Abstract
Hybrid rice breeding offers an important strategy to improve rice production, in which the cultivation of a male sterile line is the key to the success of cross-breeding. CRISPR/Cas9 systems have been widely used in target-site genome editing, whereas their application for crop genetic improvement has been rarely reported. Here, using the CRISPR/Cas9 system, we induced specific mutations in TMS5, which is the most widely applied thermo-sensitive genic male sterility (TGMS) gene in China, and developed new “transgene clean” TGMS lines. We designed 10 target sites in the coding region of TMS5 for targeted mutagenesis using the CRISPR/Cas9 system and assessed the potential rates of on- and off-target effects. Finally, we established the most efficient construct, the TMS5ab construct, for breeding potentially applicable “transgene clean” TGMS lines. We also discussed factors that affect the editing efficiency according to the characteristics of different target sequences. Notably, using the TMS5ab construct, we developed 11 new “transgene clean” TGMS lines with potential applications in hybrid breeding within only one year in both rice subspecies. The application of our system not only significantly accelerates the breeding of sterile lines but also facilitates the exploitation of heterosis.
Similar content being viewed by others
Introduction
Rice (Oryza sativa L.) is one of the most important staple foods worldwide, providing almost one-quarter of the global dietary energy supply for humans1,2. The demand for food continues to rise as the population rapidly grows, necessitating the production of 40% more rice in 2030, and arable land is limited together with environmental degradation3. Hybrid rice, which has been developed in more than 40 countries worldwide and plays a key role in the global food supply4, has a 10–20% yield advantage over conventional rice and occupies approximately 60% of the total rice area in China5,6. The hybrid rice developed using the three- and two-line hybrid breeding systems dominates hybrid rice production in China7,8. The three-line system utilizes a cytoplasmic male-sterile (CMS) line, a restorer line and a maintainer line to produce hybrid seeds and to maintain the CMS line8,9. The restorer line carries special CMS-restorer genes to restore the fertility of the special CMS line. The germplasm resources related to cytoplasmic sterility are limited: only 1% of rice germplasms can serve as maintainer lines in China, and only 5% of rice germplasms carry CMS-restorer genes in Southeast Asia10. The limited genetic resources of the restorer lines and the low genetic biodiversity of CMS and restorer lines in the three-line system have prevented further developments. The two-line breeding system uses either photoperiod-sensitive genic male-sterile (PGMS) or thermo-sensitive genic male-sterile (TGMS) lines as sterility lines under the restrictive condition or maintainer lines under the permissive condition. Almost all normal rice cultivars can restore the male fertility of the PGMS and TGMS lines, thus providing broader genetic resources to better exploit heterosis in rice11,12,13,14,15. Therefore, compared with the three-line system, the advantages of the two-line system include labour and time savings, better grain quality and higher yields, greater effectiveness and economical use of simpler procedures for breeding and hybrid seed production. Although the two-line hybrid breeding system was developed relatively late, it provides essential advantages over the three-line system and occupies approximately one-third of the total hybrid rice planting area in China16.
In recent years, great progress has been achieved in the understanding of P/TGMS, and several genes that control P/TGMS traits have been cloned in rice. Nongken58S, the first PGMS rice identified in 1973, is characterized by complete male sterility under long-day conditions and fertility recovery under short-day conditions. Its PGMS trait is determined by pms1, pms2 and pms3 (refs 12 and 17). pms3 encodes a long non-coding RNA (lncRNA) called long day-specific male fertility-associated RNA (LDMAR)18. Peiai64S is a TGMS indica rice line that was developed by transferring P/TGMS genes from Nongken58S. Its TGMS trait is conferred by p/tms12-1. We found P/TMS12-1 encodes a unique noncoding RNA that produces a 21-nucleotide small RNA13. Carbon Starved Anther (csa) is a reverse photoperiod-sensitive genic male sterile (rPGMS) rice that is sterile under short day and fertile under long day conditions. CSA encodes a R2R3 MYB transcription regulator that mediates sugar partitioning14. In addition, temperature-sensitive splicing of UDP-glucose pyrophosphorylase1 (Ugp-1) caused by Ugp-1 overexpression leads to TGMS in rice19.
Annong S-1 (AnS-1) was the first indica TGMS rice identified in 1987. In a previous work15, we demonstrated that the TGMS gene tms5 encodes the endonuclease RNase ZS1 in AnS-1. RNase ZS1 controls the TGMS trait by degrading the temperature-sensitive ubiquitin fusion ribosomal protein L40 (UbL40) mRNA. The tms5-determined TGMS lines play a major role in the two-line hybrid breeding system in China20. Mutations in TMS5 were found in 24 of 25 commercial TGMS lines that were randomly detected15.
While the development of a new commercial male sterile line using traditional breeding systems usually takes several years, sometimes more than a decade, the breeding time can be dramatically reduced using modern techniques of genetic engineering21,22,23,24. Sequence-specific nucleases (SSNs) can induce targeted DNA double-stranded breaks (DSBs) at specific genomic loci and promote endogenous pathways of DNA damage repair, finally leading to sequence-specific genome editing25,26. As a type of SSN, the CRISPR/Cas9 editing system has been harnessed to knock out the targeted gene in many species, including plants25,26,27,28,29,30,31,32,33,34,35,36,37,38. Nevertheless, its application for crop genetic improvement remains rare39.
In this study, we established a simple and efficient rice TGMS cultivating system using CRISPR/Cas9 editing technology to knock out TMS5, which is of great value in new commercial TGMS line applications. Based on this system, we developed 11 commercial “transgene clean” TGMS rice lines within only one year. This work accelerates TGMS line breeding and lays the foundation for large-scale applications in two-line hybrid rice breeding.
Results
Development of TGMS lines using CRISPR/cas9 editing of TMS5
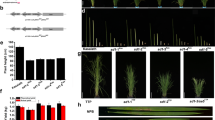
The CRISPR/Cas9 editing system mediates targeted genome editing through the complex of Cas9 endonuclease and guide RNA, which provides a simple and efficient technique compared with other genome engineering technologies, such as zinc-finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs)39,40. CRISPR/Cas9-induced editing events take place mainly in the T-DNA transformed callus cells before regeneration in rice26. Therefore, homozygous mutants with high mutation frequencies can be obtained in the T0 generation, and mutants without exogenous T-DNA can be isolated after segregation of the T1 generation. These mutants are also named “transgene clean” plants41. We used the CRISPR/cas9 editing system, which is able to target multiple sites via a single construct26, to create TMS5-targeted mutations to develop commercial TGMS lines. To obtain target sequences with high editing and low off-target efficiencies, 10 target sites (Supplementary Figure 1) screened from the TMS5 coding region were randomly divided into five groups, and the two target sequences in one group were cloned into the same vector to transform the ZH11 callus (see Methods). Transgenic plants were obtained based on five vectors. To evaluate the editing efficiencies of each target sequence, the target sites in the leaves of T0 plants were sequenced. Mutations were detected in 9 of 10 target sites. Among these nine mutation sites, the single mutation frequencies of the target sequence ranged from 46.2% to 88.2% and from 69.2% to 94.1% T0 plants contained mutations in at least one of the two target sites (Fig. 1A). Notably, the homozygotic mutation frequency for single target sequences reached 29.4% and the plants carrying homozygous mutations for at least one of two target sequences reached 32.4% (Fig. 1A). Furthermore, individuals with pollen sterility were obtained for all T0 plants under high-temperature (HT) treatment. The percentages of pollen-sterile individuals among the T0 plants ranged from 30.0% to 85.3% (Fig. 1A). TMS5b showed the highest editing, homozygous mutation, and pollen sterile frequencies (Fig. 1A). The above results indicated that the binary construct for targeting the TMS5a and TMS5b sites (named the TMS5ab construct) had the highest targeted editing efficiency.
Mutation types and frequencies in T0 plants with 10 target sites.
(A) Mutation and pollen sterility frequencies in T0 plants with 10 target sites. The target site of TMS5b displayed the highest mutation frequency. The TMS5ab construct gave rise to a higher percentage of plants with pollen sterility compared to the other constructs. The number preceding “,” is the total number or frequency of mutations, whereas the number following “,” is the number or ratio of homozygote plants. (B) CRISPR/Cas9-induced mutation types and frequencies. In all types of induced mutations, single-nucleotide insertions were most frequently detected. The largest deletion was 253 base pairs long. (C) TMS5ab construct-induced mutation frequencies in 11 cultivars of two different rice subspecies. Excluding YNSM, the mutation frequencies of TMS5a in the other ten cultivars were all higher than 72.72%.
To obtain TGMS plants, T0 sterile plants were treated at a low temperature (~22 °C, daily average temperature [DAT]). Plants with restored fertility were obtained from tranformants of each construct. Furthermore, we obtained “transgene clean” TGMS lines in T1 plants with each construct (Figs 2 and 3A and Supplementary Figure 2A). Taken together, we obtained “transgene clean” TGMS lines through the CRISPR/Cas9-mediated TMS5 site-directed mutagenesis system.
Pollen fertility of TGMS lines in the ZH11 background induced by CRISPR/Cas9 systems at high and low temperatures.
(A,B) Pollen fertility of ZH11. Normal pollen (A) in ZH11 at the HT. Normal pollen (B) in ZH11 at the LT. (C–M) Pollen fertility of TGMS plants induced by the CRISPR/Cas9-mediated TMS5 editing system. Abnormal pollen (C,E,G,I and K) in TMS5abS-12-3, TMS5cdS-1-8, TMS5efS-16-2, TMS5ghS-7-5 and TMS5ijS-7-9 at the HT. Normal pollen (D,F,H,J and M) in TMS5abS-12-3, TMS5cdS-1-8, TMS5efS-16-2, TMS5ghS-7-5 and TMS5ijS-7-9 at the LT. TMSabS-12-3, TMS5cdS-1-8, TMS5efS-16-2, TMS5ghS-7-5 and TMS5ijS-7-9 are TGMS plants induced by the TMS5ab, TMS5cd, TMS5ef, TMS5gh and TMS5ij constructs in the ZH11 background, respectively. The growth temperatures and plant names of the pollen are indicated on the left and top of the figure, respectively. HT, high temperature; LT, low temperature. Scale bars, 100 μm.
Plant morphologies of wild-type and TGMS plants at HT.
(A) Plant morphologies of ZH11 at the HT and TMS5abS-12-3 plants at the HT and LT. (B) Plant morphologies of ReB and ReBS-6-3 plants at the HT. (C) Plant morphologies of WSSM and WSSMS-2-5 plants at the HT. (D) Plant morphologies of TFB and TFBS-5-7 plants at the HT. (E) Plant morphologies of ZZB and ZZBS-10-2 plants at the HT. While exhibiting pollen sterility, these “transgene clean” TGMS plants had no obvious phenotypic variation from their hosts. HT, high temperature; LT, low temperature. Scale bars, 20 cm.
Previous reports have shown that CRISPR/Cas9 technology has a relatively high potential for off-target activity, which affects its application. The specificity of the seed sequence, including the 10–12 nucleotides close to the protospacer adjacent motif (PAM), has a significant impact on the off-target activity42. The TMS5ab construct has the highest efficiency among the constructs used for CRISPR/Cas9-mediated TMS5 editing; therefore, we analysed the off-target efficiency of TMS5ab. We predicted five and nine potential off-target sites for TMS5a and TMS5b, respectively. After sequencing those potential off-target sites, no genome editing was detected in any examined samples (Table 1). These results indicated that the TMS5ab construct had the highest editing efficiency and an undetectable off-target effect. The target sequences with high editing and low off-target efficiencies will improve the breeding efficiency and reduce the breeding cost associated with the development of commercially applied TGMS lines. Therefore, we exploited the TMS5ab construct for the transformation of other rice varieties for commercially applied TGMS lines.
Editing characteristic of TMS5 and factors influencing editing efficiency
Next, we analysed the mutation types and frequencies of the nine effective target sequences (Fig. 1B). Among all types of induced mutations, single-nucleotide insertions were most frequently detected, with a frequency up to 51.96%, consistent with previous reports25,26. The second highest frequency of mutation was nucleotide deletion, in which single-nucleotide deletions showed the highest mutation rate of 10.29%. Together with the increased number of deleted nucleotides, the mutation frequency of the target site became gradually reduced. The longest deletion fragment was 253 bp in length. Nucleotide substitutions were also observed, accounting for only 2.94% of all mutations.
Next, we analysed the factors that affect editing efficiency. First, we analysed the GC content of the target sequences because targeted genome editing relies on the binding of the single-guide RNA (sgRNA) containing the target sequence to the target sites. By comparing the GC contents in the target sequences and the corresponding editing efficiency, we found that a high GC content is correlated with a high target sequence editing efficiency (Fig. 1A and Supplementary Figure 3). We then proceeded to test the sgRNA and Cas9 expression levels from the leaves of T0 plants carrying different target sequences (Fig. 4). The Cas9 expression levels were lowest in TMS5abS-4 plants, which displayed modified TMS5b target sites and no modified TMS5a sites. In contrast, the Cas9 expression levels in TMS5efS-2, TMS5ghS-2 and TMS5ijS-1 plants were 14.1, 15.3 and 58.1 times larger than that in TMS5abS-4, respectively. However, no genome editing occurred in any of the target sites in these three lines. A similar phenomenon was also observed for the relationship between the target and the sgRNA and the editing efficiency. This finding suggested that once the sgRNA and Cas9 expression levels reach a certain threshold value, there is no obvious correlation between their expression levels and the editing efficiencies. Furthermore, because PAM is essential for identification of the target site by the target sequence-sgRNA, we analysed the effects of different PAMs on the editing efficiencies of the target sites (Fig. 1A and Supplementary Table 1). The average editing efficiencies of the target site PAMs with AGG (TMS5c, d and f), TGG (TMS5b, g, h and j), and CGG (TMS5a and i) were 80.6%, 68.8%, and 70%, respectively. Although AGG displayed a higher average editing efficiency compared with TGG and CGG, TMS5b, which contained TGG as the PAM, displayed the highest editing efficiency among all nine target sequences. These findings indicated that the type of target sequence PAMs had no significant effect on the editing efficiency. We also found that the TMS5e target sequence was highly complementary to the sgRNA, forming a secondary structure that was likely to inhibit binding of the target sequence to the genome (Supplementary Figure 4). This was the plausible cause of the genome editing failure and is supported by previous studies25,26.
Effects of Cas9 and target-sgRNA expression levels on target site editing.
(A) Cas9 expression levels in T0 plants (TMS5abS-1, TMS5abS-2, TMS5abS-4, TMS5abS-5, TMS5cdS-1, TMS5cdS-11, TMS5cdS-12, TMS5cdS-13, TMS5efS-1, TMS5efS-2, TMS5efS-7, TMS5efS-9, TMS5ghS-1, TMS5ghS-2, TMS5ijS-1 and TMS5ijS-11) assessed by real-time PCR analysis. (B) Target-sgRNA expression levels in T0 plants (TMS5abS-4, TMS5efS-7, TMS5efS-9, TMS5ghS-2, TMS5ijS-1 and TMS5ijS-11) assessed by real-time PCR analysis. The target sequence names are indicated above the column. (C) The mutation types in T0 plants (TMS5abS-1, TMS5abS-2, TMS5abS-4, TMS5abS-5, TMS5cdS-1, TMS5cdS-11, TMS5cdS-12, TMS5cdS-13, TMS5efS-1, TMS5efS-2, TMS5efS-7, TMS5efS-9, TMS5ghS-1, TMS5ghS-2, TMS5ijS-1 and TMS5ijS-11). He, heterozygote; Ho, homozygote; WT, wild type.
Development of commercial TGMS lines in rice
The TMS5ab construct established in our study showed promising utilization potential. To develop the applied TGMS lines, we selected 11 fertile elite cultivars of two different rice subspecies (including seven maintainer lines of the three-line system, three indica and one japonica conventional variety) as the hosts for the targeted genome editing using the TMS5ab construct over the course of only one year (Fig. 1C). We obtained various numbers of T0 transgenic plants in all 11 cultivars. Excluding the identification of only one YNSMS (Yuenongsimiao (YNSM) transformed with TMS5ab) plant carrying the edited TMS5a, the editing efficiencies among the other cultivars ranged from 72.72% to 100%. While no homozygous TMS5a mutation was found in YNSMS, the homozygous mutation rates of the other cultivars ranged from 11.1% to 54.55%. These results indicated that we had produced TMS5-targeted mutation plants in all 11 cultivars and that the editing efficiency of the CRISPR/Cas9 system may vary depending on the genetic backgrounds of the different cultivars. After LT treatment of the T0 sterile individuals, plants with restored fertility were selected.
To obtain TGMS plants without exogenous T-DNA, plants with restored fertility from 11 cultivars were planted under high temperature (HT) conditions. Additionally, sterile plants without hygromycin phosphotransferase (HPT) and Cas9 were obtained by PCR and Southern blot detection (Supplementary Figure 2B,C and Supplementary Table 2). These “transgene clean” sterile plants were grown under LT conditions, and T2 seeds were obtained. The “transgene clean” TGMS lines segregated in the T1 generation. While exhibiting similar pollen sterility, these “transgene clean” TGMS plants displayed no obvious phenotypic changes compared with their hosts (Fig. 3B–E and Supplementary Figure 5B). The phenotypes of these TGMS lines as a consequence of genome editing were stable in the T2 and T3 generations. To obtain TGMS lines that were applicable to two-line hybrid breeding, we next treated these TGMS lines with different temperature gradients and found that fertility was gradually reduced with increasing temperature (Fig. 5). TGMS lines of ZZBS and YJSMS showed relatively low critical sterility-inducing temperature (CSIT), i.e., below 24 °C (DAT), while in the ReBS, TFBS and WSSMS TGMS lines, the CSIT were all about 24 °C (DAT). The CSIT of the GAZS TGMS lines was approximately 26 °C (DAT), and the CSIT of ZS97BS exceeded 26 °C (DAT). ZZBS, ReBS, TFBS, WSSMS and YJSMS exhibited a pollenless sterile type, whereas ZS97BS and GAZS displayed completely abortive pollen at 28 °C (DAT). These results demonstrated that CSIT was regulated by the rice genetic background. The populations of “transgene clean” YJSMS were grown in the experimental plots of the China National Hybrid Rice R&D Center (Changsha, China) in the summer of 2015 and they displayed pollenless male sterility (Supplementary Figure 5). To confirm that TMS5 editing would not lead to female organ abortion, GAZS and YJSMS were pollinated with normal pollen. The sterile plants of GAZS and YJSMS yielded seed settings via artificial pollination (Supplementary Figure 6), indicating normal female organ development and fertilization functions. These results indicated that these lines could be used as male sterile lines, which are almost indistinguishable from traditional male sterile lines. Taken together, the findings suggest that practical TGMS lines with good agronomic characteristics can be developed by targeted editing of TMS5 using the CRISPR/Cas9 system.
Pollen fertility of TGMS lines induced by the TMS5ab construct at different temperatures.
ZZBS-10-2, ReBS-6-3, TFBS-5-7, WSSMS-2-5, YJSMS-2-4, ZS97BS-8-7 and GAZS-9-9 are the TGMS plants induced by the TMS5ab construct in the ZZB, ReB, TFB, WSSM, YJSM, ZS97B and GAZ backgrounds, respectively. These TGMS plants were grown at 22 °C, 24 °C, 26 °C, and 28 °C under photoperiod conditions of 13.5 h of light and 10.5 h of darkness. ZZBS-10-2 and YJSMS-2-4 plants were completely sterile at 24 °C (DAT); ReBS-6-3, TFBS-5-7 and WSSMS-2-5 plants were almost sterile at 24 °C (DAT); GAZS-9-9 plants were sterile at approximately 26 °C (DAT); and ZS97BS-8-7 plants were sterile at greater than 26 °C (DAT). The TGMS plant names and growth temperatures of the pollen are indicated at the left and top of the figure, respectively. Scale bars, 100 μm.
To test the combinatorial capacity of the obtained TGMS lines, we crossed HNBS with Huahang1179 (HH1179), YJSMS with R94, YJSMS with R340, YJSMS with R297, YJSMS with R173, WSSMS with R364, WSSMS with R173, WSSMS with R340, and WSSMS with R297. Subsequently, we obtained the F1 generation of HNBS and HH1179 and found that the offspring were stronger and provided a greater yield than those of the control variety (Supplementary Figure 7). In addition, the hybrid demonstrated a 31.4% increase in the main panicle length, a 47.9% increase in grain number per main spike, a 19.8% increase in thousand seed weight, a 14.4% increase in single plant yield, and a 15.9% increase in plot yield (Supplementary Table 3). Taken together, these results indicated that TGMS lines with TMS5 targeted editing using the CRISPR/Cas9 system could be applied to improve production in hybrid rice breeding.
Discussion
In recent years, RNA interference (RNAi) and antisense approaches have become more commonly used in reverse genetics systems40. In our previous work, we knocked down TMS5 expression using RNAi. Although homozygous TGMS plants were generated, sterile heterozygotes under high temperatures were simultaneously obtained15. Therefore, sterile heterozygous TGMS plants cannot be used for hybrid rice breeding. We then knocked down TMS5 expression using an antisense RNA technique and obtained very low rates of TGMS plants with a relatively high CSIT (Supplementary Figure 8), potentially due to incomplete suppression of TMS5. This result suggested that both RNAi and antisense RNA technology provide sequence-dependent post-transcriptional inhibition of gene expression, resulting in incomplete depletion of target genes40. Nevertheless, TGMS lines with low CSIT guarantee seed production in two-line hybrid rice breeding43. Importantly, using a modified sgRNA to cleave a target gene exon via DSBs and nonhomologous end-joining (NHEJ) repair in the CRISPR/Cas9 system, knockout genotypes can be deliberately generated in plants. CRISPR/Cas9-mediated site-directed mutagenesis produced a loss of function of TMS5 and generated TGMS rice with low and stable CSIT (Fig. 5). Moreover, regardless of the presence of the CRISPR/Cas9 transgene, the target gene mutations could be stably transmitted to future generations41. Therefore, “transgene clean” mutants could be obtained in the T1 generation (Figs 2, 3 and 5 and Supplementary Figure 2). In summary, the mutations generated using this strategy are essentially similar to the spontaneous or induced mutations that occur in conventional breeding, in addition to the advantage that the objective trait of CRISPR/Cas9 editing in plants can be deliberately improved39.
Currently, hybrid breeding is an important approach for the improvement of rice yield in China6. Several years or decades were required to obtain and select sterile lines under specific temperature and light conditions in the two-line breeding system21,22,23,24. In contrast, only one year was required to produce “transgene clean” tms5 TGMS plants in the T1 generation using the CRISPR/Cas9-mediated site-directed mutagenesis system. Commercial sterile lines are mainly produced using the available sterile lines as the sterile gene donor (female parent) in traditional cross and backcross breeding programs21. However, there are limited genetic resources for the intensive use of a single source of male-sterile cytoplasm, and variety collapse of a source of sterile cytoplasm is considered to be disastrous in developing hybrids44. Therefore, by developing TGMS lines in different genetic backgrounds, the genetic diversity—especially the cytoplasm diversity—can be increased, improving the efficiency of the exploitation of heterosis. The methodology applied in our work to produce TGMS lines will mitigate limitations related to traditional hybrid breeding.
The production of hybrid rice using the three-line system involves CMS, maintainer and restorer lines. Commercial CMS lines can be mainly categorized into three groups: wild-abortive (WA), Honglian (HL) and Baotai (BT)45. The utilization of different types of three-line CMS lines is restricted because special CMS-restorer genes are required from the male parents, and the germplasms of maintainers and restorers are not abundant. For example, approximately 5% of the rice germplasms can serve as restorer lines in China, Southeast Asia and America10,46,47. The CMS and maintainer lines possess the same nucleus but different cytoplasms8. In the present study, we used CRISPR/Cas9-mediated editing of nuclear genes encoding TMS5 in maintainer lines to obtain TGMS lines with extremely similar characteristics to their corresponding CMS lines. Almost all of the normal varieties with or without specific CMS-restorer genes could restore the fertility of TGMS lines12,14. This method retains all of the merits of CMS lines, avoids the problems caused by limited genetic resources and improves the heterosis utilization efficiency. Therefore, we termed this direct and efficient TGMS breeding method as “turning three-line into two-line”.
The breeding of japonica hybrid rice has not been widely applied due to a lack of male-sterile lines with high agronomic value. The japonica hybrid rice occupies approximately only 3% of the japonica rice planting area in China10,48. The promotion of japonica hybrid rice breeding will be a breakthrough in rice production in China. The sterility gene in japonica P/TGMS lines has been derived mainly from Nongken58S by employing a single genetic background, and it possesses the characteristics of high CSIT43. Nevertheless, it will be very convenient to use CRISPR/Cas9-mediated TMS5 editing to cultivate sterile lines of japonica or wide-compatibility sterile line breeding to improve japonica or japonica-indica hybrid rice breeding48.
In two-line hybrid breeding, the cultivation of sterile lines with low CSIT is the key for ensuring the purity of hybrid seeds43. We tested CSITs of 7 TGMS lines generated using the TMS5ab editing system (Fig. 5) and found that varieties with different genetic backgrounds possessed different CSITs and that most of them met the requirements of breeding programs. These results suggested that the CSIT of tms5 TGMS lines was determined according to their genetic backgrounds but not tms5. Furthermore, TGMS lines with higher CSITs could be crossed with lower CSIT lines to select new TGMS lines with lower CSITs.
In the present study, we selected 10 target sequences in the TMS5 coding region to examine the efficiency of inducing target site editing and found that the efficiency of TMS5b was highest (up to 88.2%). Among the five constructs, the editing efficiency of TMS5ab was as high as 94.1%, and the homozygous rate of one of the target sequences was as high as 32.4%. These findings showed that the production of “transgene clean” TGMS plants was very convenient using the CRISPR/Cas9 system in the T1 generation and that the breeding efficiency could be markedly improved. Furthermore, the editing efficiency of the CRISPR/Cas9 system was affected by several factors. As previously reported25,26, targets with higher GC contents demonstrated relatively higher editing efficiencies (Fig. 1A and Supplementary Figure 3). In addition, as previously described26,49, a more stable secondary structure resulted in imperfect complementary base pairing of the target sequence and the target site (Fig. 1A and Supplementary Figure 4). As a result, the target sequence with a less stable secondary structure with sgRNAs tends to be selected in this system49. We also noted that the editing efficiency of the same target sequence was altered in different rice varieties (Fig. 1C). This observation may be due to the reduced activity of the CRISPR/Cas9 editing system in some varieties or to the higher percentage of faultless repairs after DSB25. The sgRNA expression level might be the limiting factor for target site editing in Arabidopsis26. However, when the sgRNA and Cas9 expression levels reached certain thresholds, their expression may not be relevant to the target site editing efficiency in rice.
In the present study, we designed 10 target sequences in the TMS5 gene and selected the most optimal TMS5ab construct for the development of commercial TGMS lines by comparing the editing efficiency and analysing the off-target efficiency of the target sequences. Using the TMS5ab construct, the TMS5 gene was edited in 11 elite ice lines to develop the “transgene clean” TGMS lines within only one year. This method can shorten the breeding period, reduce the working complexity and cost, and facilitate the exploitation of heterosis. Our work will greatly promote the breeding process of TGMS lines and will maximize the number of cross combinations of three-line sterile lines, which will facilitate the development of hybrid rice breeding. This work can be applied for not only hybrid rice breeding but also potentially breeding of other hybrid crops.
Methods
Plant materials, growth conditions and generation of transgenic plants
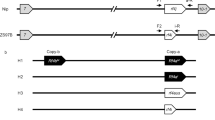
TMS5as was derived from zhonghua11 (ZH11) by knocking down TMS5 using anti-sense RNA technology. TMS5abS, TMS5cdS, TMS5efS, TMS5ghS and TMS5ijS were derived from ZH11 by the transformation of TMS5ab, TMS5cd, TMS5ef, TMS5gh, and TMS5ij vectors, respectively, via Agrobacterium-mediated transformation. ZS97BS, HNBS, TFBS, YXBS, ReBS, HHBS, ZZB, YJSMS, YNSMS, WSSMS, and GAZS were derived from the transformation of the TMS5ab vector using Zhenshan97B (ZS97B), HuanongB (HNB), TianfengB (TFB), YixiangB (YXB), ReB, HuahuiB (HHB), ZhongzheB (ZZB), Yuejingsimiao (YJSM), Yuenongsimiao (YNSM), Wushansimiao (WSSM), and GAZ as hosts, respectively. ZH11 and GAZ are conventional Japonica rice; Zhenshan97B (ZS97B), HuanongB (HNB), TianfengB (TFB), YixiangB (YXB), ReB, HuahuiB (HHB) and ZhongzheB (ZZB) are “three-line” indica maintainer lines. Yuejingsimiao (YJSM), Yuenongsimiao (YNSM), and Wushansimiao (WSSM) are conventional indica rice. Transgenic plants were generated using Agrobacterium-mediated transformation as previously described13,15. Unless indicated, all rice plants were grown in the field at the South China Agricultural University, Guangzhou (23_N, 113_E), during normal rice growing seasons, or in a growth chamber at DATs of 22, 24, 26, 28 and 30 °C with three replicates as previously described15.
Vector constructs
The TMS5as vector was constructed as previously described50. An antisense fragment from TMS5 was amplified and cloned into pYLox vector15. CRISPR/Cas9 was constructed as previously described26. Briefly, according to the design principles of the target sequences in the CRISPR/Cas9 system, 19 to 20 bases upstream of the PAM motif were selected as candidate target sequences, and target sequences in TMS5 were determined for TMS5a, TMS5b, TMS5c, TMS5d, TMS5e, TMS5f, TMS5g, TMS5h, TMS5i and TMS5j (Supplemental Table 4) by Blast analysis of the rice genome (http://blast.ncbi.nlm.nih.gov/Blast.cgi) to ensure that no other genes were targeted. Equivalent amounts of forward and reverse primers (0.05–0.1 μM) for one target were mixed and incubated at 90 °C for 1 min, followed by a gradual cool-down to room temperature to form the target adaptor. Approximately 1 μg each of the pYLgRNA-OsU6a and pYLgRNA-OsU3 vectors was digested using 10 U Bsa I (NEB), respectively. The ligation reaction (10 μl) contained 1 μ1 of 10 x T4 DNA ligase buffer, 20 ng of pYLgRNA-OsU6a or pYLgRNA-OsU3 vector, 1 μ1 of the target sequence adaptor, and 35 U of T4 DNA ligase (Takara, Dalian, China). The target sequence adaptors TMS5a, TMS5c, TMS5e, TMS5g and TMS5i, respectively, were ligated into the linearized pYLgRNA-OsU6a vector (digested with 10 U of Bsa I), and TMS5b, TMS5d, TMS5f, TMS5h, and TMS5j, respectively, were cloned into pYLgRNA-OsU3 to generate sgRNA expression cassettes with target sequences. The sgRNA expression cassettes with the target sequences TMS5a, TMS5c, TMS5e, TMS5g and TMS5i were amplified from pYLgRNA-OsU6a using primers Pps-GGL and Pgs-GG2 and the sgRNA expression cassettes with target sequences TMS5b, TMS5d, TMS5f, TMS5h, and TMS5j were amplified from pYLgRNA-OsU3 using primers Pps-GG2 and Pgs-GGR with High-Fidelity DNA polymerase KOD-Plus (Toyobo, Osaka, Japan) over 30 cycles (94 °C, 30 s; 58 °C, 30 s; 68 °C, 20 s). The sgRNA expression cassettes with target sequences and binary plasmid pYLCRISPR/Cas9Pubi-H were digested with Bsa I and subsequently purified. Ten microliters of the reaction solution consisted of 1 μl of 10 x T4 DNA ligase buffer, 15 ng of purified sgRNA expression cassettes of TMS5a and TMS5b, 60 ng of pYLCRISPR/Cas9Pubi-H, and 70 U of T4 DNA ligase (Takara, Dalian, China). The TMS5ab vector was constructed by ligating sgRNA expression cassettes of TMS5a and TMS5b to the binary plasmid pYLCRISPR/Cas9Pubi-H. The same procedures were used for the TMS5cd, TMS5ef, TMS5gh, and TMS5ij constructs as for TMS5ab. The relevant PCR primers are listed in Supplementary Table 4.
Mutation detection and analysis of transgenic plant lines
To determine the mutation of target sites, genomic DNA from the leaves of transgenic rice plants was extracted using the sodium dodecyl sulphate method51. PCR amplifications were performed using primer pairs (Supplemental Table 4) surrounding the designed target sites. The PCR products were directly sequenced or cloned into the pEASY-Blunt (TransGen Biotech, Beijing, China) vector and sequenced using the Sanger method. Mutations were identified by comparing the sequences of transgenic and WT plants. The sequencing chromatograms from mutations were analysed, and mutations containing normal sequencing chromatograms were considered homozygotes. Mutations containing superimposed sequencing chromatograms were considered heterozygous or biallelic mutations. Heterozygous sequences from direct sequencing were decoded using the degenerate sequence decoding method52. The relevant PCR primers are listed in Supplementary Table 4.
The identification of “transgene clean” plants was conducted using T1 progeny of T0 homozygous mutations. The transgenic lines were analysed by PCR using HPT and Cas9 primers and agarose gel electrophoresis. The pYLCRISPR/Cas9Pubi-H plasmids and ZH11 DNA were selected as positive and negative controls, respectively. HPT- and Cas9-negative plants were considered “transgene clean” plants. Some “transgene clean” plants identified by PCR were further confirmed by Southern blotting. Southern blots were performed using a DIG-High Prime DNA Labeling and Detection Starter Kit I (Roche Diagnostics, Mannheim, Germany) according to a standard program supplied by the manufacturer. In brief, PCR products of HPT were labelled with GIG-dUTP by random priming (Roche Diagnostics) and used as hybridization probes. Genomic DNA from the leaves of rice plants was extracted using the Cetyltrimethylammonium bromide (CTAB) method. Genomic DNA was digested overnight using EcoRI (Takara, Dalian, China) and then separated by 0.8% agarose electrophoresis and transferred onto Hybond N+ nylon membranes (Amersham Bioscience, Bucks, UK) using the capillary transfer method. Hybridization was performed at 65 °C in DIG Easy Hyb buffer. The membranes were washed twice with wash buffer (2 x SSC, and 0.1% SDS) at 68 °C and then blocked with blocking solution and incubated with anti-digoxigenin-AP solution for 30 min. Subsequently, the membranes were washed twice with wash buffer (0.1 M maleic acid, 0.15 M NaCl, 0.3% (v/v) Tween 20, pH 7.5) at room temperature for 15 min each, followed by equilibration with detection buffer (100 mM Tris-Cl, 100 mM NaCl, pH 9.5) for 2–5 min. One millilitre of CSPD ready-to-use solution was added to the membranes to stimulate chemiluminescence. The chemiluminescence signals were detected and analysed using X-ray films.
Expression analyses
Total RNA was isolated from the leaves or panicles of T0 rice plants using TRIzol (Invitrogen, Carlsbad, CA, USA). DNase I-treated total RNA (2.0 μg) was used for reverse transcription using an M-MLV-RT kit (Takara, Dalian, China) with a mixture of oligo(dT) and sgRNA RT primers (Supplemental Table 4). Real-time quantitative PCR was performed with gene- and target-specific primers (Supplemental Table 3) using the SsoFast™ EvaGreen Supermix kit (Bio-Rad, USA) and CFX96 Real-Time PCR Detection System (Bio-Rad) as previously described15. Three replicates were evaluated for each gene. The internal standard gene OsActin1 was used to normalize the cDNA levels of the target genes. Total protein was extracted from rice panicles, and immunoblot analysis was performed using a previously described method15. HSP8253 (Code: AbM51099-31-PU; BPI, China) was used as the internal standard protein. The relevant PCR primers are listed in Supplementary Table 4.
RNA secondary structure analysis
Secondary structure analysis of target-sgRNA sequences was performed with the RNA Folding Form (http://unafold.rna.albany.edu/?q=mfold/RNA-Folding-Form).
Characterization of phenotypes
Rice florets were photographed using an OLYMPUS DP70 digital camera under an OLYMPUS SZX10 dissecting microscope. Mature pollen grains were stained using 1% I2–KI solution and photographed using an OLYMPUS BX51 or Leica DNRXA microscope.
Additional Information
How to cite this article: Zhou, H. et al. Development of Commercial Thermo-sensitive Genic Male Sterile Rice Accelerates Hybrid Rice Breeding Using the CRISPR/Cas9-mediated TMS5 Editing System. Sci. Rep. 6, 37395; doi: 10.1038/srep37395 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Miura, K. et al. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat Genet 42, 545–549 (2010).
Kusano, M. et al. Using metabolomic approaches to explore chemical diversity in rice. Mol Plant 8, 58–67 (2015).
Khush, G. S. What it will take to feed 5.0 billion rice consumers in 2030. Plant Mol Biol 59, 1–6 (2005).
Su, N. et al. Disruption of a rice pentatricopeptide repeat protein causes a seedling-specific albino phenotype and its utilization to enhance seed purity in hybrid rice production. Plant Physiol 159, 227–238 (2012).
Normile, D. Agricultural research. Reinventing rice to feed the world. Science 321, 330–333 (2008).
Yuan, L. Development of hybrid rice to ensure food security. Rice Science 21, 1–2 (2014).
Cheng, S. H., Zhuang, J. Y., Fan, Y. Y., Du, J. H. & Cao, L. Y. Progress in research and development on hybrid rice: a super-domesticate in China. Ann Bot 100, 959–966 (2007).
Chen, L. & Liu, Y. G. Male sterility and fertility restoration in crops. Annu Rev Plant Biol 65, 579–606 (2014).
Lin, S. C. & Yuan, L. P. Hybrid rice breeding in China, In Innovative Approaches to Rice Breeding (Manila, Philippines: International Rice Research Institute), 35–51 (1980).
Deng, X. W. et al. Hybrid rice breeding welcomes a new era of molecular crop design. Sci Sin: Life Sci 864–868 (2013).
Yuan, L. P. Progress of two-line system hybrid rice breeding. Sci Agr Sin 23, 1–6 (2013).
Zhang, Q. et al. Using bulked extremes and recessive class to map genes for photoperiod-sensitive genic male sterility in rice. Proc Natl Acad Sci USA 91, 8675–8679 (1994).
Zhou, H. et al. Photoperiod- and thermo-sensitive genic male sterility in rice are caused by a point mutation in a novel noncoding RNA that produces a small RNA. Cell Res 22, 649–660 (2012).
Zhang, H. et al. Mutation in CSA creates a new photoperiod-sensitive genic male sterile line applicable for hybrid rice seed production. Proc Natl Acad Sci USA 110, 76–81 (2013).
Zhou, H. et al. RNase ZS1 processes UbL40 mRNAs and controls thermosensitive genic male sterility in rice. Nat Commun 5, 4884 (2014).
Huang, J. Z., E., Z. G., Zhang, H. L. & Shu, Q. Y. Workable male sterility systems for hybrid rice: Genetics, biochemistry, molecular biology, and utilization. Rice 7, 13 (2014).
Liu, N. et al. Identification of an 85-kb DNA fragment containing pms1, a locus for photoperiod-sensitive genic male sterility in rice. Mol Genet & Genomics 266, 271–275 (2001).
Ding, J. et al. A long noncoding RNA regulates photoperiod-sensitive male sterility, an essential component of hybrid rice. Proc Natl Acad Sci USA 109, 2654–2659 (2012).
Chen, R. et al. Rice UDP-glucose pyrophosphorylase1 is essential for pollen callose deposition and its cosuppression results in a new type of thermosensitive genic male sterility. Plant Cell 19, 847–861 (2007).
Zhang, H. L. et al. Identification and transition analysis of photo-/thermo-sensitive genic male sterile genes in two-line hybrid rice in China. Sci Agr Sin 48, 1–9 (2015).
Luo, X. H. & Qiu, Z. Z. Pei-Ai64S-a dual-purpose sterile line whose sterility is induced by low critical temperature. Hybrid Rice 7, 27–29 (1992).
Yang, Z. et al. Breeding of fine quality PTGMS line Guangzhan 63S in medium indica rice. Hybrid Rice 17, 4–6 (2002).
Deng, Q. Y. Breeding of the PTGMS line Y58S with wide adaptability in rice. Hybrid Rice 20, 15–18 (2005).
LU, Z. M. Highlights and difficulties in hybrid rice breeding in China. Chinese Journal of Rice Science 25, 231–235 (2011).
Zhang, H. et al. The CRISPR/Cas9 system produces specific and homozygous targeted gene editing in rice in one generation. Plant Biotechnol 12, 797–807 (2014).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8, 1274–1284 (2015).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Feng, Z. et al. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res 23, 1229–1232 (2013).
Hwang, W. Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol 31, 227–229 (2013).
Jiang, W. et al. Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res 41, e188 (2013).
Li, J. F. et al. Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31, 688–691 (2013).
Miao, J. et al. Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res 23, 1233–1236 (2013).
Shan, Q. et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 31, 686–688 (2013).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Feng, Z. et al. Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proc Natl Acad Sci USA 111, 4632–4637 (2014).
Shan, Q., Wang, Y., Li, J. & Gao, C. Genome editing in rice and wheat using the CRISPR/Cas system. Nat Protoc 9, 2395–2410 (2014).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2013).
Li, Q. L. et al. Development of japonica photo-sensitive genic male sterile rice lines by editing carbon starved anther using CRISPR/Cas9. J Genet Genomics doi: 10.1016/j.jgg.2016.04.011.
Huang, S., Weigel, D., Beachy, R. N. & Li, J. A proposed regulatory framework for genome-edited crops. Nat Genet 48 109–111 (2016).
Boettcher, M. & McManus, M. T. Choosing the right tool for the Job: RNAi, TALEN, or CRISPR. Mol Cell 58, 575–585 (2015).
Xu, R. F. et al. Generation of inheritable and “transgene clean” targeted genome-modified rice in later generations using the CRISPR/Cas9 system. Sci Rep 5, 11491 (2015).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat Biotechnol 31, 822–826 (2013).
Lei, D., Tang, W., Xie, Z., Liu, H. & Chen, L. Solutions to insecurity problems in seed production of two-line hybrid rice. Agr Sci & Technol 15, 1160–1166 1187 (2014).
Levings, C. R. The Texas cytoplasm of maize: cytoplasmic male sterility and disease susceptibility. Science 250, 942–947 (1990).
Huang, Q., He, Y., Jing, R., Zhu, R. & Zhu, Y. Mapping of the nuclear fertility restorer gene for HL cytoplasmic male sterility in rice using microsatellite markers. Chinese Sci Bull 45, 430–432 (2000).
Zhong, B. Q., Yang, Z. L. & He, G. H. Analysis of restoring and maintaining ability of thirty-three American rice varieties (Oryza Sativa L.). Journal of Southwest Agricultural University 26, 554–555 559 (2004).
Ruan, R. C., Chen, H. C., Yang, Y. S., Jin, T. Y. & You, J. M. Screening of rice (Oryza sativa) germplasm with restoring genes to CMS from local varieties collected in Guizhou. Guizhou Agr Sci 27, 34–37 (1999).
Deng, H. F. et al. Status and technical strategy on development of japonica hybrid rice in China. Hybrid Rice 21, 1–6 (2006).
Makarova, K. S. et al. Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol 9, 467–477 (2011).
Li, J. et al. Expression of RNA-interference/antisense transgenes by the cognate promoters of target genes is a better gene-silencing strategy to study gene functions in rice. PLoS One 6, e17444 (2011).
Dellaporta, S. L., Wood, J. & Hicks, J. B. A plant DNA minipreparation: Version II. Plant Mol Biol Rep 1, 19–21 (1983).
Ma, X., Chen, L., Zhu, Q., Chen, Y. & Liu, Y. G. Rapid decoding of sequence-specific nuclease-induced heterozygous and biallelic mutations by direct sequencing of PCR products. Mol Plant 8, 1285–1287 (2015).
Li, X. et al. Identification and validation of rice reference proteins for western blotting. J Exp Bot 62, 4763–4772 (2011).
Acknowledgements
We thank Prof Yao-Guang Liu (South China Agriculture University, Guangzhou) for kindly providing the plant binary vector pYLCRISPR/Cas9Pubi-H and the pYLgRNA plasmids. This work was supported by the National Basic Research Program of China (grant nos 2013CB126905 and 2013CBA01401to C.Z.), Genetically Modified Breeding Major Projects (grant nos 2016ZX08001-004-001-003 to C.Z.), the National Natural Science Foundation of China (grant nos 31301287 and 31571255 to H.Z.) and the Scientific Research Project of Guangzhou City (grant nos 201504010004 to C.Z.).
Author information
Authors and Affiliations
Contributions
H.Z. performed most of the experiments; M.H. performed the field experiments and DNA sequencing; J.L. organized the figures; L.C. constructed some vectors; Z.H., S.Z., L.Z., D.J., E.N., and B.Z. participated in the vector construction, plant transformation and transgenic plant analysis; and C.Z. conceived the experiments, designed the study and wrote the paper. All authors commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhou, H., He, M., Li, J. et al. Development of Commercial Thermo-sensitive Genic Male Sterile Rice Accelerates Hybrid Rice Breeding Using the CRISPR/Cas9-mediated TMS5 Editing System. Sci Rep 6, 37395 (2016). https://doi.org/10.1038/srep37395
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep37395
This article is cited by
-
CRISPR-Cas9 based molecular breeding in crop plants: a review
Molecular Biology Reports (2024)
-
A natural allele of OsMS1 responds to temperature changes and confers thermosensitive genic male sterility
Nature Communications (2022)
-
CRISPR/Cas tool designs for multiplex genome editing and its applications in developing biotic and abiotic stress-resistant crop plants
Molecular Biology Reports (2022)
-
Salinity tolerance mechanisms and their breeding implications
Journal of Genetic Engineering and Biotechnology (2021)
-
Genome editing with CRISPR/Cas9 in Pinus radiata (D. Don)
BMC Plant Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.