Abstract
Bioprospecting for novel microalgal strains is key to improving the feasibility of microalgae-derived biodiesel production. Tetraselmis sp. CTP4 (Chlorophyta, Chlorodendrophyceae) was isolated using fluorescence activated cell sorting (FACS) in order to screen novel lipid-rich microalgae. CTP4 is a robust, euryhaline strain able to grow in seawater growth medium as well as in non-sterile urban wastewater. Because of its large cell size (9–22 μm), CTP4 settles down after a six-hour sedimentation step. This leads to a medium removal efficiency of 80%, allowing a significant decrease of biomass dewatering costs. Using a two-stage system, a 3-fold increase in lipid content (up to 33% of DW) and a 2-fold enhancement in lipid productivity (up to 52.1 mg L−1 d−1) were observed upon exposure to nutrient depletion for 7 days. The biodiesel synthesized from the lipids of CTP4 contained high levels of oleic acid (25.67% of total fatty acids content) and minor amounts of polyunsaturated fatty acids with ≥4 double bonds (<1%). As a result, this biofuel complies with most of the European (EN14214) and American (ASTM D6751) specifications, which commonly used microalgal feedstocks are usually unable to meet. In conclusion, Tetraselmis sp. CTP4 displays promising features as feedstock with lower downstream processing costs for biomass dewatering and biodiesel refining.
Similar content being viewed by others
Introduction
Microalgal biomass has drawn increasing attention for different biotechnological applications over the last few years, as shown by the significant developments in terms of funding allocated for microalgal research. The establishment of several start-up companies and commercial products from microalgae (e.g. DSM Life’s® and Qualitas Health® nutraceuticals as well as Encapso® drilling oil) have confirmed that these microorganisms can be important feedstocks in different markets. Nevertheless, recent achievements observed in the field of microalgal biotechnology were mainly due to research and innovation efforts towards the development of microalgae-based biofuels1. Although the technology to produce biofuels (e.g. biodiesel) from microalgal biomass has been successfully demonstrated, several techno-economical reports concluded that the biofuels obtained from microalgal feedstocks are still unable to compete with fossil fuels2,3,4. To overcome the current constraints for the commercialization of microalgae-based biofuels, the optimization of the whole production pipeline is required. An important first step is strain selection, which has to take into account later steps, such as its robustness, easy and low cost downstream processing and the effective development of a biorefinery.
During the last decades, aquaculture has been the main market for microalgal biomass, particularly for rearing bivalves and enhancing the nutrition of live prey5,6. Since unsaturated lipids are essential metabolites for the proper development of fish larvae and bivalve growth7, microalgal strains with high contents of polyunsaturated fatty acids (PUFA) have been selected and commercialized. However, microalgal feedstocks with high amounts of PUFA are not suitable for biodiesel production, because the large number of double bonds present in these fatty acids decrease the oxidation stability of the end product8,9,10. In fact, both European (EN14214) and American (ASTM D6751) specifications impose strict limits concerning the presence of PUFA in biodiesel. Therefore, screening for strains containing high lipid content with low levels of unsaturated fatty acids is crucial to enhance the productivity and quality of the feedstock used for biodiesel production10,11,12 and decreases the costs of biodiesel refining11.
Flow cytometry coupled to fluorescence activated cell sorting (FACS) is a powerful high-throughput technique for bioprospecting microalgae present in environmental samples, because this technology enables the screening of thousands of cells in a short period of time for a specific purpose (e.g. high lipid contents). This can be accomplished by acquiring different signals, such as complexity and relative cell size coupled with the autofluorescence of photosynthetic pigments and the fluorescence of solvatochromic dyes13,14,15. Depending on the final product, bioprospection using FACS enables the isolation of strains with a desired biochemical profile through different selection approaches (e.g. dyes that only emit fluorescence in the presence of lipids), narrowing down the number of strains that have indeed a high potential for a given biotechnological application16,17. Concomitantly, selection procedures must also contemplate the growth performance and robustness of a given strain, as both features are crucial for the up-scaling and effective production in large-scale systems18.
This work aimed to characterize and evaluate the potential of a novel euryhaline microalga isolated from a salt marsh near a wastewater stream in the south of Portugal as a feedstock for biodiesel production. Tetraselmis sp. CTP4 was selected from 96 isolates as a result of a screening effort using FACS to bioprospect for novel microalgal strains with biotechnological potential.
Results
Microalgae isolation and identification
Tetraselmis sp. CTP4 was isolated by FACS using the sorting procedure shown in Fig. 1. Figure 1A presents the two-dimensional plot combining the SSC (side angle light scatter) and FL3 (fluorescence emission at 695 nm) signals, relating the relative inner cell complexity with chlorophyll autofluorescence, respectively. Through the combination of these signals, the first sorting trait was established in order to differentiate non-photosynthetic from photosynthetic cells; in this way, further analyses focusing only on photosynthetic cells were carried out17. Figure 1B shows the combination of the allophycocyanin autofluorescence signal (FL4) and the emission of BODIPY 505/515 (FL1), a lipid-staining solvatochromic dye. With this combination of signals, three clusters of cells representing three different microalgal species could be clearly distinguished. Two (P3 and P4) of the three clusters displayed higher values of BODIPY fluorescence. The gates used in the sorting procedure were effective for the isolation of cells belonging to three different species as verified by microscopy upon sorting directly onto microscope slides (Fig. 1C–E). The isolates were entitled CTP3, CTP4 and CTP5, according to the numbering of the clusters obtained in the cytometer. Interestingly, among all isolates, CTP4 presented the most dense cluster of events as compared with other microalgae found in the environmental sample (Fig. 1B). This was an important first indicator that CTP4 was able to compete with other microalgae that were co-cultivated during the pre-enrichment step of the isolation process. Moreover, this microalga also showed the highest levels of BODIPY fluorescence (FL1), which strongly suggested that this microalga contained significant amounts of lipids. As a result, CTP4 was selected for further study.
Cell sorting procedure used to isolate CTP4 strain by means of fluorescence activated cell sorting (A,B), and different clusters sorted directly onto microscope slides observed with differential interference contrast (C–E). The first sorting trait (Chl gate) was applied through the combination of inner cell complexity, SSC-A, with chlorophyll autofluorescence, FL3 (A). The final gates used to isolate the CTP4 strain combined BODIPY fluorescence (FL1) with the signal of the allophycocyanin autofluorescence channel (FL4). Strains isolated from clusters P4 (C), P2 (D) and P3 (E). The first strain was named as CTP4 (C), whereas the second corresponded to an unidentified strain with apparent cell disruption (D) and the third strain was an unidentified diatom (E). Scale bar = 5 μm.
Chlorodendrophyceae 18S rDNA sequences were analysed by Bayesian (BI) and Maximum Likelihood (ML) inference and the consensus tree is shown in Fig. 2 using equivalent sequences from Trebouxiophyceae algae as the outgroup. Topology of the BI and ML consensus trees indicates that Tetraselmis sp. CPT4, isolated in this study, belongs to the T. striata/convolutae clade with a posterior probability of 0.98 and a bootstrap value of 82%, respectively (Fig. 2).
Bayesian inference tree of species of the Chlorodendrophyceae class inferred using 18S rDNA sequences.
Maximum-likelihood bootstrap values (>50) and Bayesian inference posterior probabilities (>0.70) are indicated at the branches, respectively. The CTP4 strain, isolated in this study, clustered with the striata/convolutae clade of the Tetraselmis genus.
Culture robustness and dewatering
To further characterise the novel strain, Tetraselmis sp. CTP4 was cultivated in seawater-based Modified Algal Medium (MAM) and in a non-sterile, non-nitrified wastewater effluent (Fig. 3). Cultures were grown until a cell concentration (CC) of about 2.7 × 106 cells mL−1 was reached. Both batch cultures displayed similar growth curves, reaching stationary phase at day 8. Cultures were monitored daily by bright field microscopy and flow cytometry. Although bacterial and microalgal contaminants were frequently observed in the non-sterile wastewater culture, Tetraselmis sp. CTP4 remained the dominant specie throughout.
The natural sedimentation of cultures, presented as volume of settled culture per litre of culture, was investigated during the course of 8 hours using Imhoff cones (Fig. 4). The settling of the cultures revealed a decreasing logarithmic curve, converging towards 18% of the initial volume over 6 hours.
Biomass growth and lipid induction
The growth curves of cultures grown under nutrient repletion (N+) and nutrient starvation (N−) in a two-stage growth system are shown in Fig. 5A. Both cultures grew exponentially until the end of the 1st stage (day 10), reaching a CC of 2.9 × 106 cells mL−1. The specific growth rate (μ) during this stage was similar in both culture conditions (0.29–0.31 d−1). In the 2nd stage, N+ cultures continued to grow, although at a slower rate, reaching a final concentration of 3.3 × 106 cells mL−1, whereas the N− cultures plateaued at approximately 3.0 × 106 cells mL−1. At the beginning of the 1st stage the lipid content of both cultures was approximately 10% of DW (Fig. 5B). During the exponential phase (between day 4 and 8), a decrease in the lipid content (5–8% of DW) was observed. During the 2nd stage (lipid accumulation stage), cultures supplemented with nutrients (N+) maintained the same lipid content, displaying a final lipid content of 10% of DW. However, the N− cultures reached a significantly higher lipid content: approximately 33% of DW. These results were confirmed by staining with BODIPY 505/515; microalgae grown under nutrient starvation contained a significantly higher amount of lipid bodies that stained positively for the solvatochromic dye (Fig. 6). The biomass and lipid productivities obtained in the present study and previous reports with other Tetraselmis strains are presented in Table 1. The N+ cultures displayed higher biomass productivity compared to those in the N− treatment, yielding 0.29 and 0.25 g L−1 d−1, respectively. On the other hand, the final lipid productivity doubled under the N− conditions (52.1 mg L−1 d−1) when compared to N+ cultures (24.5 mg L−1 d−1).
Cultures grown in a two-stage system under nutrient repletion (N+) and nutrient depletion (N−) conditions.
The growth curves (A) and the corresponding mean lipid content (B) of cultures exposed to both culture conditions is shown (n = 3). Dashed line shows the addition of nutrients to the N+ cultures (A).
Biochemical profile and properties of CTP4 biodiesel
Upon synthesis of biodiesel derived from the lipids extracted from CTP4 cells, the fatty acid methyl esters (FAME) profile of the biofuel was determined (Table 2). Palmitic (C16:0), oleic (C18:1) and linoleic (C18:2) acids were the major FAME detected, representing approximately 75% of the total fatty acids (TFA) in the biodiesel mixture. Other FAME also found at relatively high amounts were palmitoleic (C16:1) and hexadecatrienoic (C16:3) acids (10% of TFA), whereas only minor levels of hexadecadienoic (C16:2), linolenic (C18:3) and eicosapentaenoic (C20:5n-3, EPA) acids were detected. The properties of the synthesised biodiesel were then determined (Table 3) and compared to the limits established by the international biodiesel specifications (EN14214 and ASTM D6751). All FAME related properties, namely total FAME, cetane number (CN), iodine value (IV), cold filter plugging point (CFPP), linolenic acid and PUFA ≥4 double bonds (db) contents, were within or close to the values established in the EN14214 specification. The measured density, kinematic viscosity and oxidation stability were 0.85 Kg L−1, 3.64 mm2 s−1 and 4.74 h, respectively. Produced biodiesel was devoid of glycerol (total and free) and acylglycerols (mono-, di- and triacylglycerols). The levels of group I (Na + K) and group II (Ca + Mg) metals were 0.45 and 0.05 mg Kg−1, respectively. Hence, both values were below the maximum limits specified by both standards. The phosphorous content was, however, higher than the specified limits.
Discussion
Microalgae intended for large-scale production need to be robust and present high growth rates in order to withstand wide environmental conditions and outgrow competitors and predators. In this sense, the enrichment step carried out before the FACS isolation step promoted the isolation of strains able to outcompete other cells also found in environmental samples. This step is thus crucial for the selection of robust microalgae that are able to become dominant even under challenging conditions and in the presence of contaminants17,19. Strains of interest for biodiesel production should develop clusters with a higher number of events during FACS isolation, combined with a higher lipid-BODIPY signal. From an initial pool of 96 isolates obtained by authors’ FACS-based methodology, Tetraselmis sp. CTP4 was selected as a promising biodiesel feedstock due to the combination of dominance over contaminants and lipid content. Strain identity was confirmed by phylogenetic analysis showing that Tetraselmis sp. CTP4 belongs to the striata/convolutae clade, in accordance with data reported by Arora et al.20.
The robustness of this euryhaline microalga was evaluated by growing cultures in non-sterile urban wastewater. Tetraselmis sp. CTP4 displayed similar growth curves in both the control MAM and the wastewater and, most importantly, it dominated over the microorganisms naturally present in the wastewater. Strains isolated in areas of wastewater discharges usually show high tolerance to oxidative stress and are often well suited for wastewater treatment21. Moreover, recent trials have shown that Tetraselmis sp. CTP4 can grow at salinities ranging from ~1 to 100‰ (data not shown). Halotolerant strains display a key advantage for large-scale production, since the manipulation of the salinity (high vs. low salt shifts) in the culture medium can manage and contain possible contaminants, without affecting significantly the biomass productivity of the cultures. This feature is also a key feature to recycle the marine culture medium after dewatering, upon which wide variations of salinity can occur, as recently reported by Fon-Sing et al.22 in pilot scale open raceways used to grow a euryhaline Tetraselmis strain.
Harvesting/dewatering of cultures is a main constraint in the whole microalgal production pipeline, due to the high-energy demands associated with biomass recovery from massive amounts of water23,24. In this sense, the ability of Tetraselmis sp. CTP4 to settle down naturally is another crucial advantage, as it allows the removal of 80% of the culture medium after a 6-h sedimentation step without the addition of flocculants or the use of a pre-concentration procedure. Through this approach only 20% of the culture volume needs to be harvested using common methods (e.g. centrifugation and filtration), having thus the potential of decreasing significantly the costs of biomass dewatering.
To compare the lipid production of cultures, cells were cultured under nutrient repletion (N+) and depletion (N−) using a two-stage growth system. Results showed that cells grown under nutrient depletion yielded a 3-fold increase in the total lipid content and significantly higher cell size (15–22 μm) than the N+ treatment (9–12 μm). Lipid accumulation was only triggered upon nutrient starvation, which might affect amino acid levels needed for protein synthesis, providing a higher number of carbon skeletons that will be available to triacylglycerol biosynthesis25,26. The two-stage approach used for inducing lipid accumulation in Tetraselmis sp. CTP4 effectively improved the lipid productivity of this microalgae, leading to a 2-fold increase in cultures exposed to nutrient depletion. The results of this two-stage system are in accordance with Gouveia et al.27 and Campenni’ et al.28. However, they do not match the results recently published by Kim et al.29, where N+ cultures displayed higher lipid productivity. Although a strain-dependent response cannot be excluded, such difference may be explained by the short induction period of the 2nd stage (36 hours) used by these authors, which probably did not enable an effective lipid induction in the N− cultures. In the present work, the two-stage approach only promoted lipid accumulation in Tetraselmis sp. CTP4 after 48 hours of nutrient starvation.
The overall biomass and lipid productivities established in the present work for Tetraselmis sp. CTP4 matched those from previous reports on other Tetraselmis strains (Table 1). Moreover, chlorophytes and particularly those of the Tetraselmis genus have already been shown to be able to grow in outdoor systems, and can be promising feedstocks for the production of microalgae-based biofuels30,31,32. The latter conclusion is confirmed by the fact that biodiesel synthesised from wet biomass of Tetraselmis sp. CTP4 displayed values within or close to the limits defined by the EN14214 and ASTM D6751 specifications (Table 3). From all properties investigated, the phosphorus content was the only parameter clearly outside the limits described in both specifications. However, this result was expected since, generally, microalgae oils present a significant amount of phospholipids33,34 that are co-extracted with the triacylglycerols. Therefore, removal of phospholipids (e.g. degumming) from the microalgae oil is essential in order to reduce the content of phosphorus in microalgal biodiesel and fulfil the limits of both specifications34,35.
Most microalgae strains fail to address the properties related with the saturation of the lipid profile, namely, the content of linolenic acid, PUFA ≥4 db, IV, CN, and most importantly, the oxidation stability8,10,36. In fact, the oxidative stability of the produced biodiesel (4.74 h) is to the authors’ knowledge the highest value reported for B100 microalgae-based biodiesel, except for that of Scenedesmus sp. (5.42 h)37. Perrier et al.11 and Chen et al.37,38 produced B100 biodiesel from Chlorella protothecoides, Nannochloropsis sp. and a dinoflagellate with induction periods of 4.52, 0.8–1.93 and 1.02 h, respectively. The oxidative stability of CTP4 biodiesel was also significantly higher than the values previously reported for the biodiesel produced from other vegetable sources, such as soybean (3.9 h), palm (3.52 h), rice bran (1.7 h) and sunflower (0.4 h) oils39,40,41,42. Such oxidative stability is probably related with the FAME profile of CTP4 and consequently of the produced biodiesel that revealed only trace values of long-chain PUFA, and high contents of palmitic, oleic and linoleic acids, accounting for nearly 75% of the TFA. The FAME profile of the biodiesel produced from CTP4 presents a lower degree of unsaturation than those of other Tetraselmis strains previously published in the literature10,31,43,44. The same is observed when the lipid profile of CTP4 is compared with that of most common commercial strains of microalgae, such as Nannochloropsis oculata and Phaeodactylum tricornutum10. The low unsaturation degree of Tetraselmis sp. CTP4 is a crucial advantage for biodiesel production, as recently highlighted in several reports8,10,11,12.
In conclusion, Tetraselmis sp. CTP4 displays several promising features as a biodiesel feedstock, including robustness, high biomass and lipid productivities, and potential for reduced downstream costs related with biodiesel refining and biomass dewatering.
Methods
Microalgae isolation and culture scale-up
Tetraselmis sp. CTP4 was isolated near a wastewater treatment plant in Ria Formosa, a coastal lagoon located in the south of Portugal (Algarve), by a microplate-based high throughput screening procedure described in Pereira et al.17. Briefly, water samples were supplemented with concentrated MAM17 and left exposed to indirect sunlight for approximately 1–2 weeks. Afterwards, aliquots were taken and stained with BODIPY 505/515 (4,4-difluoro-1, 3, 5, 7-tetramethyl-4-bora-3a,4a-diaza-s-indacene; Life Technologies Europe BV, Porto, Portugal) as described in Cooper et al.45 to prepare for flow cytometry. Stained samples were acquired in a Becton Dickinson FACS Aria II (BD Biosciences, Erembodegem, Belgium) equipped with a blue and red laser (488 and 633 nm, respectively) and FACSDiva (version 6.1.3) software. Four channels were used to record the fluorescence signal, namely FL1, FL2, FL3 and FL4 centred at 530/30, 585/42, 695/40 and 660/20 nm, respectively, after excitation with the blue (FL1-FL3) or red (FL4) laser. Cells emitting higher levels of fluorescence due to chlorophyll pigments and lipids stained with BODIPY were sorted directly onto 96-well microplates containing 250 μL of solid (agar) MAM and onto microscope slides. Colonies growing in the wells of the microplates were transferred to Petri dishes containing agar supplemented with MAM. The biomass growing on the Petri dishes was scrapped and transferred to 100-mL Erlenmeyer flasks with sterilized seawater and MAM and later transferred to 1-L photobioreactors with aeration.
Microscopy
Microscopic images were acquired in a Zeiss AXIOMAGER Z2 microscope, with a coollSNApHQ2 camera and AxioVision software version 4.8 (Carl Zeiss MicroImaging GmbH, Göttingen, Germany), using the 100 × lens. Brightfield microscopy was carried out using differential interference contrast (DIC), while Zeiss 38 He filter set (Carl Zeiss MicroImaging GmbH, Gõttingen, Germany) for fluorescein isothiocyanate (FITC) was used to acquire the fluorescence images. Samples used for fluorescence microscopy were stained with BODIPY 505/515 as described for the flow cytometry analysis. Images were treated using Image J software (Research Service Branch, NIH, Bethesda, MD).
Taxonomic identification
This strain was identified by means of 18S rDNA sequencing. DNA extraction was performed with the EZNA DNA plant extraction kit (Omega Bio-Tek, Norcross, GA) according to the manufacturer’s guidelines. The obtained DNA was amplified by PCR with the primers 18SUnivFor (5′-ACCTGGTTGATCCTGCCAGT-3′) and 18SUnivRev (5′-TCAGCCTTGCGACCATAC-3′) as described in Pereira et al.17,19 and sequenced at an in-house DNA sequencing facility equipped with an Applied Biosystems 3130XL DNA sequencer (Life Technologies BV, Porto, Portugal). The obtained sequence was deposited in GenBank with the accession number KX278369, and compared with the GenBank database using BLASTn (https://blast.ncbi.nlm.nih.gov). The sequences were aligned and visually inspected using CLC Sequence Viewer (v. 7.6.1, Quiagen) and curated with Gblocks v. 0.91b software46. Curation was performed allowing gap positions within the final blocks and a maximum of 8 contiguous nonconserved positions and a minimum block length of 5 nucleotides. Phylogenetic analysis was performed using Maximum-likelihood (ML) and Bayesian inference (BI). The substitution models that best fit the data set were selected using MrModeltest2 v.2.347 and PAUP* v.4.0b1048 applying the Akaike information criterion (AIC; Akaike 1974). ML analysis was performed using RaxML v. 7.0.449, assuming a GTR + I + G substitution model with 400 bootstrap replicates. Posterior probabilities were determined by Markov Chain Monte Carlo (MCMC) sampling in MrBayes v. 3.1.250,51. MrBayes analyses were also conducted using the model GTR + I + G with 6 chains for 10,000,000 MCMC generations, sampling every 1,000th generation and using the default for all the other settings. The MCMC runs convergence and burn-in were determined through the analysis of the generations vs. log probability plot using the trace analysis tool TRACER v1.652. The final tree was drawn with FigTree v.1.3.153.
Microalgae growth
All experiments were performed in a specialized growth chamber (Aralab Fitoclima S 600 PL clima plus 400), at 20 ± 0.5 °C, under continuous lighting (100 μmol m−2 s−1). Cultures were grown using 100 mL glass reactors, aerated continuously with filtered (0.2 μm) compressed air. Standard culture medium was seawater (salinity = 36‰) supplemented with MAM. For the wastewater experiment, non-nitrified sewage effluent was supplied by Quinta do Lago wastewater treatment plant (WWTP, Quinta do Lago, Algarve, Portugal).
Lipid induction
Lipid induction assays were carried out using a two-stage growth system. Cultures were grown until day 10, under controlled conditions, as described in the previous section to allow the optimal growth of cultures reaching a high cell concentration (1st stage). At this stage, the nitrate content of the growth medium was completely depleted as determined by spectrophotometric methods described in APHA54. At day 10, the produced inoculum was either supplemented with concentrated MAM (N+, nutrient replete) or was left without nutrients (N−, nutrient depleted) to promote lipid induction (2nd stage). All the experiments were carried out in triplicate and average values are reported. Results were statistically analysed using SPSS (release 15.0, SPSS Inc., Chicago, IL) software, using analysis of variance (one-way ANOVA) and Tukey HSD post-hoc test with a confidence interval of 95%.
Growth evaluation and chemical analysis of cultures
Determination of algal growth
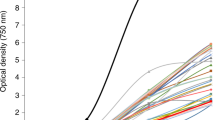
Microalgal biomass concentration was determined by measuring the optical density of the cell culture in a 96-well plate spectrophotometer (Biotek Synergy 4) at 750 nm. CC was measured using a Neubauer counting chamber according to the manufacturer indications and through flow cytometry using CountBright™ absolute counting beads. For biomass concentration, expressed in a dry weight (DW) basis, 10 mL of algal suspension was filtered through a 0.45 μm cellulose acetate filter, washed with ammonium formate (37 g L−1) and dried in an oven with forced air circulation at 60 °C until constant weight.
Total lipid determination
Total lipid content was determined following the Bligh & Dyer method55 with a few modifications as described in Pereira et al.17. Briefly, biomass was extracted with a mixture of chloroform, methanol and water (2:2:1), and homogenised with an IKA Ultra-Turrax disperser (IKA-Werke GmbH, Staufen, Germany) for 2 minutes. Phase separation was achieved by centrifugation, and the chloroform phase was transferred to new vessels with a Pasteur pipette. Afterwards, a known volume of chloroform (0.5–1 mL) was pipetted to pre-weighed tubes and evaporated overnight. The resulting dried residue was weighed and compared with the obtained DW to allow an accurate determination of the lipid fraction.
Biodiesel synthesis
Lipids were extracted directly from wet biomass as described by Yang et al.56 with modifications. Briefly, 100 g of wet microalgae paste were dispersed in 250 mL of absolute ethanol at reflux temperature for 120 minutes (EtOH-1). Afterwards, the crude ethanol extract was separated from the remaining biomass by centrifugation (4000 g, 10 minutes). The biomass was further extracted using the same aforementioned conditions for 60 (EtOH-2, 200 mL) and 30 (EtOH-3, 150 mL) minutes. All extracts were pooled and the ethanol was evaporated from the mixture using a rotatory evaporator.
Extracted lipids were converted to biodiesel by acid catalysed transesterification using the method described in Gangadhar et al.10 with modifications. Briefly, a solution of methanol and concentrated sulphuric acid (2% H2SO4 in methanol) was added to a round bottom flask containing the extracted lipids. The reaction mixture was stirred at reflux temperature for approximately 4 hours. The conversion of triacylglycerols into FAME was followed by thin-layer chromatography, using hexane and ethyl acetate (95:5 v/v) as mobile phase. Upon reaction completion, the solvent was evaporated using a rotatory evaporator and the fatty acids were sequentially extracted three times with hexane. The resulting fractions were pooled and washed with distilled water to neutralize the acid.
Fatty acid methyl esters profile
Produced biodiesel was analysed on a Bruker GC-MS (Bruker SCION 456/GC, SCION TQ MS) equipped with a ZB-5MS (30 m × 0.25 mm internal diameter, 0.25 μm film thickness, Phenomenex) capillary column using helium as carrier gas. The temperature program was 60 °C (1 min), 30 °C min−1 to 120 °C, 5 °C min−1 to 250 °C, and 20 °C min−1 to 300 °C (2 min). Injection temperature was 300 °C. For identification and quantification of the FAME total ion mode was used. Because of differences in the response factors, for each FAME separate calibration curves were determined in triplicate, using the Supelco® 37 Component FAME Mix (Sigma-Aldrich, Sintra, Portugal) commercial standard. In the case where no standard was available, the response factor of the most similar FAME, in terms of structure, was used. Results are expressed as a percentage of total FAME content.
Biodiesel properties
The density of the produced biodiesel was determined at 15 °C using a certified Lenz pycnometer. Biodiesel kinematic viscosity was measured at 40 °C using a micro Ubbelohde viscometer in accordance with ISO 3105. Glycerol and acylglycerols contents were determined as per EN14105 method. Group I and II metals, as well as the phosphorous content, were determined by a microwave plasma atomic emission spectrometry (MP-AES 4200, Agilent technologies) according to Agilents’ technical note (5990-9005EN).
The CN of the FAME mix (CNmix) was estimated using the equation described in Knothe57, relating the CN (CNc) and relative amount (Ac) of each FAME in the biodiesel mixture:

The oxidation stability was estimated using a Rancimat (model 743) according to the standard EN 14112:2003.
IV was calculated using the factors estimated for different FAME according to the EN14214.
The CFPP was calculated using the equations proposed by Ramos et al.58. This model relies on the estimation of the CFPP through the determination of the long chain saturated factor (LCSF) in accordance with the following equations:


Additional Information
How to cite this article: Pereira, H. et al. Isolation of a euryhaline microalgal strain, Tetraselmis sp. CTP4, as a robust feedstock for biodiesel production. Sci. Rep. 6, 35663; doi: 10.1038/srep35663 (2016).
References
Borowitzka, M. A. High-value products from microalgae-their development and commercialisation. J. Appl. Phycol. 25, 743–756 (2013).
Chisti, Y. Biodiesel from microalgae. Biotechnol. Adv. 25, 294–306 (2007).
Wijffels, R. H. & Barbosa, M. J. An outlook on microalgal biofuels. Science 329, 796–799 (2010).
Davis, R., Aden, A. & Pienkos, P. T. Techno-economic analysis of autotrophic microalgae for fuel production. Appl. Energ. 88, 3524–3531 (2011).
Benemann, J. R. Microalgae aquaculture feeds. J. Appl. Phycol. 4, 233–245 (1992).
Borowitzka, M. A. Microalgae for aquaculture: opportunities and constraints. J. Appl. Phycol. 9, 393–401 (1997).
Adarme-Vega, T. C. et al. Microalgal biofactories: a promising approach towards sustainable omega-3 fatty acid production. Microb. Cell Fact. 11, 96 (2012).
Knothe, G. Will biodiesel derived from algal oils live up to its promise? A fuel property assessment. Lipid Technol. 23, 247–249 (2011).
Knothe, G. Fuel properties of highly polyunsaturated fatty acid methyl esters. Prediction of fuel properties of algal biodiesel. Energ. Fuels 26, 5265–5273 (2012).
Gangadhar, K. N. et al. Assessment and comparison of the properties of biodiesel synthesized from three different types of wet microalgal biomass. J. Appl. Phycol. 28, 1571–1578 (2016).
Perrier, B. et al. Production of a methyl ester from the microalgae Nannochloropsis grown in raceways on the French west coast. Fuel 153, 640–649 (2015).
Piligaev, A. V. et al. Isolation of prospective microalgal strains with high saturated fatty acid content for biofuel production. Algal Res. 12, 368–376 (2015).
Acreman, J. Algae and cyanobacteria: isolation, culture and long-term maintenance. J. Ind. Microbiol. 13, 193–194 (1994).
Reckermann, M. Flow sorting in aquatic ecology. Sci. Mar. 64, 235–246 (2000).
Sinigalliano, C. D. et al. Viable cell sorting of dinoflagellates by multiparametric flow cytometry. Phycologia 48, 249–257 (2009).
Doan, T. Y., Sivaloganathan, B. & Obbard, J. P. Screening of marine microalgae for biodiesel feedstock. Biomass Bioenerg. 35, 2534–2544 (2011).
Pereira, H. et al. Microplate-based high throughput screening procedure for the isolation of lipid-rich marine microalgae. Biotechnol. Biofuels 4, 1–12 (2011).
Mutanda, T. et al. Bioprospecting for hyper-lipid producing microalgal strains for sustainable biofuel production. Bioresour. Technol. 102, 57–70 (2011).
Pereira, H. et al. Isolation and fatty acid profile of selected microalgae strains form the Red sea for biofuel production. Energies 6, 2773–2783 (2013).
Arora, M., Anil, A. C., Leliaert, F., Delany, J. & Mesbahi, E. Tetraselmis indica (Chlorodendrophyceae, Chlorophyta), a new species isolated from salt pans in Goa, India. Eur. J. Phycol. 41, 61–78 (2013).
Osundeko, O., Davies, H. & Pittman, J. K. Oxidative stress-tolerant microalgae strains are highly efficient for biofuel feedstock production on wastewater. Biomass Bioenerg. 56, 284–294 (2013).
Fon-Sing, S., Isdepsky, A., Borowitzka, M. A. & Lewis, D. M. Pilot-scale continuous recycling of growth medium for the mass culture of a halotolerant Tetraselmis sp. in raceway ponds under increasing salinity: a novel protocol for commercial microalgal biomass production. Bioresour. Technol. 161, 47–54 (2014).
Uduman, N., Qi, Y. & Danquah, M. K. Dewatering of microalgal cultures: A major bottleneck to algae-based fuels. J. Renew. Sustain. Energ. 2, 1–15 (2010).
Sharma, K. K., Garg, S., Li, Y., Malekizadeh, A. & Schenk, P. M. Critical analysis of current microalgae dewatering techniques. Biofuels 4, 304–397 (2013).
Converti, A., Casazza, A. A., Ortiz, E. Y., Perego, P. & Del Borghi, M. Effect of temperature and nitrogen concentration on the growth and lipid content of Nannochloropsis oculata and Chlorella vulgaris for biodiesel production. Chem. Eng. Process. 48, 1146–1151 (2009).
Procházková, G., Brányiková, I., Zachleder, V. & Brányik, T. Effect of nutrient supply status on biomass composition of eukaryotic green microalgae. J. Appl. Phycol. 26, 1359–1377 (2014).
Gouveia, L., Marques, A., Silva, T. L. & Reis, A. Neochloris oleoabundans UTEX # 1185: a suitable renewable lipid source for biofuel production. J. Ind. Microbiol. Biotechnol. 36, 821–826 (2009).
Campenni’, L., Nobre, B. P., Santos, C. A., Oliveira, A. C., Aires-Barros, M. R., Palavra, A. F. & Gouveia, L. Carotenoids and lipids production of autotrophic microalga Chlorella protothecoides under nutritional, salinity and luminosity stress conditions. Appl. Microbiol. Biotechnol. 97, 1383–1393 (2013).
Kim, G., Bae, J. & Lee, K. Nitrate repletion strategy for enhancing lipid production from marine microalga Tetraselmis sp. Bioresour. Technol. 205, 274–279 (2016).
Hu, Q. et al. Microalgal triacylglycerols as feedstocks for biofuel production: perspectives and advances. Plant J. 54, 621–639 (2008).
Huerlimann, R., de Nys, R. & Heimann, K. Growth, lipid content, productivity, and fatty acid composition of tropical microalgae for scale-up production. Biotechnol. Bioeng. 107, 245–257 (2010).
Zhou, W. et al. Local bioprospecting for high-lipid producing microalgal strains to be grown on concentrated municipal wastewater for biofuel production. Bioresour. Technol. 102, 6909–6919 (2011).
Lu, S. H. et al. Phospholipid metabolism in an industry microalga Chlorella sorokiniana: the impact of inoculum sizes. PLoS ONE 8, 11 (2013).
Iyer, R. The issue of reducing or removing phospholipids from total lipids of a microalgae and an oleaginous fungus for preparing biodiesel. Biofuels 7, 55–72 (2016).
Dibenedetto, A., Collucci, A. & Pastore, C. Heterogenous Catalysis Applied To Conversion Of Biogenic Substances, Platform Molecules And Oils. In: Michele, A., Dibendetto, A. & Dumeignil, F. editors. Biorefinery: From Biomass to Chemicals and Fuels. Berlin/Boston: Walter de Gruyter GmbH & Co, 279–295 (2012).
Stansell, G. R., Gray, V. M. & Sym, S. D. Microalgal fatty acid composition: implications for biodiesel quality. J. Appl. Phycol. 24, 791–801 (2012).
Chen, L., Liu, T., Zhang, W., Chen, X. & Wang, J. Biodiesel production from algae oil high in free fatty acids by two-step catalytic conversion Bioresour. Technol. 111, 208–214 (2012).
Chen, Y. H., Huang, Y. B., Chiang, H. T. & Tang, C. T. Fuel properties of microalga Chlorella protothecoides oil biodiesel and its blends with petroleum diesel. Fuel 94, 270–273 (2012).
Mittelbach, M. & Schober, S. The influence of antioxidants on the oxidation stability of biodiesel. J. Am. Oil Chem. Soc. 80, 817–823 (2003).
Knothe, G. Some aspects of biodiesel oxidative stability. Fuel Process. Technol. 88, 669–677 (2007).
El Boulifi, N., Bouaid, A., Martinez, M. & Aracil J. Optimization and oxidative stability of biodiesel production from rice bran oil. Renew. Energ. 53, 141–147 (2013).
Botella, L., Bimbela, F., Martín, L., Arauzo, J. & Sánchez, J. L. Oxidation stability of biodiesel fuels and blends using the Rancimat and PetroOXY methods. Effect of 4-allyl-2,6-dimethoxyphenol and catechol as biodiesel additives on oxidation stability. Front. Chem. 2, 1–9 (2014).
Molina Grima, E., Garcia Camacho, F., Sánchez Perez, J. A. & Garcia Sánchez, J. L. Biochemical productivity and fatty acid profiles of Isochrysis galbana Parke and Tetraselmis sp. as a function of incident light intensity. Process Biochem. 29, 119–126 (1994).
Montaini, E., Chini Zittelli, G., Tredici, M. R., Molina Grima, E., Fernández Sevilla, J. M. & Sánchez Perez, J. A. Long-term preservation of Tetraselmis suecica: influence of storage on viability and fatty acid profile. Aquaculture 134, 81–90 (1995).
Cooper, M. S., Hardin, W. R., Petersen, T. W. & Cattolico, R. A. Visualizing “green oil” in live algal cells. J. Biosci. Bioeng. 109, 198–201 (2010).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol. 56, 564–577 (2007).
Nylander, J. A. A. MrModeltest 2.0. Program distributed by the author. Evolutionary Biology Centre, Uppsala University. Norbyvagen 18 D. SE-752 36, Uppsala, Sweden (2004).
Swofford, D. L. PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. Sinauer Associates, Inc. Publishers. Sunderland, Massachusetts. Available at http://paup.csit.fsu.edu/ (2003).
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics. 22, 2688–2690 (2006).
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian inference of phylogenetic trees under mixed model. Bioinformatics 19, 1572–1574 (2003).
Rambaut, A., Suchard, M. A., Xie, D. & Drummond, A. J. TRACER v1.6, Available at: http://beast.bio.ed.ac.uk/Tracer (2014).
Rambaut, A. FigTree v1.4.2, Available at: http://tree.bio.ed.ac.uk/software/figtree (2006).
APHA, AWWA, WEF. American Public Health Association, American Water Works Association, and Water Environment Federation In Standard Methods for the Examination of Water and Wastewater 20th edn (eds Clesceri, L. S., Greenberg, A. E., Eaton, A. D. ) Part 4000 (APHA, 1999).
Bligh, E. G. & Dyer, W. J. A rapid method for the total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
Yang, F. et al. Extracting lipids from several species of wet microalgae using ethanol at room temperature. Energ. Fuels 29, 2380–2386 (2015).
Knothe, G. A comprehensive evaluation of the cetane numbers of fatty acid methyl esters. Fuel 119, 6–13 (2014).
Ramos, M. J., Fernández, C. M., Casas, A., Rodriguez, L. & Perez, A. Influence of fatty acid composition of raw materials on biodiesel properties. Bioresour. Technol. 100, 261–268 (2009).
Rodolfi, L. et al. Microalgae for oil: strain selection, induction of lipid synthesis and outdoor mass cultivation in a low-cost photobioreactor. Biotechnol. Bioeng. 102, 100–112 (2009).
Fon Sing, S. & Borowitzka, M. A. Isolation and screening of euryhaline Tetraselmis spp. suitable for large-scale outdoor culture in hypersaline media for biofuels. J. Appl. Phycol. 28, 1–14 (2016).
Acknowledgements
The authors would like to thank Cristina Oliveira (PhD, LNEG, Portugal) for the kind determination of the oxidative stability of the biodiesel samples. The present work was funded by the Portuguese national budget and the CCMAR/Multi/04326/2013 grant of the Foundation for Science and Technology (FCT). H.P. (SFRH/BD/105541/2014) and C.B.S. (SFRH/BD/78062/2011) are FCT doctoral research fellows. K.N.G. (SFRH/BPD/81882/2011) and L.C. (IF/00049/2012) were supported by FCT as a post-doctoral research fellow and as FCT Investigator Programme recipient, respectively.
Author information
Authors and Affiliations
Contributions
H.P. conceived, designed and performed the experiments; prepared the figures/tables and wrote the manuscript. K.N.G. and F.X.M. designed the experiments, performed the lipid chemistry and drafted the manuscript. P.S.C.S. and T.S. performed the assay with the wastewater, prepared figures/tables and drafted the manuscript. C.B.d.S. and L.M.S. carried out the DNA isolation and amplification, as well as the phylogenetic analysis and drafted the manuscript. L.C., L.G., J.C.S.V. and L.B. conceived and designed the experiments, contributed to the funding and writing of the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Pereira, H., Gangadhar, K., Schulze, P. et al. Isolation of a euryhaline microalgal strain, Tetraselmis sp. CTP4, as a robust feedstock for biodiesel production. Sci Rep 6, 35663 (2016). https://doi.org/10.1038/srep35663
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep35663
This article is cited by
-
Bioprospecting of Ten Microalgae Species Isolated from Saline Water Lake for Evaluation of the Biodiesel Production
BioEnergy Research (2023)
-
Counting microalgae cultures with a stereo microscope and a cell phone using deep learning online resources
Bulletin of the National Research Centre (2022)
-
Intensive production of the harpacticoid copepod Tigriopus californicus in a zero-effluent ‘green water’ bioreactor
Scientific Reports (2022)
-
Current perspective on wastewater treatment using photobioreactor for Tetraselmis sp.: an emerging and foreseeable sustainable approach
Environmental Science and Pollution Research (2022)
-
Improvement of carotenoid extraction from a recently isolated, robust microalga, Tetraselmis sp. CTP4 (chlorophyta)
Bioprocess and Biosystems Engineering (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.