Abstract
In many marine invertebrates, larval metamorphosis is induced by environmental cues that activate sensory receptors and signalling pathways. Nitric oxide (NO) is a gaseous signalling molecule that regulates metamorphosis in diverse bilaterians. In most cases NO inhibits or represses this process, although it functions as an activator in some species. Here we demonstrate that NO positively regulates metamorphosis in the poriferan Amphimedon queenslandica. High rates of A. queenslandica metamorphosis normally induced by a coralline alga are inhibited by an inhibitor of nitric oxide synthase (NOS) and by a NO scavenger. Consistent with this, an artificial donor of NO induces metamorphosis even in the absence of the alga. Inhibition of the ERK signalling pathway prevents metamorphosis in concert with, or downstream of, NO signalling; a NO donor cannot override the ERK inhibitor. NOS gene expression is activated late in embryogenesis and in larvae, and is enriched in specific epithelial and subepithelial cell types, including a putative sensory cell, the globular cell; DAF-FM staining supports these cells being primary sources of NO. Together, these results are consistent with NO playing an activating role in induction of A. queenslandica metamorphosis, evidence of its highly conserved regulatory role in metamorphosis throughout the Metazoa.
Similar content being viewed by others
Introduction
Most marine invertebrate phyla include species with a biphasic life cycle comprised of a small, planktonic larval form that settles onto the benthos and metamorphoses into a morphologically distinct juvenile1,2,3. Settlement and metamorphosis typically requires that the larva first obtains “competence” to be able to detect and respond to a local inductive environmental cue4,5. Acquisition of competence may require that the larva first encounters a broad-scale environmental condition indicative of a generally appropriate habitat6,7. A competent larva in the presence of an appropriate local inductive cue will generally settle and initiate metamorphosis. In eumetazoan marine invertebrate larvae, this is achieved with the aid of neurosensory systems that are finely tuned to detect precise locations most likely to ensure successful postlarval development, juvenile growth and ultimately adult reproduction5,8. Typically, activation of neurosensory cells by a specific environmental cue results in the release of neuroactive chemicals in the nervous system, which in turn activate a cascade of intercellular signalling events leading initially to behavioural changes at settlement, and subsequently to both morphogenetic and further behavioural changes at metamorphosis9,10,11,12,13.
Of the signalling systems so far identified as regulators of settlement and metamorphosis, many are not conserved beyond the taxonomic level of family, or sometimes even genus14,15,16,17,18,19. An exception is nitric oxide (NO), a gaseous signalling molecule that has been implicated in repressing settlement and metamorphosis in a wide range of marine bilaterians, including ascidians20,21, polychaetes22, echinoderms23, crustaceans24 and gastropods25,26,27,28. NO activity in settling sea urchin larvae appears to be localised to neurons in the post-oral ciliary band that have a chemosensory function, consistent with the conserved role for NO in animal chemosensory pathways more generally12. In all of these bilaterian cases, a reduction in the level of endogenously synthesised NO appears sufficient to either alone initiate metamorphosis, or at least to enhance the response to an inductive exogenous cue. The phylogenetic diversity of taxa in which this has been experimentally observed has led to the proposition that NO maintains larvae in a prepared state for metamorphosis25 and that this role for NO may be a highly conserved feature of marine invertebrate settlement and metamorphosis29,30. However, there are documented cases in ascidians and a gastropod of NO playing a positive role in regulating the initiation and rate of metamorphosis31,32,33,34, raising the possibility that differing roles of NO may be associated with different animal life histories and ecologies30.
NO is an ancient signalling molecule used not only by animals, but also by bacteria and non-metazoan eukaryotes35,36,37. In animals, NO is primarily synthesised by nitric oxide synthase (NOS)38. It mediates a wide range of physiological responses, often via the activation of its primary receptor, soluble guanylyl cyclase (sGC)39,40,41,42, although it can also directly activate other signalling pathways by protein S-nitrosylation43 and nitration44. Importantly, NO appears to have a very deep ancestral role in regulating the timing of life cycle transitions in response to changes in environmental conditions. This role has been documented across the tree of life, in bacteria, slime moulds, fungi, plants and animals45,46,47,48,49,50, which clearly raises the possibility that NO was involved in regulating the major life cycle transition from pelagia to benthos even in the earliest animals. To investigate this, we explore here for the first time the role of NO in metamorphosis in a non-bilaterian animal, namely the haplosclerid demosponge, Amphimedon queenslandica51 that has a typical pelagobenthic life cycle3,52.
A. queenslandica naturally release larvae on a daily basis3,53,54. These larvae acquire competency by 4–6 hour post emergence (hpe)3,53,54, at which time they readily settle and metamorphose on the articulated coralline algae (CA) Amphiroa fragilissima55. Although sponges lack neurons56,57, they have the ability to sense their surroundings and to coordinate behaviours in response to those surroundings58,59,60. The presence of NOS or NO activity has been reported previously in demosponges, although not in relation to settlement and metamorphosis61,62,63. Here, we provide evidence that NO does indeed regulate settlement and the initiation of metamorphosis in A. queenslandica by demonstrating the effects of small molecules that (i) inhibit NOS activity, (ii) scavenge NO and (iii) generate exogenous NO.
We find that NO appears to induce, rather than represses, settlement and metamorphosis as has been observed in some ascidians and molluscs31,32,33. As has been documented in the ascidian Ciona33,44,64, NO signalling appears to work via – or in concert with – the ERK signalling pathway in A. queenslandica. Consistent with this positive regulation of metamorphosis, we find that NOS gene expression increases in epithelial, globular and sub-epithelial cells as larvae develop competence to settle and metamorphose. NO is enriched in these cells in larvae and this enrichment continues into the start of metamorphosis.
Results
Nitric oxide induces metamorphosis of Amphimedon queenslandica larvae
At 25 °C, Amphimedon queenslandica larvae become competent to respond to an inductive cue associated with the coralline algae Amphiroa fragilissima within 4–6 hours after emerging from their parent3. A. fragilissima is a potent inducer of settlement and metamorphosis, with over 90% of competent larvae typically settling on its surface just 1–2 hours after first contact55. In contrast, larvae maintained in 0.2 μm filtered sea water (FSW) – in the absence of any inductive cues – rarely settle, even when maintained for 24 hours or more3,55. These contrasting effects on competent A. queenslandica larvae are ideal for investigating mechanisms underlying the induction and inhibition of settlement and metamorphosis.
Competent larvae exposed to 10 mM L-nitroarginine methyl ester (L-NAME), which is a competitive inhibitor of NOS, 15 minutes prior to being exposed to A. fragilissima were significantly inhibited from initiating and undergoing metamorphosis, even by 24 hours post-induction (hpi) (Fig. 1A). These L-NAME-treated larvae remained in a normal free-swimming state; no mortality was observed. Likewise, the NO scavenger 2-(4-carboxyphenyl)-4,5-dihyrdo-4,4,5,5-tetramethyl-1H-imidazolyl-1-oxy-3-oxide (PTIO) inhibited larvae from initiating metamorphosis in a concentration-dependent manner, with all concentrations over 100 μM significantly reducing the number of individuals metamorphosing in the presence of the inductive coralline algae (Fig. 1B). PTIO-treated larvae also displayed normal swimming behaviour and survival.
Effect of L-NAME and PTIO on metamorphosis of Amphimedon queenslandica.
(A) Larvae subjected to 0.1, 1 and 10 mM L-NAME at competence (4–6 hpe) and then either reared in FSW or in the presence of A. fragilissima (+CA). 10 mM L-NAME significantly inhibits A. fragilissima-induced metamorphosis. (B) Larvae were subject to 100–250 μM PTIO at competence and then reared in the presence of A. fragilissima (+CA). All concentrations of PTIO significantly reduce A. fragilissima-induced rates of metamorphosis. Data are presented as the mean percentage of larval metamorphosis ± SEM. Letters above error bars indicate statistically significant differences (P < 0.05), as determined by one-way analysis of variance and Tukey’s HSD post hoc testing.
Application of the exogenous NO donor sodium nitroprusside (SNP; used at 2.5 μM) induced significantly higher levels of metamorphosis than the FSW control, and in fact induced metamorphosis equivalent to A. fragilissima (Fig. 2). In contrast, a different exogenous NO donor, namely S-Nitroso-N-acetylpenicillamine (SNAP, used at 2.5, 12.5 and 25 μM) did not induce metamorphosis above levels in the DMSO control, although its presence had no effect on the ability of A. fragilissima to induce high rates of metamorphosis (Fig. 2). In all treatments, larvae and postlarvae behaved and developed normally.
Effect of SNAP and SNP on metamorphosis of Amphimedon queenslandica.
(A) Larvae subjected to 2.5–25 μM SNAP at competence and reared in FSW undergo a significantly higher rate of metamorphosis than the FSW control but not higher than the DMSO control. In the presence of A. fragilissima (+CA), rates of SNAP-treated larvae do not differ from +CA controls. (B) Larvae exposed to 2.5 μM SNP at competence and reared in FSW have significantly higher rates of metamorphosis than the FSW control, similar to the A. fragilissima controls. Data are presented as the mean percentage of larval metamorphosis ± SEM. Letters above error bars indicate statistically significant differences (P < 0.05), as determined by one-way analysis of variance and Tukey’s HSD post hoc testing.
Nitric oxide signalling activates the MAPK pathway in A. queenslandica
The MAPK (mitogen-activated protein kinase)/ERK (extracellular signal-regulated kinases) pathway is involved in a wide range of developmental and physiological signalling events and is known both to activate NO production and to be activated by NO, including in ascidian metamorphosis33,44,64. ERK can be selectively inhibited by U0126. In A. queenslandica, we found that experimental application of 10 μM U0126 to competent larvae completely inhibits settlement and metamorphosis, even in the presence of A. fragilissima (Fig. 3A). This strongly suggests that the MAPK/ERK pathway is necessary for the initiation of metamorphosis. To address whether ERK signalling potentially lies upstream or downstream of NO signalling, we combined U0126 treatment with 25 μM SNP, which is a source of exogenous NO. We found that U0126-treated larvae were unable to settle and metamorphose in the presence of SNP alone, A. fragilissima alone, or both together (Fig. 3B). These results are consistent with ERK signalling being necessary for settlement and metamorphosis, and potentially occurring downstream of the metamorphosis-inducing NO signal. Further, when U0126-treated larvae were subsequently washed with FSW and re-exposed to the coralline algae cue, they settled at rates similar to normal untreated larvae indicating the U0126 is not acting as a general toxin (Fig. 3B).
Inhibition of MAPK signalling prevents A. queenslandica metamorphosis.
(A) Larvae subjected to 10 μM U0126 are inhibited from initiating metamorphosis, even in the presence of the inductive coralline alga; an equivalent concentration of 0.1% DMSO does not inhibit metamorphosis. (B) Larvae subjected to NO donor, 2.5 μM SNP, undergo metamorphosis at rates similar to coralline alga controls. MAPK inhibitor U0126 inhibits metamorphosis when larvae are in the presence of SNP, coralline algae or both. Larvae removed from U0126, washed in FSW and subjected to the inductive coralline alga undergo high rates of normal metamorphosis.
A. queenslandica has a single NOS gene that is structurally similar to other metazoan NOS
A single NOS gene, hereafter referred to as AqNOS, is present in the A. queenslandica genome65,66. Both the protein domain organisation and the intron/exon organisation of AqNOS are highly similar to that of other metazoan NOSs, in accordance with the high degree of conservation discussed in ref. 67. Indeed many of exon-intron boundaries are conserved between A. queenslandica and bilaterians, although, as is true for this sponge genome in general, intron sizes are comparatively small, with a mean intron size of just 242 bp (Supplementary Fig. 1). AqNOS has a conserved domain architecture that includes an N-terminal PDZ domain as is present in human nNOS, and a split NADPH-ribose binding site as is present in the gastropod mollusc Lymnaea stagnalis (Supplementary Fig. 2); specifically, there is a 129 residue insert in the NADPH-ribose binding site compared to the 84 amino acid insert in the L. stagnalis gene68. The C-terminal end of AqNOS is relatively long compared with other animal NOS proteins (Supplementary Fig. 2).
AqNOS gene expression supports a positive regulatory role of NO in A. queenslandica metamorphosis
As measured by quantitative RT-PCR, AqNOS transcript levels increase continuously through embryogenesis, eventually peaking in free-swimming larvae at 1 hpe (Fig. 4). Transcript abundance then gradually decreases towards competency (5 hpe), but still remains well above embryonic levels. In larvae that are not induced to settle, the transcript abundance increases after competency, reaching a second peak of expression at 19 hpe, and then decreases again by 29 hpe (Fig. 4). In postlarval samples (those induced to settle and metamorphose), peak AqNOS expression occurs by 4 hpi, at the very start of metamorphosis, and is followed by a gradual decrease during postlarval development.
AqNOS expression during Amphimedon queenslandica development.
AqNOS transcript abundance measured by qRT-PCR on pools of ~40 embryos still in brood chambers, larvae that are still swimming (red circles) and postlarvae that have settled and initiated metamorphosis (blue diamonds). White stage embryos are undergoing cleavage; brown embryos are commencing cell migration leading to two cell layers; spot stage embryos have two distinct layers and an obvious anteroposterior polarity; 1 and 3 hpe (hours post-emergence) are precompetent swimming larvae; 5+ hpe are competent swimming larvae; and 4, 14 and 124 hpi (hours post-induction) are metamorphosing postlarvae (for further details on developmental stages see ref. 70). Results are presented as log-transformed mean ± SEM of three technical replicates; the SEM was so small for most samples that it is not visible at the scale depicted.
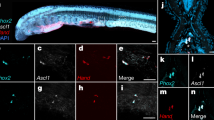
Using whole mount in situ hybridisation, we detect AqNOS transcripts in multiple larval cell types, specifically (i) globular cells that are interspersed on the larval surface between columnar epithelial cells69; (ii) the columnar epithelial cells themselves, which form the outer surface layer of the sponge larva; and (iii) spherulous cells that mainly comprise the sub-epithelial layer but also are sparsely distributed in the inner cell mass of the larva (Fig. 5). The expression of AqNOS in the epithelium is broad but does not include anterior cuboidal cells, for which a function has yet to be ascribed, or the posterior pigment ring cells and adjacent photosensory cells with long-cilia70 (Fig. 5B–E). In the columnar epithelium, AqNOS transcripts are enriched in the basal part of the cells; it is unclear if flask cells, which are interspersed amongst the columnar cells in the anterior third of the larva55, express AqNOS. In the inner cell mass, which is comprised of a number of described and undescribed cell types71, we see only two cell types with detectable levels of AqNOS transcript; these are spherulous cell types similar in morphology to the globular cells, and subepithelial spherulous cells (Fig. 5D,F).
Localised expression of AqNOS and endogenous nitric oxide in Amphimedon queenslandica larvae.
(A–H) AqNOS gene expression detected by in situ hybridisation: (A–C) whole mounts, (C–H) sections. (I) DAF-FM stained larva. Posterior to left except in panels B and C, which show posterior and anterior views, respectively. (A) Lateral view showing posterior pigment ring (pr) and anterior cuboidal cells (cc). The ‘salt-and-pepper’ staining indicates localisation of AqNOS transcripts to globular cells embedded in outer epithelial layer, and in underlying cell layers. (B) Posterior view with staining inside the pigment ring (asterisk) and in a ‘salt-and-pepper’ pattern in the lateral epithelium, but not in the pigment ring itself, or in the adjacent photosensory epithelium (arrow). (C) Anterior view showing AqNOS transcripts enriched in globular cells but not in anterior cuboidal cells inside the dotted line; there is staining in cells underlying cuboidal cells. (D) Longitudinal section through a larva revealing AqNOS expression in spherulous cells in inner cell mass (icm) and subepithelial layer (sel), in columnar epithelial and globular cells (gc) in the epithelial layer (el), and in globular cells inside the pigment ring (asterisk), but not in epithelial cells outside the pigment ring (arrow). (E–H) Higher magnifications of sections: (E) Larval anterior showing AqNOS expression primarily in epithelium and spherulous cells in the subepithelial layer, bordered by dashed lines, but not in anterior cuboidal cells; (F) Larval inner cell mass showing AqNOS transcripts enriched in a subset of spherulous cells but not in other cell types; (G) Lateral section showing AqNOS expression in columnar epithelial and globular cells in epithelial layer, and spherulous cells in subepithelium, bordered by dashed lines; (H) Higher magnification of a more heavily stained larval epithelium showing AqNOS transcripts localised to globular cells and to basal portion of columnar epithelium. (I) Endogenous nitric oxide, detected using the NO-specific indicator DAF-FM, enriched in globular cells (large, bright green cells dotting the larval surface) and less so in epithelial cells, in a larva just commencing metamorphosis; posterior pigment ring is bordered by dashed lines. Scale bars: (A–D), I, 100 μm; E-G, 20 μm; F, 10 μm.
To confirm that AqNOS-expressing cells are the primary source of NO in larvae and at the initiation of metamorphosis, we stained larvae with 4-amino-5-methylamino-2′,7′-difluoro-fluorescein diacetate (DAF-FM), which detects endogenous NO. The resultant DAF-FM staining pattern (Fig. 5I) followed closely the AqNOS in situ hybridisation pattern (Fig. 5A–H); signals from cells in the inner cell mass or subepithelial layer could not be resolved with this technique. The DAF-FM signal was brightest in globular cells dotting the surface of the larva (Fig. 5I). As in the AqNOS in situ hybridisation, there was no apparent difference in staining intensity of these cells along the anterior-posterior axis (Fig. 5I). NO was also detected in epithelial cells, although DAF-FM staining was markedly less in these cells, again reflecting the AqNOS in situ hybridisation pattern (Fig. 5). Tracing DAF-FM staining in larvae over the first hour of metamorphosis, during which time the larva attaches to the substratum and begins to flatten into the juvenile body plan, revealed no changes in the overall pattern of NO; that is, through the first hour of metamorphosis, NO appeared to remain highest in globular cells and detectable in the epithelium (Supplementary Movie 1).
In situ hybridisation of larvae revealed AqNOS-expressing globular cells apparently in the process of migrating out of the sub-epithelial layer – where they are classified as spherulous cells – to the outer epithelial layer, a migration that has been observed previously for globular cells during larval development69,70,71,72. The detection of AqNOS and NO in these cells, at a time coincident with larval responsiveness to NO, implies that globular cells are a source of endogenous NO, and involved in the process of larval settlement and metamorphosis. In their final location around the periphery of the larva, globular cells are tightly packed with large electron-dense granules, basal nuclei and apices that protrude outwards above the columnar epithelium73. Based on this distinctive morphology, we were able to trace the ontogeny of globular cells using toluidine blue (Fig. 6). At the spot stage of development (Supplementary Fig. 3), a population of spherulous cells that display similar staining characteristics to the larval globular cells are throughout the embryo (Fig. 6D1,E1). By the ring stage (Supplementary Fig. 3), the most outer layer of the embryo becomes clear of the spherulous cell population (Fig. 6A2–E2), which is now more densely visible at the interface between inner and outer layers, effectively forming a third layer in the embryo (Fig. 6D2). As the ring continues to form and the late embryo starts to elongate into the larval form (Fig. 6A3–E3), the spherulous cells migrate from the sub-epithelial layer and either intercalate into the ciliated columnar epithelium or localise at the posterior pole. Once they are in their final location, we now consider these cells to be globular cells. At hatching, globular cells are evident around the entire periphery of the A. queenslandica larva (Fig. 6A4–E4), except at the very anterior pole (Fig. 6E4) and amongst the pigmented cells at the posterior pole (Fig. 6C4). In the swimming larva, spherulous cells continue to migrate to the surface, and thus continue to increase the total number of externally-localised globular cells. By the time the larva attains competence, globular cells appears to be at higher density in the anterior larval epithelium (Fig. 6A4–B4).
Ontogeny of spherulous and globular cells indicated by toluidine blue staining.
(A1–E1) Spot stage embryo. (A2–E2) Early ring stage embryo. (A3–E3) Late ring (pre-hatching) stage embryo. (A4–E4) Pre-competent larva. (A1–A4) Whole mount embryos and larvae. (B1–B4) Sectioned embryos and larvae. Boxes correspond to respective C –E micrographs. (C1–C4) Higher magnification of posterior portion of embryos and larvae. (D1–D4) Higher magnification of lateral portion of embryos and larvae. (E1–E4) Higher magnification of anterior portion of embryos and larvae. ap, anterior pole; pr, pigment ring; ps, pigment spot; dashed line denotes boundary between outer epithelial layer and subepithelial layer; stars denote representative spherulous and globular cells; arrows in D2, E2 and E4 denote flask cells.
Discussion
Inhibiting NOS activity by the application of 10 mM L-NAME, or by the scavenging of released NO with PTIO, prevents Amphimedon queenslandica larvae from initiating metamorphosis even in the presence of the highly inductive coralline algae Amphiroa fragilissima. Consistent with this, the artificial elevation of NO levels in FSW, via the application of SNP, induces high rates of A. queenslandica larval settlement and metamorphosis, even in the absence of A. fragilissima. Together, these results indicate that NO signalling is necessary – and perhaps also sufficient – for the induction of settlement and initiation of metamorphosis of A. queenslandica larvae. The activating role of NO in A. queenslandica settlement and metamorphosis contrasts with observations in a range of bilaterian larvae where NO appears to play a repressive or inhibitory role20,21,22,23,24,25,26,27,28, but is similar to other species where it appears to play an inductive or stimulatory role30,31,32,33,34.
A. queenslandica has a single NOS gene (AqNOS) that encodes a protein domain architecture that is deeply conserved with the rest of the animal kingdom, yet includes a number of uncommon metazoan features, including a N-teminal PDZ domain and a split NADPH-ribose binding domain. AqNOS transcript abundance increases during embryogenesis to reach maximum levels in larvae, in which expression is spatially enriched in the ciliated columnar epithelium, and in globular and spherulous cells. These temporal and spatial expression profiles support a role for NOS in regulating settlement and metamorphosis in A. queenslandica. Corroborating these gene expression analyses is the detection, using the DAF-FM indicator, of comparable relative levels of NO in columnar epithelial and globular cells. Globular cells, which appear in A. queenslandica to be derived from spherulous cells, intercalate into the larval columnar epithelium and have apical protrusions that extend beyond the larval surface. These cells, along with the larval flask cells55, have been implicated as having a sensory role, based on their expression of postsynaptic genes, neurogenic factors and signals, and genes in the Toll pathway74.
The common morphology of AqNOS-expressing spherulous cells in the inner cell mass and subepithelial layer, and the globular cells on the larval surface, suggests that these cells are ontogenetically related and have a similar role in NO signalling. However, prior direct evidence of a role for globular or spherulous cells in controlling metamorphosis has not existed, and recently another larval cell type – the flask cell – has been implicated in contributing to the regulation of this process55. Regardless, the requirement of NO signalling implicates these NOS-expressing cells in the regulation of settlement and metamorphosis. Interestingly, globular cells accumulate differentially in the epithelial layer of the anterior half of larvae as they develop competence to settle and metamorphose. This anterior region is also where the flask cells are primarily located, and in fact the anterior third of the larva appears to play a critical role in the induction of metamorphosis55. Inhibition of intracellular calcium signalling in flask cells also inhibits metamorphosis55, further supporting a functional link between these cells and NOS-expressing cells. Together, these results support a model where the anterior part of A. queenslandica larvae contacts an environmental cue, which induces an interplay of intra- and intercellular signals in flask and globular cells that results in the release of NO and the initiation of metamorphosis (Fig. 7).
A model for the role of nitric oxide in the induction of larval settlement and metamorphosis.
(A,B) A schematic summarising the potential role of nitric oxide in larval settlement of Amphimedon queenslandica. (A) The competent A. queenslandica larva settles onto a coralline alga, anterior side touching the substratum. (B) A close-up of the boxed anterior region shown in (A). Exogenous signals associated with the algal surface activate a calcium signalling in flask cells (fc)55, which in turn signals to the surrounding cellular environment via membrane bound (e.g. Notch-Delta signalling71 or locally diffusible signals (dashed arrows). In A. queenslandica, this includes columnar epithelial and globular cells (gc), which express high levels of AqNOS (brown). These cells are induced to release NO (arrows) at a rate to induce morphogenetic and physiological changes in other larval cells, including other NOS-expressing cells (brown), leading to larval settlement and the initiation of metamorphosis. MAPK signalling is also required for A. queenslandica settlement and metamorphosis (not shown). (C) A general model for nitric oxide signalling in induction of larval settlement and metamorphosis. Receptor-activated calcium signalling occurs in sensory cells upon contact with an inductive exogenous cue, resulting in the release of a signalling ligand. This is detected by NOS-expressing cells, possibly by calcium-calmodulin binding to NOS, activating the synthesis of NO. The NO signal can induce cellular changes, including by binding to its receptor soluble guanylyl cyclase (sGC) and S-nitrosylation of members of the Ras-GTPase super family; these may result in the activation of the MAPK/ERK signalling pathway. In bilaterian larvae where NO inhibits or represses the settlement and metamorphosis20,21,22,23,24,25,26,27,28, a different model of NO and NOS regulation is required. icm, inner cell mass.
NO is a short-lived, highly diffusible molecular messenger75. In poriferans, which lack a nervous system, NO may allow for a coordinated and relatively rapid response to an external signal, as required for the initiation of metamorphosis. Settlement and metamorphosis in A. queenslandica is accompanied by a diversity of cellular changes, including widespread apoptosis and transdifferentiation76. The highly organised and patterned larva transforms within hours into a less structured “mat” of cells that adhere both to each other and to the substrate76. There is a rapid decrease in the number of cell types at this transition, with some larval cell types transdifferentiating into a common pluripotent stem cell type, the archeocyte53,76. There is no evidence of metamorphosis commencing in a particular part of the larva or via a morphogenetic wave, although the anterior of the larva usually makes first contact with the inductive substratum. Thus, the widespread morphogenetic transformation of the larva that rapidly follows settlement is consistent with the broad expression pattern of AqNOS, with the presence of NO in epithelial and globular cells along the entire anteroposterior axis, and with the rapid diffusion of a gaseous molecule such as NO (Fig. 7).
Inhibition of metamorphosis by treatment with U0126, a MAPK inhibitor, is consistent with the ERK pathway also being necessary for the induction of settlement and metamorphosis in A. queenslandica. This inhibition is maintained in the presence of the NO donor SNP or the inductive coralline algae A. fragilissima, or both, suggesting that the ERK pathway is downstream of the NO signalling event that initiates settlement and metamorphosis. However, we cannot exclude the possibility that this activation of MAPK/ERK occurs separately from the NO signalling pathway. NO signalling is often mediated by the binding of NO to the heme moiety of soluble guanylyl cyclase (sGC), thus activating this enzyme and resulting in an increase in the synthesis of cyclic guanosine 5′ monophosphate (cGMP)75. Cyclic GMP in turn (i) activates cGMP-dependent protein kinases (PKG), (ii) activates or inhibits cAMP-specific phosphodiesterases (PDEs), and (iii) opens cyclic nucleotide-gated cation channels, leading to a wide range of context-specific physiological and developmental outcomes in animals, some of which include ERK signalling29,41,77,78,79. In addition, NO can impact on cell state directly by S-nitrosylation of cysteine residues of signalling pathway components and transcription factors43,80,81, including members of the Ras-GTPase super family82,83. Thus it is possible that ERK activation may occur via the S-nitrosylation of Ras, or an alternative mechanism, in A. queenslandica (Fig. 7). For example, in Ciona, an elevation of NO under oxidative-stress-like conditions results in protein nitration, including tyrosine residues in ERK44. In this ascidian, NO also down-regulates MAP kinase phosphatases, which results in a increase in ERK signalling, and thus indicates yet another possible means by which NO can increase the signalling activity of this pathway.
Conclusions and hypothesis
We have provided multiple lines of evidence that support NO being both necessary and sufficient to activate larval settlement and metamorphosis in a marine sponge, A. queenslandica. Known NOS inhibitors and NO scavengers inhibit metamorphosis, and an NO donor induces metamorphosis. The localised expression of AqNOS and concomitant accumulation of detectable NO together suggest that larval globular cells and columnar epithelial cells are likely sources of NO in the larvae of this sponge (Figs 5 and 7). These cells may be interacting with sensory flask cells55 in the larval anterior to control settlement and metamorphosis (Fig. 7). NO typically represses the initiation of metamorphosis in multiple bilaterian marine invertebrates20,21,22,23,24,25,26,27,28,29, although there are other reported cases where NO acts as an activator of metamorphosis30,31,32,33,34. The results presented here for the demosponge A. queenslandica phylogenetically extend the role of NO in animal metamorphosis beyond the bilaterians, thus raising the possibility that its action harks back to the last common animal ancestor, and that it has been able to switch between being an activator and repressor of life cycle transitions over the course of animal evolution.
Using the results presented herein to build on previously published studies, we hypothesise more generally that settlement and metamorphosis in marine invertebrates requires first the binding of an environmentally-derived ligand to cell-surface sensory receptors (potentially G-protein coupled receptors - GPCRs) that in turn initiates intracellular calcium signalling (Fig. 7). Since NO is a critical modulator of chemosensory stimuli, especially in olfactory circuits84, it has long been proposed that the regulation of NOS can be linked to GPCRs activated by external chemical ligand binding85, although it is noteworthy that studies to date have failed to find support for a role of GPCRs in one annelid86 and two cnidarian87 species.
A current model of GPCR-NOS signalling is that a ligand-bound GPCR complex activates the signalling cascade of phospholipase C-inositol-1,4,5-triphosphate, resulting in the release of Ca2+ from the endoplasmic reticulum into the cytosol85. In A. queenslandica, this may be occurring in globular cells by induction by an exogenous ligand or from a signal originating in flask cells. In support of the latter, there is evidence of calcium signalling in globular cells after an initial release of intracellular calcium in flask cells55. Both flask cells and globular cells express Notch and a unique suite of Delta ligands71. In addition, flask cells extend filopodial processes above and within the epithelium55, raising the possibility that these cell types can interact via this signalling pathway. Regardless of whether direct or indirect, calcium signalling likely allows the activation of NOS, and thus NO production, via the binding of calmodulin42,88.
As a gaseous molecule, NO can freely and rapidly diffuse across cells and the cellular membrane, and can quickly modify its target molecules by covalent binding89. Evidence to date suggests that this target could be either Ras or sGC, both of which can result in activation of the ERK pathway and, ultimately, regulation of the comprehensive morphogenetic changes that constitute metamorphosis (Fig. 7); recent analyses in the ascidian Ciona indicate that NO may affect ERK signalling also by direct nitration or inhibition of MAP kinase phosphatases33,44. Indeed, application of an sGC inhibitor, ODQ (1H-[1,2,4]oxadiazolo[4,3-a]quinozalin-l-one), significantly increases the induction of metamorphosis in several marine invertebrates20,23,24,25,26,27. We suggest that the model outlined in Fig. 7 could provide a framework for future experiments, exploiting available pharmaceutical reagents to manipulate each step, and thus to test the role of NO and its upstream and downstream components in the metamorphosis of a wide range of marine invertebrate species.
Methods
Amphimedon queenslandica larval cultivation
Adult specimens of the sponge Amphimedon queenslandica were collected from Heron Island Reef, Great Barrier Reef, Australia (23° 26′S, 151° 55′E). All embryological material was procured from dissected brood chambers as described in Leys et al.90. For larval collection, adult specimens were maintained in tanks with gentle flow of ambient seawater. Larvae released naturally during the day91 were collected in containers with 100 μm nitex mesh bottoms placed at the outflow. Larvae were collected every hour, allowing the determination of larval ages within 1 hour post emergence (hpe); that is, a 1 hpe sample contains larvae that had emerged between 0 (larvae collected immediately after emergence from its maternal adult) and 1 h (larvae that emerged from the maternal adult shortly after the previous collection). The collected larvae were maintained in 500 mL ambient seawater at 25 ± 0.5 °C for at least 4–5 h to allow them to reach competency3 (Supplementary Fig. 3).
Assessment of NO-manipulating small molecules on metamorphosis
The following small molecules that influence NO levels were used in this study: NOS inhibitor L-nitroarginine-methyl-ester (L-NAME; Sigma), NO scavenger 2-(4-carboxyphenyl)-4,5-dihyrdo-4,4,5,5-tetramethyl-1H-imidazolyl-1-oxy-3-oxide (PTIO; Sigma), and NO donors S-nitroso-N-acetyl-penicillamine (SNAP; Sapphire Bioscience) and sodium nitroprusside (SNP; Sigma). All these chemicals have demonstrated significant effects on marine invertebrate metamorphosis20,21,22,23,24,25,26,27.
Stock solutions of 0.5 M L-NAME, 2.5 mM PTIO and 2.5 mM SNP were prepared in 0.2 μm filtered sea water (FSW), stored at 4 °C and diluted just prior to the experiment; SNP was prepared just prior to its use. The stock solution of 50 mM SNAP was prepared in dimethyl sulfoxide (DMSO) diluted immediately prior to the experiment. To maintain the steady concentration of NO delivered by SNAP, the solution was renewed every 6 h25.
All settlement assays were performed in 6-well 35-mm diameter sterile polycarbonate tissue culture dishes, with 10 ml of FSW per well, and were initiated at 5 hpe, by which time all larvae would be competent (Supplementary Fig. 3). Three replicates of 15 larvae were incubated continuously in wells containing either (1) 10 ml FSW only, (2) 10 ml FSW with freshly prepared shards of coralline algae A. fragilissima, which were collected from the Heron Island reef flat and cleaned just prior to each experiment, covering approximately 25% of the bottom of the well or (3) 10 ml FSW containing a one of the above mentioned small molecules either with or without coralline algal shards. Inhibitory molecules L-NAME and PTIO were added to the wells with larvae 15 minutes prior to the addition of A. fragilissima. The number of larvae initiating metamorphosis was scored after incubating the culture dishes for 24 h post induction with A. fragilissima (24 hpi) in the dark at 25 °C. Postlarvae were deemed to have initiated metamorphosis only if they met all three of the following criteria: (i) were attached to a substrate; (ii) had begun flattening along the anteroposterior axis; and (iii) had begun resorbing their pigment ring53. Because of natural biological variation in larval competence and the A. fragilissima inducer, all controls were run in every experiment to allow treatments to be related to controls in every cohort.
Settlement and metamorphosis data were analysed by one-way analysis of variance (ANOVA) by testing treatment as a factor. Significant differences among treatments were detected by Tukey’s HSD post hoc testing. Prior to the analyses, all data were arcsine-transformed to improve the normal distribution of samples. Levene’s test was performed to assure the homogeneity of variance among treatments. All statistical analyses were performed in R (R Foundation for Statistical Computing). An alpha value of 0.05 was used to determine a significant difference92.
U0126 treatments
A 10 mM solution of the MAPK inhibitor U0126 (Promega, V1121) was prepared in DMSO and further diluted in (FSW) to a final concentration of 10 μM. Competent larvae (5 hpe; Supplementary Fig. 3) were transferred into FSW containing either 10 μM U0126 or 0.1% DMSO controls as described above for NO small molecule treatments. Treated and control larvae were subject to coralline algae or SNP or both 15 minutes later, as described above. To ensure the inhibitory effect of U0126 was not related to toxicity, U0126-treated larvae were washed twice in FSW after a 15 min treatment, and then were subject to coralline algae as described above.
Analysis of the A. queenslandica NOS gene
The nucleotide (XM_011411881) and derived amino acid (XP_011410183) sequences of A. queenslandica NOS are available in the NCBI database, where the gene is annotated as an inducible NOS, and at http://metazoa.ensembl.org/Amphimedon_queenslandica/Info/Index where the gene is annotated as Aqu1.226552 based on genome version 165. Using these databases, as well as our most recent in-house automated gene model prediction (Aqu2.1 gene models; see ref. 66), we manually curated a NOS gene model, evaluated intron–exon boundaries and domain organization as previously described for other genes93, and compared these to the metazoan-wide survey of NOS gene structure by Andreakis et al.67. Herein we refer to this gene as AqNOS.
Quantitative RT-PCR analysis of AqNOS expression
RNA was purified from embryonic, larval, and postlarval stages as previously described93,94; each developmental stage was represented by a single RNA sample purified from a pool of approximately 40 individual embryos, larvae or postlarvae that were sourced from a pool of 10 different adult sponges. Embryonic staging followed established morphological criteria: white (early blastula), brown (late blastula), spot (concentrated pigment cells are visible at the posterior end), and ring (appearance of conspicuous pigmented ring formation at the posterior end) (Supplementary Fig. 3)53,70. Larval age was determined by duration (in hours) post emergence from the maternal brood chamber (hpe) as described above. To ensure non-overlap of larval age between samples, only 1, 3, and 5 hpe samples were assayed (Supplementary Fig. 3). To collect postlarval samples, competent larvae (5 hpe) were exposed to live A. fragilissima shards in 12 cm petri dishes to induce metamorphosis, and settled postlarvae that had initiated metamorphosis were then sampled at 4, 14 and 24 hpi. Non-induced, still-swimming larval samples of the same developmental age were also collected at 9, 19 and 29 hpe for comparison (Supplementary Fig. 3).
AqNOS transcript abundance during normal development was determined by qRT-PCR using the following oligonucleotide primer set: forward 5′-TACGAGGAGCTCACCTACGG-3′ and reverse 5′-TCCTTGGTTTGTGGCATAGC-3′. The geometric mean of two reference genes –succinate dehydrogenase complex, subunit A (SDHA) (forward 5′-CGGGGAGTGGTAGCTATGAA-3′ and reverse 5′-TGAAACTGTACAAACTCCATGTCT-3′) and tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide (YWHAZ) (forward 5′- AGCGGAGGGCAAAAAGAA-3′ and reverse 5′-CCTCGGCAAGATAGCGATAGTAGT-3′) was used to normalise the level of transcription for obtaining relative gene expression values. Their stability within and between embryonic, larval, and postlarval stages was confirmed using Genorm software95.
The following qRT-PCR reaction parameters were used: initial denaturation 95 °C for 10 min (ramp rate 4.4 °C/sec), and 40–50 cycles of 95 °C for 5 sec (ramp rate 4.4 °C/sec), 58 °C for 10 sec (ramp rate 2.2 °C/sec), and 72 °C for 20 sec (ramp rate 4.4 °C/sec). Melt curve data acquisition was from 55–95 °C with continuous measurement (acquisition/°C = 5) to confirm the purity of PCR product by the presence of a single peak in the temperature melt curve. All samples were run in triplicate to calculate the average values to take technical variations into account. For each primer pair, a standard curve was generated to calculate the efficiency of qRT-PCR using a dilution series from the calibrator sample, which was a mixture of 4 μl (normal development and small molecule exposure) of all undiluted cDNA samples of each experimental set. In addition to the developmental stage cDNAs, a no-template (H2O) control and the calibrator sample were included for each qRT-PCR run and for each primer pair. The efficiencies of each primer pair and the cycle threshold of each sample were calculated by the second derivative method using Roche Light Cycler 480 software program. Relative expression ratios were calculated as the ratio of gene expression between the gene of interest and the geometric mean of the two reference genes (normalisation factor) relative to their calibrator sample. The expression ratios and standard errors were calculated using REST-RG beta software version 396.
Whole mount in situ hybridisation analysis
AqNOS gene expression patterns were assessed in precompetent (3 hpe) and competent (5 hpe) A. queenslandica larvae (Supplementary Fig. 3). Fixation and storage of specimens, and whole mount in situ hybridisation (WMISH) were performed as described in Larroux et al.93,94. 798 and 1649 bp fragments of the AqNOS cDNA were used as probes (Supplementary Fig. 4). WMISH images were captured with Olympus BX60 using a Nikon Digital Sight DS-U1 camera.
Detection of endogenous nitric oxide
NO was detected in vivo using 4-amino-5-methylamino-2′,7′-difluoro- fluorescein diacetate (DAF-FM) (Molecular Probes) as previously described for the ascidian Ciona21. Competent larvae (6–8 hpe) were incubated in the dark with 5 μM DAF-FM in FSW for 30 min, and then washed twice and maintained in sea water. Larvae were subsequently viewed and photographed in the process of settling and initiating metamorphosis (Supplementary Movie 1; Fig. 5I)
Tracing the ontogeny of spherulous and globular cells by staining with toluidine blue
WMISH analysis highlighted the particular relevance to this study of two cell types, namely spherulous and globular cells, that are know from previous work to share an ontogenetic relationship69,70,71,72. The distinctive morphology of globular cells70,73 facilitates the tracing of their ontogeny using toluidine blue. To do so, embryos from spot stage through to pre-competent larvae were fixed in 4% paraformaldehyde + 0.05% glutaraldehyde/1XMOPS buffer, dehydrated, then embedded in Epon epoxy resin and cut into 1 um sections. Sections were stained with a standard toluidine blue solution (0.5% toluidine blue, 1% borax).
Additional Information
How to cite this article: Ueda, N. et al. An ancient role for nitric oxide in regulating the animal pelagobenthic life cycle: evidence from a marine sponge. Sci. Rep. 6, 37546; doi: 10.1038/srep37546 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Hadfield, M. G. Why and how marine-invertebrate larvae metamorphose so fast. Semin. Cell Dev. Biol. 11, 437–443 (2000).
Pechenik, J. A. Larval experience and latent effects-metamorphosis is not a new beginning. Integr. Comp. Biol. 46, 323–333 (2006).
Degnan, S. M. & Degnan, B. M. The initiation of metamorphosis as an ancient polyphenic trait and its role in metazoan life-cycle evolution. Philos. Trans. R. Soc. London. B 365, 641–651 (2010).
Jackson, G. A. & Strathmann, R. R. Larval mortality from offshore mixing as a link between precompetent and competent periods of development. Am. Nat. 118, 16–26 (1981).
Pawlik, J. R. Chemical ecology of the settlement of benthic marine invertebrates. Ocean. Mar. Biol. Annu. Rev 30, 273–335 (1992).
Gaylord, B., Hodin, J. & Ferner, M. C. Turbulent shear spurs settlement in larval sea urchins. Proc. Natl. Acad. Sci. USA 110, 6901–6906 (2013).
Hodin, J., Ferner, M. C., Ng, G., Lowe, C. J. & Gaylord, B. Rethinking competence in marine life cycles: ontogenetic changes in the settlement response of sand dollar larvae exposed to turbulence. R. Soc. Open Sci. 2, 150114, doi: 10.1098/rsos.150114 (2015).
Hadfield, M. & Paul, V. Natural chemical cues for settlement and metamorphosis of marine invertebrate larvae. In Marine chemical ecology (eds. McClintock, J. B. & Baker, B. J. ) 431–461 (CRC press, 2001).
Cloney, R. A. Ascidian larvae and the events of metamorphosis. Am. Zool. 22, 817–826 (1982).
Hadfield, M. G. The D P Wilson lecture. research on settlement and metamorphosis of marine invertebrate larvae: past, present and future. Biofouling 12, 9–29 (1998).
Leise, E. M. & Hadfield, M. G. An inducer of molluscan metamorphosis transforms activity patterns in a larval nervous system. Biol. Bull. 199, 241–250 (2000).
Bishop, C. D. & Brandhorst, B. P. Development of nitric oxide synthase-defined neurons in the sea urchin larval ciliary band and evidence for a chemosensory function during metamorphosis. Dev. Dyn. 236, 1535–1546 (2007).
Conzelmann, M. et al. Conserved MIP receptor-ligand pair regulates Platynereis larval settlement. Proc. Natl. Acad. Sci. USA 110, 8224–8229 (2013).
Baxter, G. & Morse, D. E. G protein and diacylglycerol regulate metamorphosis of planktonic molluscan larvae. Proc. Natl. Acad. Sci. USA 84, 1867–1870 (1987).
Degnan, B. M. & Morse, D. E. Developmental and morphogenetic gene regulation in Haliotis rufescens larvae at metamorphosis. Am. Zool. 35, 391–398 (1995).
Biggers, W. J. & Laufer, H. Settlement and metamorphosis of Capitella larvae induced by juvenile hormone-active compounds is mediated by protein kinase C and ion channels. 196, 187–198 (1999).
Eri, R. et al. Hemps, a novel EGF-like protein, plays a central role in ascidian metamorphosis. Development 126, 5809–5818 (1999).
Amador-Cano, G., Carpizo-Ituarte, E. & Cristino-Jorge, D. Role of protein kinase C, G-protein coupled receptors, and calcium flux during metamorphosis of the sea urchin Strongylocentrotus purpuratus. Biol. Bull. 210, 121–131 (2006).
Heyland, A., Degnan, S. M. & Reitzel, A. M. Emerging patterns in the regulation and evolution of marine invertebrate settlement and metamorphosis. In Mechanisms of life history evolution (eds. Flatt, T. & Heyland, A. ) 29–42 (Oxford University Press, 2011).
Bishop, C. D., Bates, W. R. & Brandhorst, B. P. Regulation of metamorphosis in ascidians involves NO/cGMP signaling and HSP90. J. Exp. Zool. 289, 374–384 (2001).
Comes, S. et al. Regulatory roles of nitric oxide during larval development and metamorphosis in Ciona intestinalis. Dev. Biol. 306, 772–784 (2007).
Biggers, W. J. et al. Inhibitors of nitric oxide synthase induce larval settlement and metamorphosis of the polychaete annelid Capitella teleta. Invertebr. Reprod. Dev. 56, 1–13 (2011).
Bishop, C. D. & Brandhorst, B. P. NO/cGMP signaling and HSP90 activity represses metamorphosis in the sea urchin Lytechinus pictus. Biol. Bull. 201, 394–404 (2001).
Zhang, Y. et al. The regulatory role of the NO/cGMP signal transduction cascade during larval attachment and metamorphosis of the barnacle Balanus (=Amphibalanus) amphitrite. J. Exp. Biol. 215, 3813–3822 (2012).
Froggett, S. J. & Leise, E. M. Metamorphosis in the marine snail Ilyanassa obsoleta, yes or NO? Biol. Bull. 196, 57–62 (1999).
Pechenik, J. A. et al. Nitric oxide inhibits metamorphosis in larvae of Crepidula fornicata, the slippershell snail. Biol. Bull. 213, 160–171 (2007).
Bishop, C. D. et al. Analysis of nitric oxide-cyclic guanosine monophosphate signaling during metamorphosis of the nudibranch Phestilla sibogae Bergh (Gastropoda: Opisthobranchia). Evol. Dev. 10, 288–299 (2008).
Romero, M. R., Phuong, M. A., Bishop, C. D. & Krug, P. J. Nitric oxide signaling differentially affects habitat choice by two larval morphs of the sea slug Alderia willowi: mechanistic insight into evolutionary transitions in dispersal strategies. J. Exp. Biol. 216, 1114–1125 (2013).
Bishop, C. D. & Brandhorst, B. P. On nitric oxide signaling, metamorphosis, and the evolution of biphasic life cycles. Evol. Dev. 5, 542–550 (2003).
Bishop, C. D. & Biggers, W. J. In Haliotis, NO means YES. Front. Mar. Sci. 1, 51 (2014).
Ueda, N. & Degnan, S. M. Nitric oxide acts as a positive regulator to induce metamorphosis of the ascidian Herdmania momus. PLoS One 8, e72797, doi: 10.1371/journal.pone.0072797 (2013).
Ueda, N. & Degnan, S. M. Nitric oxide is not a negative regulator of metamorphic induction in the abalone Haliotis asinina. Front. Mar. Sci. 1, 21, doi: 10.3389/fmars.2014.00021 (2014).
Castellano, I., Ercolesi, E. & Palumbo, A. Nitric Oxide Affects ERK Signaling through Down-Regulation of MAP Kinase Phosphatase Levels during Larval Development of the Ascidian Ciona intestinalis. PLoS One 9, 102907, doi: 10.1371/journal.pone.0102907 (2014)
Castellano, I. et al. The diatom-derived aldehyde decadienal affects life cycle transition in the ascidian Ciona intestinalis through nitric oxide/ERK signalling. Open Biol. 5, 140182, doi: 10.1098/rsob.140182 (2015).
Feelisch, M. & Martin, J. F. The early role of nitric oxide in evolution. Trends Ecol. Evol. 10, 496–499 (1995).
Durner, J., Gow, A. J., Stamler, J. S. & Glazebrook, J. Ancient origins of nitric oxide signaling in biological systems. Proc. Natl. Acad. Sci. USA 96, 14206–14207 (1999).
Murad, F. & Barber, R. A hypothesis about cellular signaling with nitric oxide in the earliest life forms in evolution. Free Radic. Biol. Med. 47, 1325–1327 (2009).
Stuehr, D. J. Enzymes of the L-arginine to nitric oxide pathway. J. Nutr. 134, 2748–2751 (2004).
Jacklet, J. W. Nitric oxide signaling in invertebrates. Invertebr. Neurosci. 3, 1–14 (1997).
Colasanti, M. & Venturini, G. Nitric oxide in invertebrates. Mol. Neurobiol. 17, 157–174 (1998).
Palumbo, A. Nitric oxide in marine invertebrates: a comparative perspective. Comp. Biochem. Physiol. A. Mol. Integr. Physiol. 142, 241–248 (2005).
Daff, S. N. O synthase: structures and mechanisms. Nitric oxide 23, 1–11 (2010).
Sha, Y. & Marshall, H. E. S-nitrosylation in the regulation of gene transcription. Biochim. Biophys. Acta - Gen. Subj. 1820, 701–711 (2012).
Ercolesi, E. et al. Protein nitration as footprint of oxidative stress-related nitric oxide signaling pathways in developing Ciona intestinalis. Nitric Oxide 27, 18–24 (2012).
Tao, Y. P., Misko, T. P., Howlett, a C. & Klein, C. Nitric oxide, an endogenous regulator of Dictyostelium discoideum differentiation. Development 124, 3587–3595 (1997).
Wilken, M. & Huchzermeyer, B. Suppression of mycelia formation by NO produced endogenously in Candida tropicalis. Eur. J. Cell Biol. 78, 209–213 (1999).
Golderer, G., Werner, E. R., Leitner, S., Gröbner, P. & Werner-Felmayer, G. Nitric oxide synthase is induced in sporulation of Physarum polycephalum. Genes Dev. 15, 1299–1309 (2001).
He, Y. et al. Nitric oxide represses the Arabidopsis floral transition. Science 305, 1968–1971 (2004).
Thompson, S. E. M., Callow, M. E. & Callow, J. A. The effects of nitric oxide in settlement and adhesion of zoospores of the green alga Ulva. Biofouling 26, 167–178 (2010).
Cáceres, L. et al. Nitric oxide coordinates metabolism, growth, and development via the nuclear receptor E75. Genes Dev. 25, 1476–1485 (2011).
Hooper, J. N. A. & Van Soest, R. W. A new species of Amphimedon (Porifera, Demospongiae, Haplosclerida, Niphatidae) from the Capricorn-Bunker group of islands, Great Barrier Reef, Australia: target species of the ‘sponge genome project’. Zootaxa 1314, 31–39 (2006).
Degnan, S. M. & Degnan, B. M. The origin of the pelagobenthic metazoan life cycle: what’s sex got to do with it? Integr. Comp. Biol. 46, 683–690 (2006).
Leys, S. P. & Degnan, B. M. Embryogenesis and metamorphosis in a haplosclerid demosponge: gastrulation and transdifferentiation of larval ciliated cells to choanocytes. Invertebr. Biol. 121, 171–189 (2002).
Maritz, K., Calcino, A., Fahey, B., Degnan, B. & Degnan, S. M. Remarkable consistency of larval release in the spermcast-mating demosponge Amphimedon queenslandica (Hooper and van Soest). Open Mar. Biol. J. 4, 57–64 (2010).
Nakanishi, N., Stoupin, D., Degnan, S. M. & Degnan, B. M. Sensory flask cells in sponge larvae regulate metamorphosis via calcium signaling. Integr. Comp. Biol. 55, 1018–1027 (2015).
Leys, S. & Meech, R. Physiology of coordination in sponges. Can. J. Zool. 84, 288–306 (2006).
Nickel, M. Evolutionary emergence of synaptic nervous systems: what can we learn from the non-synaptic, nerveless Porifera? Invertebr. Biol. 129, 1–16 (2010).
Mackie, G. O. Is there a conduction system in sponges? In Levi C., Boury-Esnault N., editors Colloques internationaux du centre national de la recherche scientifique, Paris, CNRS (1979).
Elliott, G. R. D. & Leys, S. P. Coordinated contractions effectively expel water from the aquiferous system of a freshwater sponge. J. Exp. Biol. 210, 3736–3748 (2007).
Leys, S. P. Elements of a ‘nervous system’ in sponges. J. Exp. Biol. 218, 581–591 (2015).
Giovine, M. et al. Heat stress-activated, calcium-dependent nitric oxide synthase in sponges. Nitric Oxide 5, 427–431 (2001).
Ellwanger, K. & Nickel, M. Neuroactive substances specifically modulate rhythmic body contractions in the nerveless metazoon Tethya wilhelma (Demospongiae, Porifera). Front. Zool. 3, 7, doi: 10.1186/1742-9994-3-7 (2006).
Elliott, G. R. D. & Leys, S. P. Evidence for glutamate, GABA and NO in coordinating behaviour in the sponge, Ephydatia muelleri (Demospongiae, Spongillidae). J. Exp. Biol. 213, 2310–2321 (2010).
Chambon, J.-P., Nakayama, A., Takamura, K., McDougall, A. & Satoh, N. ERK- and JNK-signaling regulate gene networks that stimulate metamorphosis and apoptosis in tail tissues of ascidian tadpoles. Development 134, 1203–1219 (2007).
Srivastava, M. et al. The Amphimedon queenslandica genome and the evolution of animal complexity. Nature 466, 720–726 (2010).
Fernandez-Valverde, S. L., Calcino, A. D. & Degnan, B. M. Deep developmental transcriptome sequencing uncovers numerous new genes and enhances gene annotation in the sponge Amphimedon queenslandica. BMC Genomics 16, 387, doi: 10.1186/s12864-015-1588-z (2015).
Andreakis, N. et al. Evolution of the nitric oxide synthase family in metazoans. Mol. Biol. Evol. 28, 163–179 (2011).
Korneev, S. A. et al. Molecular characterization of NOS in a mollusc: expression in a giant modulatory neuron. J. Neurobiol. 35, 65–76 (1998).
Richards, G. S. et al. Sponge genes provide new insight into the evolutionary origin of the neurogenic circuit. Curr. Biol. 18, 1156–1161 (2008).
Degnan, B. M. et al. Porifera In Evolutionary developmental biology of invertebrates (ed. Wanninger, A. ) 65–106 (Springer, 2015).
Richards, G. S. & Degnan, B. M. The expression of Delta ligands in the sponge Amphimedon queenslandica suggests an ancient role for Notch signaling in metazoan development. Evodevo 3, 15, doi: 10.1186/2041-9139-3-15 (2012).
Gauthier, M. E. A., Du Pasquier, L. & Degnan, B. M. The genome of the sponge Amphimedon queenslandica provides new perspectives into the origin of Toll-like and interleukin 1 receptor pathways. Evol. Dev. 12, 519–533 (2010).
Leys, S. P. & Degnan, B. M. Cytological basis of photoresponsive behavior in a sponge larva. Biol. Bull. 201, 323–338 (2001).
Sakarya, O. et al. A post-synaptic scaffold at the origin of the animal kingdom. PLoS One 2, e506, doi: 10.1371/journal.pone.0000506 (2007).
Garthwaite, J. Dynamics of cellular NO-cGMP signaling. Front. Biosci. 10, 1868–1880 (2005).
Nakanishi, N., Sogabe, S. & Degnan, B. M. Evolutionary origin of gastrulation: insights from sponge development. BMC Biol. 12, 26, doi: 10.1186/1741-7007-12-26 (2014).
Hofmann, F., Ammendola, A. & Schlossmann, J. Rising behind NO: cGMP-dependent protein kinases. J. Cell Sci. 113, 1671–1676 (2000).
Derbyshire, E. R. & Marletta, M. A. Biochemistry of soluble guanylate cyclase In Handbook of experimental pharmacology (eds. Schmidt Harald, H. H. W., Hofmann, F. & Stasch, J.-P. ) 17–31 (Springer, 2009).
Colasanti, M., Persichini, T. & Venturini, G. Nitric oxide pathway in lower metazoans. Nitric Oxide 23, 94–100 (2010).
Hess, D. T., Matsumoto, A., Kim, S.-O., Marshall, H. E. & Stamler, J. S. Protein S-nitrosylation: purview and parameters. Nat. Rev. Mol. Cell Biol. 6, 150–166 (2005).
Daaka, Y. S-nitrosylation-regulated GPCR signaling. Biochim. Biophys. Acta - Gen. Subj. 1820, 743–751 (2012).
Oliveira, C. J. R. et al. The low molecular weight S-nitrosothiol, S-nitroso-N-acetylpenicillamine, promotes cell cycle progression in rabbit aortic endothelial cells. Nitric Oxide 18, 241–255 (2008).
Eller-Borges, R. et al. Ras, Rac1, and phosphatidylinositol-3-kinase (PI3K) signaling in nitric oxide induced endothelial cell migration. Nitric Oxide 47, 40–51 (2015).
Breer, H. & Shepherd, G. M. Implications of the NO/cGMP system for olfaction. Trends Neurosci. 16, 5–9 (1993).
Christopoulos, A. & El-Fakahany, E. E. The generation of nitric oxide by G protein coupled receptors. Life Sci. 64, 1–15 (1999).
Holm, E. R., Nedved, B. T., Carpizo-Ituarte, E. & Hadfield, M. G. Metamorphic signal transduction in Hydroides elegans (Polychaeta: Serpulidae) is not mediated by a G protein. Biol. Bull. 195, 21–29 (1998).
Tran, C. & Hadfield, M. G. Are G-protein-coupled receptors involved in mediating larval settlement and metamorphosis of coral planulae? Biol. Bull. 222, 128–136 (2012).
Kone, B. C., Kuncewicz, T., Zhang, W. & Yu, Z.-Y. Protein interactions with nitric oxide synthases: controlling the right time, the right place, and the right amount of nitric oxide. Am. J. Physiol. Renal Physiol. 285, F178–F190 (2003).
Moroz, L. L. & Kohn, A. B. On the comparative biology of nitric oxide (NO) synthetic pathways: parallel evolution of NO-mediated signaling. Adv. Exp. Biol. 01, 1–44 (2007).
Leys, S. P. et al. Isolation of Amphimedon developmental material. CSH Protoc. 2008, pdb.prot5095 (2008).
Maritz, K., Calcino, A., Fahey, B., Degnan, B. & Degnan, S. Remarkable consistency of larval supply in the spermcast-mating demosponge Amphimedon queenslandica (Hooper and van Soest). Open Mar. Biol. 4, 57–64 (2010).
Zar, J. Biostatistical analysis, 5th edition (Prentice Hall, 2010).
Larroux, C. et al. Whole-mount in situ hybridization In Amphimedon. Cold Spring Harb. Protoc. 2008, pdb.prot5286 (2008).
Larroux, C. et al. Developmental expression of transcription factor genes in a demosponge: insights into the origin of metazoan multicellularity. Evol. Dev. 8, 150–173 (2006).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, research0034.0031–0034.0011 (2002).
Pfaffl, M. W., Horgan, G. W. & Dempfle, L. Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res. 30, e36 (2002).
Acknowledgements
We acknowledge the University of Queensland’s Heron Island Research Station for providing facilities for biological collection and experiments. We thank Carmel McDougall, Nagayasu Nakanishi, and Claire Larroux for helpful advice in WMISH analysis, Melanie Havler for assistance with WMISH sample sectioning, and Daniel Stoupin for DAF-FM imagery. This research was conducted while NU held an Endeavour International Postgraduate Research Scholarship provided by Australian Government at the University of Queensland, and was supported by grants from the Australian Research Council to SMD.
Author information
Authors and Affiliations
Contributions
N.U., G.R., R.C., B.M.D., and S.M.D. designed and conducted metamorphosis assays. NU conducted qRT-PCR analysis. N.U., A.K. and M.A. conducted in situ hybridisation analysis. G.R. conducted the cell ontogeny analysis. N.U., S.M.D., B.M.D. and G.R. conceived the study and wrote the manuscript. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ueda, N., Richards, G., Degnan, B. et al. An ancient role for nitric oxide in regulating the animal pelagobenthic life cycle: evidence from a marine sponge. Sci Rep 6, 37546 (2016). https://doi.org/10.1038/srep37546
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep37546
This article is cited by
-
Taurine as a key intermediate for host-symbiont interaction in the tropical sponge Ianthella basta
The ISME Journal (2023)
-
Chemical cognition: chemoconnectomics and convergent evolution of integrative systems in animals
Animal Cognition (2023)
-
Symbiont transmission in marine sponges: reproduction, development, and metamorphosis
BMC Biology (2022)
-
Microbes from Mum: symbiont transmission in the tropical reef sponge Ianthella basta
ISME Communications (2022)
-
Bivalves are NO different: nitric oxide as negative regulator of metamorphosis in the Pacific oyster, Crassostrea gigas
BMC Developmental Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.