordered populations relax towards δ* = γ1/N.
An intermittent region ultimately ends with a particular order winning over. After this all populations slowly relax back to the steady state with equal ratios δ* (curved lines ending in horizontal plateaus in Fig. 4B). The exact form of the relaxation to the steady state is derived in supplementary materials. While δ(t) ≫ δ* the relaxation is proportional to 1/(t/N). The expected number of collapse events for iδ(t) to go from ~1 to ~δ* is ~N/δ* or about 300 for the parameters used in Fig. 4. When  the relaxation crosses over to δ(t) − δ* ~ exp(−δ*t).
the relaxation crosses over to δ(t) − δ* ~ exp(−δ*t).
Discussion
Here and before27 we investigated the impact of severe and sudden population collapses on ecosystem composition and diversity. This approach is complementary to a more traditional description of ecosystem dynamics at or around the steady state solution2,5,9,28. The emergent cyclic dynamic in our model is entirely collapse-driven and thus distinct from either stable or transient periodic oscillations present in predator-prey ecosystems described by the Lotka-Volterra equations10,23,24,28,29,30.
The key assumption used in our study is that larger populations are more exposed to sudden collapses than the smaller populations. This is the foundation of “Kill-the-Winner” (KtW) principle proposed in ref. 5. The resulting negative (or stabilizing) frequency-dependent selection promotes the ecosystem diversity even in the simplest case considered above, where species interact with each other only via competition for a single rate-limiting resource. This KtW bias is very important as it shifts the collapse-driven dynamics away from “diversity waves” we reported before27 towards population cycles investigated in this study. Indeed, as demonstrated in ref. 27 a version of collapse-driven dynamics in which the likelihood of a collapse is uncorrelated with population size or even biased towards smaller populations (Kill-the-Looser model) results in ebbing and flowing species diversity and bi-modal distribution of species abundances. This should be contrasted with constant diversity predicted in the KtW, KtWT, and KtK models studied here. In our scenario diversity is maintained as population reaches dominance and collapses in a particular order. The species abundance distribution in the here presented models is not bi-modal but uniform on the logarithmic scale (data not shown).
The model in this paper focus on one particularly lethal aspect of density dependent selection, and analyze it in detail. A key result is eq. 4, that quantify a dynamical maintenance of diversity, through frequent collapses of the largest populations. The falsifiable prediction is that higher flux of new phages makes well mixed microbial ecosystem more diverse. The obtained diversity is obtained by preferential attack on the largest population, whereas the traditional steady state KtW emphasize the coexistence of slow and fast growing bacteria in presence of phage5. Our argument for targeting the largest population preferentially is that new virulent phages tend to induce larger collapses for host population that facilitate more effective spreading of the phage. In effect our scenario is similar to the assumption that larger population densities have larger effective R0-factors for an “epidemic”.
Our approach is a simplified approach to analyze an open, yet well defined system. Apart from assuming the preferential targeting of large populations, it is formulated in the limit where the total population reach carrying capacity before new invasions take place. This last assumption can easily be relaxed by assigning a collapse rate/collapse size that depend on population sizes also before the sum of all populations reaches carrying capacity. We tested that such increasingly frequent collapses also allowed for higher microbial diversity, and found that it open for even higher diversity than predicted by eq. 4. This is because the bacteria are then so violently exposed to phages that they never sense their mutual competition at saturation.
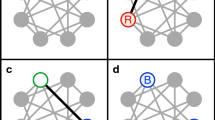
To test how sensitive are our results with respect to introduction of other types of interactions between bacterial species as well as to a more branched topology of Phage-Bacterial Infection Networks (PBIN) we simulated a variant of our model where in addition to abundant (KtW) species the infecting phage results in collapse of a constant number K of other bacterial species. This version of the model is reminiscent of the Bak-Sneppen model of species co-evolution31. We tested this model for K = 1 and K = 25 (out of N = 500). In the first case we observed no impact on diversity, while in the second case the diversity saturated at lower values of σΔt. All together we can conclude that the diversity profile shown in Fig. 1B remains qualitatively (and sometimes even quantitatively) unaffected by additional interactions between microbial species or more interconnected PBINs.
According to our results the principal determinant of the ecosystem diversity D is the width σ of the distribution of logarithmic growth rates of individual bacterial strains or species. This difference is amplified during the average time Δt between population collapses. Thus the overall frequency of collapses is a very important parameter with more frequent collapses counter-intuitively resulting in more diverse ecosystems. That is because in our scenario frequent collapses weaken the effect of competitive exclusion ultimately driving the diversity down to no more than single species per rate-limiting nutrient. Larger magnitude of collapses also promotes higher diversity but its impact increases only weakly (logarithmically) with the collapse ratio γ.
It is instructive to compare the determinants of microbial diversity in the static, steady state KtW model and in our more dynamic, collapse-driven variant. In the static KtW model2,5 the steady state population size of each of the bacteria B* = δ/βη is determined exclusively by parameters of the phage to which it is susceptible: its burst size (β), death (or dispersal and dilution) rate (δ), and its infection rate (η) at a density equal the bacterial carrying capacity. This steady state population of a phage-controlled bacterium is usually much lower than the carrying capacity of the environment: B* ≪ 1. Thus a large number of bacteria each susceptible to its unique phage predator can coexists with each other5. Higher diversity can subsequently be achieved by carefully adding pairs of bacteria and phages, latter possibly supplemented by their epigenetic variants32, each consuming a small fraction of the carrying capacity5,9. Substantial diversity is found to be fragile to new invaders, in the form of bacteria that grow faster than resident ones or phages that prey on several bacteria at once9.
In contrast to this the diversity in our model is determined by both statistics of collapses as well as the spread of growth rates of resident bacterial species. In case of mild or infrequent collapses and large disparity in bacterial growth rates competitive exclusion principle is restored within our model as it then predicts an ecosystem dominated by just one fastest growing bacterium. When collapses are frequent (short Δt) and severe (large |log γ|), while growth rates of individual bacterial strains or species are close to each other (small σ), Eq. 4 predicts high diversity of co-existing bacterial species. This prediction is robust with respect to exact causes of collapses, including the relatively frequent16 invasion of phages that are capable of infecting several bacterial species.
Overall, the falsifiable (and counter-intuitive) prediction of the collapse-driven “Kill-the-Winner” model differentiating it from its stationary counterpart, is that by increasing frequency (and to a smaller extent severity) of collapses one could support higher diversity of microorganisms. In real world ecosystems, this particular aspect of density dependent selection is to be supplemented by more classical engines of microbial diversity2,5,9,32 and in addition be modulated in case spatial heterogeneity33,34 reduce the assumed collapse sizes.
Additional Information
How to cite this article: Maslov, S. and Sneppen, K. Population cycles and species diversity in dynamic Kill-the-Winner model of microbial ecosystems. Sci. Rep. 7, 39642; doi: 10.1038/srep39642 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Hardin, G. The competitive exclusion principle. Science 131, 1292–1297 (1960).
Campbell, A. Conditions for the existence of bacteriophage. Evolution 15, 153–165 (1961).
Levin, B. R. Frequency-dependent selection in bacterial populations. Philosophical transactions–Royal Society. Biological sciences 319(1196), 459–472 (1988).
Rodriguez-Valera, F. et al. Explaining microbial population genomics through phage predation. Nature Reviews Microbiology 7(11), 828–836 (2009).
Thingstad, T. F. Elements of a theory for the mechanisms controlling abundance, diversity, and biogeochemical role of lytic bacterial viruses in aquatic systems. Limnology and Oceanography 45(6), 1320–1328 (2000).
Moebus, K. & Nattkemper, H. Bacteriophage sensitivity patterns among bacteria isolated from marine waters. Helgoländer Meeresuntersuchungen 34(3), 375–385 (1981).
Flores, C. O., Meyer, J. R., Valverde, S., Farr, L. & Weitz, J. S. Statistical structure of host–phage interactions. Proceedings of the National Academy of Sciences, USA 108(28), E288–E297 (2011).
Weitz, J. S. et al. Phage–bacteria infection networks. Trends in microbiology 21(2), 82–91 (2013).
Haerter, J. O., Mitarai, N. & Sneppen, K. Phage and bacteria support mutual diversity in a narrowing staircase of coexistence. The ISME journal 8(11), 2317–2326 (2014).
Jover, L. F., Cortez, M. H. & Weitz, J. S. Mechanisms of multi-strain coexistence in host–phage systems with nested infection networks. Journal of theoretical biology 332, 65–77 (2013).
Chao, L., Levin, B. R. & Stewart, F. M. A complex community in a simple habitat: an experimental study with bacteria and phage. Ecology 58, 369–378 (1977).
Campbell, B. J., Yu, L., Heidelberg, J. F. & Kirchman, D. L. Activity of abundant and rare bacteria in a coastal ocean. Proceedings of the National Academy of Sciences, USA 108(31), 12776–12781 (2011).
Fernández, A. et al. How stable is stable? Function versus community composition. Applied and Environmental Microbiology 65(8), 3697–3704 (1999).
Hantula, J., Kurki, A., Vuoriranta, P. & Bamford, D. H. Ecology of bacteriophages infecting activated sludge bacteria. Applied and environmental microbiology 57, 2147–2151 (1991).
Shapiro, O. H., Kushmaro, A. & Brenner, A. Bacteriophage predation regulates microbial abundance and diversity in a full-scale bioreactor treating industrial wastewater. The ISME journal 4, 327–336 (2010).
Shapiro, O. H. & Kushmaro, A. Bacteriophage ecology in environmental biotechnology processes. Current opinion in biotechnology 22(3), 449–455 (2011).
Sun, C. L. et al. Phage mutations in response to CRISPR diversification in a bacterial population. Environmental microbiology 152, 463–470 (2013).
Labrie, S. J., Samson, J. E. & Moieau, S. Bacteriophage resistance mechanisms. Nature Reviews, Microbiology 8, 317–327 (2010).
Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Comparative genomics of defense systems in archaea and bacteria. Nucleic acids research 4(18), 4360–4377 (2013).
Wommack, K. E. & Colwell, R. R. Virioplankton: viruses in aquatic ecosystems. Microb Molec Biol Rev 64, 69–114 (2000).
Wigington, C. H. et al. Re-examining of the relationship between marine virus and microbial cell abundance. bioRxiv p. 025544 (2015).
Jones, L. E. & Ellner, S. P. Effects of rapid prey evolution on predator–prey cycles. Journal of mathematical biology 55(4), 541–573 (2007).
Yoshida, T. et al. Cryptic population dynamics: rapid evolution masks trophic interactions. PLoS Biol 5(9), e235 (2007).
Wang, Z. & Goldenfeld, N. Fixed points and limit cycles in the population dynamics of lysogenic viruses and their hosts. Physical Review E 82, 011918 (2010).
Burgers, J. M. Application of a model system to illustrate some points of the statistical theory of free turbulence. Proc. Acad. Sci. Amsterdam 43 no 2 (1940).
Hopf, E. The partial differential equation y t + yy x = μ xx . Communications on Pure and Applied Mathematics 3(3), 201–230 (1950).
Maslov, S. & Sneppen, K. Diversity waves in collapse-driven population dynamics. Plos Computational Biology 11 e1004440 (2015).
Levin, B. R., Stewart, F. M. & Chao, L. Resource-limited growth, competition, and predation: a model and experimental studies with bacteria and bacteriophage. American Naturalist 111(977), 3–24 (1977).
Weitz, J. S., Hartman, H. & Levin, S. A. Coevolutionary arms races between bacteria and bacteriophage. Proceedings of the National Academy of Sciences, USA 102(27), 9535–9540 (2005).
Shih, H. Y. & Goldenfeld, N. Path-integral calculation for the emergence of rapid evolution from demographic stochasticity. Physical Review E 90(5), 050702 (2014).
Bak, P. & Sneppen, K. Punctuated equilibrium and criticality in a simple model of evolution. Physical review letters 71, 4083–4086 (1993).
Sneppen, K., Semsey, S., Seshasayee, A. S. N. & Krishna, S. Restriction modification systems as engines of diversity. Front. Microbiol. 6, 528, doi: 10.3389/fmicb.2015.00528 (2015).
Heilmann, S., Sneppen, K. & Krishna, S. Coexistence of phage and bacteria on the boundary of self-organized refuges. Proceedings of the National Academy of Sciences, USA 109(31), 12828–12833 (2012).
Haerter, J. O. & Sneppen, K. Spatial structure and Lamarckian adaptation explain extreme genetic diversity at CRISPR locus. MBio 3(4), e00126–12 (2012).
Author information
Authors and Affiliations
Contributions
S.M. and K.S. contributed equally to the work and ideas presented in this manuscript. Both authors wrote the manuscript text and reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Maslov, S., Sneppen, K. Population cycles and species diversity in dynamic Kill-the-Winner model of microbial ecosystems. Sci Rep 7, 39642 (2017). https://doi.org/10.1038/srep39642
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep39642
This article is cited by
-
Nutrients strengthen density dependence of per-capita growth and mortality rates in the soil bacterial community
Oecologia (2023)
-
Mutualistic interplay between bacteriophages and bacteria in the human gut
Nature Reviews Microbiology (2022)
-
Shotgun metagenomics evaluation of soil fertilization effect on the rhizosphere viral community of maize plants
Antonie van Leeuwenhoek (2022)
-
Evolutionary dynamics of predator in a community of interacting species
Nonlinear Dynamics (2022)
-
Inter-phylum negative interactions affect soil bacterial community dynamics and functions during soybean development under long-term nitrogen fertilization
Stress Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.