Abstract
The conversion of normal prion protein (PrP) into pathogenic PrP conformers is central to prion disease, but the mechanism remains unclear. The α-helix 2 of PrP contains a string of four threonines, which is unusual due to the high propensity of threonine to form β-sheets. This structural feature was proposed as the basis for initiating PrP conversion, but experimental results have been conflicting. We studied the role of the threonine string on PrP conversion by analyzing mouse Prnpa and Prnpb polymorphism that contains a polymorphic residue at the beginning of the threonine string, and PrP mutants in which threonine 191 was replaced by valine, alanine, or proline. The PMCA (protein misfolding cyclic amplification) assay was able to recapitulate the in vivo transmission barrier between PrPa and PrPb. Relative to PMCA, the amyloid fibril growth assay is less restrictive, but it did reflect certain properties of in vivo prion transmission. Our results suggest a plausible theory explaining the apparently contradictory results in the role of the threonine string in PrP conversion and provide novel insights into the complicated relationship among PrP stability, seeded conformational change, and prion structure, which is critical for understanding the molecular basis of prion infectivity.
Similar content being viewed by others
Introduction
Transmissible spongiform encephalopathies (TSEs), or prion diseases, are a group of fatal neurodegenerative diseases that includes Creutzfeldt-Jakob disease (CJD) in humans, scrapie in sheep and goats, bovine spongiform encephalopathy (BSE) in cattle, and chronic wasting disease (CWD) in cervids1. The prion hypothesis posits that the conversion of normal host-encoded prion protein (PrP) is central to the pathogenesis of these diseases2,3. The normal PrP isoform (PrPC) is a cell surface glycoprotein that is soluble, protease-sensitive, and rich in α-helices. During the disease, some PrPs convert to the aberrantly folded isoform (PrPSc), which is aggregated, resistant to protease digestion, and rich in β-sheets3,4,5. The pathogenic PrPSc is able to seed a conformational change of normal PrPC into PrPSc, resulting in PrPSc accumulation and neurodegeneration. Decades of research have provided compelling evidence supporting the hypothesis6,7.
The normal isoform, PrPC, contains a long unfolded N-terminal region and a C-terminal globular domain consisting of three α-helices and two short anti-parallel β-strands8,9,10,11. Although small changes in the amino acid sequence do not significantly alter the overall structure of PrPC9,10, they can severely alter prion transmissibility12,13,14,15,16. PrP sequence variations in different species create strong barriers against inter-species prion transmission17,18. Within the same species, PrP polymorphisms also greatly influence the susceptibility to intra-species prion transmission12,15,19,20. A classic example of an intra-species barrier is the mouse PrP polymorphism of allele a (Prnpa; Leu at residue 108 and Thr at residue 189) and allele b (Prnpb; Phe at residue 108 and Val at residue 189), which drastically alter the length of the incubation period of prion disease12,20,21. Elegant studies with knock-in mice revealed that the most efficient prion transmission requires both residues in the host PrP to match those of the inoculum PrP20. While both residues 108 and 189 affect the incubation time, residue 189 plays a major role in determining the efficiency of prion transmission20.
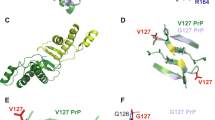
Residue 189 is within the highly conserved string of 4 threonines located at the C-terminus of α-helix 2 of PrP (Fig. 1)22. Because threonine tends to form β-sheet structures23, such a sequence of β-sheet-prone threonines in a helix is highly unusual. This local instability has been proposed as the underlying structural basis for initiating the PrPC-to-PrPSc conversion24. Consistent with this hypothesis, the C-terminus of α-helix 2 has been found to be unstable25,26, and the peptide corresponding to helix 2 is able to form amyloid fibrils that can act as seeds for full-length wild-type PrP27. Replacing threonines with α-helix-prone alanines stabilizes the PrPC structure and prevents in vitro PrP aggregation28. A thermodynamic study of the α-helix 2 peptide showed that the free energy separating the α-helix and β-sheet conformations of this peptide is very small, indicating that the structural behavior of this peptide is likely determined by the environment29. Indeed, compounds binding to this region have been shown to prevent PrP conversion and increase the survival of diseased animals30,31,32.
Schematic representation of murine PrP.
PrP consists of a long N-terminal disordered region and a C-terminus that has three α-helices and two anti-parallel β-strands. The sequences of PrPa, PrPb, two derivative mutants (L108F and T189V), and three residue-191 mutants are shown; mutations are indicated in red.
Interestingly, the insertion of extra amino acids in the C-terminus of α-helix 2 does not affect the ability of mutant ovine PrPs to convert to the infectious PrPSc conformation22. Further, a comprehensive mutagenesis study of mouse PrP showed that individually replacing the amino acids in the C-terminus of α-helix 2 with any other amino acid does not significantly influence the formation of infectious prions in scrapie-infected N2a cells33. Very recently, Munoz-Montestino reported that deleting 4 or 5 amino acids in this region (including all four threonines) does not affect the conversion of ovine PrP to infectious prions in RK13 cells34. These data apparently contradict the observations from prion transmission studies in Prnpbknock-in mice, which showed that the threonine 189 (mouse numbering) significantly affects disease incubation times20.
The above studies were performed on cultured cells or living animals, and multiple factors might alter the efficiency of PrP conversion in these complicated systems (such as cell metabolic state, PrP presentation on the cell surface, the presence of multiple cofactors, etc.). Therefore, we chose two relatively simple in vitro assays that recapitulate many characteristics of PrPC-to-PrPSc conversion to study the influence of the threonine string in prion conversion. The first is protein misfolding cyclic amplification (PMCA), which consists of successive sonication and incubation steps35. We previously showed that this technique can convert bacterially expressed recombinant PrP (recPrP) into aggregated and proteinase K (PK)-resistant conformer in the presence of cofactors36,37. This conformational change produces recPrion, the pathogenic conformer of recPrP, that causes bona fide prion disease in wild-type mice with pathological properties identical to those of naturally occurring prions36,38. In addition to PMCA, we also used a denaturant-induced recPrP amyloid fibril formation assay39,40,41, which is commonly used to model PrP conversion.
Using these two assays, we compared recombinant PrPa and PrPb, two derivatives of PrPa bearing either Phe at residue 108 (108 F) or Val at residue 189 (189 V), and three PrPa mutants that replace Thr 191 with either valine, alanine, or proline. Our results offer new insights into the conversion of PrP into an infectious prion.
Results
PrPb polymorphism compromises recPrP conversion by PMCA
The presence of Phe 108 and Val 189 in PrPb creates a strong barrier to the transmission of PrPa-based prion strains20,42. The recPrion was generated de novo by PMCA from recPrPa protein36 and it has been propagated with recPrPa for years. To determine whether there is a transmission barrier between recPrion and recPrPb, we propagated recPrion with recPrPb or with the PrPa-derived mutants recPrPL108F and recPrPT189V. PMCA was carried out as described previously38,43; the PMCA substrate consisted of recPrP plus two cofactors, synthetic POPG and total RNA isolated from normal mouse liver38,43. For each recPrP variant, PMCA was carried out for six rounds, and the generation of the C-terminal PK-resistant PrP band at round 6 was considered a successful propagation. As a positive control, recPrPa was successfully propagated (Fig. 2 and Table 1). In contrast, PrPb and the two derivative mutants failed to propagate in all 20 trials (Fig. 2 and Table 1). These data suggested that, similar to the barrier observed in animal study20, there is a barrier for propagating recPrPa-adapted recPrion to recPrPb and that both polymorphic residues 108 and 189 contribute to the barrier.
PMCA propagation of recPrion.
Recombinant PrPa, PrPb, PrPL108F, or PrPT189V was used as the substrate for PMCA. Each PMCA reaction was carried out for 6 rounds and PMCA products were digested with 50 μg/mL PK at 37 °C for 30 min. PrP was detected by immunoblot analysis with POM1 anti-PrP antibody55. “C” indicates undigested recPrP as a control. The original immunoblot images are in the Supplementary Figure S3.
PrPb polymorphism at residues 108 and 189 differently affect recPrP amyloid fibril growth
Next, we analyzed the influence of PrPb polymorphism on PrP conversion with the less stringent but commonly used in vitro recPrP amyloid fibril growth assay39. In unseeded reactions, the amyloid fibril formation by recPrP was highly variable (Fig. 3a). Within the 96-h experiment, 4 out of 5 recPrPa replicates formed amyloid fibrils de novo after a lag time of about 56 h. For recPrPb, only 1 out of 5 replicates formed amyloid fibrils de novo after a lag time of about 79 h, and none of the other 4 recPrPb replicates formed fibrils within the 96-h experiment (Fig. 3a). These results were consistent with a previous report showing that recPrPb has a reduced capability to form amyloid fibrils44.
Amyloid fibril growth with recombinant PrPa, PrPb, PrPL108F, or PrPT189V.
Thioflavin-T (ThT) kinetics of recombinant PrPa, PrPb, and two derivative mutants (L108F or T189V) in the absence (a) or in the presence (b) of seeds. The seeds used were pre-formed recPrPa amyloid fibrils. Shaded areas are error bars, which indicate standard deviation.
Surprisingly, when the two PrPa-derived mutants recPrPL108F and recPrPT189V were tested, they behaved differently. Within the 96-h experiment, 4 out of 5 recPrPL108F replicates formed amyloid fibrils de novo with a lag time similar to that of recPrPa (Fig. 3a). The recPrPT189V however, behaved similar to recPrPb (Fig. 3a), that is, only 2 out 5 recPrPT189V replicates formed amyloid fibril de novo within 96 h, and the average lag phase was around 91 h. The same trend was observed in seeded reactions, in which all four recPrPs were seeded with the same recPrPa fibril seed and amyloid fibril growth was detected in all samples with less variation. The recPrPL108F behaved like recPrPa, both showing a rapid growth of amyloid fibrils (Fig. 3b). In contrast, recPrPT189V and recPrPb amyloid fibril growth had a longer lag phase, and the intensity of ThT fluorescence signal was much lower (Fig. 3b). Since these reactions were monitored at the same time, in the same plate, and under identical condition, the difference in amyloid fibril growth is likely resulted from the different amino acid sequences of recPrP.
Overall, these results supported the idea that the recPrPb protein has a reduced capability to form amyloid fibrils, which is largely due to the influence of residue 189 of PrP. These findings are consistent with the observations in animal studies that residue 189 is the major determinant for the transmission barrier between PrPa and PrPb. However, the amyloid fibril growth results regarding residue 108 were different from in vivo findings20.
The influence of mutations at residue 191 on recPrP conversion
Because residue 189 has a larger influence on the PrPa/PrPb transmission barrier and it is at the beginning of the 4-threonine string of α-helix 2, it is reasonable to ask if changing another threonine residue within that string would have similar effects. Thus, we mutated Thr-191 in recPrPa to Val to mimic the recPrPT189V change in recPrPb. We also mutated Thr-191 to α-helix-prone Ala or to the secondary structure breaker Pro. All three mutants were expressed, purified, and subject to PMCA.
Unlike recPrPT189V, recPrPT191V was able to propagate recPrion by PMCA, but the efficiency was lower (about 60%) and the PK-resistant band at the 6th round was generally weaker than that of recPrPa (Fig. 4 and Table 1). The recPrPT191A mutant had an even lower efficiency, only around 25% of reactions completed six rounds of propagation, and the PK-resistant band was also weaker than that of recPrPa (Fig. 4 and Table 1). On replacing Thr 191 with Pro, none of the 20 trials yielded successful propagation (Fig. 4 and Table 1). Thus, relative to Thr-189, Thr-191 is more tolerant to amino acid replacement (with the exception of Pro): both Val and Ala at this position can be tolerated but with less-efficient recPrion propagation by PMCA.
PMCA propagation of recPrion with recombinant PrPT191A, PrPT191V, or PrPT191P as substrate.
Each reaction was run for 6 rounds and PMCA products were digested with 50 μg/mL PK at 37 °C for 30 min. PrP was detected by immunoblot analysis with POM1 anti-PrP antibody. “C” indicates undigested recPrP as a control. The original immunoblot images are in the Supplementary Figure S3.
All residue-191 mutants were assayed for amyloid fibril growth. In unseeded reactions, 4 our 5 recPrPT191A or recPrPT191V were able to form fibrils de novo, which were similar to wild-type recPrPa (4 out 5 replicates formed amyloid fibrils) except for the longer lag phases (Fig. 5a). For recPrPT191P however, only 1 out of 5 replicates formed fibril de novo within the 96-h experimental period, and the lag phase was 94 h (Fig. 5a). When fibril growth was seeded with recPrPa fibril seed, all three residue-191 variants grew amyloid fibrils robustly (Fig. 5b). Thus, except for the reduced ability of recPrPT191P to form amyloid fibrils de novo, no significant difference in amyloid fibril growth was found when Thr-191 was replaced with Val, Ala, or Pro.
Detecting prion infectivity of converted recPrP variants using cell infection assay
The Elispot plate–based cell infection assay is a sensitive and quantitative prion infectivity measure45,46. We used CAD5 cells, a clonally selected cell line that is susceptible to infection by most mouse prion strains45,46, to test PMCA products for two purposes. First, we wanted to determine whether the in vitro converted recPrP variants were infective. Second, because the endogenous PrP in CAD5 cells is wild-type PrPa, using converted recPrP variants to infect the cells would allow us to evaluate the effect of amino acid mismatch on PrP conversion.
The round-6 PMCA products were used for the assay. As shown in Fig. 6, no infectivity was detected in the recPrP variants that did not produce a detectable C-terminal PK-resistant band, i.e., recPrPb, recPrPL108F, recPrPT189V, and recPrPT191P (Fig. 6). This result supports the notion that PrP conformation is the main determinant of prion infectivity. The three recPrP variants that generated the PK-resistant conformer at round 6 all had prion infectivity (Fig. 6). Although the PK-resistant bands of recPrPT191V and recPrPT191A were much weaker than that of recPrPa (Figs 2 and 4), the infectivity of recPrPT191V was comparable to that of wild-type recPrPa and the infectivity of T191A mutant was lower by about one order of magnitude. Nonetheless, the undiluted samples of recPrPa, recPrPT191V, and recPrPT191A produced similar numbers of cells carrying PK-resistant PrPSc (Fig. 6), suggesting that the amino acid mismatch at 191 did not significantly affect the seeded conversion process.
Elispot cell culture assay of prion infectivity.
CAD5 cells were infected with serial 10-fold dilutions of each PMCA product as indicated. Elispot assay was performed by filtering 20,000 third-passage cells onto the membranes of Elispot plates, and the proportion of cells contained PK-resistant PrP was identified by enzyme-linked immunosorbent assay (ELISA).
Thermal stability of recPrP variants
The sequence of four threonines at the C-terminus of α-helix 2 is unstable, which has led to it being proposed as a possible initiator of PrP conversion24,25,26. To determine the change in thermal stability among different recPrP variants generated in this study, we used a CPM (7-diethylamino-3-(4-maleimidophenyl)-4-methylcoumarin) thermal stability assay47,48. Among all the recPrPs, recPrPa had the highest melting temperature (Tm = 78.8 °C) (Fig. 7, Table 1). The Tm for recPrPb was 74.1 °C, suggesting it is less thermally stable than recPrPa. All single amino acid variants except for recPrPT191P had melting temperatures around 70 °C, and recPrPT191A had the highest Tm among this group (70.3 °C), consistent with the idea that the α-helix prone alanine stabilizes the helical structure of recPrP. Interestingly, recPrPT191P had the highest melting temperature (74.3 °C) among all recPrP variants, suggesting that this recPrP, with a shortened α-helix 2, is quite stable; this result is consistent with a recent report34. Importantly, there was no correlation between thermal stability and PrP conversion, either by PMCA or by amyloid fibril growth assay.
Discussion
In this study, we investigated the influence of the string of four threonines on PrP conversion by using two in vitro PrP conversion assays. The PMCA assay showed that the transmission barrier between murine PrPa and PrPb can be recapitulated in recPrion propagation and that both residues 108 and 189 contribute to the barrier. The amyloid fibril growth assay revealed a strong influence of residue 189 on fibril growth, but residue 108 had little effect. Moreover, our results also showed a position effect in the threonine string. When Thr-191 was mutated to Val or Ala, the mutants converted to the infectious prion conformation with a reduced efficiency. But when Thr-191 was mutated to Pro, it could not be converted by PMCA, suggesting that this residue is likely to be part of the β-sheet core, but not the loop region of in the final infectious conformation. Finally, we have shown that the variations in thermal stability caused by these PrP polymorphisms or mutations do not correlate with the ability of these recPrP to convert to either amyloid fibrils or infectious prions.
The lack of correlation between PrP thermal stability and convertibility is not particularly surprising. The instability of PrP will lead to misfolding or aggregation, but infectious prion conformations and/or amyloid fibril structures are ordered protein aggregates with defined structures, which require the normal PrP not only to unfold, but also to gain the defined structure of infectious prions or amyloid fibrils. Since unfolded PrP does not spontaneously convert to ordered aggregates, simply increasing the instability of PrP will increase PrP misfolding, but not necessarily result in increased production of infectious prions or amyloid fibrils.
The PrPa/PrPb transmission barrier was originally identified through genetic analyses of prion disease susceptibility in mouse strains49,50 and elegantly dissected by using knock-in mouse technology20,21. Here we used PMCA to show that the difference of two amino acids between recPrPa and recPrPb created a barrier to propagating recPrPa-adapted recPrion, and both residues contributed to this barrier. Note that by changing the PMCA parameters, it is possible to adapt the prion propagation to various types of prions, as illustrated by propagating mouse prion to hamster PrPC via PMCA51. In order to study the PrPa/PrPb transmission barrier, we did not make any adjustment and used PMCA parameters that propagate recPrion with recPrPa to test all recPrP variants, which allowed us to detect differences in transmission efficiency. The compromised PMCA propagation with recPrPb or the derivative mutants recPrPL108F and recPrPT189V was consistent with in vivo results20.
Relative to PMCA, the amyloid fibril growth assay is less restrictive, and all recPrP variants grew amyloid fibrils (Suppl. Fig. 1, Table 1). Similar to the report by Cortez et al.44, we found that recPrPb was less efficient in forming amyloid fibrils. Unexpectedly, recPrPL108F behaved similarly to recPrPa in both unseeded and seeded reactions (Figs 3 and 5), suggesting that the properties of recPrPb in amyloid fibril growth are largely determined by residue 189. This finding is consistent with observations from animal study20, but the lack of influence of residue 108 is not. Unlike PMCA products, none of the amyloid fibrils could infect cultured CAD5 cells (data not shown) or seed the PMCA reaction (Suppl. Fig. 2). These observations suggest that the amyloid fibril growth assay is a less faithful PrP-conversion assay than PMCA. Notably, it has been suggested that recPrP has the freedom to oligomerize to a large number of assemblies in the presence of 2 M GuHCl (a condition for growing PrP amyloid fibrils) and recPrP amyloid fibrils are able to adapt to the structure that is most suitable for the buffer conditions44,52. Thus, the misfolded recPrP has more freedom to adopt various structures under amyloid fibril growth condition, which may account for the less faithfulness by the amyloid fibril growth assay. Nonetheless, our finding that residue 189 is the major determinant of the fibril-growing property of recPrPb reveals that the amyloid fibril growth assay does recapitulate certain aspects of in vivo PrP conversion.
Thr-189 is at the beginning of the unusual four-threonine sequence in the α-helix 2 of PrP (Fig. 1), and the role of this string in PrP conversion is controversial22,24,25,26,33,34. In our PMCA study, all variants in this string had a reduced capability to form infectious prions, which could be due to the influence on three aspects of PrP conversion: (1) the seeding process, (2) the PrP conformational change, and (3) the final prion structure. Given that recPrion formed by recPrPT191V or recPrPT191A efficiently seeds wild-type PrP in CAD5 cells (Fig. 6), the amino acid mismatch between recPrP variants and wild-type PrP does not appear to significantly affect the seeding process. Because a single amino acid change is unlikely to alter the protein conformational change process, the most plausible explanation is that the threonine string forms part of the β-sheeted core of recPrion. This theory is consistent with our observation that replacing Thr-191 with Pro, a secondary structure breaker, strongly inhibits the propagation of recPrion by PMCA.
Although our data is consistent with the threonine string being a part of the β-sheeted core of recPrion, it does not suggest that this sequence is an obligatory part of the core in all prion strains. The finding by Munoz-Montesino et al. that deleting all four threonines does not affect the conversion of ovine PrP in RK13 cells34 indicates that this stretch is in the non-essential part in certain prion strains. Removing the threonine string may reduce the restraint of the final infectious prion structure of some prion strains, which would allow the PrP to better accommodate the incoming prion structure and propagate prion structures that it normally has difficulty propagating. This would explain the more efficient propagation of ovine PrP-adapted CJD strains by threonine deletion mutant34. Overall, our theory that the threonine string constitutes the β-sheeted core in certain prion strains, but is in the non-essential region in other strains explains the seemingly conflicting results, particularly the strong influence of residue 189 on prion transmission20 and the efficient prion formation by the threonine deletion mutant34.
It is important to note that the recPrion was formed de novo in a test tube with recPrP36, which might be different from native prions formed in animals. The PMCA and amyloid fibril growth are both in vitro assays, which may only capture certain aspects of the in vivo prion propagation process. Notwithstanding the limitations, these in vitro assays offer relatively pure systems allowing us to assess the direct effects of these amino acids on PrP conversion, which is difficult to achieve using in vivo systems. Moreover, the relevance of our findings to in vivo prion biology is supported by the facts that the recPrion causes bona fida prion disease in wild-type animals36,38. Further, our PMCA results are fully consistent, and our findings from amyloid fibril growth assay are partially consistent, with the results of in vivo prion transmission in knock-in mice20.
In summary, we compared the PMCA propagation of recPrion with the amyloid fibril growth assay; analyzed the complicated relationships among PrP thermal stability, PrP conversion, and prion infectivity, which have resulted in novel insights into the role of the threonine string in forming infectious prions. Confirmation that the threonine string can be part of the β-sheeted core or non-essential region in different prion strains may offer a novel view of the structural differences among various strains and help elucidate the molecular structure of infectious prions.
Methods
Site-directed mutagenesis
The mutant variants of the mouse PrP were generated from pPROEX HTb vector containing mouse PrP23–230 using the QuikChange site-directed mutagenesis kit (Stratagene). The primers are listed in the online supplementary information, which were synthesized by Integrated DNA Technologies, Inc. (IDT). All mutations were confirmed by DNA sequencing.
Expression and purification of recPrPs
All recPrP proteins were expressed in E. coli BL21 (DE3) and purified according to a previously reported protocol53 with minor modifications. Briefly, inclusion bodies were collected (15,000 × g, 30 min, 4 °C), resuspended in 6 M Guanidine hydrochloride (GuHCl), 10 mM Tris-HCl, 100 mM Na-PO4 buffer, 10 mM β-mercaptoethanol (βME), pH 8.0, sonicated on ice until completely solubilized, and applied to a Ni-NTA column (Qiagen). The recPrP was refolded on the column by running a GuHCl gradient (from 6 M GuHCl, 10 mM Tris-HCl, 10 mM βME, pH 8.0 to 10 mM Tris-HCl, 100 mM Na-PO4 buffer, pH 8.0) at 1 mL/min. After wash, recPrP was eluted with 10 mM Tris-HCl, 100 mM Na-PO4 buffer, and 500 mM imidazole, pH 5.8. All eluted fractions were combined and dialyzed using 7,000 MWCO, SnakeSkinTM Dialysis Tubing (Thermo Scientific) for 1.5 h against 10 mM Na-PO4 buffer, pH 5.8, and then dialyzed overnight against deionized water. Protein samples were lyophilized (FreeZone 4.5, LABCONCO) and stored at −80 °C for further use.
Protein misfolding cyclic amplification (PMCA)
The PMCA reaction was carried out as previously described43,54. Each PMCA round consisted of 48 cycles of sonication (0.5 min) and incubation (29.5 min), and 1/10 of the reaction mixture was transferred to a fresh substrate mixture to start a new round.
Amyloid fibril formation assay
For amyloid fibril growth, 0.5 mg/mL of recPrP was incubated in 2 M GuHCl, 100 mM K-PO4 buffer, pH 6.5, and 20 μM thioflavin T (ThT). The reaction volume was 200 μL per well in 96-well plates (Corning, Lot No. 065514030). In the seeded reaction, seeding was achieved by adding 1 μL of mouse recPrP fibrils (0.5 mg/mL) to each well. The plate was incubated at 37 °C with continuous shaking on a mcrioplate reader (SYNERGY2, BioTek). The fibril kinetics was monitored by measuring ThT fluorescence intensity every 15 min using 440-nm excitation and 480-nm emission. The amyloid-formation kinetics was calculated from five replicates using GraphPad Prism (GraphPad Software, Inc.).
The enzyme-linked immunospot (Elispot) cell infection assay
The Elispot cell infection assay was adapted from previous studies45,46 with minor modifications. Briefly, 200 μL of PMCA products at round 6 were collected and centrifuged at 100,000 × g, 4 °C for 1 hour and the pellets were washed twice, followed by centrifugation at 100,000 × g, 4 °C for 1 hour after each wash. After the final wash, the pellets were resuspended in 200 μL of CAD5 growth media (OPTI-MEM, 5% BGS and 1% penicillin and streptomycin) and sonicated for 30 seconds with 50% output (Misonic Sonicator XL2020). Each sample was serially diluted 10, 100, and 1,000 times and 60 μL of undiluted and diluted samples were used to infect CAD5 cells. After two 1:10 splits, 20,000 CAD5 cells/well were transferred to the Millipore 96-well Elispot plates (MSIPN4W) and subjected to the Elispot assay45,46. The images were taken by S6 Micro Analyzer (CTL Analyzers, LLC) and processed by the ImmunoSpot software (CTL Analyzers, LLC). The graph was generated using GraphPad Prism (GraphPad Software, Inc.).
Thermal unfolding using CPM fluorescence dye
The thermal stability assay was performed as previously described48. Briefly, 4 mg of 7-diethylamino-3-(4-maleimidophenyl)-4-methylcoumarin (CPM) (Sigma) was dissolved in 1 mL of DMSO. The solution was diluted 20-fold with water to make a working solution, which was kept on ice during the experiment. Diluted dye (40 μL) was mixed with 480 μL of 2 mg/mL recPrP and used immediately while protected from light to reduce photobleaching. The thermal denaturation assay was performed in a total volume of 130 μL of reaction mixture, and the mixture was transferred to a sub-micro quartz fluorometer cuvette and heated at a rate of 2 °C/min in a Cary Eclipse fluorescence spectrophotometer (Agilent Technologies). The fluorescence signal was monitored as the temperature increased (excitation 387 nm, emission 463 nm). The assay was performed over the temperature range of 20–95 °C. Data were fitted using the GraphPad Prism program (GraphPad Software, Inc.), and the melting temperature (Tm) was calculated from the Boltzmann sigmoidal equation. Each experiment’s calculations were based on an average of 4 replicates.
Additional Information
How to cite this article: Abskharon, R. et al. The role of the unusual threonine string in the conversion of prion protein. Sci. Rep. 6, 38877; doi: 10.1038/srep38877 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Aguzzi, A. Prion diseases of humans and farm animals: epidemiology, genetics, and pathogenesis. J Neurochem 97, 1726–1739 (2006).
Prusiner, S. B. Novel proteinaceous infectious particles cause scrapie. Science 216, 136–144 (1982).
Prusiner, S. B. Prions. Proc Natl Acad Sci USA 95, 13363–13383 (1998).
Aguzzi, A., Baumann, F. & Bremer, J. The prion’s elusive reason for being. Annu Rev Neurosci 31, 439–477, doi: 10.1146/annurev.neuro.31.060407.125620 (2008).
Caughey, B. & Chesebro, B. Prion protein and the transmissible spongiform encephalopathies. Trends in Cell Biol. 7, 56–62 (1997).
Soto, C. Prion hypothesis: the end of the controversy? Trends in biochemical sciences 36, 151–158, doi: 10.1016/j.tibs.2010.11.001 (2011).
Ma, J. & Wang, F. Prion disease and the ‘protein-only hypothesis’. Essays Biochem 56, 181–191, doi: 10.1042/bse0560181 (2014).
Riek, R. et al. NMR structure of the mouse prion protein domain PrP(121–321). Nature 382, 180–182 (1996).
Calzolai, L., Lysek, D. A., Perez, D. R., Guntert, P. & Wuthrich, K. Prion protein NMR structures of chickens, turtles, and frogs. Proc Natl Acad Sci USA 102, 651–655, doi: 10.1073/pnas.0408939102 (2005).
Lysek, D. A. et al. Prion protein NMR structures of cats, dogs, pigs, and sheep. Proc Natl Acad Sci USA 102, 640–645, doi: 10.1073/pnas.0408937102 (2005).
Knaus, K. J. et al. Crystal structure of the human prion protein reveals a mechanism for oligomerization. Nature structural biology 8, 770–774 (2001).
Westaway, D. et al. Distinct prion proteins in short and long scrapie incubation period mice. Cell 51, 651–662 (1987).
Barron, R. M. et al. Changing a single amino acid in the N-terminus of murine PrP alters TSE incubation time across three species barriers. The EMBO journal 20, 5070–5078 (2001).
Hizume, M. et al. Human prion protein (PrP) 219K is converted to PrPSc but shows heterozygous inhibition in variant Creutzfeldt-Jakob disease infection. J Biol Chem 284, 3603–3609, doi: 10.1074/jbc.M809254200 (2009).
Asante, E. A. et al. A naturally occurring variant of the human prion protein completely prevents prion disease. Nature 522, 478–481, doi: 10.1038/nature14510 (2015).
Collinge, J. & Clarke, A. R. A general model of prion strains and their pathogenicity. Science 318, 930–936 (2007).
Beringue, V., Vilotte, J. L. & Laude, H. Prion agent diversity and species barrier. Veterinary research 39, 47, doi: 10.1051/vetres:2008024 (2008).
Scott, M. et al. Transgenic mice expressing hamster prion protein produce species-specific scrapie infectivity and amyloid plaques. Cell 59, 847–857 (1989).
Wadsworth, J. D. et al. Human prion protein with valine 129 prevents expression of variant CJD phenotype. Science 306, 1793–1796 (2004).
Barron, R. M. et al. Polymorphisms at codons 108 and 189 in murine PrP play distinct roles in the control of scrapie incubation time. The Journal of general virology 86, 859–868, doi: 10.1099/vir.0.80525-0 (2005).
Moore, R. C. et al. Mice with gene targetted prion protein alterations show that Prnp, Sinc and Prni are congruent. Nature genetics 18, 118–125, doi: 10.1038/ng0298-118 (1998).
Salamat, K. et al. Integrity of helix 2-helix 3 domain of the PrP protein is not mandatory for prion replication. J Biol Chem 287, 18953–18964, doi: 10.1074/jbc.M112.341677 (2012).
Minor, D. S. et al. Evaluation of blood pressure measurement and agreement in an academic health sciences center. Journal of clinical hypertension 14, 222–227, doi: 10.1111/j.1751-7176.2012.00599.x (2012).
Ji, H. F. & Zhang, H. Y. beta-sheet constitution of prion proteins. Trends in biochemical sciences 35, 129–134, doi: 10.1016/j.tibs.2009.12.002 (2010).
Hosszu, L. L. et al. Definable equilibrium states in the folding of human prion protein. Biochemistry 44, 16649–16657, doi: 10.1021/bi051277k (2005).
Bjorndahl, T. C. et al. Detailed biophysical characterization of the acid-induced PrP(c) to PrP(beta) conversion process. Biochemistry 50, 1162–1173, doi: 10.1021/bi101435c (2011).
Yamaguchi, K., Matsumoto, T. & Kuwata, K. Critical region for amyloid fibril formation of mouse prion protein: unusual amyloidogenic properties of the helix 2 peptide. Biochemistry 47, 13242–13251, doi: 10.1021/bi801562w (2008).
Singh, J., Kumar, H., Sabareesan, A. T. & Udgaonkar, J. B. Rational stabilization of helix 2 of the prion protein prevents its misfolding and oligomerization. Journal of the American Chemical Society 136, 16704–16707, doi: 10.1021/ja510964t (2014).
Tizzano, B. et al. The human prion protein alpha2 helix: a thermodynamic study of its conformational preferences. Proteins 59, 72–79, doi: 10.1002/prot.20395 (2005).
Kuwata, K. et al. Hot spots in prion protein for pathogenic conversion. Proc Natl Acad Sci USA 104, 11921–11926, doi: 10.1073/pnas.0702671104 (2007).
Ronga, L. et al. Does tetracycline bind helix 2 of prion? An integrated spectroscopical and computational study of the interaction between the antibiotic and alpha helix 2 human prion protein fragments. Proteins 66, 707–715, doi: 10.1002/prot.21204 (2007).
Forloni, G. et al. Tetracyclines affect prion infectivity. Proc Natl Acad Sci USA 99, 10849–10854 (2002).
Shirai, T. et al. Evaluating prion models based on comprehensive mutation data of mouse PrP. Structure 22, 560–571, doi: 10.1016/j.str.2013.12.019 (2014).
Munoz-Montesino, C. et al. Generating Bona Fide Mammalian Prions with Internal Deletions. J Virol 90, 6963–6975, doi: 10.1128/JVI.00555-16 (2016).
Morales, R., Duran-Aniotz, C., Diaz-Espinoza, R., Camacho, M. V. & Soto, C. Protein misfolding cyclic amplification of infectious prions. Nature protocols 7, 1397–1409, doi: 10.1038/nprot.2012.067 (2012).
Wang, F., Wang, X., Yuan, C. G. & Ma, J. Generating a prion with bacterially expressed recombinant prion protein. Science 327, 1132–1135, doi: 10.1126/science.1183748 (2010).
Zhang, Z. et al. De novo generation of infectious prions with bacterially expressed recombinant prion protein. FASEB J 27, 4768–4775, doi: 10.1096/fj.13-233965 (2013).
Wang, X. et al. Intraperitoneal Infection of Wild-Type Mice with Synthetically Generated Mammalian Prion. PLoS Pathog 11, e1004958, doi: 10.1371/journal.ppat.1004958 (2015).
Cobb, N. J., Apostol, M. I., Chen, S., Smirnovas, V. & Surewicz, W. K. Conformational stability of mammalian prion protein amyloid fibrils is dictated by a packing polymorphism within the core region. J Biol Chem 289, 2643–2650, doi: 10.1074/jbc.M113.520718 (2014).
Bocharova, O. V., Breydo, L., Parfenov, A. S., Salnikov, V. V. & Baskakov, I. V. In vitro conversion of full-length mammalian prion protein produces amyloid form with physical properties of PrP(Sc). J Mol Biol 346, 645–659 (2005).
Legname, G. et al. Synthetic mammalian prions. Science 305, 673–676 (2004).
Cortez, L. M. & Sim, V. L. Implications of prion polymorphisms. Prion 7, 276–279, doi: 10.4161/pri.25566 (2013).
Wang, F., Wang, X. & Ma, J. Conversion of bacterially expressed recombinant prion protein. Methods 53, 208–213, doi: 10.1016/j.ymeth.2010.12.013 (2011).
Cortez, L. M., Kumar, J., Renault, L., Young, H. S. & Sim, V. L. Mouse prion protein polymorphism Phe-108/Val-189 affects the kinetics of fibril formation and the response to seeding: evidence for a two-step nucleation polymerization mechanism. J Biol Chem 288, 4772–4781, doi: 10.1074/jbc.M112.414581 (2013).
Mahal, S. P. et al. Prion strain discrimination in cell culture: the cell panel assay. Proc Natl Acad Sci USA 104, 20908–20913, doi: 10.1073/pnas.0710054104 (2007).
Li, J., Browning, S., Mahal, S. P., Oelschlegel, A. M. & Weissmann, C. Darwinian evolution of prions in cell culture. Science 327, 869–872, doi: 10.1126/science.1183218 (2010).
Alexandrov, A. I., Mileni, M., Chien, E. Y., Hanson, M. A. & Stevens, R. C. Microscale fluorescent thermal stability assay for membrane proteins. Structure 16, 351–359, doi: 10.1016/j.str.2008.02.004 (2008).
Wang, Z., Ye, C., Zhang, X. & Wei, Y. Cysteine residue is not essential for CPM protein thermal-stability assay. Analytical and bioanalytical chemistry 407, 3683–3691, doi: 10.1007/s00216-015-8587-4 (2015).
Dickinson, A. G., Meikle, V. M. & Fraser, H. Identification of a gene which controls the incubation period of some strains of scrapie agent in mice. Journal of comparative pathology 78, 293–299 (1968).
Carlson, G. A. et al. Linkage of prion protein and scrapie incubation time genes. Cell 46, 503–511 (1986).
Castilla, J. et al. Crossing the species barrier by PrP(Sc) replication in vitro generates unique infectious prions. Cell 134, 757–768, doi: 10.1016/j.cell.2008.07.030 (2008).
Alvarez-Martinez, M. T. et al. Dynamics of polymerization shed light on the mechanisms that lead to multiple amyloid structures of the prion protein. Biochim Biophys Acta 1814, 1305–1317, doi: 10.1016/j.bbapap.2011.05.016 (2011).
Zahn, R., von Schroetter, C. & Wuthrich, K. Human prion proteins expressed in Escherichia coli and purified by high-affinity column refolding. FEBS Lett 417, 400–404 (1997).
Wang, F. et al. Genetic informational RNA is not required for recombinant prion infectivity. J Virol 86, 1874–1876, doi: 10.1128/JVI.06216-11 (2012).
Polymenidou, M. et al. The POM monoclonals: a comprehensive set of antibodies to non-overlapping prion protein epitopes. PLoS One 3, e3872, doi: 10.1371/journal.pone.0003872 (2008).
Acknowledgements
Authors would like to thank Drs Kuntal Pal, Karsten Melcher, and Eric Xu at Van Andel Research Institute for advices on the thermal stability assay and access to the fluorescence spectrophotometer, Dr. Charles Weissmann at Scripps Florida for the CAD5 cell line, and David Nadziejka at Van Andel Research Institute for editing the manuscript. This work was supported by NIH grants R01NS060729, R01NS071035, and P01AI106705.
Author information
Authors and Affiliations
Contributions
J.M. designed the study, analyzed data, and wrote the main manuscript text; R.A. generated recPrP variants, performed PMCA, amyloid fibril growth, and thermal stability analyses, and analyzed data; F.W. initiated the study of PrP variants within the threonine string, performed PMCA and Elispot assay, and analyzed data; K.J.V.S., K.S. performed AFM analyses of amyloid fibrils and analyzed data. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Abskharon, R., Wang, F., Vander Stel, K. et al. The role of the unusual threonine string in the conversion of prion protein. Sci Rep 6, 38877 (2016). https://doi.org/10.1038/srep38877
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep38877
This article is cited by
-
Prion protein amino acid sequence influences formation of authentic synthetic PrPSc
Scientific Reports (2023)
-
In Vitro Seeding Activity of Glycoform-Deficient Prions from Variably Protease-Sensitive Prionopathy and Familial CJD Associated with PrPV180I Mutation
Molecular Neurobiology (2019)
-
Prion infectivity is encoded exclusively within the structure of proteinase K-resistant fragments of synthetically generated recombinant PrPSc
Acta Neuropathologica Communications (2018)
-
Soluble polymorphic bank vole prion proteins induced by co-expression of quiescin sulfhydryl oxidase in E. coli and their aggregation behaviors
Microbial Cell Factories (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.