Abstract
Brown planthopper (BPH) is the most destructive pest of rice in Asia. To date 29 BPH resistance genes have been identified, but only a few genes are being used in breeding due to inefficient markers for marker-assisted selection (MAS) and little knowledge of the real effects of the genes. In this study we individually transferred 13 genes or QTLs (Bph14, QBph3, QBph4, Bph17, Bph15, Bph20, Bph24, Bph6, Bph3, Bph9, Bph10, Bph18 and Bph21) into cultivar 9311 by marker assisted backcross breeding (MABB). Through positive and negative selection we narrowed the segments from donors containing Bph14, Bph15, Bph6 and Bph9 to 100–400 kb. Whole-genome background selection based on a high resolution SNP array was performed to maximize reconstitution of the recurrent parent genome (RPG 99.2–99.9%). All genes reduced BPH growth and development and showed antibiotic responses in seedlings. Based on genetic effects and amino acid sequences of genes in three clusters we inferred that Bph10 and Bph21 might be identical to Bph26, whereas Bph9 and Bph18 were different. Bph15 might be same with Bph17, but QBph4, Bph20 and Bph24 might be different. We believe that these NILs will be useful in rice BPH resistance research and breeding.
Similar content being viewed by others
Introduction
Brown planthopper (BPH; Nilaparvata lugens Stål), a monophagous sucking insect, is the most damaging insect pest of rice in Asia1,2. After BPH feeding at the tiller base host plants generally appear yellow and withered3, with heavy infestation causing “hopperburn”4,5. BPH also transmits viral pathogens such as grassy stunt virus (RGSV) and ragged stunt virus (RRSV)6. Conventional methods of controlling BPH depend on insecticides that are costly in terms of labor, money, and the environment. Furthermore, overuse of insecticides may affect populations of natural enemies of BPH and lead to resistance/tolerance to insecticides, and resurgence of the BPH problem7. Therefore, breeding for host resistance is considered the most economical and environmentally friendly strategy for BPH control8.
At least 29 resistance genes have been identified in Oryza sativa spp. indica and wild rice species9. Among them, 14 genes have been fine-mapped to specific regions on chromosomes 3, 4, 6 and 1210 and five genes were cloned. Cloned genes Bph14, Bph18 and Bph26 (or bph2) encode coiled-coil, nucleotide-binding, leucine-rich repeat (CC-NB-LRR) proteins, whereas Bph17 consists of three clustered genes encoding lectin-receptor kinases, and Bph29 encodes a B3 DNA-binding domain8,11,12,13,14.
Molecular marker-assisted backcross breeding (MABB) has greatly improved the efficiency and effectiveness of rice breeding. There are three aspects of MABB, namely, positive selection for the target gene using linked markers, negative selection of alleles from the donor parent surrounding the target gene, and background selection for the maximum recovery of the recurrent parent genome using polymorphic markers covering the whole genome15,16. MABB has been widely used in rice breeding for disease and insect resistance16,17,18. Likewise, MABB has been used to develop multiple BPH-resistant introgression lines (ILs) and near-isogenic lines (NILs). Linkage drag between Bph3 and Wxa alleles was successfully broken by negative selection allowing development of ILs with broad-spectrum resistance to BPH and good quality19. By using marker 7312. T4A that co-segregated with Bph18 in positive selection, and 260 SSR markers for background selection a group of ILs carrying Bph18 were developed20. However, only a few genes have been exploited successfully in breeding for BPH resistance due to inefficient markers and inadequate knowledge of the actual effects of the resistance genes.
In our study, 13 BPH resistance genes from nine donors [RH (Bph3 and Bph17), B5 (Bph14 and Bph15), IR54751-1-2-44 (QBph3 and QBph4), IR65482-4-136 (Bph10), IR71033-121 (Bph20 and Bph21), IR65482-7-216 (Bph18), Swarnalata (Bph6), Pokkali (Bph9) and BR96 (Bph24)]11,21,22,23,24,25,26,27,28,29 were individually incorporated into cultivar (cv.) 9311 (an elite restorer parent for hybrids in China) by MABB, and 13 monogenic NILs were developed with enhanced BPH resistance. Some NILs carry chromosomal fragments from the donor parents of less than 100 kb, and have reconstituted recurrent parent genomes (RPG) exceeding 99.5% as a result of using a breeding chip with high-density SNP markers for negative and background selection. The conserved amino acid sequences of gene clusters on chromosomes 3, 4 and 12 were compared with those of Bph14, Bph17 and Bph26, respectively.
Results
Development of monogenic NILs
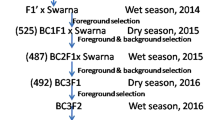
To develop monogenic NILs, 13 BPH resistance genes or QTLs, identified in nine donor accessions were individually incorporated into 9311 by MABB (Fig. 1). The entire scheme took 9 crosses, 4 generations of backcrossing and one generation of selfing. In each backcross generation, individuals heterozygous at the target locus were further backcrossed to the recurrent parent 9311.
Positive and negative selection
Among the 13 genes, four (Bph14, Bph15, Bph6 and Bph9) were transferred by positive and negative selection (Fig. 2). Taking Bph6 as an example, closely linked markers Y37 and RM17008 were used for positive selection during the introgression process. Two other markers, J6-7 and J6-10, approximately 28 kb upstream of Y37 and 32 kb downstream of RM17008, respectively, were used in negative selection. Among 2,000 BC1F2 progenies (segregating at the Bph6 locus), by selection on one side of the Bph6 locus, one plant heterozygous in the region near the Bph6 allele and homozygous for the 9311 region at the J6-7 locus, was selected and then backcrossed to 9311 to produce the BC2F1. Then five selected plants heterozygous at Bph6 were selfed to produce a BC2F2 population. Among 3,000 BC2F2 progenies, two plants heterozygous for Bph6, and homozygous for the 9311 allele at the J6-10 locus, were selected. Finally, the linked segments around Bph14, Bph15, Bph6 and Bph9 were narrowed to less than 100 kb, 400 kb, 100 kb and 200 kb, respectively (Fig. 2).
For the remaining genes, flanking marker pairs closely linked to the target genes were used in positive selection, including XC4-9/IN15-6 for Bph17, RM586/RM589 for Bph3, XC3-14/IN76-2 for QBph3, XY4-17/XC4-27 for QBph4, XY4-17/HJ28 for Bph20, HJ12/HJ9 for Bph21, XC12-2/HJ12 for Bph10, HJ12/XC18-7 for Bph18, and J22/16717 for Bph24 (Supplementary Table S1).
Background selection
The selected recombinants B14-2, B15-2, B6-3 and B9-3 heterozygous at the Bph14, Bph15, Bph6 and Bph9 loci, respectively, were each backcrossed to 9311 to produce advanced backcross (BC) generation materials. In the BC3 to BC4 generations, the 6 K SNP chip30 was used to select individuals with the highest RPG. Selected individuals were self-pollinated to produce the BC4F2 generations. As shown in Fig. 3 individuals B14, B15, B6 and B9 not only had small donor segments linked to the target genes, but also carried few additional donor segments on other chromosomes. RPG recoveries were 99.7, 99.2, 99.8 and 99.9% in the monogenic NILs carrying Bph14, Bph15, Bph6 and Bph9, respectively (Table 1). The other 9 genes were likewise, transferred by MABB using positive selection and background selection; RPG recoveries were 98.5, 92.9, 93.2, 98.5, 95.3, 98.4, 98.8, 97.8 and 97.5% in the NILs harboring Bph17, Bph3, QBph3, QBph4, Bph10, Bph18, Bph20, Bph21 and Bph24, respectively (Supplementary Fig. S1, Table 1). Finally, 13 monogenic NILs were selected and designated as Bph14-NIL, QBph3-NIL, QBph4-NIL, Bph17-NIL, Bph15-NIL, Bph20-NIL, Bph24-NIL, Bph6-NIL, Bph3-NIL, Bph9-NIL, Bph10-NIL, Bph18-NIL and Bph21-NIL.
Agronomic performance of NILs and their hybrids
A number of agronomic traits were measured, including days to heading (DTH), plant height (PH), panicle number (PN), number of grains (NG), number of grains per panicle (NPG), spikelet fertility (SF), 1000-grain weight (GW), and yield per plant (YD). Bph3-NIL and Bph21-NIL showed significant decreases in GW compared to 9311, QBph3-NIL had significantly higher SF and GW, resulting in increased yield (Table 1). The improved hybrids H2613S/QBph4-NIL and H2613S/Bph15-NIL had significantly higher SF compared to the conventional hybrid H2613S/9311. H2613S/Bph24-NIL showed significantly higher NG and NPG, but lower GW, leading to an equal yield compared to the conventional hybrid (Table 2). However, for most traits comparisons, there were no significant differences between the NILs and 9311, improved hybrids and conventional hybrids (Tables 1 and 2).
Seedling response
After about 14 days of BPH infestation in the greenhouse the 9311 control showed 100% wilting whereas the NILs were surviving. In short, the NILs harboring single genes or QTL on chromosome 4 (QBph4, Bph17, Bph15, Bph20, Bph24 and Bph6) had higher resistance than those on chromosome 12 (Bph10, Bph18 and Bph21), except for Bph9 (Fig. 4A). The Bph24-NIL had the lowest response score (1.3), representing the highest level of seedling resistance among the 13 NILs. To explore the potential usefulness of these genes in hybrids, 13 monogenic hybrids between H2613S and the NILs were also evaluated. The hybrid response closely paralleled those of corresponding NILs (Fig. 4B). Hybrids carrying Bph14, QBph4, Bph17, Bph6, Bph3, Bph9 and Bph10 showed lower levels of resistance than the corresponding NILs, indicating incomplete dominance of these genes. However, there were no significant differences between the response scores of the remaining six monogenic NILs and their hybrids, indicating complete dominance of those genes (Fig. 4C).
Seedling responses of NILs and their hybrids.
(A and B) Response scores of NILs and their hybrids, respectively. (C) Comparison of the responses of NILs and corresponding hybrids. (D) Representative images of Bph14-NIL, Bph6-NIL, Bph9-NIL and Bph17-NIL damaged by BPH infestation. The sample sizes of each NIL and hybrid were 48 and 24, respectively. Uppercase letters above the error bars indicate significantly differences in ranking by Duncan’s test at P < 0.01. Error bars, SEM. * and ** significantly different at P < 0.05 and P < 0.01, respectively.
Honeydew excretion and survival rate of BPH on NILs
To determine whether the presence of resistance genes affected BPH growth and development, we compared the areas of honeydew excretion for BPH feeding on each NIL. The results were in accordance with the seedling responses (Fig. 5A,B), suggesting that honeydew deposition is a simple measurable indicator of BPH fitness on the lines. The excretion areas were grouped into four classes: the smallest, QBph3, Bph9, Bph15 and Bph20 (approximate area 10 mm2); small, QBph4, Bph3, Bph6, Bph14, Bph17 and Bph24 (20 mm2); higher, Bph10, Bph18, and Bph21 (50 mm2); the highest, 9311 (130 mm2) (Fig. 5A).
To test whether antibiosis was a factor in BPH resistance, BPH survival rates were measured on the monogenic NILs every day for 9 days after infestation (DAI). Mean BPH survival rates were lowest on Bph15-NIL, Bph17-NIL, Bph20-NIL and Bph14-NIL, and decreased more rapidly over the 9 days than on the other NILs (survival rate at 9 DAI: 28–43%) (Fig. 5C). Survival rates decreased less quickly on Bph6-NIL, Bph9-NIL, Bph3-NIL, QBph3-NIL and Bph10-NIL (45–55% at 9 DAI), and least slowly on Bph18-NIL, QBph4-NIL, Bph21-NIL and Bph24-NIL (65–75% at 9 DAI). The average survival rate on the control (9311) showed the slowest reduction (83% of the BPHs were alive at 9 DAI).
BPH survival rates on the NILs mostly paralleled the data for honeydew accumulation and seedling response. The lower effectiveness of the NILs with QBph4 and Bph24 compared to those with Bph17, Bph15 and Bph20 suggests these two genes might have different mechanisms of resistance.
Sequence comparison of genes in three clusters
Genes for BPH resistance in rice were reported to cluster on chromosomes 3, 4 and 12, among which Bph14 on chromosome 3, Bph17 on chromosome 4, Bph29 on chromosome 6, and Bph26 and Bph18 on chromosome 12 have been cloned8,11,12,13,14. To explore whether the genes in the clusters surrounding Bph14, Bph26 and Bph17 were the same, we sequenced and compared the amino acid sequence of the Bph26 alleles from Bph9-NIL, Bph10-NIL and Bph21-NIL, the Bph14 allele from the QBph3-NIL and Bph14-NIL, and the Bph17 allele from the QBph4-NIL, Bph15-NIL, Bph17-NIL, Bph20-NIL and Bph24-NIL. The proteins encoded by Bph26 alleles from the monogenic NILs carrying Bph10 and Bph21 were identical in amino acid sequence as the cloned Bph26. Bph18 and Bph26 are different alleles with many amino acid substitutions12. However, due to inability to obtain the PCR product of the third exon of the Bph26 allele from the Bph9-NIL using six pairs of specific primers, only the first and second exons were compared and a few nucleotide polymorphisms causing amino acid substitutions were detected (Fig. 6A). Compared to Bph14, the Bph14 allele from QBph3-NIL had a number of amino acid substitutions in the LRR domain (Fig. 6B). Compared to the amino acid sequence of the cloned Bph17, that from Bph15-NIL was the same, while that from NILs of QBph4, Bph20 and Bph24 showed several substitutions (Fig. 6C).
Sequence comparison of conserved amino acid sequences of cloned genes.
(A) Comparison of the genes/QTLs on chromosome12 with Bph26. The gene sequence of Bph18 was reported in Ji et al.12. (B) Comparison of QBph3 and Bph14. (C) Comparison of the genes/QTLs on chromosome 4 with Bph17. The sequence information for 9311 is from the GenBank database (http://rise.genomics.org.cn/rice/index2.jsp).
Discussion
To achieve improvement in target traits by MABB, breeders aim to minimize the introgressed segments from donors in order to reduce linkage drag, and to maximize reconstitution of recurrent parent genomes. Previously, improved lines contained large fragments (>1,000 kb) of the target gene regions, due to lack of suitable closely linked molecular markers and limited knowledge of the actual chromosomal locations of the resistance genes14. To reduce linkage drag, we performed positive and negative selection to obtain resistant recombinants between flanking markers in target regions based on high resolution physical maps of BPH resistance genes in two large backcross populations (BC1F2 and BC2F2). Introgressed segments containing four genes (Bph14, Bph15, Bph6 and Bph9) were finally narrowed to less than 400 kb (Fig. 2). In contrast, the introgressed segments for the remaining nine genes exceeded 1,000 kb as negative selection was not employed (Supplementary Fig. S1).
In previous MABB programs, background selection was based on RFLP and SSR markers which had low resolution and inadequate whole genome coverage15,21. With development of next generation sequencing technology, large numbers of SNPs became available, and two breeding chips RICE6K and RiceSNP50 with high-throughput SNP arrays were developed in China30,31, making efficient whole-genome background selection a reality. Background selection with high resolution SNP markers was used in rice breeding to improve blast resistance and wide compatibility16,32. In the present study, the breeding chip RICE6K was employed in background selection for improving BPH resistance. Undesirable donor segments in each NIL could be viewed on the haplotype map produced by the chip. For example, after backcrossing and background selection, Bph6-NIL and Bph9-NIL only had four and three short segments from donors, and the RPG recoveries were 99.8 and 99.9%, respectively (Fig. 3). Our work demonstrated that positive and negative selection of target loci and whole genome background selection based on the 6 K SNP chip was a powerful way of developing monogenic NILs.
To date, eight BPH resistance genes, Bph1, bph2, Bph3, Bph6, Bph14, Bph15, Bph18 and Bph27 (t) have been individually incorporated into indica or japonica varieties by MABB18,19,20,21,33,34,35. These NILs or ILs carrying single resistance genes have been reported as resistant to one or more BPH biotypes predominating in various countries. However, the real effects of these genes cannot be compared using the original source genotypes due to the diverse genetic backgrounds in which additional resistance QTLs may be present. In the present study, 13 BPH resistance genes were separately introduced into cv. 9311 by MABB. Generally, the NILs carrying QBph4, Bph15, Bph17, Bph20, Bph24 and Bph6 on chromosome 4 showed significantly higher resistance than those carrying Bph10, Bph18 and Bph21 on chromosome 12, indicating that the gene cluster on chromosome 4 was more effective in conferring BPH resistance. The response scores of most NILs consistently matched seedling response, honeydew deposition area, and BPH survival rate scores, indicating a similar resistance mechanism. However, the Bph24-NIL showed the highest level of seedling resistance and the least honeydew excretion, but a higher BPH survival rate, indicated that Bph24 might mediate a resistance mechanism than antibiosis (Figs 4 and 5). However, the RPG coverage differs among the 13 NILs and this may have some effect on the agronomic performance and BPH responses (Table 1 and Supplementary Fig. S1). Previously, Hu et al.18 proved that improved hybrid rice containing Bph14 and Bph15 showed enhanced resistance compared to a conventional line. In our study, hybrid F1 descendants of H2613S and NILs showed higher resistance than conventional hybrid rice that lacked resistance genes. Bph14, QBph4, Bph17, Bph6, Bph3, Bph9 and Bph10 showed significantly less resistance in hybrids than corresponding NILs, indicating incomplete dominance (Fig. 4).
It is noteworthy that multiple BPH resistance genes cluster together on rice chromosomes; eight genes (Bph1, bph2, Bph9, Bph10, Bph18, Bph19, Bph21 and Bph26) are clustered on chromosome 12 L, and six (QBph4, Bph15, Bph12, Bph17, Bph20 and Bph24) on chromosome 4S36. These gene clusters might involve different genes, different alleles at a single locus, or even the same gene with different haplotypes21. Based on response phenotypes of the NILs and comparisons of amino acid sequence with the cloned Bph26, Bph14 and Bph17, some aspects were resolved. The amino acid sequences of Bph10 and Bph21 were identical to Bph26 and the response phenotypes of the two monogenic NILs were similar to that of Bph26-NIL, whereas Bph9 and Bph18 were different (Figs 4, 5 and 6). We inferred that Bph10, Bph21 and Bph26 might be the same gene, but Bph9 and Bph18 were likely different alleles in this locus. QBph3 has a number of amino acid substitutions compared with Bph14. Hu et al.22 reported that QBph3 and Bph14 were tightly linked on chromosome 3 L, but the QBph3-NIL showed a higher degree of resistance than Bph14-NIL. Thus, they might be alleles or linked genes that mediate different resistance mechanisms. Bph15 shared the same amino acid sequence as Bph17, and the Bph15-NIL had a similar BPH response phenotype to the Bph17-NIL with resistance scores of 2.4 versus 2.7, honeydew deposition area of 11.8 vs 18.1, and survival rates 33% vs 38%. However, there were differences between Bph17 and the alleles from the QBph4, Bph20 and Bph24 NILs in amino acid sequence and response phenotype. These results indicated that Bph15 was likely to be identical to Bph17, whereas QBph4, Bph20 and Bph24 might be different. Lv et al.35 and Hu et al.22 reported that Bph15 and QBph4 were located proximally to Bph17, thus differing from the present findings. One possible reason is that besides Bph17 there is another resistance gene/QTL in the donor B5.
Cv. 9311 is an elite restorer line for two-line hybrid rice and HL CMS three-line hybrid rice because of its good adaptation, ideal plant type, good grain quality and high yield potential. However, due to the absence of disease (bacterial blight, blast) and insect (stem borer, BPH) resistance, the commercial application of 9311 is limited. Our 9311 NILs with high BPH resistance will provide a choice of parent lines for use in producing hybrid rice.
Generally pyramiding major resistance genes would enhance the resistance level of rice plant. However, whether the pyramided resistances will also improve the durability of BPH resistance is still unknown37. Furthermore, gene pyramiding might increase the risk of linkage drag, and higher resistance levels might lead to stronger selection pressure on BPH, thus accelerating evolution of the pest and resulting in failure of the resistance genes. In order to improve the durability of BPH resistance, 10 BPH resistance genes were transferred into 9311 individually and the corresponding multiline hybrid combinations developed in this study could be a possible strategy to prolong their effectiveness. Moreover, since these genes are from different indica cultivars and wild relatives with different resistance mechanisms, they may suppress the development of a harmful dominant race.
Methods
Plant materials and insects
Nine donor parents (DPs) were used to develop a set of monogenic NILs carrying genes for BPH resistance in the genetic background of cultivar (cv.) 9311. The DPs [RH (Bph3 and Bph17), B5 (Bph14 and Bph15), IR54751-1-2-44 (QBph3 and QBph4), IR65482-4-136 (Bph10), IR71033-121 (Bph20 and Bph21), IR65482-7-216 (Bph18), Swarnalata (Bph6), Pokkali (Bph9) and BR96 (Bph24)] were obtained from IRRI, Wuhan University and Guangxi Academy of Agricultural Sciences. Hua 2613S (H2613S) is an improved two-line thermo-sensitive genic male sterile line with blast resistance gene R6 added by molecular marker assisted selection to the genetic background of Guangzhan 63S (data not shown). Both cv. 9311 and Guangzhan 63S are leading male and female parents for a number of commonly used two line hybrids in China, including Yangliangyou 6, the most widely cultivated hybrid in central and southern China during the last five years.
The BPH insects (predominantly biotype-2) used for infestation were collected from rice fields in Wuhan, and maintained on TN1 plants under greenhouse conditions at Huazhong Agricultural University.
DNA extraction and marker analysis
Genomic DNA was extracted from fresh seedling leaves by the CTAB method38 with minor modifications. The PCR protocol was as follows: 2 μl (20 ng/μl) DNA in an 8 μl reaction mixture [2.0 μl 10 × buffer, 1.6 μl dNTP (2 mM), 1.4 μl MgCl2 (25 mM), 0.4 μl each primer (50 ng/μl), 2 μl ddH2O, and 0.2 μl Taq E (5 U/μl)], 10 μl ddH2O, and 20 μl mineral oil. The cycling regime was: 94 °C for 4 min, followed by 32 cycles of 94 °C/30 s, 55 °C/30 s, and 72 °C/40 s, and a final extension at 72 °C for 7 min. PCR products were separated on 4% non-denaturing polyacrylamide gels and detected by silver staining. SSR sequences were identified in Gramene (www.gramene.org), and Indel markers were designed based on sequence alignments of the Nipponbare and 9311 reference genomes in Rice Var Map (http://ricevarmap.ncpgr.cn/) (Table 1). For genome-wide genotyping based on the RICE6K SNP array, DNA amplification, fragmentation, chip hybridization, single base extension, staining and scanning were conducted by the Life Science and Technology Center, China National Seed Co. LTD, Wuhan, China30.
Markers used for positive and negative selection
Positive selection for the presence of genes Bph17, Bph3, QBph3, QBph4, Bph6, Bph9, Bph10, Bph14, Bph15, Bph18, Bph20, Bph21 and Bph24 was conducted using gene-linked marker pairs XC4-9/IN15-6, RM586/RM589, C3-14/IN76-2, XY4-17/XC4-27, Y37/RM17008, J18-7/HJ12, XC12-2/HJ12, C3-14/IN76-2, RM261/HJ16, HJ12/J18-7, XY4-17/HJ28, HJ12/HJ9, XC12-2/HJ12 and HJ22/RM16717, respectively. Flanking marker pairs for negative selection were J64/J6-7/6-10 (Bph6), RM570/J14-8/J14-12 (Bph14), IN15-6/J23/MS5 (Bph15) and J18-15/RM3331/HJ9 (Bph9) (Supplemental Table S1).
Collection of agronomic trait data from field experiments
The 26 NILs and hybrids containing the BPH resistance genes were planted in a randomized complete block design at Wuhan in the summer of 2015. Plots of each line consisted of two rows with 10 plants per row at a planting density of 17 cm between plants in a row and 27 cm between rows. The central eight plants from each plot, were used to measure agronomic traits including plant height (PH), days to heading (DTH), panicle number (PN), number of grains (NG), number of grains per panicle (NPG), spikelet fertility (SF), 1,000 grain weight (GW), and yield per plant (YD). There were three replications for each NIL and hybrid combination.
Seedling response assays
The BPH bioassay was performed as a modified bulk seedling test in the greenhouse, following the method of Pathak et al.39. Seeds of 9311 and test lines were sown as random groups in 50 cm × 30 cm × 10 cm (height) plastic trays. Seedlings were thinned to 12 plants per line at the three-leaf stage and infested with second and third instar nymphs at a density of 15 insects per seedling. When all 9311 seedlings (control) had died [10–12 days after infestation (DAI)] the plants in other lines were examined, and each seedling was given a score of 1, 3, 5, 7 or 9 according to the criteria described by Huang et al.40 where higher scores indicate greater susceptibility to BPH. These tests were performed in three replications.
Honeydew area
Determination of areas of honeydew deposition followed the method of Du et al.11. Thirty-day-old NILs and 9311 (control) were covered by inverted transparent plastic cups placed over a filter paper resting on plastic Petri dishes. After starving for 2 h, five fifth instar BPH nymphs were placed in each cup. Two days later, the filter papers were oven dried for 30 min at 60 °C and treated with 0.1% solution of ninhydrin in acetone. Areas of ninhydrin-positive deposits were measured using Image J software. Tests were conducted in eight replications.
BPH survival rates
Survival rates were calculated following the method of Du et al.11. To determine nymph survival rates on rice lines, each plant was infested with 20 second and third instar nymphs and covered with a cylindrical plastic cup. Survival rates calculated as percentages of surviving nymphs divided by the total number of nymphs released at the beginning were recorded daily for 9 days.
Statistical analysis
Mean phenotypic values of plants were compared using one-way ANOVA. Duncan’s multiple range and t tests were used for multiple mean comparisons. All statistical analyses were conducted using SPSS 7 for Windows version 16.0 (SPSS Inc., USA).
Additional Information
How to cite this article: Xiao, C. et al. Development and evaluation of near-isogenic lines for brown planthopper resistance in rice cv. 9311. Sci. Rep. 6, 38159; doi: 10.1038/srep38159 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Nasu, S. Rice leafhoppers. In The Major Insect Pests of the Rice Plant. 493–523 (Johns Hopkins Press, Baltimore, Maryland, 1964).
Jena, K. K. & Kim, S. M. Current status of brown planthopper (BPH) resistance and genetics. Rice 3, 161–171 (2010).
Watanabe, T. & Kitagawa, H. Photosynthesis and translocation of assimilates in rice plants following phloem feeding by the planthopper Nilaparvata lugens (Homoptera: Delphacidae). J. Econ. Entomol. 93, 1192–1198 (2000).
Sogawa, K. The rice brown planthopper: feeding physiology and host plant interactions. Ann. Rev. Entomol. 27, 49–73 (1982).
Matsumura, M. et al. Current status of insecticide resistance in rice planthoppers in Asia. In Planthoppers: New threats to the sustainability of intensive rice production systems in Asia. 233–244 (International Rice Research Institute, Manila, The Philippines, 2009).
Dyck, V. A. & Thomas, B. The brown planthopper problem. In Brown planthopper: Threat to Rice Production in Asia. 3–17 (International Rice Research institute, Manila, The Philippines, 1979).
Heinrichs, E. & Mochida, O. From secondary to major pest status: the case of insecticide-induced rice brown planthopper, Nilaparvata lugens, resurgence. Protection Ecol. 7, 201–218 (1984).
Wang, Y. et al. Map-based cloning and characterization of BPH29, a B3 domain-containing recessive gene conferring brown planthopper resistance in rice. J. Exp. Bot. 66, 6035–6045 (2015).
Ling, Y. & Weilin, Z. Genetic and biochemical mechanisms of rice resistance to planthopper. Plant Cell Rep. 35, 1559–1572 (2016).
Hu, J. et al. A new finely mapped Oryza australiensis-derived QTL in rice confers resistance to brown planthopper. Gene 561, 132–137 (2015).
Du, B. et al. Identification and characterization of Bph14, a gene conferring resistance to brown planthopper in rice. Proc. Natl. Acad. Sci. USA 106, 22163–22168 (2009).
Ji, H. et al. Map-based cloning and characterization of the BPH18 gene from wild rice conferring resistance to brown planthopper (BPH) insect pest. Sci. Rep. 6, 3476 (2016).
Tamura, Y. et al. Map-based cloning and characterization of a brown planthopper resistance gene BPH26 from Oryza sativa L. ssp. indica cultivar ADR52. Sci. Rep. 4, 5872 (2014).
Liu, Y. et al. A gene cluster encoding lectin receptor kinases confers broad-spectrum and durable insect resistance in rice. Nat. Biotechnol. 33, 301–305 (2015).
Chen, S., Lin, X., Xu, C. & Zhang, Q. Improvement of bacterial blight resistance of ‘Minghui 63′, an elite restorer line of hybrid rice, by molecular marker-assisted selection. Crop Sci. 40, 239–244 (2000).
Khanna, A. et al. Development and evaluation of near-isogenic lines for major blast resistance gene(s) in Basmati rice. Theor. Appl. Genet. 128, 1243–1259 (2015).
Jiang, H. et al. Improving blast resistance of Jin 23B and its hybrid rice by marker-assisted gene pyramiding. Mol. Breeding 30, 1679–1688 (2012).
Hu, J. et al. Pyramiding and evaluation of three dominant brown planthopper resistance genes in the elite indica rice 9311 and its hybrids. Pest Manag. Sci. 69, 802–808 (2013).
Jairin, J. et al. Development of rice introgression lines with brown planthopper resistance and KDML105 grain quality characteristics through marker-assisted selection. Field Crops Res. 110, 263–271 (2009).
Suh, J. P. et al. Development of elite breeding lines conferring Bph18 gene-derived resistance to brown planthopper (BPH) by marker-assisted selection and genome-wide background analysis in japonica rice (Oryza sativa L.). Field Crops Res. 120, 215–222 (2011).
Qiu, Y., Guo, J., Jing, S., Zhu, L. & He, G. High-resolution mapping of the brown planthopper resistance gene Bph6 in rice and characterizing its resistance in the 9311 and Nipponbare near isogenic backgrounds. Theor. Appl. Genet. 121, 1601–1611 (2010).
Hu, J. et al. Fine mapping and pyramiding of brown planthopper resistance genes QBph3 and QBph4 in an introgression line from wild rice O. officinalis. Mol. Breeding 35, 3, 10.1007/s11032-015-0228-2 (2015).
Jairin, J., Phengrat, K., Teangdeerith, S., Vanavichit, A. & Toojinda, T. Mapping of a broad-spectrum brown planthopper resistance gene, Bph3, on rice chromosome 6. Mol. Breeding 19, 35–44 (2006).
Sun, L., Su, C., Wang, C., Zhai, H. & Wan, J. Mapping of a major resistance gene to the brown planthopper in the rice cultivar Rathu Heenati. Breeding Sci. 55, 391–396 (2005).
Yang, H. et al. High-resolution genetic mapping at the Bph15 locus for brown planthopper resistance in rice (Oryza sativa L.). Theor. Appl. Genet. 110, 182–191 (2004).
Ishii, T., Brar, D. S., Multani, D. S. & Khush, G. S. Molecular tagging of genes for brown planthopper resistance and earliness introgressed from Oryza australiensis into cultivated rice, O. sativa. Genome, 37, 217–221 (1994).
Rahman, M. L. et al. High-resolution mapping of two rice brown planthopper resistance genes, Bph20(t) and Bph21(t), originating from Oryza minuta. Theor. Appl. Genet. 119, 1237–1246 (2009).
Jena, K. K., Jeung, J. U., Lee, J. H., Choi, H. C. & Brar, D. S. High-resolution mapping of a new brown planthopper (BPH) resistance gene, Bph18(t), and marker-assisted selection for BPH resistance in rice (Oryza sativa L.). Theor. Appl. Genet. 112, 288–297 (2006).
Murata, K. et al. Mapping of a brown planthopper (Nilaparvata lugens Stål) resistance gene Bph9 on the long arm of rice chromosome 12. Cereal Res. Commun. 29, 245–250 (2001).
Yu, H., Xie, W., Li, J., Zhou, F. & Zhang, Q. A whole-genome SNP array (RICE6K) for genomic breeding in rice. Plant Biotechnol. J. 12, 28–37 (2014).
Chen, H. et al. A high-density SNP genotyping array for rice biology and molecular breeding. Mol. Plant 7, 541–553 (2014).
Mi, J. et al. Stacking S5-n and f5-n to overcome sterility in indica-japonica hybrid rice. Theor. Appl. Genet. 129, 563–575 (2016).
Sharma, P., Torii, A., Takumi, S., Mori, N. & Nakamura, C. Marker-assisted pyramiding of brown planthopper (Nilaparvata lugens Stål) resistance genes Bph1 and Bph2 on rice chromosome 12. Hereditas 140, 61–69 (2004).
Liu, Y. et al. Marker assisted pyramiding of two brown planthopper resistance genes, Bph3 and Bph27(t), into elite rice cultivars. Rice 9, 27, 10.1186/s12284-016-0096-3 (2016).
Lv, W. et al. BAC and RNA sequencing reveal the brown planthopper resistance gene BPH15 in a recombination cold spot that mediates a unique defense mechanism. BMC Genomics 15, 674 (2014).
Hu, J., Xiao, C. & He, Y. Recent progress on the genetics and molecular breeding of brown planthopper resistance in rice. Rice 9, 30, 10.1186/s12284-016-0099-0 (2016).
Qiu, Y., Guo, J., Jing, S., Zhu, L. & He, G. Development and characterization of japonica rice lines carrying the brown planthopper-resistance genes BPH12 and BPH6. Theor. Appl. Genet. 124, 485–494 (2012).
Murray, M. G. & Thompson, W. F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8, 4321–4325 (1980).
Pathak, M., Cheng, C. & Fortuno, M. Resistance to Nephotettix impicticeps and Nilaparvata lugens in varieties of rice. Nature 223, 502–504 (1969).
Huang, Z., He, G., Shu, L., Li, X. & Zhang, Q. Identification and mapping of two brown planthopper resistance genes in rice. Theor. Appl. Genet. 102, 929–934 (2001).
Acknowledgements
We are very grateful to Professor D.S. Brar, IRRI, for providing seeds, Professor Rongbai Li, Guangxi University, for providing seeds carrying Bph24, and Professor Hongxia Hua and students for technical assistance in the BPH bioassays. This work was supported in part by grants from the National High Technology Research (2014AA10A604), the National Program on Research and Development of Transgenic Plants (2016ZX08001002-002), the National Natural Science Foundation (31371701) and the Foundation of the Ministry of Agriculture of China (CARS-01-03).
Author information
Authors and Affiliations
Contributions
C.X., J.H., Y.T.A., M.X.C., G.J.G. and Q.L.Z. performed the experiments. C.X. and J.H. analyzed the data. Y.Q.H. conceived and designed the experiments. G.C.H. provided seeds of some resistant donors. C.X. and Y.Q.H. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Xiao, C., Hu, J., Ao, YT. et al. Development and evaluation of near-isogenic lines for brown planthopper resistance in rice cv. 9311. Sci Rep 6, 38159 (2016). https://doi.org/10.1038/srep38159
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep38159
This article is cited by
-
Dissection and validation of quantitative trait loci (QTLs) conferring grain size and grain weight in rice
Euphytica (2024)
-
Effect of nitrogen fertilizer on the resistance of rice near-isogenic lines with BPH resistance genes
Botanical Studies (2022)
-
Development of InDel markers of Bph3 and pyramiding of four brown planthopper resistance genes into an elite rice variety
Molecular Breeding (2020)
-
Genomic Breeding of Green Super Rice Varieties and Their Deployment in Asia and Africa
Theoretical and Applied Genetics (2020)
-
Current understanding of the genomic, genetic, and molecular control of insect resistance in rice
Molecular Breeding (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.