Abstract
In Arabidopsis, the 24-nucleotide (nt) small interfering RNAs (siRNAs) mediates RNA-directed DNA methylation (RdDM) and transcriptional gene silencing (TGS) of transposable elements (TEs). In the present study, we examined genome-wide changes in DNA methylation and siRNA accumulation in Arabidopsis induced by expression of the Cucumber mosaic virus silencing suppressor protein 2b known to directly bind to both the 21/24-nt siRNAs as well as their associated Argonaute proteins. We demonstrated a genome-wide reduction of CHH and CHG methylation in the 2b-transgenic plants. We found that 2b suppressed RdDM not only at the previously annotated loci directed by 24-nt siRNAs but also a new set of loci associated with 21/22-nt siRNAs. Further analysis showed that the reduced methylation of TEs and coding genes targeted by 21/22-nt siRNAs was associated with sequestration of the duplex siRNAs by the 2b protein but not with changes in either siRNA production or transcription. Notably, we detected both the deletion and/or the transposition of multicopy TEs associated with 2b-induced hypomethylation, suggesting potential TE reactivation. We propose that the silencing of many TEs in Arabidopsis is controlled by the 24- and 21-nt endogenous siRNAs analogous to Drosophila TE silencing by PIWI-interacting RNAs and siRNAs.
Similar content being viewed by others
Introduction
The methylation of cytosines in nuclear DNA is a conserved epigenetic silencing mechanism and controls many important biological processes, including defense against transposon proliferation, control of genomic imprinting, and the regulation of gene expression, which imparts an additional layer of heritable information upon the DNA code1,2,3,4. In plants, three main DNA methylation pathways mediate the methylation of CG, CHG, and CHH (where H = A/T/C) sequence contexts. METHYLTRANSFERASE 1 (MET1) is responsible for the maintenance of CG methylation, and CHG methylation is maintained by CHROMOMETHYLASE 3 (CMT3). DOMAINS REARRANGED METHYLTRANSFERASES 1 (DRM1) and 2 (DRM2) are responsible for CHH methylation through the RNA-directed DNA methylation (RdDM) pathway5,6. RdDM triggered by 24-nucleotide (24-nt) small interfering RNAs (siRNAs) has been studied extensively7,8. Although recent data demonstrated that DNA methylation is not always associated with the accumulation of corresponding siRNAs, transgenerational maitenance of gene body CHG and CHH methylation required RdDM9,10,11.
In plants, the RdDM pathway involves a plant-specific RNA polymerase, Pol IV, which transcribes hetero-chromatic regions into non-coding transcripts. These transcripts are converted into double-stranded (ds)RNA precursors by RNA-dependent RNA polymerase 2 (RDR2). Dicer-like enzyme 3 (DCL3) acts on these precursors and processes them into 24-nt siRNAs. The resulting 24-nt siRNAs are loaded into ARGONAUTE 4 (AGO4), AGO6, or AGO9 complexes to target Pol V-dependent nascent scaffold transcripts and recruit the DRM2 to guide de novo cytosine DNA methylation and maintain transcriptional gene silencing (TGS) at Pol V-transcribed loci12,13.
In addition to the canonical RdDM mediated by 24-nt siRNAs (herein referred to as 24-nt-siRdDM), several recent studies support a role for 21-22-nt siRNAs in directing DNA methylation and maintenance of the silencing of a few selected loci. Several silencing-related factors that have been previously implicated only in post-transcriptional gene silencing (PTGS), including RDR1, RDR6, AGO1, AGO2, and DCL2, are involved in 21-22-nt siRNA-mediated RdDM of some transposable elements (TEs) and several intergenic regions14,15. Recently, 15 TE subfamilies with the individual loci undefined, were reported to accumulate RDR6-dependent 21-22-nt siRNAs in a ddm1 mutant background15. In addition to the TEs, two endogenous TAS loci (trans-acting small interfering RNAs, ta-siRNAs) also display a high cytosine methylation status at ta-siRNA-generating regions, and 21-22-nt ta-siRNAs are required to guide the DNA methylation of TAS loci14.
Comprehensive methylome analyses of many Arabidopsis silencing mutants have shown that among all of the mutants that are not involved in canonical 24-nt-siRdDM, the rdr1 mutant was even stronger than the rdr6 mutant in reducing the methylation of all CG, CHG, and CHH contexts in chromosome 116 in regions that were more likely to be associated with genes and 21-22-nt siRNAs in wild-type Arabidopsis17. These data suggest that proteins other than RDR6, such as RDR1, that generate 21-22-nt siRNAs play a role and affect more loci in DNA methylation. Thus, genome-wide 21-22-nt siRNA-related methylation loci (herein referred to as 21/22-nt-siRdDM to distinguish the canonical 24-nt-siRdDM) remain to be elucidated.
Plant DNA viruses encode viral suppressors of RNA silencing (VSRs) that interfere with DNA methylation by mainly disrupting the methylcycle. For example, geminiviral VSRs such as AC2/AL2, C2/L2 and βC1 act by inhibiting the adenosine kinase (ADK) activity that plays a crucial role in adenosine and methylcycle maintenance or cytokinin regulation18,19,20, by interacting with S-adenosylmethionine decarboxylase 1 (SAMDC1), which catalyzes the conversion of S-adenosylmethionine (SAM) to decarboxylated SAM (dcSAM)21, or by inhibiting the activity of S-adenosyl-homocysteine-hydrolase (SAHH), which is also involved in the methylcycle22. Recently studies reported that potyviral helper component proteinase (HCPro) and Crinivirus coat protein also suppress antiviral RNA silencing through disruption of the methylcycle23,24. In contrast, VSR protein 2b encoded by the positive-strand RNA virus Cucumber mosaic virus (CMV) suppresses the DNA methylation associated with PTGS25,26 and binds to duplex siRNAs in 21-, 22- and 24-nt classes as well as AGO proteins27,28,29. The suppression of PTGS and DNA methylation in a few selected 24-nt-siRdDM loci (e.g. MEA-ISR and SN1) by CMV 2b protein requires siRNA-binding activity but is independent of its ability to interact directly with AGOs28. However, the 2b-AGO interaction may facilitate the 2b suppression function because 2b-AGO interactions in the nucleolus redistribute both the 2b and AGO proteins in the nucleus28 and inhibit AGO1 slicer activity in vivo30. However, it remains unknown if 2b binds to siRNAs in vivo and interferes with genome-wide DNA methylation.
In the present study, we discovered a genome-wide reduction of CHH and CHG methylation in 2b-transgenic Arabidopsis plants (line 2b-3) expressing the 2b protein encoded by the severe Shan-Dong (SD) isolate from CMV sub-group I. We also found that the 2b protein co-immunoprecipitated 21-, 22- and 24-nt duplex siRNAs in vivo, with a preference for 21- and 22-nt siRNAs. By characterizing the differential methylation regions (DMRs) associated the total and 2b-bound siRNAs, we discovered a subset of endogenous 21/22-nt-siRdDM loci, including TEs and coding genes, in both gene bodies and upstream regions. The transcriptome analysis revealed that the DNA methylation status at most 21/22-nt-siRdDM loci did not impact their expression. However, DNA blotting hybridization detected deletion and/or transposition of multicopy TEs with reduced methylation in the presence of the 2b protein, suggesting potential reactivation of multicopy TEs by the 2b protein.
Results
Identification of differential methylation regions (DMRs) between wild type and 2b transgenic plants
The 2b-transgenic Arabidopsis plants created by transformation of the full-length 2b coding sequence from the severe Shan-Dong (SD)CMV isolate28 was used in this study. The accumulation of the 2b protein in 2b-transgenic line 2b-3 was confirmed (Figure S1A). We first compared genome-wide DNA methylation patterns in Col-0 and 2b-3 by bisulfite sequencing. We found that tandem repeat, non-coding (nc)RNA, pseudogenes and transposons were highly methylated in both Col-0 and 2b-3 plants (Fig. 1A). The DNA methylation levels in CG, CHG and CHH of the expressed genes showed a tendency to decrease at transcription start sites (TSS), but the methylation level of CG to increase at gene bodies (Fig. 1B), which was consistent with previous reports for the whole genome methylation sequencing results in Col-031,32 and 2b-3 (Figures S1 and S2). Next, we identified sequences that were differentially methylated in Col-0 and 2b-3 by calculating methylation levels in the context of each cytosine and defined DMRs in 2b-3 plants in which the DNA methylation levels varied significantly compared with Col-0 plants. This led to the identification of 23,552 hypomethylated DMRs (hypo-DMRs) and 5,093 hypermethylated DMRs (hyper-DMRs) (Fig. 1C), which made up 3.31% of the genomic sequence in total. Separation into each cytosine context revealed a high proportion of CG in hyper-DMRs (2,729/5,093) compared with hypo-DMRs (4,298/23,552); however, a remarkable amount (15,141) of CHH in hypo-DMRs was obtained (Fig. 1C), in agreement with the observation of a subtle genome-wide decrease in CHH content in 2b-3 compared with Col-0 (Fig. 1B) (P > 0.05, t-test).
Bisulfite sequencing profiling of methylation in wild type Col-0 and 2b transgenic plants.
(A) The average genome-wide methylation level of tandem repeats, ncRNA, pseudogenes and transposons. (B) The average methylation level of coding genes genome-wide. The mean methylation level of CG (red line) refers to the y-axis on the left, and that of CHG and CHH (blue and green lines) refers to the y-axis on the right. (C) Comparison of the levels of hyper- and hypo-DMRs in CG, CHG and CHH cytosine contexts between Col-0 and 2b-3 plants. (D) The average methylation level of coding genes in DMRs. (E) The average methylation level of tandem repeats, ncRNA, pseudogenes and transposons in DMRs.
Decreased CHH/CHG methylation of genomic sequences in 2b transgenic plants
The identified DMRs were located not only in coding genes but also in tandem repeats, ncRNA, pseudogenes and TEs (Fig. 1D,E). For the coding genes, while the average methylation in the CG context was slightly affected by 2b, the average methylation of CHG and especially of CHH was greatly impaired either in the gene body or upstream and downstream of genes in 2b-3 (Fig. 1D). Note that the number on the y-axis for DMR coding genes was 0.6 and 0.3 for CG and CHG/CHH, respectively (Fig. 1D), but the number for coding genes in the whole genome was 0.3 and 0.08 for CG and CHG/CHH, respectively (Fig. 1B), showing that the average methylation level for coding region DMRs was much higher than that of the total coding genes in the whole genome. These data suggested that coding genes with a high density of methylation were targeted by the 2b protein, leading to the decreased CHH and CHG methylation observed in 2b-3 plants (Fig. 1D). A high density of methylation was also detected for DMRs located in tandem repeats, ncRNA, pseudogenes and TEs compared to that in the whole genome (compare Fig. 1A,E). Again, the average methylation level for the DMRs in those loci was also greatly reduced, especially in CHH, in 2b-3 (Fig. 1E). These data together indicated that stable expression of the 2b protein reduced genome-wide CHH and CHG methylation in Arabidopsis plants, providing a distinct in vivo model to investigate the mechanism of the RdDM6,28.
Identification and characterization of small RNAs in 2b transgenic plants
The 2b protein has been shown to be capable of binding to 21-, 22-, and 24-nt siRNAs28. Therefore, we investigated the profiles of siRNAs in Col-0 and 2b-3 plants by sequencing the total small RNAs from Col-0 and 2b-3 plants, as well as those co-immunoprecipitated with the 2b protein from 2b-3 plants (Table S1).
Consistent with a previous study33, total siRNAs extracted from Col-0 were dominated by 21- and 24-nt species, with a slightly greater population of 24-nt siRNAs compared with 21-nt siRNAs (Fig. 2A). However, 21-nt siRNAs became much more abundant in 2b-3 plants, while the levels of 24-nt siRNAs remained comparable to those in Col-0 plants (P < 0.001, proportion test) (Fig. 2A). The endogenous siRNAs co-immunoprecipitated with the 2b protein from 2b-3 plants (referred to as 2b-coIPed siRNAs) was similar in the length distribution to the total siRNAs of 2b-3 plants sequenced without co-immunoprecipitation (Fig. 2A). Indeed, perfectly matched siRNA duplexes represented 0.71% of the Col-0 siRNA library, but increased to 1.15% and 1.75%, respectively, in the 2b-3 and 2b-coIPed siRNA libraries (Fig. 2B). In addition to the perfect duplexes, the 2b protein also exhibited a high affinity for imperfect siRNA duplexes that had formed between micro(mi)RNA/miRNA*28. The miRNAs* were scoped by matching siRNAs to miRNA precursor hairpin sequences, and the levels of miRNAs* increased in 2b-3 (Fig. 2C), which may account for the stabilization and increased levels of miRNAs in 2b-3 plants. A large amount of miRNAs and miRNAs* in the 2b-coIPed siRNA library suggested that the miRNA/miRNAs* duplexes were likely co-precipitated by the 2b protein (Fig. 2C). Previous in vitro reciprocal-competing binding assays showed that 2b possessed a similar binding affinity for the 21- and 24-nt siRNA duplexes28. Thus, the ratio of each length of perfectly paired siRNA duplex (not including the miRNA/miRNA* duplexes) was counted. Despite a large amount of 24-nt duplexes in Col-0 and 2b-3 plants, the 21-nt siRNA duplexes were markedly increased in the 2b-coIPed siRNA library (P < 0.001, chi-square test) (Fig. 2D). This finding suggests an in vivo preference for the 2b protein to bind 21-nt siRNA duplexes, which may explain why 21-nt siRNAs were more abundant in 2b-3 plants than Col-0 plants.
Deep sequencing profiling of small RNAs in wild type Col-0 and 2b transgenic plants.
(A) Proportions of different lengths of total siRNAs from Col-0, 2b-3 plants and 2b-coIPed complexes. (B) The ratio of perfectly matched siRNA duplexes in each siRNA library. (C) The numbers of miRNAs and potential miRNA/miRNA* duplexes (labeled as hairpins) by small RNAs that matched with the annotated miRNA precursor sequences in each siRNA library. (D) The proportion of each length of perfectly paired siRNA duplex. (E) The 5′ terminal nucleotide compositions of total siRNA and siRNA duplexes in different libraries (left columns for total siRNAs,middle for 21/22-nt and right for 24-nt siRNAs).
Next, we assessed the features of the 2b-coIPed siRNAs by comparing the 5′-terminal nucleotides of the siRNA duplexes in the Col-0, 2b-3 and 2b-coIPed libraries. Figure 2E shows that, while the proportion of the siRNAs duplexes with 5′-terminal adenine (1A) from 2b-3 plants both before (39.7%) and after 2b co-immunoprecipitation (36.5%) was slightly lower than that in Col-0 (45.0%) (Fig. 2E), the proportion of 1U siRNA duplexes was markedly increased in the 2b-coIPed siRNAs, especially for 24-nt, compared with the Col-0 and 2b-3 siRNA duplexes (Fig. 2E) (P < 0.001, proportion test). The previous studies showed that the in vitro 2b binding siRNA assay had no obvious binding preference for siRNAs beginning with different nucleotides at the 5′ terminus28, and that the 2b protein potentially interacted directly with the AGO1 and AGO4 proteins27,28,29. It is known that AGO1 mainly binds to 21-nt 1U miRNAs and siRNAs, but also approximately 30% of 24-nt 1U siRNAs species and that 2b interacts with AGO1 in vivo3,28,33,34,35. Thus, our data together suggest that the binding of the 2b protein directly to siRNA duplexes and indirectly to siRNAs in AGO1 complex may contribute to the population structure of the endogenous siRNAs found in 2b-3 plants.
Correlation between DMRs and siRNAs in wild type and 2b transgenic plants
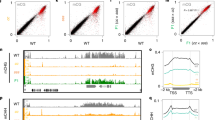
Next, we analyzed the correlation between the identified DMRs and the siRNAs in the three libraries. 13.28% of the total siRNAs found in Col-0 were mapped to the identified DMRs, with 13% localized to the hypo-DMRs and the remaining 0.28% to the hyper-DMRs. 10% and 11% of the total and 2b-coIPed siRNAs from 2b-3 plants were mapped to the identified DMRs, and again, mostly to hypo-DMRs in both cases, and more than 50% of DMR-associated 2b-coIPed siRNAs overlapped with DMR-associated siRNAs from 2b-3 plants (Fig. 3A). When these siRNA-targeted DMRs were divided into the three context sequences, it became apparent that the DMR-associated siRNAs found in 2b-coIPed siRNA library were preferentially mapped to CHH hypo-DMRs (Fig. 3C, Table S2). Figure 3B shows examples of hyper- and hypo-DMRs corresponding to 2b-coIPed siRNAs in different chromosome regions. We further examined the relative abundance of the DMR-associated siRNAs in different sizes mapped to hypo- and hyper-DMRs in the three context sequences (Fig. 3D–F). We found that a high density of siRNAs aggregated with hypo-DMRs, especially with CHG and CHH in hypo-DMRs (Fig. 3D–F, solid lines), whereas a very small portion of the siRNAs in the three libraries were mapped to hyper-DMRs (Fig. 3D–F, dashed lines). As shown in Fig. 3D, a high proportion of 24-nt siRNAs from Col-0 plants were mapped to CHG and CHH hypo-DMRs, consistent with the known role of the 24-nt siRNAs involved in canonical RdDM processes7,8. The DMR-associated siRNAs in both the total and 2b-coIPed siRNA libraries from 2b-3 plants also aggregated with CHG and CHH hypo-DMRs (Fig. 3E,F); however, the relative abundance of the DMR-associated 24-nt siRNAs in both libraries was significantly reduced compared to that from Col-0 plants (P < 0.001, proportion test). These findings indicate that expression of the 2b protein in Arabidopsis plants specifically reduced the levels of the 24-nt siRNAs associated with the identified hypo-DMRs (Fig. 3D–F) without altering the accumulation of the total 24-nt siRNAs (Fig. 2A).
Analysis of the correlation between DMRs and siRNAs in wild type Col-0 and 2b transgenic plants.
(A) The ratio of siRNAs that matched with hyper- and hypo-DMRs from the Col-0, 2b-3 and 2b-coIPed siRNA libraries. Slash populated areas mark the overlap siRNAs between 2b-3 and 2b-coIPed siRNA libraries. (B) Examples of hyper- and hypo-DMRs corresponding to 2b-coIPed siRNAs in different chromosome regions. Vertical lines show the methylation level of each cytosine in Col-0 and 2b-3. Horizontal lines show the 2b-coIPed siRNAs that matched with the DMRs. (C) The numbers of 2b-coIPed siRNAs that matched with the hyper- or hypo-DMRs in CG, CHG and CHH contexts. (D–F) Proportion of different lengths of siRNAs from each siRNA library that matched with the hypo-DMRs (solid lines) and hyper-DMRs (dotted lines) in CG, CHG and CHH contexts.
By comparison, the accumulation of the 21-nt siRNAs associated with the identified hypo-DMRs in 2b-3 plants was similar to that in Col-0 plants (Fig. 3E) unlike the total 21-nt siRNAs, which were much more abundant in 2b-3 plants than Col-0 plants (Fig. 2A). However, we detected a modest enrichment of these hypo-DMRs-associated 21-nt siRNAs in the 2b-coIPed siRNAs library (Fig. 3F), suggesting in vivo sequestration of the DMR-associated 21-nt siRNAs by the 2b protein. Therefore, it is likely that both the reduced production of the DMR-associated 24-nt siRNAs and the sequestration of the DMR-associated 21-nt siRNAs play a role in the genome-wide reduced DNA methylation detected in 2b-3 plants.
Categorization of hypo-DMR loci with various lengths of 2b-coIPed siRNAs
To examine the hypothesis, we divided the identified hypo-DMR loci into five groups according to the relative abundance of the DMR-associated 24-nt and 21-/22-nt siRNAs in the 2b-coIPed siRNA library. The hypo-DMR loci in group I and V were associated only with 24-nt siRNAs (marked as 2b-24only) and 21-22-nt siRNAs (2b-21/22only), respectively. Group II hypo-DMRs were targeted by 24-nt siRNAs that were at least two times in excess of the 21- and 22-nt siRNAs (2b-24mainly) whereas group IV hypo-DMRs were targeted by 21- and 22-nt siRNAs that were at least two times in excess of the 24-nt siRNAs (2b-21/22mainly). Hypo-DMR loci that were associated with an equivalent amount of 24-nt and 21-22-nt siRNAs were included in group III (2b-24≈21/22).
2,863 of the 18,383 hypo-DMR loci (15.57%) were associated only with 24-nt siRNAs (2b-24only), and included 458 CG loci, 322 CHG loci and 2083 CHH loci. As expected, mainly 24-nt 1A siRNAs from Col-0 plants were mapped to these loci (Fig. 4A and S3), despite the overall low density of siRNAs in Col-0 (Fig. 4B). The levels of DMR-associated 24-nt siRNAs were greatly reduced in 2b-3 plants (P < 0.001, Mann-Whitney U test) (Fig. 4A), indicating that group I loci corresponded to those loci targeted by the canonical RdDM mediated by AGO4-loaded 24-nt siRNAs13.
Categorization of hypo-DMR loci with various lengths of 2b-coIPed siRNAs.
(A) Heatmap of the siRNA density within each hypo-DMR in CG, CHG and CHH contexts. Columns represent data for 21/22- and 24-nt siRNAs from Col-0, 2b-3 and 2b-coIPed siRNA libraries that matched to the five groups of hypo-DMR loci. The numbers of hypo-DMR loci in each group are shown (n). The arrows mark the siRNAs in the 2b-coIPed library. The thickness of the arrows represents the abundance of 2b-coIPed siRNAs. The density of matched siRNAs to each hypo-DMR locus (row) coincided with the degree in red for each individual heatmap, but they are not comparable among each other. (B) Box plots of the siRNA density for each group of hypo-DMRs in Col-0, 2b-3 and 2b-coIPed siRNA libraries.
63.59% of the identified hypo-DMR loci belonged to groups II and III (Fig. 4A). Group II hypo-DMRs (2b-24mainly) included 364 for CG, 1374 for CHG and 4353 for CHH while those in group III (2b-24≈21/22) contained 523 for CG, 1319 for CHG and 3756 for CHH (Fig. 4A), indicating enrichment for CHH (69.37%) and CHG (23.04%) hypo-DMRs. Loci in these two groups were associated with a high density of siRNAs in Col-0 plants (Fig. 4B). There was a large amount of 24-nt 1A siRNAs mapped to hypo-DMRs, especially CHH, in Col-0 plants, and the amount was significantly reduced in 2b-3 plants (P < 0.001, Mann-Whitney U test) (Fig. 4A and S3), suggesting that hypomethylation in the loci of groups II and III resulted from the reduced production of the 24-nt siRNAs in 2b-3 plants. However, the involvement of the 21-22-nt siRNAs in RdDM at these loci cannot be excluded since their sequestration in the 2b complex.
2b-21/22mainly (group IV) contained 1,816 hypo-DMR loci (9.88%), which included 210 loci for CG, 387 (21.3%) for CHG and 1,219 (67.1%) for CHH (Fig. 4A). Group IV loci were associated with a high density of siRNAs and a large amount of 24-nt 1A siRNAs in Col-0 plants (Fig. 4B and S3), suggesting that they play a role in RdDM at these loci. A reduction of the 24-nt siRNAs in 2b-3 plants might account for the hypomethylation at these loci; however, because the majority of the 2b-coIPed siRNAs were 21-22 nt in length, we also propose a role for 21-22-nt siRNAs in RdDM at these loci in Col-0 plants (see below section). Indeed, 2b-21/22mainly included the TAS1C and TAS3 loci, in which DNA methylation has been found to be directly guided by 21-22-nt siRNAs despite the presence of existing 24-nt tasiRNAs at these loci14.
2,015 of the 18,383 hypo-DMR loci (10.96%) was associated only with 21- and 22-nt siRNAs and these group V loci included 668 for CG, 242 for CHG and 1105 for CHH (Fig. 4A). The density of siRNAs in these loci was low (Fig. 4B), but notably, the levels of 24-nt siRNAs was still much higher compared with the 21-22-nt siRNAs in Col-0 plants and were greatly reduced in 2b-3 plants, especially with respect to the CHG and CHH hypo-DMRs (P < 0.001, Mann-Whitney U test) (Fig. 4A). The majority of the 1A 24-nt siRNAs at these loci in Col-0 (Figure S3) were suggested to a play role in RdDM. However, despite potential interference with RdDM by the interaction with 2b-AGO4, 2b-coIPed only 21-22-nt siRNAs at these hypo-DMR loci, and the decreases in 24-nt siRNAs in 2b-3 indicated that, like the above hypo-DMR group IV, the 21-22-nt siRNAs probably played a role in RdDM at these endogenous loci.
Taken together, our data for the hypo-DMRs from group IV and V suggest that 21-22-nt siRNAs participate, by either directly guiding or alongside 24-nt siRNAs, in DNA methylation at many endogenous loci.
Identification of hundreds of TE loci targeted by highly abundant 21- and 22-nt siRNAs
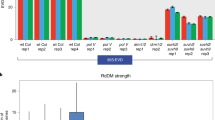
A total of 8468 individual TE elements from all 18 superfamilies (including all 318 subfamilies) contained the identified hypo-DMRs (Figure S4), which is consistent with TEs as major targets of RdDM36. Most (79.7%) of these TEs were targeted by the hypo-DMRs in groups I, II and III; however, the hypo-DMRs in groups IV and V were identified in a large number (1719) of the targeted TE elements (Figure S4). We next calculated the total number of the 24-nt siRNAs and 21-/22-nt siRNAs from Col-0 and 2b-3 plants mapped to the TEs in each of the five groups (Fig. 5A). High levels of 24-nt siRNAs were mapped to the TEs in each of the five groups in Col-0 plants and their accumulation was greatly reduced in 2b-3 plants (Fig. 5A), supporting a major role for 24-nt siRNAs to target TEs for RdDM.
Numbers of various lengths of siRNAs that matched with TEs and coding genes in each hypo-DMR group.
(A,B) The numbers of siRNAs that matched with all TE loci (A) or only non-miRNA-target TE loci (B) in the indicated hypo-DMR groups. (C,D) The numbers of siRNAs that matched with the hypo-DMRs located in upstream regions of coding genes (C) and gene bodies (D) in five hypo-DMR groups. (E) The numbers of siRNAs that matched with the TAS genes in hypo-DMR group IV.
Notably, we found that the 21-/22-nt siRNAs mapped to the TEs with group IV hypo-DMRs of either Col-0 or 2b-3 plants were distinctly much more abundant than both the 24-nt siRNAs mapped to the same group of TEs and the 21-/22-nt siRNAs mapped to the TEs of the remaining four groups (Fig. 5A). This group contained 960 TEs in total from each of the 18 superfamilies (Figure S4). Recently studies reported that miRNAs-directed secondary 21/22-nt siRNAs biogenesis from many TEs when they are epigenetically reactivated (named as easiRNAs)37,38. To rule out the group IV possible contains miRNAs-targeted TEs, we analyzed all known miRNA-targeted TEs in Arabidopsis and found the group IV contained 139 loci in miRNA-targeted TEs38. However, more than 90% of the 21-/22-nt siRNAs mapped to the group IV TEs remained highly abundant (P < 0.001, chi-square test) after removing 21-/22-nt siRNAs mapped to the miRNA-targeted TE loci (Fig. 5B).
We further examined the 15 TE subfamilies identified previously as potential targets of RdDM by RDR6-dependent 21-22-nt siRNAs15. We found that 829 TE elements in the 15 subfamilies from superfamilies LTR/Copia, RC/Helitron, LTR/Gypsy and DNA/En-Spm contained hypo-DMRs in groups IV and V. However, most (3267 loci) of the TEs in the 15 subfamilies were targeted by hypo-DMRs in groups I, II and III (Figure S4), and were associated with highly abundant 24-nt siRNAs (Fig. 5A). Thus, it is unlikely that RdDM targeting all of the TEs in these 15 subfamilies relied on 21-22-nt siRNAs.
Identification of a subset of coding genes targeted by highly abundant 21-22-nt siRNAs
Fewer hypo-DMRs (4483 loci) from the five groups were localized to the coding genes compared to the hypo-DMRs specific to TEs, including 2867 loci at upstream promoters and 1616 loci in coding sequence regions (CDS). Mapping of the total 24-nt siRNAs and 21-/22-nt siRNAs from Col-0 and 2b-3 plants to the upstream promoters and CDS in each of the five groups revealed shared features with the siRNAs associated TEs (Fig. 5C,D). The levels of 24-nt siRNAs mapped to the coding genes in each group in Col-0 were greatly reduced in 2b-3 (P < 0.001, chi-square test) (Fig. 5D,E). We also found that the 21-/22-nt siRNAs mapped to 172 group IV CDS loci were more abundant than the 21-/22-nt siRNAs mapped to the CDS loci of the remaining four groups in both Col-0 and 2b-3 plants (Fig. 5D), suggesting possible involvement of 21-/22-nt siRNAs in regulating the methylation of these CDS loci. Interestingly, we detected an increased accumulation of the 21-/22-nt siRNAs mapped to this group of CDS loci in 2b-3 plants than Col-0 plants, in particular for 5 of the 7 Arabidopsis TAS loci identified in this group (Fig. 5E). In Col-0 plants, TAS3B and TAS4 contained no hypo-DMRs and exhibited very low levels of DNA methylation or siRNAs (Figure S5). In contrast, TAS1A, TAS1B, TAS1C, TAS2, and TAS3 were heavily methylated and targeted by abundant siRNAs predominantly 21- and 22-nt in length in Col-0 plants. The hypomethylation of the five TAS loci in 2b-3 plants was associated with the strong sequestration of the 21-/22-nt siRNAs mapped to the same loci (Figure S5), indicating a role for TAS-specific 21-22-nt siRNAs in the DNA methylation of TAS loci as reported previously for TAS1C and TAS314.
The involvement of 21-22-nt siRNAs in RdDM does not impact the expression of the target loci
We next compared the transcription levels of the coding genes containing hypo-DMRs in groups IV and V in Col-0 and 2b-3 plants by transcriptome sequencing. 126 of the 172 CDS loci in group IV were detected in either Col-0 or 2b-3 plants and the expression levels of these genes were generally low. Only 11 of the 126 CDS loci and 27 of the 665 promoter loci in group IV showed a significant increase in the transcription in 2b-3 (P < 0.05, FDR < 0.001) (Figure S6A,B) and the expression levels of the five TAS genes did not exhibit significant differences between Col-0 and 2b-3 plants (Figure S6C, green dots). Similarly, only 26 of the 521 CDS loci and 30 of the 601 promoter loci in group V displayed a significant increase in transcription in 2b-3 plants compared with Col-0 plants (P < 0.05, FDR < 0.001) (Figure S6D,E). Taken together, our data suggest that neither DNA methylation nor the sequestration of the 21-/22-nt siRNAs by the 2b protein has an obvious effect on the transcription of the majority coding genes in groups IV and V, consistent the earlier observations14.
2b-mediated hypomethylation destabilizes multicopy numbers of TE loci
To evaluate the impact of the hypomethylation of TEs induced by the 2b protein, Southern blots were performed to examine the copy number of several TEs containing hypo-DMRs associated with siRNAs co-immunoprecipitated with the 2b protein. These included four loci in the superfamily LTR/Gypsy (AT2TE18830, AT3TE60465, AT3TE67540, and AT1TE46265, which were classified in the ATLANTYS1, ATLANTYS2, ATGP3 and ATHILA6B families, respectively), two loci in the DNA/MuDR superfamily (AT1TE68310 and AT5TE04360 in the VANDAL2N1 and ATMU10 families, respectively), and one locus, AT3TE48455, in the LINE/L1 superfamily. As showed in Fig. 6, three TEs with one hybridization band in Col-0 (AT3TE67540, AT1TE46265 and AT5TE04360) remained unaltered in the third generation (F3) and F4 of the 2b-3 plants. Four hybridization bands of AT3TE48455 also remained unchanged in the 2b-3 plants. However, loss of one or more hybridization bands was observed for the TEs (AT2TE18830, AT3TE60645, and AT1TE68310) with high copy numbers in F4 of the 2b-3 plants compared with Col-0 (Fig. 6). A clear lack of one hybridization band was detected for AT3E60465 (group IV) in both F3 and F4 of 2b-3 plants and for AT1TE68310 (group II) in F4 but not in F3 of 2b-3 plants, and the absence of at least three hybridization bands was detected for AT2TE18830 (group IV) in F4 with two of them absent in F3 of 2b-3 plants (Fig. 6). We also detected one additional band for AT2TE18830 in both F3 and F4 of 2b-3 plants (Fig. 6). The possibility of an incomplete restriction enzyme digestion was ruled out because three biological replicates were performed and similar results were obtained. These results suggested that 2b-mediated hypomethylation caused the destabilization of TEs, resulting in potential TE deletion and/or transposition in 2b-3 transgenic plants.
Detection of TE transposition in the third and fourth generations of 2b transgenic plants.
DNA blotting analysis of seven TEs that overlapped with hypo-DMRs. The DNA samples were extracted from wild type Col-0, and the third (F3) and fourth (F4) generations of 2b-3 plants. Hollow arrow indicate additional hybridization bands in F3 and F4 compared with Col-0; solid arrows indicate the lack of a hybridization band in (F3) and/or (F4) compared with Col-0.
Discussion
The CMV silencing suppressor 2b protein exhibits complex biochemical and subcellular targeting activities, including the suppression of RNA silencing in the cytoplasm and DNA methylation in the nucleus. Our bisulfite sequencing analysis result revealed a genome-wide reduction of CHG and, especially, CHH methylation in 2b-transgenic Arabidopsis plants, and identified the DMRs between wild-type and 2b-transgenic plants, consistent with the previous notion that the 2b protein plays a role in the suppression of RdDM required for constant de novo CHH methylation6.
Most of the previous researches investigating RdDM have constructed a canonical RdDM that de novo DNA methylation is mediated by 24-nt siRNAs (24-nt-siRdDM). In the present study, aided by the capability of the 2b protein to bind both the 21-22-nt and 24-nt ds-siRNAs as well as their associated AGO proteins and suppress DNA methylation28, we was able to characterize DMRs associated the total and 2b-bound siRNAs. We found that 2b suppressed DNA methylation not only at the previously annotated 24-nt-siRdDM loci but also a new set of endogenous 21/22-nt-siRdDM loci, including TEs and coding genes, in both gene bodies and upstream regions.
We found that a large amount of 21-22-nt siRNAs were matched with the endogenous 21/22-nt-siRdDM loci (group IV) in Col-0 and were co-IPed by 2b protein. Interestingly, the increase in 1C in 2b-bound siRNAs was associated with group IV CHH hypo-DMRs (Figure S3D). The 2b-mediated stabilization of siRNA duplexes might account for the increase in 1C due to antisense siRNAs. A previous study reported that AGO5 predominantly binds siRNAs with 1C34, and these results suggested that AGO5 might have a functional role in regulating 21/22-nt-siRdDM. Nevertheless, in DMRs in group IV and V, 2b-coIPed siRNAs are only 21-22-nt (group V) or largely 21-22-nt (group IV) in length, suggesting that DNA methylation of this set of endogenous loci might be guided directly by 21-22-nt siRNAs (e.g., the group V loci), and 21-22-nt siRNAs might also be required and/or act upstream of 24-nt siRNAs-guided RdDM.
In general, body-methylated genes are constitutively expressed at a higher level, and promoter-methylated genes tend to be expressed in a tissue-specific manner39. Pol V was found to preferentially associate at regions in which promoters and TEs overlap40. RNA-sequence datasets have shown that the active targeting of promoters and TEs by Pol V has functional implications for transcription40. We found that the majority of hypo-DMRs in upstream (promoter) regions contain TEs (Figure S6). More than 20% of 2b-coIPed siRNAs were matched with 88.6% of the previously reported potentially Pol V-dependent siRNA loci as well as more than 90% of Pol IV- and Pol IV + Pol V-dependent siRNAs loci41 (Figure S7). Transcriptome sequencing of 21/22-nt-siRdDM loci revealed that only a small amount of hypo-DMR loci in both promoter regions and gene bodies displayed a two-fold increase in transcription in the presence of 2b protein (Figure S6). Together with the finding that more than 90% of Pol V-dependent siRNAs are 21-nt33, these results suggest a functional role for Pol V in the transcription of 21/22-nt-siRdDM. However, the expression levels of the majority 21/22-nt-siRdDM loci were generally low, and the DNA methylation status at most of them did not impact the expression of these coding genes (Figure S6). Consistent with the previous finding that a high level of DNA methylation had no obvious effect on the regulation of transcription of the two TAS genes (TAS1C and TAS3)14, the transcripts of the 5 TAS genes in these 21/22-nt-siRdDM loci also displayed no significant changes in 2b-3 plants. Taken together, our data suggest that 21-22-nt siRNAs derived from 21/22-nt-siRdDM loci do not result from changes in transcription and siRNA biogenesis associated with the loss of DNA methylation in the presence of 2b protein, and they further support the notion that 21-22-nt siRNAs play roles in the direct guidance of, as well as alongside 24-nt siRNA in, DNA methylation in a subset of endogenous loci.
The ability of TE to mobilize and insert anywhere in the genome to cause mutations makes them key targets for silencing. Reactivation of TEs causes transposition is previously reported in mutants that have lost both heterochromatic histone modifications and DNA methylation, such as ddm1. The EVD locus has been shown to have lost methylation and transposition with an increase in copy number in epi15RIL lines derived from the met1 or ddm1 background42,43. We found that TEs with multicopies (either in group IV (AT2TE18830 and AT3TE60465) or group II (AT1TE68310)) were likely more susceptible to induced deletions and/or proliferation following 2b-mediated hypomethylation. The inactivation of siRNAs by 2b protein without affecting MET1 or DDM1 may maintain most TEs in an inactive state and retain the low copy numbers.
It has been reported that the accumulation of 21-nt siRNAs from Athila TEs prevents transposition and promotes transgenerational TE silencing44. It has been suggested that TE reactivation in the vegetative nucleus (VN) may result in the silencing of Athila TEs in sperm cells via Athila 21-nt siRNAs generated from the reactivated Athila TE transcripts from the VN44. This model of TE reactivation and potential epigenetic reprogramming is likely to be conserved for reprogramming in the germline in animals to reveal intact TEs in the genome and regulate their activity in gametes. In animals such as Drosophila, 25-29-nt PIWI-interacting RNAs (piRNAs) are generated in germline cells and are one means to attack TEs45,46,47. During germline reprogramming, epigenetic marks are first lost and then robustly reset in each generation, resulting in transient, and paradoxical, TE expression48,49,50,51,52,53,54,55. We found that Athila TEs are classified into five hypo-DMR groups (Figure S8). It remains elusive whether the silencing and methylation of 21/22-nt-siRdDM Athila TEs in the plant body occurred similarly to pollen and were also mediated by 21/22-nt siRNAs derived from reactivated canonical 24-nt-siRdDM Athila TEs. Alternatively, but not mutually exclusive, in the plant body, 21/22-nt-siRdDM TE loci might also be directly targeted by Pol IV, which transcribes hetero-chromatic regions into non-coding transcripts for siRNA generation and feedback reinforcement of DNA methylation and silencing13. Nevertheless, the inactivation of siRNAs by 2b protein caused hypomethylation but not necessarily transposition in most of the 21/22-nt-siRdDM TE loci, suggesting that these 21/22-nt-siRdDM TEs might be under rigorous control. Therefore, the loss of siRNAs alone, at least in the plant body, likely does not result in the transposition of TEs with low copy numbers.
Nevertheless, the deletion or proliferation of TEs might result in a decrease and increase in TE copy numbers, leading to the contraction or expansion of TEs in 2b-3 plants. Overexpression of the 2b in plants mainly causes hypomethylation of large amount of genomic sequences that might include genes functions in mediating DNA methylation. We cannot rule out that this might result in enhancing methylation of certain sequences, as about 5000 loci hyper-DMRs were also found in 2b-3 plants. Our recent small RNA deep sequencing and blotting analysis indicate a repetitive DNA origin for CMV satellite RNA in the Nicotiana plant genome, a small noncoding subviral RNA pathogen that depends on CMV as a helper virus for its replication and spread in plants56. The “cut-off” TE segments are no longer “junk DNA” and could also be the potential origin of the noncoding subviral DNA/RNA pathogen.
In summary, by classifying the various lengths of 2b-coIPed siRNAs in parallel with hypo-DMRs in 2b transgenic plants, we found that, approximately 10% of the 2b-coIPed siRNAs were mapped to the hypo-DMRs. The DNA methylation of a subset of these hypo-DMRs was either directly guided by 21-22-nt siRNAs or required 21-22-nt siRNAs to function upstream of the 24-nt siRNAs-guided RdDM. Similarly to the canonical 24-nt-siRdDM, the majority of the 21/22-nt-siRdDM loci were enriched for CHH and CHG. The expression levels of the 21/22-nt-siRdDM coding genes were generally low, and their DNA methylation status did not impact the expression of these genes. The 2b-mediated hypomethylation of TEs potentially resulted in the deletion and transposition of TEs with multicopies.
Methods
Bisulfite Sequencing and Bioinformatic Analyses
Genomic DNA was extracted from Arabidopsis leaves via the CTAB method57. Library construction and bisulfite sequencing were accomplished by BGI (Table S3). SOAP58 was used to align the clean reads against reference genome (Supplemental Methods).
DMRs were identified by comparison of the Col-0 and 2b-3 methylomes using windows that contained at least five cytosines sliding at one cytosine, and CG, CHG and CHH were analyzed separately. A DMR was identified if the P-value from Chi-squared test was ≤0.05.
Small RNA sequencing and data Analysis
Total RNA was isolated from wide-type Col-0 and 2b-3 plants using hot-phenol extraction59. For 2b-associated sRNAs sequencing, aerial tissues from 6myc-SD2b transgenic plants were subjected to immunoprecipitation using anti-α-myc Agarose (Abmart). After washed with IP buffer twice, the immunoprecipitates was incubated in 600 μL Trizol (Invitrogen). After extraction with 0.2 (v/v) chloroform, sRNAs were precipitated with an equal volume of isopropanol.
Small RNAs library construction and Illumina sequencing was performed by BGI (Supplemental Methods). sRNA northern blot was used to verify the miRNA expression (Figure S9).
sRNAs map to DMRs
The sRNAs that perfect matched to genome were included in our analysis. Blastn was used to search the sRNAs perfect match to DMRs or overlap with DMRs, allowing for both unique and multiple mapping reads. sRNAs density was the RPM normalized by DMRs length. Heatmap and boxplot of sRNAs density were performed by R program.
Transcriptome Sequencing
Library construction and Illumina sequencing was performed by BGI. All clean reads were aligned against both Arabidopsis genomic and cDNA (TAIR10) (Supplemental Methods). The gene expression level is calculated by using RPKM60 method (Reads Per kb transcriptome per Million reads). FDR (False Discovery Rate)61 were used to correct the diversity test P-value.
Southern blot
Ten μg of DNA was digested overnight with endonuclease (Thermo Scientific), separated by a 1% agarose gel and transferred to HyBond-NX membrane and cross-linked with UV irradiation. The DNA probes were generated by PCR, and the primers specific for each TEs were listed in the Table S4. The probes were radioactively labelled by [α-32P]-dCTP using a RediprimeTM II Random prime Labelling System (Amersham Pharmacia Biotech). Hybridization was performed in hybridization buffer at 65 °C overnight. After washed with 2 × SSC/0.2% SDS twice, the blots were exposed to a storage phosphor screen for 10 h. The hybridizing signals were visualized using a phosphorimager.
Additional Information
How to cite this article: Zhao, J.-H. et al. Genome-wide identification of endogenous RNA-directed DNA methylation loci associated with abundant 21-nucleotide siRNAs in Arabidopsis. Sci. Rep. 6, 36247; doi: 10.1038/srep36247 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Bird, A. DNA methylation patterns and epigenetic memory. Genes & development 16, 6–21, doi: 10.1101/gad.947102 (2002).
Chan, S. W., Henderson, I. R. & Jacobsen, S. E. Gardening the genome: DNA methylation in Arabidopsis thaliana. Nature reviews 6, 351–360, doi: nrg1601 (2005).
Zhang, X. et al. Cucumber mosaic virus-encoded 2b suppressor inhibits Arabidopsis Argonaute1 cleavage activity to counter plant defense. Genes & development 20, 3255–3268, doi: 20/23/3255 (2006).
Lister, R. et al. Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 133, 523–536, doi: S0092-8674(08)00448-0 (2008).
Lindroth, A. M. et al. Requirement of CHROMOMETHYLASE3 for maintenance of CpXpG methylation. Science (New York, N.Y.) 292, 2077–2080, doi: 10.1126/science.1059745 (2001).
Cao, X. & Jacobsen, S. E. Locus-specific control of asymmetric and CpNpG methylation by the DRM and CMT3 methyltransferase genes. Proc Natl Acad Sci USA 99 Suppl 4, 16491–16498, doi: 10.1073/pnas.162371599162371599 (2002).
Matzke, M., Kanno, T., Daxinger, L., Huettel, B. & Matzke, A. J. RNA-mediated chromatin-based silencing in plants. Current opinion in cell biology 21, 367–376, doi: 10.1016/j.ceb.2009.01.025 (2009).
Zhang, H. & Zhu, J.-K. RNA-directed DNA methylation. Current opinion in plant biology 14, 142–147 (2011).
Dalakouras, A. & Wassenegger, M. Revisiting RNA-directed DNA methylation. RNA biology 10, 453–455, doi: 10.4161/rna.23542 (2013).
Zhao, M., San Leon, D., Delgadillo, M. O., Garcia, J. A. & Simon-Mateo, C. Virus-induced gene silencing in transgenic plants: transgene silencing and reactivation associate with two patterns of transgene body methylation. The Plant journal: for cell and molecular biology 79, 440–452, doi: 10.1111/tpj.12579 (2014).
Dalakouras, A., Dadami, E., Zwiebel, M., Krczal, G. & Wassenegger, M. Transgenerational maintenance of transgene body CG but not CHG and CHH methylation. Epigenetics 7, 1071–1078, doi: 10.4161/epi.21644 (2012).
Wierzbicki, A. T., Haag, J. R. & Pikaard, C. S. Noncoding transcription by RNA polymerase Pol IVb/Pol V mediates transcriptional silencing of overlapping and adjacent genes. Cell 135, 635–648, doi: S0092-8674(08)01192-6 (2008).
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nature reviews 11, 204–220 (2010).
Wu, L., Mao, L. & Qi, Y. Roles of dicer-like and argonaute proteins in TAS-derived small interfering RNA-triggered DNA methylation. Plant Physiol 160, 990–999, doi: 10.1104/pp.112.200279 (2012).
Nuthikattu, S. et al. The initiation of epigenetic silencing of active transposable elements is triggered by RDR6 and 21-22 nucleotide small interfering RNAs. Plant Physiol 162, 116–131, doi: 10.1104/pp.113.216481 (2013).
Stroud, H., Greenberg, M. V., Feng, S., Bernatavichute, Y. V. & Jacobsen, S. E. Comprehensive analysis of silencing mutants reveals complex regulation of the Arabidopsis methylome. Cell 152, 352–364, doi: 10.1016/j.cell.2012.10.054 (2013).
Lee, T. F. et al. RNA polymerase V-dependent small RNAs in Arabidopsis originate from small, intergenic loci including most SINE repeats. Epigenetics 7 (2012).
Wang, H., Hao, L., Shung, C. Y., Sunter, G. & Bisaro, D. M. Adenosine kinase is inactivated by geminivirus AL2 and L2 proteins. Plant Cell 15, 3020–3032, doi: 10.1105/tpc.015180 (2003).
Wang, H., Buckley, K. J., Yang, X., Buchmann, R. C. & Bisaro, D. M. Adenosine kinase inhibition and suppression of RNA silencing by geminivirus AL2 and L2 proteins. Journal of virology 79, 7410–7418, doi: 10.1128/JVI.79.12.7410-7418.2005 (2005).
Buchmann, R. C., Asad, S., Wolf, J. N. & Mohannath, G. & Bisaro, D. M. Geminivirus AL2 and L2 proteins suppress transcriptional gene silencing and cause genome-wide reductions in cytosine methylation. Journal of virology 83, 5005–5013, doi: 10.1128/JVI.01771-08 (2009).
Zhang, Z. et al. BSCTV C2 attenuates the degradation of SAMDC1 to suppress DNA methylation-mediated gene silencing in Arabidopsis. Plant Cell 23, 273–288, doi: 10.1105/tpc.110.081695 (2011).
Yang, X. et al. Suppression of methylation-mediated transcriptional gene silencing by betaC1-SAHH protein interaction during geminivirus-betasatellite infection. PLoS pathogens 7, e1002329, doi: 10.1371/journal.ppat.1002329 (2011).
Ivanov, K. I. et al. Molecular insights into the function of the viral RNA silencing suppressor HCPro. The Plant journal: for cell and molecular biology 85, 30–45, doi: 10.1111/tpj.13088 (2016).
Canizares, M. C. et al. Effects of the crinivirus coat protein-interacting plant protein SAHH on post-transcriptional RNA silencing and its suppression. Molecular plant-microbe interactions: MPMI 26, 1004–1015, doi: 10.1094/mpmi-02-13-0037-r (2013).
Li, H. W. et al. Strong host resistance targeted against a viral suppressor of the plant gene silencing defence mechanism. The EMBO journal 18, 2683–2691, doi: 10.1093/emboj/18.10.2683 (1999).
Guo, H. S. & Ding, S. W. A viral protein inhibits the long range signaling activity of the gene silencing signal. The EMBO journal 21, 398–407 (2002).
Gonzalez, I. et al. Cucumber mosaic virus 2b protein subcellular targets and interactions: their significance to RNA silencing suppressor activity. Molecular plant-microbe interactions: MPMI 23, 294–303, doi: 10.1094/MPMI-23-3-0294 (2010).
Duan, C. G. et al. Suppression of Arabidopsis ARGONAUTE1-Mediated Slicing, Transgene-Induced RNA Silencing, and DNA Methylation by Distinct Domains of the Cucumber mosaic virus 2b Protein. Plant Cell, doi: tpc.111.092718 (2012).
Hamera, S., Song, X., Su, L., Chen, X. & Fang, R. Cucumber mosaic virus suppressor 2b binds to AGO4-related small RNAs and impairs AGO4 activities. Plant J 69, 104–115, doi: 10.1111/j.1365-313X.2011.04774.x (2012).
Feng, L., Duan, C.-G. & Guo, H.-S. Inhibition of in vivo Slicer activity of Argonaute protein 1 by the viral 2b protein independent of its dsRNA-binding function. Molecular plant pathology 14, 617–622, doi: 10.1111/mpp.12033 (2013).
Cokus, S. J. et al. Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452, 215–219, doi: 10.1038/nature06745 (2008).
Li, Y. et al. Identification of microRNAs involved in pathogen-associated molecular pattern-triggered plant innate immunity. Plant Physiol 152, 2222–2231, doi: 10.1104/pp.109.151803 (2010).
Wang, H. et al. Deep sequencing of small RNAs specifically associated with Arabidopsis AGO1 and AGO4 uncovers new AGO functions. Plant J 67, 292–304, doi: 10.1111/j.1365-313X.2011.04594.x (2011).
Mi, S. et al. Sorting of small RNAs into Arabidopsis argonaute complexes is directed by the 5′ terminal nucleotide. Cell 133, 116–127 (2008).
Takeda, A., Iwasaki, S., Watanabe, T., Utsumi, M. & Watanabe, Y. The mechanism selecting the guide strand from small RNA duplexes is different among argonaute proteins. Plant & cell physiology 49, 493–500, doi: 10.1093/pcp/pcn043 (2008).
Girard, A. & Hannon, G. J. Conserved themes in small-RNA-mediated transposon control. Trends in cell biology 18, 136–148, doi: 10.1016/j.tcb.2008.01.004 (2008).
Sarazin, A. & Voinnet, O. Exploring new models of easiRNA biogenesis. Nature genetics 46, 530–531, doi: 10.1038/ng.2993 (2014).
Creasey, K. M. et al. miRNAs trigger widespread epigenetically activated siRNAs from transposons in Arabidopsis. Nature, doi: 10.1038/nature13069 (2014).
Zhang, X. et al. Genome-wide high-resolution mapping and functional analysis of DNA methylation in arabidopsis. Cell 126, 1189–1201 (2006).
Zhong, X. et al. DDR complex facilitates global association of RNA polymerase V to promoters and evolutionarily young transposons. Nature structural & molecular biology 19, 870–875, doi: 10.1038/nsmb.2354 (2012).
Mosher, R. A., Schwach, F., Studholme, D. & Baulcombe, D. C. PolIVb influences RNA-directed DNA methylation independently of its role in siRNA biogenesis. Proc Natl Acad Sci USA 105, 3145–3150 (2008).
Mirouze, M. et al. Selective epigenetic control of retrotransposition in Arabidopsis. Nature 461, 427–430, doi: 10.1038/nature08328 (2009).
Mari-Ordonez, A. et al. Reconstructing de novo silencing of an active plant retrotransposon. Nature genetics 45, 1029–1039, doi: 10.1038/ng.2703 (2013).
Slotkin, R. K. et al. Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell 136, 461–472, doi: 10.1016/j.cell.2008.12.038 (2009).
Han, B. W., Wang, W., Li, C., Weng, Z. & Zamore, P. D. Noncoding RNA. piRNA-guided transposon cleavage initiates Zucchini-dependent, phased piRNA production. Science (New York, N.Y.) 348, 817–821, doi: 10.1126/science.aaa1264 (2015).
Mohn, F., Handler, D. & Brennecke, J. Noncoding RNA. piRNA-guided slicing specifies transcripts for Zucchini-dependent, phased piRNA biogenesis. Science (New York, N.Y.) 348, 812–817, doi: 10.1126/science.aaa1039 (2015).
Siomi, H. & Siomi, M. C. RNA. Phased piRNAs tackle transposons. Science (New York, N.Y.) 348, 756–757, doi: 10.1126/science.aab3004 (2015).
Calvi, B. R. & Gelbart, W. M. The basis for germline specificity of the hobo transposable element in Drosophila melanogaster. The EMBO journal 13, 1636–1644 (1994).
Dupressoir, A. & Heidmann, T. Germ line-specific expression of intracisternal A-particle retrotransposons in transgenic mice. Molecular and cellular biology 16, 4495–4503 (1996).
Pasyukova, E., Nuzhdin, S., Li, W. & Flavell, A. J. Germ line transposition of the copia retrotransposon in Drosophila melanogaster is restricted to males by tissue-specific control of copia RNA levels. Molecular & general genetics: MGG 255, 115–124 (1997).
Ostertag, E. M. et al. A mouse model of human L1 retrotransposition. Nature genetics 32, 655–660, doi: 10.1038/ng1022 (2002).
Lau, N. C. et al. Characterization of the piRNA complex from rat testes. Science (New York, N.Y.) 313, 363–367, doi: 10.1126/science.1130164 (2006).
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103, doi: 10.1016/j.cell.2007.01.043 (2007).
Brower-Toland, B. et al. Drosophila PIWI associates with chromatin and interacts directly with HP1a. Genes & development 21, 2300–2311, doi: 10.1101/gad.1564307 (2007).
Carmell, M. A. et al. MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Developmental cell 12, 503–514, doi: 10.1016/j.devcel.2007.03.001 (2007).
Zahid, K. et al. Nicotiana small RNA sequences support a host genome origin of cucumber mosaic virus satellite RNA. PLoS genetics 11, e1004906, doi: 10.1371/journal.pgen.1004906 (2015).
Allen, G. C., Flores-Vergara, M. A., Krasynanski, S., Kumar, S. & Thompson, W. F. A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nature protocols 1, 2320–2325, doi: 10.1038/nprot.2006.384 (2006).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967, doi: 10.1093/bioinformatics/btp336 (2009).
Pathak, R. R. & Lochab, S. A method for rapid isolation of total RNA of high purity and yield from Arthrospira platensis. Canadian journal of microbiology 56, 578–584, doi: 10.1139/w10-045 (2010).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nature methods 5, 621–628, doi: 10.1038/nmeth.1226 (2008).
Kim, K. I. & van de Wiel, M. A. Effects of dependence in high-dimensional multiple testing problems. BMC bioinformatics 9, 114, doi: 10.1186/1471-2105-9-114 (2008).
Acknowledgements
This study was supported by the Ministry of Science and Technology (2014CB138402) and the Natural Science Foundation of China (Nos 91519327 and 31600124).
Author information
Authors and Affiliations
Contributions
H.-S.G., J.-H.Z. and S.-W.D. conceived and designed the study. J.-H.Z., Y.-Y.F. and C.-G.D. performed the experiment and the analysis. R.-X.F. discussed the results. H.-S.G., J.-H.Z. and S.-W.D. wrote the paper. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhao, JH., Fang, YY., Duan, CG. et al. Genome-wide identification of endogenous RNA-directed DNA methylation loci associated with abundant 21-nucleotide siRNAs in Arabidopsis. Sci Rep 6, 36247 (2016). https://doi.org/10.1038/srep36247
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36247
This article is cited by
-
Designing Climate-Resilient Crops for Sustainable Agriculture: A Silent Approach
Journal of Plant Growth Regulation (2023)
-
Multiple MicroRNAs are Involved in Regulating Peanut (Arachis hypogaea L.) Resistance to Sclerotium rolfsii at the Early Stage
Tropical Plant Biology (2022)
-
Environmental RNAi pathways in the two-spotted spider mite
BMC Genomics (2021)
-
Phytopathogen-induced changes to plant methylomes
Plant Cell Reports (2018)
-
Methods for discovering genomic loci exhibiting complex patterns of differential methylation
BMC Bioinformatics (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.