Abstract
High abundance proteins like protease inhibitors of plasma display a multitude of interactions in natural environments. Quantitative analysis of such interactions in vivo is essential to study diseases, but have not been forthcoming, as most methods cannot be directly applied in a complex biological environment. Here, we report a quantitative microscale thermophoresis assay capable of deciphering functional deviations from in vitro inhibition data by combining concentration and affinity measurements. We obtained stable measurement signals for the substrate-like interaction of the disease relevant inhibitor α-1-antitrypsin (AAT) Z-variant with catalytically inactive elastase. The signal differentiates between healthy and sick AAT-deficient individuals suggesting that affinity between AAT and elastase is strongly modulated by so-far overlooked additional binding partners from the plasma.
Similar content being viewed by others
Introduction
One of the fundamental regulatory mechanisms in cell biology is the finely tuned interaction between enzymes and their substrates1. Binding between an enzyme and a substrate is most likely strongly influenced by the composition of the biological buffer system, for example by various plasma components2. However, functional assessment of protein-protein interactions in native environments has been a major challenge to date3,4.
Immunoassays and surface plasmon resonance are the current standard approaches to quantitate concentrations in plasma and to determine affinities of potential known targets5,6. However, both methods require surface immobilization of an antigen or an antibody, potentially imposing steric hindrance and molecular activity problems. Immobilization-free approaches based on fluorescence resonance energy transfer (FRET) require availability and labeling of both interacting partners7,8. Therefore, FRET-based methods cannot determine the affinity of a person-specific endogenous protein to a known target of interest in a plasma sample.
Microscale thermophoresis (MST) is a recently established immobilization-free affinity measurement technique that has been applied to characterize ligand-binder interactions and affinity constants ranging from pM to mM under challenging conditions9,10,11. MST employs the physical phenomenon of thermophoresis – movement of molecules in a temperature gradient. Biomolecules tend to move against the temperature gradient causing the depletion of a biomolecule in the heated spot. In the MST binding measurement, the depletion of the fluorescently labeled binder changes upon ligand binding. Measurement of depletion at increasing ligand concentrations results in a binding curve, which can be used to derive the affinity.
In our previous work using an MST-based approach, we showed how the affinity as well as the concentration of the ligand can potentially be determined in complex matrices such as blood serum12. The first limitation of this method is the sensitivity as only an additionally spiked but not initially present ligand could be detected in serum. Secondly, low affinity interactions remain a challenge, as they often require unrealizable high concentrations of the unlabeled binder to establish a complete binding curve.
An important clinically relevant example of protein interactions in plasma is the inhibition of the enzyme neutrophil elastase (NE) by α-1-antitrypsin (AAT). Low plasma levels of AAT predispose to emphysema and chronic obstructive pulmonary disease13,14. Decreased AAT levels are often caused by the relatively common Z-mutation (Glu342Lys)15 which accounts for 95% of patients with clinically relevant manifestations. Although all individuals with the Z-mutation have reduced AAT levels, development of lung emphysema is highly variable among ZZ carriers and some are minimally affected13,14,16.
The interaction between NE and AAT has been well studied in vitro using the two purified interacting proteins in standard buffers17. AAT inhibits NE irreversibly in a two-step reaction18. The first and rate-determining step is the formation of a reversible encounter complex between NE and AAT, where AAT mimics a substrate19,20. Previous approaches focused only on the concentration of AAT5,13,15,16, although kinetically efficient formation of this encounter complex also depends on the dissociation constant between NE and AAT19,20.
In this work, we developed a novel assay that overcomes the limitations of the previous approaches and allowed us to characterize the formation of the NE-AAT encounter complex at the given AAT level of a blood plasma sample under constant equilibrium conditions.
Results
Method development: combining concentration and affinity
In our assay, a low affinity inhibitor I1 (here: AAT) competes with a high affinity inhibitor I2 (here: elafin) for a catalytically inactive labeled enzyme E (here: NE) (Fig. 1a). Inactivation of E was important to prevent the removal of I1 from the equilibrium by the irreversible formation of a covalent complex, thereby allowing us to analyze the encounter reaction in the plasma environment.
Dependency of the amplitude on the AAT concentration and the dissociation constant of NE-AAT binding.
(a) Describes the theoretical background of the method. At each data point, the inhibitor I1 (here: AAT) competes with I2 (here: elafin) for the enzyme E (here: labeled inactive NE). The affinity between E and I2 is much higher than the affinity between E and I1. (b) To a constant amount of plasma (7.5%) containing I1 and a constant amount of E (500 pM) increasing concentrations of I2 were added and the thermophoretic depletion measured. Titration curves with theoretical fits are given for three individuals (patients 1, 8 and 10) as examples in (b). All individuals were homozygous for the Z-variant AAT. Each depletion value is given as mean ± S.D. from three technical replicates. The amplitude is defined as the difference of minimal depletion signals in the low end range (dark grey box with arrow) and maximal depletion signals in the high end range of I2 concentrations (light grey box with arrow). The thermophoretic depletion of free E is smaller than that of E bound to I1 or I2, and depletion at each I2 concentration indicates the proportion of bound E. (c,d) Are simulations according to the mass action laws. (c) Shows that the theoretical amplitude (simulated as the amount of bound E) increases when the dissociation constant between E and I1 increases. (d) On the other hand, the theoretical amplitude decreases with the increasing concentrations of I1.
The labeled E at a fixed final concentration of 500 pM was mixed with 12-fold diluted plasma containing the analyte I1. To obtain binding curves, increasing amounts of I2 were then added and thermophoretic depletion of free E and E bound to I1 (EI1) or I2 (EI2) was measured for each concentration of I2 (Fig. 1a,b). We observed that the depletion in the low end range of I2 concentrations varied between plasma samples while the depletion in the high end range of I2 concentrations did not change (Fig. 1b).
To compare the binding curves, we introduced a normalized parameter, the thermophoretic amplitude, by calculating the difference between the depletion signals in these two I2 concentration ranges (Fig. 1b, Supplementary Information 1). We observed that the thermophoretic amplitude varied between plasma samples. We hypothesized that I1 concentrations and, more importantly, I1 affinities to E are not necessarily constant in the person-specific proteomic environment of plasma samples (Fig. 1b).
To characterize the system at different affinities between I1 and E (Fig. 1c) and different I1 concentrations (Fig. 1d), we described it with the mass action law equations and simulated the corresponding binding curves. The simulation results indicate that the thermophoretic amplitude increases with an increasing KD(EI1) (Fig. 1c small insertion graph) and decreases with increasing I1 concentrations (Fig. 1d small insertion graph). Hence, the thermophoretic amplitude reports on both I1 concentration and KD(EI1) (a detailed mathematical analysis is provided in the Supplementary Information 2).
Thermophoretic amplitude is better correlated with disease status
To evaluate whether the thermophoretic amplitude that includes affinity variations represents a better lung function parameter than AAT concentration alone, we determined both parameters in plasma samples of individuals homozygous for the AAT Z-variant. We then compared these to FEV1 (the forced expiratory volume in one second) expressed as the percentage (%) predicted for age, sex, height, and race, which is widely used to assess the level of airway obstruction and bronchoconstriction in AAT-deficient individuals16,21.
In order to avoid measuring the concentration of inactive AAT, we titrated the concentration of functional plasma AAT with purified and active site-titrated NE. We observed a significantly lower thermophoretic amplitude (P = 0.0008) with plasma samples from individuals showing a high FEV1 (≥80%) than from those with low FEV1 (<80%) (Fig. 2a), while the concentration of functional AAT did not differ between the two groups (P = 0.6368). Hence, inclusion of affinity with our method resulted in a much better relationship to disease status and severity. To explain the discrepancy between the two quantification methods, we reasoned that KD(EI1) of the Z-variant in fact varied in the plasma samples.
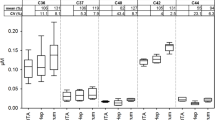
Thermophoretic amplitude, but not plasma concentration of AAT is associated with disease severity.
(a) ZZ-carriers (n = 26) were divided into two groups with high FEV1 (≥80%, n = 8) and low FEV1 (<80%, n = 18), which is a clinical parameter of airway obstruction. The thermophoretic amplitude was significantly lower (P = 0.0008) in samples with high FEV1 (***P < 0.001; ns not significant). The amplitude is given as a mean value determined from at least two independent experiments. (b) The concentration of functional AAT was not significantly different (P = 0.6368) between the groups with high FEV1 (≥80%) and low FEV1 (<80%). The AAT concentration was measured twice by elastase titration in duplicates, and is given as a mean value here.
Z-variant AAT is sensitive to plasma environments
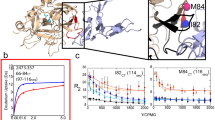
To investigate whether affinity is indeed sensitive to a given plasma environment, we determined the KD(EI1) of the AAT Z-variant in the two AAT-deficient plasma pools from persons with high (≥80%) and low FEV1 (≤50%) values (Fig. 3). To facilitate the recombinant production of the aggregation-prone Z-variant, we combined three stabilizing mutations derived from the work of Lee et al.22 – Met374Ile, Ser381Ala, Lys387Arg – and introduced them into the Z-variant. As a control, we expressed AAT without the Z-mutation but containing the same stabilizing mutations. The affinity of the Z-variant to NE was more than two times better in the plasma pool from individuals with high FEV1 (≥80%) (KD(EI1) = 500 ± 100 nM) than from individuals with low FEV1 (≤50%) (KD(EI1) = 1300 ± 250 nM), confirming the plasma-dependent change in the affinity between NE and Z-variant AAT as observed with our newly developed assay. Additionally, we found that the Z-mutation improves the encounter reaction of AAT with NE (Supplementary Fig. 1a) and Z-variant AAT is more susceptible to plasma components than control AAT (Supplementary Fig. 1b).
Plasma-dependent change in affinity of NE-AAT binding.
We compared the affinity between AAT and NE in two pools of plasma from individuals with high FEV1 (≥80%, n = 8) and low FEV1 (≤50%, n = 12). The affinity between Z-variant of AAT and NE was higher in plasma with high FEV1 (≥80%) (KD(EI1) = 500 ± 100 nM) compared to low FEV1 (≤50%) (KD(EI1) = 1300 ± 250 nM). Presented binding curves represent example measurements where each measurement point (mean ± S.D.) was derived from three technical replicates. Fitted binding curves and KD(EI1) values (mean ± S.D.) were derived from global fitting of four measurements (three independent protein expressions). The measurements were performed in 7.5% plasma and with 5 nM of labeled NE.
Discussion
By incorporating a high affinity ligand in thermophoretic measurements, we expanded the spectrum of applications towards the low affinity range where e.g. enzyme-substrate interactions take place. This new approach allowed us to detect unexpected plasma-dependent interferences with AAT, suggesting that the functional capacity of the Z-variant AAT is determined by the heterogeneity of other plasma components, which modulate the affinity of AAT towards target enzymes. For example, it was reported23,24 and has been confirmed by our group (unpublished) that lipoproteins bind to wildtype AAT, thereby influencing its inhibition of NE and thus could also affect the reactivity of the Z-variant in the plasma environment.
The analysis of the interaction between AAT and NE in plasma with our assay provides a possible explanation of the variability in disease development in ZZ carriers. Normally, the reduced AAT plasma levels in ZZ carriers are compensated by the enhanced affinity of Z-variant AAT to NE. However, the enhanced affinity between Z-variant AAT and NE can be disrupted by plasma factors, especially in individuals who have higher concentrations of interfering plasma components.
Our work underscores the importance of the endogenous personally unique and distinct environment where substrates interact with enzymes with relatively low affinity in a complex web of other proteins and lipid components. Such personalized variations and effects have often been overlooked. Our approach may open a way to study how other components from the native environments interfere with biochemical reactions.
Methods
Production of recombinant proteins
As the natural Z-variant is on a M1(Ala213) background, we amplified a cDNA fragment encoding the M1(Ala213) variant using the primer DJ3689 (5′-GACTTCCACGTGGACCAGGCGACCACCGTGAA-3′) and DJ3613 (5′-GATGACCGGTTTTTTGGGTGGGATTCACCACTT-3′) (cursive: restriction site; bold: mutation) and cloned it into the Pml I and Age I site of the previously described M1(Val213) AAT construct in pTT5 plasmid (Perera et al.25. To introduce the Z-mutation (Glu342Lys), a PCR fragment was generated with the primers DJ3557 (5′-CCACGATATCATCACCAAGTTCCT-3′) and DJ3558 (5′-GTATGGCCTCGAGGAACATGGCCCCAGCAGCTTCAGTCCCTTTCTTGTCGATGGT-3′) and inserted into the M1(Ala213) AAT pTT5 plasmid between the Eco RV and Abs I sites. The three stabilizing mutations (M374I, S381A, K387R) were introduced by PCR using the following forward and backward primers DJ 3646 (5′- CACGATATCATCACCAAGTTCCTGGAAAATGAAGACAGAAGGTCTGCCGACTTACATTTACC-3′) and DJ3696 (5′-ATGACCGGTTTTTTGGGTGGGATTCACCACTCTTCCCATGAAGAGGGGAGC CTTGGTATTTTGTTCAATAATTAAG-3′) respectively. The product was inserted into the Eco RV and Age I sites.
To produce a catalytically inactive variant of NE, we mutated the Ser195 to Ala195 by inserting an oligo duplex (DJ3532 (5′-GTGAACGTATGCACTCTGGTGCCACGTCGGCAGGCAGGCATCTGCTTCGGGGACGCT-3′) and DJ3533 (5′-CGTCCCCGAAGCAGATGCCTGCCTGCCGACGTGGCACCAGAGTGCATACGTTCACAC-3′)) into the Alf I site of the previously described wildtype mouse NE construct in pTT5 (Dau et al.26. To enable site-specific labeling of NE, we added a C-terminal short peptide with a cysteine flanked by three aspartate residues on each side by the insertion of an oligoduplex (DJ3632 (5′- CTAGCGACGACGATTG CGACGATG ATC-3′) and DJ3633 (5′- CTAGGATCATCGTCGCAATCGTCGTCG-3′)) into the Avr II site.
All proteins were expressed in HEK293 EBNA cells (Yves Durocher, National Research Council Canada, Montreal, Canada) in FreeStyleTM 293 expression medium (Thermo Fisher Scientific Inc.), 1% Pluronic and G418 (25 μg ml−1) at 37 °C and 8% CO2.
Labelling of NE
First, we added 1 mM DTT to the recombinant NE in storage buffer (20 mM Na2HPO4, 300 mM NaCl pH 7.4) and incubated it for two hours at room temperature to reduce all cysteine-tags. Then we removed DTT by precipitating NE with 75% ammonium sulfate. This reduction and precipitation step was repeated once. NE was dissolved in storage buffer and incubated at room temperature for one hour after adding a 5-fold molar excess of dye (Alexa Fluor® 647 NHS Ester, ThermoFisher Scientific). To remove the excess dye, the solution was added to PD MiniTrap G-10 column (GE Healthcare) according to the manufacturers instruction.
Determination of AAT concentration in blood plasma
After titrating trypsin (Sigma-Aldrich, St. Louis) with 4-nitrophenyl 4-guanidinobenzoate (Sigma-Aldrich, St Louis, USA) in veronal buffer, AAT (Athens Research & Technology, Athens, GA, USA) was titrated against trypsin. A dilution series of AAT was incubated with a constant amount of trypsin at 37 °C for 1 hour. The residual activity was measured using Boc-Gln-GLy-Arg-AMC. Thereafter, a dilution series of the titrated AAT was incubated with a constant volume of active human NE (Elastin Products Company, Inc., Owensville) at 37 °C for 1 hour. The remaining activity was measured with MCA-GEAIPTSIPPEVK(Dnp)-rr (EMC microcollections, Tübingen, Germany). All reactions were performed in 150 mM NaCl, 50 mM Tris, 0.01% Triton-X-100, pH 7.4.
Active site titrated neutrophil elastase was added to a dilution series of plasma and incubated at 37 °C for 1 hour. The plasma was diluted in 150 mM NaCl, 50 mM Tris, 0.01% Triton-X-100, pH 7.4. The residual activity was measured using MCA-GEAIPTSIPPEVK(Dnp)-rr (EMC microcollections, Tübingen, Germany).
Determination of the concentration of recombinant AAT
A dilution series of AAT-variants was incubated with 1.6 nM active site titrated human NE at 37 °C for 1 hour. The residual activity was measured using MCA-GEAIPTSIPPEVK(Dnp)-rr (EMC microcollections, Tübingen, Germany).
Statistical analysis
Results are given as median values. A Mann-Whitney test without assuming Gaussian distribution was applied to all studies (***P < 0.001). Statistical analyses were done with GraphPad Prism 6 software (GraphPad Software).
Development of the Binding Model for Simulations
Mass action law equations with two dissociation constants were used to describe the dependency of free NE on the concentration of AAT and on the affinity of AAT to NE in the presence of two NE ligands – AAT and elafin.


where
 – dissociation constant of NE – AAT interaction,
– dissociation constant of NE – AAT interaction,
 – dissociation constant of NE – elafin interaction,
– dissociation constant of NE – elafin interaction,
[E], [I1], [I2] – total concentrations of NE, AAT, and elafin in the reaction, respectively,
[EI1] – concentration of NE-AAT complex in the reaction,
[EI2] – concentration of NE-elafin complex in the reaction,
 – fraction of occupied NE,
– fraction of occupied NE,
Equation (2) was solved for [E], which was then applied to equation (1) to yield a cubic equation for x. The cubic equation was solved numerically in Igor Pro 5.03 and the smallest real root of the equation was then used to plot the fraction of bound NE x against different elafin concentrations I2 to simulate binding curves.
Determination of the thermophoretic amplitude
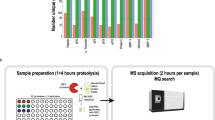
Human recombinant elafin (Sigma-Aldrich) was serially diluted (1:1) over five orders of magnitude in PBS. Separately, plasma was diluted (final concentration: 7.5%) with storage buffer substituted with 0.02% Tween 20 and anti-photobleaching enzyme and substrate components (Monolith Anti Photobleach Kit, NanoTemper Technologies) were added. This plasma solution was mixed 1:1 with PBS for background measurements. After adding fluorescently labeled NE (final concentration: 500 pM) to the plasma solution, we mixed this 1:1 with each concentration of the serial elafin dilution. The samples were loaded into the Monolith™ NT.115 premium coated capillaries (NanoTemper Technologies), and incubated at 22 °C for 2 hours. The samples were measured in the instrument (Monolith NT.115Pico, NanoTemper Technologies) at 22 °C using 60% light-emitting diode and 40% infrared laser (IR) powers with IR laser on/off times of 25/5 seconds. Each dilution point was measured in triplicates. For each plasma sample, the whole procedure was repeated three times to yield independent triplicates.
In Matlab R2014a (8.3.0.532), the time trace of the background signal was subtracted from the time traces of the sample signals. Then, the fluorescence after the temperature jump and equilibrated thermophoresis was normalized to the fluorescence before the IR laser heating yielding fluorescence depletion values. Fluorescence depletion values of three technical replicates (mean ± S.D.) were plotted in per mille units (‰) on a linear y-axis against the concentration of the serially diluted elafin on the log10 x-axis resulting in binding curves. The binding curves were shifted so that the depletion value at the baseline, where elafin concentration is saturating, was the same. In Igor Pro 5.03, a weighted fit to the quadratic solution of the mass action law was performed to yield the amplitude A of each curve:

where
KD – dissociation constant of NE – elafin interaction,
[E] – total concentration of NE,
x – total concentration of elafin,
A – the amplitude of the curve,
t – y-offset of the curve,
with A, t, and KD as free fit parameters.
Determination of dissociation constant between AAT and NE
Freshly purified AAT was serially diluted (1:1) over five orders of magnitude in storage buffer. Separately, plasma was diluted in storage buffer substituted with 0.2% Tween 20 (final concentration: 7.5%) and anti-bleaching reagents were added. This plasma solution was mixed 1:1 with storage buffer for background measurements. Fluorescently labeled NE (final concentration: 5 nM) was added to the plasma solution and mixed 1:1 with the serial dilution of AAT. The samples were incubated at 22 °C for 2 hours and measured at 22 °C in the instrument using 40% light-emitting diode and 40% infrared laser (IR) powers with IR laser on/off times of 20/5 seconds.
Binding curves were obtained as described for the determination of the thermophoretic amplitude. They were then additionally normalized to 0 at the lowest AAT concentration and to 1 at the highest AAT concentration. This corresponds to the proportion of bound NE at each titration point. The global fit of at least three replicates to the quadratic solution of the mass action law was performed to yield the dissociation constant of each curve.
Plasma samples
FEV1 was determined post bronchodilator treatment and according to the British Thoracic Society/Association of Respiratory Technicians and Physiologists (BTS/ARTP) guidelines as described previously21. All subjects provided written informed consent, and ethical approval was obtained for all aspects of this study (South Birmingham Research Ethics Committee LREC 3359).
Additional Information
How to cite this article: Dau, T. et al. Quantitative analysis of protease recognition by inhibitors in plasma using microscale thermophoresis. Sci. Rep. 6, 35413; doi: 10.1038/srep35413 (2016).
References
Overall, C. M. & Blobel, C. P. In search of partners: linking extracellular proteases to substrates. Nat. Rev. Mol. Cell Biol. 8, 245–257 (2007).
Adams, T. E. & Huntington, J. A. Thrombin-cofactor interactions: structural insights into regulatory mechanisms. Arterioscler. Thromb. Vasc. Biol. 26, 1738–1745 (2006).
Syafrizayanti, Betzen, C., Hoheisel, J. D. & Kastelic, D. Methods for analyzing and quantifying protein-protein interaction. Expert Rev. Proteomics 11, 107–120 (2014).
Sinz, A. Investigation of protein-protein interactions in living cells by chemical crosslinking and mass spectrometry. Anal. Bioanal. Chem. 397, 3433–3440 (2010).
McElvaney, N. G. Diagnosing α1-antitrypsin deficiency: how to improve the current algorithm. Eur. Respir. Rev. 24, 52–57 (2015).
Mariani, S. & Minunni, M. Surface plasmon resonance applications in clinical analysis. Anal. Bioanal. Chem. 406, 2303–2323 (2014).
Kraynov, V. S. et al. Localized Rac activation dynamics visualized in living cells. Science 290, 333–337 (2000).
Uhlik, M. T. et al. Rac-MEKK3-MKK3 scaffolding for p38 MAPK activation during hyperosmotic shock. Nat. Cell Biol. 5, 1104–1110 (2003).
Wienken, C. J., Baaske, P., Rothbauer, U., Braun, D. & Duhr, S. Protein-binding assays in biological liquids using microscale thermophoresis. Nat. Commun. 1, 1–7 (2010).
Seidel, S. A. I. et al. Microscale thermophoresis quantifies biomolecular interactions under previously challenging conditions. Methods 59, 301–315 (2013).
Jerabek-Willemsen, M. et al. MicroScale Thermophoresis : Interaction analysis and beyond. J. Mol. Struct. 1077, 101–113 (2014).
Lippok, S. et al. Direct detection of antibody concentration and affinity in human serum using microscale thermophoresis. Anal. Chem. 84, 3523–3530 (2012).
Gooptu, B., Ekeowa, U. I. & Lomas, D. A. Mechanisms of emphysema in alpha1-antitrypsin deficiency: molecular and cellular insights. Eur. Respir. J. 34, 475–488 (2009).
Sun, Z. & Yang, P. Role of imbalance between neutrophil elastase and alpha 1-antitrypsin in cancer development and progression. Lancet. Oncol. 5, 182–190 (2004).
Hutchison, D. C. Natural history of alpha-1-protease inhibitor deficiency. Am. J. Med. 84, 3–12 (1988).
Stockley, R. A. & Turner, A. M. α-1-Antitrypsin deficiency: clinical variability, assessment, and treatment. Trends Mol. Med. 20, 105–115 (2014).
Dobó, J. & Gettins, P. G. W. alpha1-Proteinase inhibitor forms initial non-covalent and final covalent complexes with elastase analogously to other serpin-proteinase pairs, suggesting a common mechanism of inhibition. J. Biol. Chem. 279, 9264–9269 (2004).
Potempa, J., Korzus, E. & Travis, J. The serpin superfamily of proteinase inhibitors: structure, function, and regulation. J. Biol. Chem. 269, 15957–15960 (1994).
Kang, U. B. et al. Kinetic mechanism of protease inhibition by alpha-1-antitrypsin. Biochem. Biophys. Res. Commun. 323, 409–415 (2004).
Gettins, P. G. W. Serpin structure, mechanism, and function. Chem. Rev. 102, 4751–4804 (2002).
Guidelines for the measurement of respiratory function. Recommendations of the British Thoracic Society and the Association of Respiratory Technicians and Physiologists. Respir. Med. 88, 165–194 (1994).
Lee, K. N., Park, S. D. & Yu, M. H. Probing the native strain iin alpha1-antitrypsin. Nat. Struct. Biol. 3, 497–500 (1996).
Gordon, S. M. et al. Rosuvastatin alters the proteome of high density lipoproteins: Generation of alpha-1-antitrypsin enriched particles with anti-inflammatory properties. Mol. Cell. Proteomics 14, 3247–3257 (2015).
Talmud, P. J. et al. Progression of atherosclerosis is associated with variation in the alpha1-antitrypsin gene. Arterioscler. Thromb. Vasc. Biol. 23, 644–649 (2003).
Perera, N. C. et al. NSP4 is stored in azurophil granules and released by activated neutrophils as active endoprotease with restricted specificity. J. Immunol. 191, 2700–2707 (2013).
Dau, T., Sarker, R. S. J., Yildirim, a. O., Eickelberg, O. & Jenne, D. E. Autoprocessing of neutrophil elastase near its active site reduces the efficiency of natural and synthetic elastase inhibitors. Nat. Commun. 6, 6722 (2015).
Acknowledgements
This work was supported by a joint grant (JE194/4-1 and BR 2152/2-1) of the Deutsche Forschungsgemeinschaft. This project has received funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement No. 668036 (RELENT). E.E.V. is supported by a DFG Fellowship through the Graduate School of Quantitative Biosciences Munich (QBM).
Author information
Authors and Affiliations
Contributions
T.D., E.V.E. and S.A.I.S. designed and performed the experiments. D.E.J. and D.B. supervised the project. R.A.S. provided plasma samples and clinical data. All authors discussed the results and implications and commented on the manuscript at all stages.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Dau, T., Edeleva, E., Seidel, S. et al. Quantitative analysis of protease recognition by inhibitors in plasma using microscale thermophoresis. Sci Rep 6, 35413 (2016). https://doi.org/10.1038/srep35413
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep35413
This article is cited by
-
PD-1/PD-L1 binding studies using microscale thermophoresis
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.