Abstract
In Oncidium, redox homeostasis involved in flowering is mainly due to ascorbic acid (AsA). Here, we discovered that Oncidium floral repression is caused by an increase in AsA-mediated NO levels, which is directed by the enzymatic activities of nitrate reductase (NaR) and nitrite reducatase (NiR). Through Solexa transcriptomic analysis of two libraries, ‘pseudobulb with inflorescent bud’ (PIB) and ‘pseudobulb with axillary bud’ (PAB), we identified differentially expressed genes related to NO metabolism. Subsequently, we showed a significant reduction of NaR enzymatic activities and NO levels during bolting and blooming stage, suggesting that NO controlled the phase transition and flowering process. Applying AsA to Oncidium PLB (protocorm-like bodies) significantly elevated the NO content and enzyme activities. Application of sodium nitroprusside (-NO donor) on Arabidopsis vtc1 mutant caused late flowering and expression level of flowering-associated genes (CO, FT and LFY) were reduced, suggesting NO signaling is vital for flowering repression. Conversely, the flowering time of noa1, an Arabidopsis NO-deficient mutant, was not altered after treatment with L-galacturonate, a precursor of AsA, suggesting AsA is required for NO-biosynthesis involved in the NO-mediated flowering-repression pathway. Altogether, Oncidium bolting is tightly regulated by AsA-mediated NO level and downregulation of transcriptional levels of NO metabolism genes.
Similar content being viewed by others
Introduction
Flowering is a complicated process coordinated by environmental and endogenous factors to ensure plant reproduction in appropriate conditions. Forward and reverse genetic tools have shown the critical role of genes in photoperiodism (responding to low temperatures), aging and phytohormones in the regulation of flowering1. Noteworthy, current evidence has suggested that several antioxidants, such as ascorbate (AsA) and glutathione, function as negative repressors of flowering time2,3,4,5,6. The AsA-mediated flowering time can be assessed by the following two aspects: AsA level and redox ratio. Arabidopsis VTC1 encodes a GDP-mannose pyrophosphatase gene in a Smirnoff-Wheeler pathway for AsA biosynthesis. The vtc1 mutant is deficient in AsA levels with 40% of the AsA amount of wild-type(wt) plants, and it displays facilitated flowering under a long-day photoperiod7,8. Other AsA-deficient mutants encoding different genes in the Smirnoff-Wheeler pathway displayed early flowering similar to that of vtc15. Furthermore, the expression levels of genes participating in the photoperiodic flowering pathway, including FT, and CO, are promoted in the AsA-deficient mutants. Various photoperiodic and autonomous pathway mutants are epistatic to vtc1, the flowering time of the vtc1 mutant growing under a short-day photoperiod is susceptible to light intensity. The endogenous AsA level prominently declines when the plants are in transition from the vegetative stage to the reproductive stage accompanied with an elevated expression level of OgLEAFY6. In contrast, exogenous application of AsA or its precursor causes delayed flowering of Brassica rapa, Arabidopsis and Oncidium6,9,10. This evidence suggests that the effect of AsA on the repression of flowering is flexible and rapid in response to environmental cues. AsA functions as a co-factor in numerous phytohormone biosynthesis pathways and regulates endogenous levels of gibberellin, abscisic acid, salicylic acid, and ethylene4. Therefore, the effect of AsA on flowering has been speculated to alter phytohormone levels. AsA-mediated flowering is primarily proposed in a mechanism independent of its antioxidant activity because of an invariable H2O2 level in the vtc1 mutant compared to wild type5. However, other reports have shown that H2O2 level increases before floral initiation of morning glory (Pharbitis nil)11, wheat12, Arabidopsis13 and Oncidium6. In addition, enervated flowering is present in several mutants lacking enzymes that oxidize AsA to dehydroascorbate (DHA), including APX and AsA oxidase14,15. Additionally, the negative role of NO on floral induction in Arabidopsis has been validated16. The nox1 mutant disrupts a chloroplast phosphoenolpyruvate/phosphate translocator to accumulate L-arginine at a higher level than wild type, thus exhibiting higher NO emission and delayed flowering16,17. NO produced from the nitrate-related system displays a 100-fold greater output than NO produced from an arginine-associated or NOS-like system, which demonstrates the crucial role for nitrate reductase (NaR) in NO synthesis in planta18,19. AsA is an important co-factor for NOS activity in mammals20, but the genes encoding mammal-like NOS in plants are still unidentified21.
The life cycle of Oncidium ‘Grower Ramsay’ starts off with its vegetative stage and may progress into two different life pathways as follows: either flowering with inflorescence (transition to reproductive phase) or regenerating a new axillary bud (retaining the vegetative stage) (Fig. 1a). The determining factors for these two phase-transitions are still unknown. Previously, we have demonstrated that endogenous AsA is essential for phase transition and the flowering process6. Similarly, the redox homeostasis of Oncidium is reliant on ambient temperature as well as phase transition signaled by the decrease in AsA levels in ‘pseudobulb with inflorescent bud’ (PIB) tissues22,23. However, the specific mechanism of AsA and NO signaling in repressing flowering is poorly understood. In the present study, we demonstrated that the repression of Oncidium flowering is determined by the coordinated action of ascorbic acid and nitric oxide. We presented evidences that the early flowering phenotype of Arabidopsis vtc1 mutant line was delayed after exogenous sodium nitroprusside (SNP) application. Our data also showed that the biochemical activities of NO-biosynthesis (NiR and NaR) in Arabidopsis vtc1 mutant lines were obviously lower than those of wild type. In contrast, the early flowering time of noa1, a NO accumulation-deficient mutant, was not altered after treatment with L-galacturonate, a precursor of AsA production. Altogether, we showed that the flowering repression mechanism is mainly dependent on NO level, which is mediated by AsA content through regulating the NO-biosynthetic enzymatic activity.
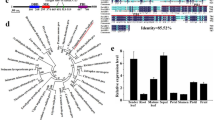
Tissue-specific sorting of Oncidium transcriptomes.
(a) Diagram of the Oncidium plant life cycle. The inflorescent bud (I) and axillary bud (AX) are formed concomitantly at each side of the pseudobulb (P) base at the vegetative stage. Either bolting or developing to next generation is regulated by a complicated genetic network (modified based upon previously published6,22). L. denotes the leaf numbering from the top to the base. (b) Heat map showing unsupervised hierarchical clustering of differential expression genes between PAB and PIB with p value <0.5 and fold change ≥5.0. Red indicates a high expression level, and green indicates a low expression level. White indicates no expression. PAB: pseudobulb with axillary bud. PIB: pseudobulb with inflorescent bud.
Results
Solexa sequencing: Statistical characterization of global gene expression
Solexa deep sequencing technology was performed to sequence the transcriptome of ‘pseudobulb with inflorescent bud’ (PIB) and ‘pseudobulb with axiliary bud’ (PAB) (Fig. 1a). After trimming adapter sequences and removing sequences shorter than 75 bases, sequencing depths of 925,937 and 665,127 contigs were achieved in PAB and PIB libraries (Supplementary Table S1) with a total of 106.1 million and 79.3 million reads, respectively. The most-aligned results displayed a total of 51,883 (47.8%) and 32,747 (30.2%) afresh-assembled unigenes, which were annotated in this manner by Nr and Swiss-Port respectively and oriented for subsequent analysis (Supplementary Table S1). The expression levels of the assembled unigenes indicated that 98,711 (90.9%) unigenes displayed similar or extremely low expression levels between the two libraries (Fig. 1b). The parallel majorities within the two categories in the PAB and PIB libraries were as follows: metabolic process and cellular process in biological processes; catalytic activity and binding in molecular functions; and organelle and cell in cellular components (Supplementary Fig. S1). Supplementary Table S2 illustrates the functional enrichment analysis of PAB library versus PIB library was done by fatiGO executed in blast2go. The most significantly identified genes were related to ribosome biogenesis (GO: 0042254), chromatin assembly or disassembly (GO: 0006333) and cell redox homeostasis (GO: 0045454) within cellular process subcategory (GO: 0009987), gluconeogenesis (GO:0006094) and ATP biosynthetic process (GO: 0006754) in metabolic process subcategory (GO: 0008152), GTPase activity (GO: 0003924) and hydrogen ion transmembrane transporter activity (GO: 0015078) within catalytic activity subcategory (GO: 0003824), GTP binding (GO: 0005525), peptide binding (GO: 0042277), magnesium ion binding (GO: 0000287), nucleic acid binding (GO: 0003676) and chromatin binding (GO: 0003682) within ‘binding’ subcategory (GO: 0005488), mitochondrial inner membrane (GO: 0005743) within ‘organelle’ subcategory (GO: 0044422) and ribosomal subunit (GO: 0033279) within ‘cell’ subcategory (GO: 00044464) (Supplementary Table S2, Supplementary Fig. S2). The majority of the mRNA transcripts from the PAB library was proposed to function in promoting next generation vegetative growth or floral repression. The genes participating in NOS orthologue proteins, GTP binding (GO: 0005525) and magnesium ion binding (GO: 0000287), were absent in the PIB libraries of inflorescence initiation (Supplementary Table S2, Supplementary Fig. S2).
Differential expression of NO- biosynthesis- related genes correlated with phase transition
The functional enrichment analysis of two transcriptomic profiles from PAB and PIB tissues were investigated. The data showed that nitrogen-associated metabolism was active in PAB and not in PIB. The genes were grouped into three clusters, namely the L-arginine-dependent pathway, the nitrite-dependent pathway and the class II non-symbiotic hemoglobin pathway (Fig. 2a). The genes participating in the L-arginine-dependent pathway displayed a consistently high expression level, while the genes in the nitrate-dependent pathway and non-symbiotic hemoglobin displayed relatively low expression levels (Fig. 2a). Genes participating in the L-arginine-dependent pathway showed little difference between the two transcriptomic libraries and they presented higher signal intensity. To further verify the Solexa analysis results, the expression levels of ten NO metabolism genes were monitored by RT-PCR (Fig. 2b–k). Nitrate reductase (NaR), nitrite reductase (NiR) and among others, displayed significantly reduced expression levels in PIB compared to those in PAB (Fig. 2d–k), similar to the patterns in heat map (Fig. 2a). In contrast, the NOS protein inhibitor (NOSPI) and NOS interacting protein (NOSIP) exhibited higher expression levels in PIB than in PAB (Fig. 2).
The expression pattern of 51 assembled unigenes involved in nitric oxide metabolism from two libraries.
(a) The pathway of NO metabolism in planta. The heatmap of expression profiling generated from two transcriptomic libraries. Red indicates a high expression level, and green indicates a low expression level. PAB: pseudobulb with axillary bud. PIB: pseudobulb with inflorescent bud. (b–k) The relative gene expression levels of the following ten genes involved in NO metabolism were monitored in PIB and PAB at the vegetative stage (□ white square), the bolting period ( gray square) and the reproductive stage (■ black square): NOS-interacting protein (NOSIP); NOS protein inhibitor (NOSPI); NOS-associated protein (NOSAP); nitric oxide synthase-like (NOS); nitrite transporter (NiT); nitrite reductase (NiR); nitrate reductase (NaR); nitrate transporter (NaT); molybdopterin synthase catalytic subunit (MPT) and non-symbiotic hemoglobin (Hb-1). Gene expression levels in each sample were compared to those in PAB at the vegetative stage. Error bars indicate standard deviation (s.d.). Asterisks represent significant differences from PAB tissue at the vegetative stage according to two-way ANOVA (*p < 0.05; **p < 0.005; ***p < 0.0005).
gray square) and the reproductive stage (■ black square): NOS-interacting protein (NOSIP); NOS protein inhibitor (NOSPI); NOS-associated protein (NOSAP); nitric oxide synthase-like (NOS); nitrite transporter (NiT); nitrite reductase (NiR); nitrate reductase (NaR); nitrate transporter (NaT); molybdopterin synthase catalytic subunit (MPT) and non-symbiotic hemoglobin (Hb-1). Gene expression levels in each sample were compared to those in PAB at the vegetative stage. Error bars indicate standard deviation (s.d.). Asterisks represent significant differences from PAB tissue at the vegetative stage according to two-way ANOVA (*p < 0.05; **p < 0.005; ***p < 0.0005).
Low NO level was associated with low NaR activities and repressing phase transition and floral initiation of Oncidium
To analyze the endogenous NO level, microscopic observation with the aid of the NO-specific fluorescent dye, 4, 5-diaminofluorescein diacetate (DAF-2DA), revealed that the NO level at the reproductive stage was decreased by approximately 60% compared to that of the vegetative stage (Fig. 3a). To further investigate NO production, we assayed the NaR activity and quantified NO level and nitrate concentration in Oncidium at the following three different stages: vegetative (V), bolting (B), and reproductive (R). Our data demonstrated that the NO level, nitrate concentration and NaR enzymatic activity were markedly decreased in the period between the vegetative stage and the reproductive stage (Fig. 3b–d), suggesting a low production of NO. Orchids at the vegetative stage were subjected to different concentrations of SNP, an effective NO donor functionally known to repress floral transition in Arabidopsis. Floral transition was repressed after treatment with 20 and 100 μM SNP (Fig. 3e). These results clearly indicated that the NO molecule is an effector that can repress floral initiation in Oncidium. Furthermore, we analyzed the following flowering-associated genes by qRT-PCR: OgFT, OgFYF, OgAP1, and OgTFL (Fig. 3f–i). Our results showed that the transcript levels of OgFT, OgFYF, and OgAP1 were lowered, but the transcript levels of OgTFL, a floral repressor, were enhanced by SNP treatment (Fig. 3f–i).
The activities of NO biosynthetic enzymes and the levels of NO in Oncidium PIB tissues at different stages.
(a) The NO intensity was detected using the green fluorescence of DAF-2T, and white light indicated the morphology and cell status in PIB tissues(scale bar = 50μM) (b–d) The nitric oxide (NO) levels (b), nitrate level (c) and nitrate reductase (NaR) enzymatic activity (d) were quantified in Oncidium PIB tissues at the vegetative (V), bolting (B) and reproductive (R) stages. (e) Photographs of Oncidium orchids treated with water (mock), 20 μM SNP or 100 μM SNP, which was applied on the juvenile inflorescent buds for one month. White arrows indicate the developing inflorescence (scale bar = 1cm). (f–i) Expression levels of floral genes in the inflorescence of mock or SNP-treated Oncidium orchids. The relative gene expression levels in SNP-treated plants were compared to those of mock plants. Error bars indicate standard deviation (s.d.). Asterisks represent significant differences compared with vegetative stage or mock respectively and were analyzed by two-way ANOVA (*p < 0.05; **p < 0.005; ***p < 0.0005).
NO level and the activity of NaR-related NO biosynthesis pathway were enhanced by ascorbate
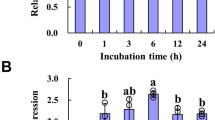
It is well-known that the reduced form of ascorbate (AsA) and reactive oxygen species are essential for phase transition in Oncidium6,22,23. To further elucidate the relationship of AsA and NO during phase transition of Oncidium, Oncidium tissue culture of a protocorm-like body (PLB) was used to monitor the effect of AsA on NO production. As shown in Fig. 4a the fluorescent intensity of Oncidium PLB, which indicated NO levels, increased under high doses (5 mM) of AsA compared to low doses of AsA (1 mM) and mock treatment. Accordingly, the NO levels and NaR enzymatic activities were elevated in response to exogenous AsA treatment (Fig. 4b,c). However, the transcript level of NaR was unaltered in response to AsA (Fig. 4d). In addition, the nitrate and arginine levels in Oncidium PLB tissues increased with AsA dosage (Fig. 4e,f). These results demonstrated that AsA is effective in elevating the precursor levels and enzyme activities for NO biosynthesis, thus increasing NO levels (Fig. 4).
NO-related metabolism in Oncidium protocorm-like body (PLB) tissues treated with ascorbate (AsA).
(a) The NO intensity of Oncidium PLB tissues grown in ½ MS without treatment (mock), or under treatment with 1 mM and 5 mM AsA for 5 days. NO intensity was indicated by a green fluorescence of DAF-2T, and white light indicated the morphology and cell status in tissues (scale bar = 50μm). (b) The relative NO intensity in AsA-treated Oncidium PLB tissues was compared to that in mock Oncidium PLB cultures. (c,d) The relative activities of nitrate reductase (NaR) and OgNaR gene expression in AsA-treated Oncidium PLB, comparing to those of mock Oncidium. (e,f) Nitrate and arginine amounts in mock or AsA-treated Oncidium PLB tissues are shown. Significant differences compared with mock were analyzed by two-way ANOVA (*p < 0.05; **p < 0.005; ***p < 0.0005).
The AsA-deficient mutant, vtc1, produces low NO, leading to early flowering
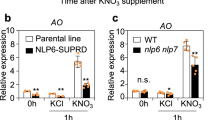
To further validate the functional effects of AsA on NO production, Arabidopsis vtc1 mutant was employed. The Arabidopsis vtc1 mutant, containing 40% of the wild type AsA level and displaying an early flowering phenotype, was shown to have low NO level (Fig. 5a,b). On contrary, there were 80% of 20 μM SNP treated-Arabidopsis vtc1mutant lines flowered with 8 rosette leaves as compared to the mock-treated Arabidopsis vtc1 mutant lines (Fig. 5b). A comparative survey of the NO- biosynthesis-related genes, such as AtNIA1 (AT1G77760), AtNIA2 (AT1G37130), AtNIR1 (AT2G15620) and AtNOS/AtNOA1 (AT3G47450), revealed similar expression levels after SNP treatment between the two plants (Fig. 5c, Supplementary Fig. S3). This indicated that SNP did not affect transcription activity of NO- biosynthesis-related genes. Furthermore, the expression patterns of flowering-associated genes, namely CONSTANS (CO), GIGANTEA (GI), Flowering locus T (FT) and LEAFY (LFY), were assayed in plants treated with SNP. The transcript levels of CO, GI, FT and LFY were lowered by SNP treatment (Fig. 5c). However, lower levels of nitrate and nitrite were discovered in vtc1 accompanied by decreased NaR and NiR enzymatic activities (Fig. 5d,e). In contrast, Arabidopsis vtc1 contained higher levels of arginine and citrulline than the wild type (Fig. 5f). No significant differences in the expression of NO- biosynthesis-related genes were observed between in Wt and vtc1 mutant. However, the levels of nitrate and nitrite, activities of NaR and NiR were lower in vtc1 due to its lower AsA level, suggesting that lower AsA level caused lower level of NO production and led to early flowering.
Endogenous NO and related compounds were analyzed in A. thaliana Col-0 (Wt) and vtc1.
(a) Photographs of the Wt and vtc1 mutant grown on the ½ MS medium under mock or 20 μM SNP for 3 weeks. It showed that vtc1 showed earlier flowering than Wt (upper panel arrow marks), but 20 μM SNP decreased percentage of early flowering plants (lower panel) scale bar = 1cm). (b) The percentage of flowering Wt or vtc1 at different development stages specified according to rosette leaf number under mock or SNP treatments (n wt and nvtc1 ≥ 30). (c) RT-PCR analysis showed the expression pattern of NO-related and flowering-associated genes (AtNIA1, AtNIA2, AtNIR1, AtNOA1, AtCO, AtFT, and AtLFY) after treatment with 20 μM SNP (actin was used as an internal control). (d) The levels of nitrate (□) and nitrite (■) in the Wt and vtc1 mutant. (e) The NaR activity (□) and NiR activity (■) in the Wt and vtc1 mutant. (f) The levels of arginine (□) and citrulline (■) in the Wt and vtc1 mutant. Error bar represents ± SE. Significant differences between samples were determined using two-way ANOVA.
CaCl2 and H2O2 have been reported to act as elicitor to induce NO production. However, our results showed that CaCl2 and H2O2 were effective to induce NO production in wild type, whereas both were ineffective to induce NO production in vtc1 (as shown by florescent intensity) (Fig. 6a). Additionally, compared to the wild type, Arabidopsis vtc1 mutant line displayed moderately lower production of NO in response to CaCl2 and H2O2 (Fig. 6b). It was further confirmed through enzymatic assay of NaR, that NO production was significantly lowered in the Arabidopsis vtc1 mutant line compared to the wild type (Fig. 6c). These data suggested that change in AsA redox ratio in Arabidopsis vtc1 line had no direct influence on endogenous NO level.
CaCl2 and H2O2 were ineffective to induce NO in Arabidopsis vtc1 mutant line.
(a) The NO intensity in leaf segments of the Wt and vtc1 mutant treated with water (mock), CaCl2 and H2O2. White light indicated the morphology and cell status in tissues (scale bar = 50μm). (b) The relative NO intensity in treated Wt and vtc1 was compared to that of mock Wt. (c) The relative activities of nitrate reductase (NaR) after treatment of CaCl2 or H2O2 in the vtc1 mutant, were compared to those of Wt.
The effect of L-galactonolactone (L-GalL) on the flowering time of A. thaliana Col-0 (Wt) and noa1
To ascertain AsA-mediated function in NO biosynthesis and the repressive role in the flowering process, exogenous L-GalL, a precursor of AsA, was applied on Arabidopsis Wt and Arabidopsis noa1 mutant, a NO-deficient mutant and displayed early flowering. As shown in the Fig. 7a, the wild type plants showed delayed flowering after L-galactonolactone (L-GalL) treatment. Comparing to the mock treatment, Arabidopsis noa1 displayed slightly delayed flowering time with 6–8 rosette leaves after L-galactonolactone (L-GalL) treatment (Fig. 7a,b). To further investigate the relationship of AsA on NO metabolism, we assayed total AsA content and the redox ratio of reduced/oxidized form of AsA in Arabidopsis Wt and noa1 mutant under SNP treatment. As shown in the Fig. 7c, the level of AsA showed an increase in noa1 mutant lines compared with Wt. This result indicated that AsA is accumulated in noa1 mutant regardless it’s low NO accumulation and early flowering phenotype, compared to the wild type. On the other hand, Wt Arabidopsis seedling treated with SNP significantly reduced in endogenous AsA level. Moreover, the redox ratio of reduced/oxidized form of AsA was found to be lowered in Arabidopsis noa1 mutant seedlings after SNP treatment (Fig. 7c). These results suggest that the flowering repression in Arabidopsis was determined by NO level, not the AsA content.
Flowering time change of A. thaliana Col-0 (Wt), and noa1 mutant in response to exogenous L-galactonolactone (L-GalL) treatment.
(a) The effect of L-galactonolactone (L-GalL) on the flowering time of A. thaliana Col-0 (Wt) and noa1 (scale bar = 1cm). (b) The percentage of flowering Wt or noa1 at different development stages specified according to rosette leaf number under mock or L-GalL treatments (n wt and nnoa1 ≥30). Error bars indicate standard deviation (s.d.). Asterisks represent significant differences between samples as determined using two-way ANOVA (Wt; *p < 0.05; **p < 0.005; ***p < 0.0005). (c) The ascorbic content and ratio of reduced/oxidized form of AsA in A. thaliana Col-0 (Wt) and noa1 grown in ½ MS without treatment (mock: ( ), Wt; (
), Wt; ( ), noal mutant), or under treatment with 20 μM SNP (
), noal mutant), or under treatment with 20 μM SNP ( ), Wt; (
), Wt; ( ), noa1 mutant) for 3 weeks. Significant differences compared with mock were analyzed by two-way ANOVA (*p < 0.05; **p < 0.005; ***p < 0.0005). (d) Working model of the Oncidium floral repression acquired by post-translational modification of NO- biosynthesis-related enzymes: During the vegetative stage, the AsA level, NO-related enzymes (NaR, NiR and NOS) and NO levels were high and synergistically repressed flowering. In the bolting stage, DHA was produced by the redox homeostasis of Oncidium, which lowered the AsA redox ratio, thus leading to deceased NO-related enzymatic activities for NO and phase transition. In the next vegetative stage, low levels of AsA and NO were triggered by the redox homeostasis, thus leading to the accumulation of GSNO to terminate the reproductive stage through floral repression and post-translational modification of APX.
), noa1 mutant) for 3 weeks. Significant differences compared with mock were analyzed by two-way ANOVA (*p < 0.05; **p < 0.005; ***p < 0.0005). (d) Working model of the Oncidium floral repression acquired by post-translational modification of NO- biosynthesis-related enzymes: During the vegetative stage, the AsA level, NO-related enzymes (NaR, NiR and NOS) and NO levels were high and synergistically repressed flowering. In the bolting stage, DHA was produced by the redox homeostasis of Oncidium, which lowered the AsA redox ratio, thus leading to deceased NO-related enzymatic activities for NO and phase transition. In the next vegetative stage, low levels of AsA and NO were triggered by the redox homeostasis, thus leading to the accumulation of GSNO to terminate the reproductive stage through floral repression and post-translational modification of APX.
Discussion
The genetic network of Oncidium that controls the flowering process and flowering time is quite complicated and not well understood. Large-scale sequencing approaches have been performed to reveal the flowering mechanism of Oncidium24,25,26. Our transcriptomic data revealed that NO-related genes were abundantly and differentially expressed in the hemisphere of pseudobulb tissue proximal to the inflorescence bud (PIB), compared to the other half proximal to the axillary bud (PAB) (Fig. 1). It is well known that the NO synthesis can occur both enzymatically and non-enzymatically in plants18,27,28. However, the source of NO production in plants has not yet been identified and is hotly debated29. In mammals, NOS enzymes are key enzymes that catalyze L-arginine to generate NO and citrulline by NADPH-dependent oxidation in a complex reaction involving Ca2+, FMD, and CaM30,31. Similar mechanisms have also been reported in plants for the synthesis of NO from L-arginine and from nitrite32.
Our study also discovered that the mechanism that lowers NO production and reduces NaR and NO level was associated with Oncidium phase transition and repression of the flowering process (Fig. 3). The reduced stalk inflorescence and lower transcript levels of flowering-associated genes (FYF, FT and AP1) in Oncidium further confirmed that NO is a repressor of floral transition and the flowering process (Fig. 3)16. This study provided evidence that the level of nitrate, nitric oxide and arginine were enhanced in the presence of AsA (Fig. 4a,b,e,f). Although AsA is known to be involved in cellular redox signaling but does not act specifically in any of the known flowering pathways, it plays a general role in responding to environmental signals5. The reduced form of AsA is essential for Oncidium phase transition and flowering process under high ambient temperature6,22,23. A plausible explanation may be plants would rather produce NO than H2O2 to balance redox homeostasis for its physiological functions, that are tampered by AsA (Fig. 4)33,34. However, increased enzymatic activity of NaR is not dosage -dependent that could support the alternative pathway for NO production from direct reaction of AsA (Fig. 4c)35.
Based on recent publications, it was stated that NO production is directly proportional to AsA, either by modulating electron flow through the NaR enzyme or by indirectly affecting nitrite concentration, under strict regulation36,37. Therefore, we hypothesize that the posttranslational modification of cytAPX1 by NaR- dependent NO production, might contribute to floral repression35,36,37. The ascorbate-deficient mutant, Arabidopsis vtc1, displayed a lower level of nitrate and nitrite, as well as decreased NaR and NiR activities, compared to Wt (Fig. 5). This may explain by fact that higher level of reduced ascorbate doesn’t react with NO and its enzymatic activities but may be with other nitrosating species (Fig. 5)34,35,36,37,38.
Moreover, Arabidopsis vtc1 had delayed flowering and decreased mRNA transcripts of flowering-associated genes (CO, GI, FT and LFY) after SNP treatment, implying that the delay flowering was due to the undesired side effects caused by SNP. In recent reports, NO donors (SNP, SNAP and SIN-1) induced flowering under non-inductive conditions in duckweed (Lemma aequinoctialis)39. However, SA-induced flowering in duckweed was significantly reduced by exogenous application of NO scavengers, nitric oxide synthase inhibitors and nitrate reductase inhibitor in duckweed, providing that NO was not involved in photoperiodic flowering pathway and acting as a stress negating agents from the SA-signaling pathway40. NO donor doesn’t necessarily mimic the functions of NO when level of reduced AsA is higher, but could rather involve in controlling the cellular redox signaling41,42. Note that, Arabidopsis vtc1 mutant displayed increased level of arginine and citrulline level was significant compared with Wt (Fig. 5). It is speculated that AsA act as a redox cofactor for NOS to catalyze the conversion of arginine/citrulline for NO production28,43. However, this hypothesis is less well understood and requires extensive investigation. This evidence from Arabidopsis vtc1 supports a notion that the relationship between AsA and NO adds some degree of specificity as follows: (1) low AsA caused low level of NO production and leads to early flowering, (2) NO, being an inducer of antioxidant buffering system can alter antioxidant gene expression (cytAPX1) by ROS–dependent post-translation modification, (3) basal NO production depends on AsA and requires L-arginine and citrulline as supplements under oxidative stress43,44,45. Thus, the repression of flowering was determined by NO through NO-related enzymatic activities in the presence of AsA.
The Arabidopsis vtc1 mutant line did not respond to hydrogen peroxide (H2O2) and calcium chloride (CaCl2) compared to wild type (Wt), suggesting the role of cytAPX1 maintained a low AsA ratio to scavenge H2O2 (Fig. 6)5,6,22,23. Molecular studies of the Arabidopsis noa1 mutant further showed that AsA could not rescue flowering time (Fig. 7). Here, we proposed the function of AsA in NO biosynthesis is acting as a cofactor for NO mediated flowering repression process46,47. The Oncidium flowering process was strictly controlled by the mechanism involved in the signaling between AsA and NO biosynthesis with various degrees of specificity as follows: (1) Oncidium floral repression is dependent upon NO production, (2) NO biosynthetic enzymatic activities are repressed during bolting period of Oncidium PIB, (3) AsA-mediated NO signaling increases NO biosynthesis and enzymatic activities, which could activate post-translational modification for flowering repression, (4) low NO derivatives prevents the scavenging of excess free radicals that accelerate flowering in Oncidium, and they are maintained through redox reaction utilizing arginine as a substrate, (5) AsA and NO both play mutual role in flowering repression by inducing NaR–mediated and L-arginine dependent NO biosynthesis48.
In the immediate future, it is better to understand the molecular mechanism of NO production through redox signaling (ascorbate-glutathione cycle) from NaR–mediated and L-arginine-dependent (NOS-like) post-translational modification (APX), which may be useful to clarify the importance of antioxidants and their role in the repression of the flowering process by NO in plants (Fig. 7d)49.
Materials and Methods
Plants materials and growth conditions
Oncidium ‘Gower Ramsey’ plants were obtained from the Shih-Dong Orchid Nursery in Taiwan. The plants were grown in 30 cm diameter pots under growth conditions of 25~32 °C and a 14 h/10 h (day/night) photoperiod in a greenhouse. Oncidium protocorm-like bodies (PLBs), Arabidopsis thaliana ecotype Col-0, vtc1 mutant (ABRC stock No.: CS8326) and noa1 (ABRC stock No: CS6511) were cultured in 1/2 Murashige and Skoog medium50 under long-day conditions (16 h light/8 h dark cycles) at 23 ± 2 °C.
RNA isolation and sequencing
Total RNA from each sample was isolated with TRIzol (Invitrogen) according to the manufacturer’s instructions. After being treated with RNase-free DNase I (New England BioLabs) for 30 min at 37 °C to remove the residual DNA, the RNA was sent to BGI (Beijing, China) where two cDNA libraries (PAB and PIB) were made and Solexa sequencing was performed. In all, 8,353,971 raw reads were obtained.
Annotation and functional categorization
All clean reads were assembled by the standard SOAP denovo assembler ver 1.0451 (http://soap.genomics.org.cn/). The assembled contigs were subjected to a N50 algorithm52 to confirm the better performance of the assembly output. We defined such processed sequences as unigenes. In the final step, a Blastx alignment was performed using the unigenes and the NCBI non-redundant protein database (Nr), Swiss-Prot, KEGG, and COG protein databases53. The best-aligning results were used to determine the sequence directions of those unigenes. We then retrieved proteins that had the highest sequence similarity with the given unigenes and determined their functional annotations. We used the fatiGO and Blast2GO programs (http://www.blast2go.org/) to obtain the GO annotations for the unigenes54,55.
Analysis of gene expressions by quantified real-time PCR
Total RNA was extracted from Oncidium, Arabidopsis thaliana ecotype Col-0, and vtc1. The mRNA in the total RNA (1 μg) was converted to first-strand cDNA using reverse transcriptase with the oligo-dT primer in accordance with the manufacturer’s instructions. qPCR was performed using Applied Biosystems 7500 Fast Real-Time PCR System and KAPA SYBR Green PCR® master mixture universal. For qPCR analyses, gene expression levels were normalized based on the Ubiquitin gene of Oncidium or Arabidopsis. Sequence information for the oligonucleotides used for the qPCR analysis in this study is provided in Supplementary Table S3.
NO detection
NO was quantified through DAF-2DA (Cayman) staining56. Oncidium PIB tissues at three different stages (V, B, and R), as well as A. thaliana Col and mutant seedlings were all stained according to the method described. Samples treated with elicitors were treated with CaCl2 for 1 h or H2O2 for 2 h and compared with mock at room temperature. Three biological replicates (3 biological replicates × 3 = 9 replicates per sample) of Oncidium PIB tissue at three different stages (V, B, and R) were used. For Arabidopsis, samples of more than twenty plants were used, and the averages of three biological replicates were tested. A two-way ANOVA analysis of variance was performed on the data to determine the significant differences and p-values utilizing GraphPad Prism® v5.0 software.
Measurement of nitrate and nitrite reductase activities
Oncidium PLB and Arabidopsis leaves and roots were used to determine the nitrate reductase (NaR) and nitrite reductase (NiR) activity as previously described57,58. Samples (100 mg) were ground in liquid N2 and then homogenized in 0.5 mL of extraction buffer containing 3 mM EDTA and 50 mM Tris-HCl (pH 8.0). The mixture was centrifuged at 13,000 g for 20 min at 4 °C. For the NaR activity assay, the reaction buffer (250 μL of 0.1 M potassium-phosphate buffer (pH 7.5), 50 μL of 0.1 M KNO3 and 50 μL of 2 mM NADH) was added to the resulting supernatant (100 μL), and the mixture was incubated in the dark at room temperature for 60 min. The nitrate consumption was quantified at A42059. For the NiR activity assay, 550 μL of reaction buffer (500 μL of 0.1 M sodium phosphate buffer (pH 7.5), 25 μL of 10 mM NaNO2 and 50 μL of 1.5% methyl viologen) and 25 μL of 5% NaHCO3 were added to the resulting supernatant (100 μL), and the mixture was incubated at room temperature for 30 min. The reaction was stopped by the addition of the Griess reagent, and nitrite production was quantified at 540 nm60. The assay of nitrate concentration was determined based on forming a nitro derivative of salicylic acid with nitrate. Nitrite should be removed from the sample by saturated sulfamic acid. Samples and different concentrations of NaNO3 (nitrate standard) were extracted with a saturated solution of sulfamic acid. After incubating the mixture for 2 min, salicylic acid was added to reacts with nitrate under acidic conditions to form nitrosalicylic acid. Then the pH values were adjusted to 12 with NaOH. Nitrate concentration was determined after plotting against a linear regression and using spectrometry with A42059. The nitrite content was quantified from the NaR or NiR enzymatic assay described previously61. The averages of three biological replicates (3 technical replicates × 3 = 9 replicates per sample) from samples of orchid PLB tissues were used. For Arabidopsis, the average of the three biological replicates of more than twenty plant leaves and roots were used for the assay. A two-way ANOVA analysis of variance was performed on the data to determine the significant differences and p-values utilizing GraphPad Prism® v5.0 software.
Quantification of arginine and citrulline by LC-ESI-MS
Samples were ground with a mortar and pestle in liquid N2, and metabolites were extracted in 80% methanol. Sample derivatization was performed with modifications62. In brief, 10 μL of metabolite samples was mixed with 10 μL of deionized water, 2.5 μL of 0.5 M Na2CO3 (pH 9.2), and 2.5 μL of 10 mg/μL dansyl chloride (freshly prepared in acetone), and the reaction was incubated at 60 °C for 1.5 h. Then, 75 μL of deionized water was added, and the mixture was further incubated at 60 °C for 30 min. Amino acid standards (physiological acids, neutrals and basics) (A9906; SIGMA) were prepared at various concentrations as described above for the derivatizing reagent and used to determine the concentrations of amino acids in the samples. The derivatizing samples were centrifuged at 14,000 rpm for 15 min, and the supernatants were subjected to LC-ESI-MS for arginine and citrulline quantification63. The average of three biological replicates (3 technical replicate × 3 = 9 replicates per sample) from samples of Orchid PLB tissues were used. For Arabidopsis, the average of three biological replicates of more than twenty plant leaves and roots were used for the assay. A two-way ANOVA analysis of variance was performed on the data to determine the significant differences and p-values utilizing Graphad prism v5.0 software®.
Analysis of ascorbate and hydrogen peroxide
The extraction and measurement of AsA and H2O2 levels were performed as previously described6 with slight modification. Oncidium PLB tissues were washed with ddH2O twice. For total AsA quantification, the reactant was mixed with 10 mM dithiothreitol to reduce the pool of oxidized AsA. In contrast, to assay reduced AsA, only deionized water was added to the reactant. All mixtures were supplemented with reaction buffer (10% trichloroacetic acid, 43% H3PO4, 4% α-α-bipyridyl and 3% FeCl3) and incubated at 37 ± 1 °C for 1 h. The amounts of total and reduced AsA were determined by spectrometry at A525, and the amount of oxidized AsA was calculated as previously described. The average of three biological replicates (3 technical replicates × 3 = 9 replicates per sample) from samples of Oncidium PLB tissues was used, and two-way ANOVA analysis of variance was performed to determine the significant differences. The reduced and oxidized forms of AsA were determined from the total pool of three independent biological replicates.
Additional Information
Accession codes: AtNIA1 (AT1G77760), AtNIA2 (AT1G37130), AtNIR1 (AT2G15620) and AtNOS/AtNOA1 (AT3G47450)
How to cite this article: Senthil Kumar, R. et al. Nitric oxide participates in plant flowering repression by ascorbate. Sci. Rep. 6, 35246; doi: 10.1038/srep35246 (2016).
References
Wellmer, F. & Riechmann, J. L. Gene networks controlling the initiation of flower development. Trends in Genetics 26, 519–527 (2010).
Ogawa, K., Tasaka, Y., Mino, M., Tanaka, Y. & Iwabuchi, M. Association of glutathione with flowering in Arabidopsis thaliana. Plant and Cell physiology 42, 524–530 (2001).
Yanagida, M., Mino, M., Iwabuchi, M. & Ogawa, K. Reduced glutathione is a novel regulator of vernalization-induced bolting in the rosette plant Eustoma grandiflorum. Plant and Cell Physiology 45, 129–137 (2004).
Barth, C., De Tullio, M. & Conklin, P. L. The role of ascorbic acid in the control of flowering time and the onset of senescence. Journal of Experimental Botany 57, 1657–1665 (2006).
Kotchoni, S. O., Larrimore, K. E., Mukherjee, M., Kempinski, C. F. & Barth, C. Alterations in the endogenous ascorbic acid content affect flowering time in Arabidopsis. Plant Physiology 149, 803–815 (2009).
Shen, C. H., Krishnamurthy, R. & Yeh, K. W. Decreased L-ascorbate content mediating bolting is mainly regulated by the galacturonate pathway in Oncidium. Plant and Cell physiology 50, 935–946 (2009).
Conklin, P. L. et al. Genetic evidence for the role of GDP-mannose in plant ascorbic acid (vitamin C) biosynthesis. Proceedings of the National Academy of Sciences of the United States of America 96, 4198–4203 (1999).
Veljovic-Jovanovic, S. D., Pignocchi, C., Noctor, G. & Foyer, C. H. Low ascorbic acid in the vtc-1 mutant of Arabidopsis is associated with decreased growth and intracellular redistribution of the antioxidant system. Plant Physiology 127, 426–435 (2001).
Attolico, A. D. & De Tullio, M. C. Increased ascorbate content delays flowering in long-day grown Arabidopsis thaliana (L.) Heynh. Plant Physiology and Biochemistry 44, 462–466 (2006).
Daniela, A. A. & De Tullio, M. C. Contrasting effects of increased ascorbate content on growth and development of long-day and short-day grown Brassica rapa. Caryologia 60, 185–187 (2007).
Hirai, N. et al. Accumulation of Ascorbic Acid in the Cotyledons of Morning Glory (Pharbitis nil) Seedlings during the Induction of Flowering by Low-Temperature Treatment and the Effect of Prior Exposure to High-Intensity Light. Plant and Cell Physiology 36, 1265–1271 (1995).
Badiani, M. et al. Seasonal Variations of Antioxidants in Wheat (Triticum aestivum) Leaves Grown Under Field Conditions. Functional Plant Biology 23, 687–698 (1996).
Lokhande, S. D., Ogawa, K., Tanaka, A. & Hara, T. Effect of temperature on ascorbate peroxidase activity and flowering of Arabidopsis thaliana ecotypes under different light conditions. Journal of Plant Physiology 160, 57–64 (2003).
Pignocchi, C. et al. Ascorbate oxidase-dependent changes in the redox state of the apoplast modulate gene transcript accumulation leading to modified hormone signaling and orchestration of defense processes in tobacco. Plant Physiology 141, 423–435 (2006).
Pnueli, L., Liang, H., Rozenberg, M. & Mittler, R. Growth suppression, altered stomatal responses, and augmented induction of heat shock proteins in cytosolic ascorbate peroxidase (Apx1)-deficient Arabidopsis plants. Plant Journal 34, 187–203 (2003).
He, Y. et al. Nitric oxide represses the Arabidopsis floral transition. Science 305, 1968–1971 (2004).
Streatfield, S. J. et al. The phosphoenolpyruvate/phosphate translocator is required for phenolic metabolism, palisade cell development, and plastid-dependent nuclear gene expression. Plant Cell 11, 1609–1622 (1999).
Besson-Bard, A., Pugin, A. & Wendehenne, D. New insights into nitric oxide signaling in plants. Annual Review of Plant Biology 59, 21–39 (2008).
Gupta, K. J., Fernie, A. R., Kaiser, W. M. & van Dongen, J. T. On the origins of nitric oxide. Trends in Plant Science 16, 160–168 (2011).
Luiking, Y. C., Engelen, M. P. & Deutz, N. E. Regulation of nitric oxide production in health and disease. Current Opinion in Clinical Nutrition and Metabolic Care 13, 97–104 (2010).
Moreau, M., Lee, G. I., Wang, Y., Crane, B. R. & Klessig, D. F. AtNOS/AtNOA1 is a functional Arabidopsis thaliana cGTPase and not a nitric-oxide synthase. Journal of Biological Chemistry 283, 32957–32967 (2008).
Chin, D. C., Shen, C. H., SenthilKumar, R. & Yeh, K. W. Prolonged exposure to elevated temperature induces floral transition via up-regulation of cytosolic ascorbate peroxidase 1 and subsequent reduction of the ascorbate redox ratio in Oncidium hybrid orchid. Plant and Cell Physiology 55, 2164–2176 (2014).
Chin, D. C., Hsieh, C. C., Lin, H. Y. & Yeh, K. W. A Low Glutathione Redox State Couples with a Decreased Ascorbate Redox Ratio to Accelerate Flowering in Oncidium Orchid. Plant and Cell Physiology 57, 423–436 (2016).
Tan, J., Wang, H. L. & Yeh, K. W. Analysis of organ-specific, expressed genes in Oncidium orchid by subtractive expressed sequence tags library. Biotechnology Letters 27, 1517–1528, 10.1007/s10529-005-1468-8 (2005).
Chang, Y. Y. et al. Characterization of Oncidium ‘Gower Ramsey’ transcriptomes using 454 GS-FLX pyrosequencing and their application to the identification of genes associated with flowering time. Plant and Cell Physiology 52, 1532–1545 (2011).
Fu, C. H. et al. OrchidBase: a collection of sequences of the transcriptome derived from orchids. Plant and Cell Physiology 52, 238–243 (2011).
Lum, H. K., Butt, Y. K. & Lo, S. C. Hydrogen peroxide induces a rapid production of nitric oxide in mung bean (Phaseolus aureus). Nitric Oxide 6, 205–213, 10.1006/niox.2001.0395 (2002).
Planchet, E. & Kaiser, W. M. Nitric oxide production in plants: facts and fictions. Plant Signaling and Behavior 1, 46–51 (2006).
Frohlich, A. & Durner, J. The hunt for plant nitric oxide synthase (NOS): is one really needed? Plant Science 181, 401–404 (2011).
Alderton, W. K., Cooper, C. E. & Knowles, R. G. Nitric oxide synthases: structure, function and inhibition. Biochemical Journal 357, 593–615 (2001).
Sang, J., Zhang, A., Lin, F., Tan, M. & Jiang, M. Cross-talk between calcium-calmodulin and nitric oxide in abscisic acid signaling in leaves of maize plants. Cell Research 18, 577–588 (2008).
Jasid, S., Simontacchi, M., Bartoli, C. G. & Puntarulo, S. Chloroplasts as a nitric oxide cellular source. Effect of reactive nitrogen species on chloroplastic lipids and proteins. Plant Physiology 142, 1246–1255 (2006).
Yamasaki, H. & Sakihama, Y. Simultaneous production of nitric oxide and peroxynitrite by plant nitrate reductase: in vitro evidence for the NR-dependent formation of active nitrogen species. FEBS Letters 468, 89–92 (2000).
Gross, F., Durner, J. & Gaupels, F. Nitric oxide, antioxidants and prooxidants in plant defence responses. Frontiers in Plant Science 4, 419 (2013).
Yamasaki, H., Sakihama, Y. & Takahashi, S. An alternative pathway for nitric oxide production in plants: new features of an old enzyme. Trends in Plant Science 4, 128–129 (1999).
Wang, X. & Hargrove, M. S. Nitric oxide in plants: the roles of ascorbate and hemoglobin. PloS One 8, e82611 (2013).
Rockel, P., Strube, F., Rockel, A., Wildt, J. & Kaiser, W. M. Regulation of nitric oxide (NO) production by plant nitrate reductase in vivo and in vitro. Journal of Experimental Botany 53, 103–110 (2002).
Scorza, G., Pietraforte, D. & Minetti, M. Role of ascorbate and protein thiols in the release of nitric oxide from S-nitroso-albumin and S-nitroso-glutathione in human plasma. Free Radical Biology & Medicine 22, 633–642 (1997).
Khurana, A., Khurana, J. P. & Babbar, S. B. Nitric Oxide Induces Flowering in the Duckweed Lemna aequinoctialis Welw. (Syn. L. paucicostata Hegelm.) Under Noninductive Conditions. Journal of Plant Growth Regulation 30, 378–385, 10.1007/s00344-011-9199-7 (2011).
Khurana, A., Kumar, R. & Babbar, S. B. Nitric oxide is involved in salicylic acid-induced flowering of Lemna aequinoctialis Welw. Acta Physiologiae Plantarum 36, 2827–2833 (2014).
Chun, H. J. et al. Constitutive expression of mammalian nitric oxide synthase in tobacco plants triggers disease resistance to pathogens. Molecules and Cells 34, 463–471 (2012).
Shi, S., Wang, G., Wang, Y., Zhang, L. & Zhang, L. Protective effect of nitric oxide against oxidative stress under ultraviolet-B radiation. Nitric Oxide 13, 1–9 (2005).
Crawford, N. M. Mechanisms for nitric oxide synthesis in plants. Journal of Experimental Botany 57, 471–478 (2006).
de Pinto, M. C., Paradiso, A., Leonetti, P. & De Gara, L. Hydrogen peroxide, nitric oxide and cytosolic ascorbate peroxidase at the crossroad between defence and cell death. Plant Journal 48, 784–795 (2006).
Corpas, F. J., Hayashi, M., Mano, S., Nishimura, M. & Barroso, J. B. Peroxisomes are required for in vivo nitric oxide accumulation in the cytosol following salinity stress of Arabidopsis plants. Plant Physiology 151, 2083–2094 (2009).
Van Ree, K., Gehl, B., Chehab, E. W., Tsai, Y. C. & Braam, J. Nitric oxide accumulation in Arabidopsis is independent of NOA1 in the presence of sucrose. Plant Journal 68, 225–233 (2011).
Xie, Y. et al. Roles of NIA/NR/NOA1-dependent nitric oxide production and HY1 expression in the modulation of Arabidopsis salt tolerance. Journal of Experimental Botany 64, 3045–3060 (2013).
Rasul, S., Wendehenne, D. & Jeandroz, S. Study of oligogalacturonides-triggered nitric oxide (NO) production provokes new questioning about the origin of NO biosynthesis in plants. Plant Signaling and Behavior 7, 1031–1033 (2012).
Begara-Morales, J. C. et al. Dual regulation of cytosolic ascorbate peroxidase (APX) by tyrosine nitration and S-nitrosylation. Journal of Experimental Botany 65, 527–538 (2014).
Murashige, T. & Skoog, F. A Revised Medium for Rapid Growth and Bio Assays with Tobacco Tissue Cultures. Physiologia Plantarum 15, 473–497 (1962).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Research 20, 265–272 (2010).
Lin, Y. et al. Comparative studies of de novo assembly tools for next-generation sequencing technologies. Bioinformatics 27, 2031–2037 (2011).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic acids Research 25, 3389–3402 (1997).
Al-Shahrour, F., Diaz-Uriarte, R. & Dopazo, J. FatiGO: a web tool for finding significant associations of Gene Ontology terms with groups of genes. Bioinformatics 20, 578–580 (2004).
Gotz, S. et al. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic acids Research 36, 3420–3435 (2008).
Guo, F. Q., Okamoto, M. & Crawford, N. M. Identification of a plant nitric oxide synthase gene involved in hormonal signaling. Science 302, 100–103 (2003).
Gupta, S. C. & Beevers, L. Synthesis and degradation of nitrite reductase in pea leaves. Plant Physiology 75, 251–252 (1984).
Stewart, G. R., Lee, J. A. & Orebamjo, T. O. Nitrogen metabolism of halopytes. II Nitrate availability and utilization. New Phytologist 72, 539–546 (1973).
Cataldo, D. A., Maroon, M., Schrader, L. E. & Youngs, V. L. Rapid colorimetric determination of nitrate in plant tissue by nitration of salicylic acid. Communications in Soil Science and Plant Analysis 6, 71–80 (1975).
Tsikas, D. Analysis of nitrite and nitrate in biological fluids by assays based on the Griess reaction: appraisal of the Griess reaction in the L-arginine/nitric oxide area of research. Journal of Chromatography. B, Analytical Technologies in the Biomedical and Life Sciences 851, 51–70 (2007).
Takahashi, M., Sasaki, Y., Ida, S. & Morikawa, H. Nitrite reductase gene enrichment improves assimilation of NO(2) in Arabidopsis. Plant Physiology 126, 731–741 (2001).
Guo, K. & Li, L. Differential 12C-/13C-isotope dansylation labeling and fast liquid chromatography/mass spectrometry for absolute and relative quantification of the metabolome. Analytical Chemistry 81, 3919–3932 (2009).
Salazar, C., Armenta, J. M. & Shulaev, V. An UPLC-ESI-MS/MS Assay Using 6-Aminoquinolyl-N-Hydroxysuccinimidyl Carbamate Derivatization for Targeted Amino Acid Analysis: Application to Screening of Arabidopsis thaliana Mutants. Metabolites 2, 398–428 (2012).
Acknowledgements
The authors are grateful to the Ministry of Science and Technology, Taiwan, for financial support granted to Dr. Kai-Wun Yeh under the project MOST-104-2311-B-002-011. We are also grateful to the staffs of TCX-D800 Metablomics Core, Technology Commons, College of Life Science, NTU for help with LC-ESI-MS analysis.
Author information
Authors and Affiliations
Contributions
R.S.K. and C.-H.S. conceived, implemented and performed the experiments. R.S.K. and C.-H.S. performed bioinformatics analysis and helped in interpreting the results. P.-Y.W. assisted confocal microscopy. S.S.K. and M.S.H. participated in biochemical assays. K.-W.Y. coordinated and participated in designing the overall experimental work. R.S.K. wrote manuscript. K.-W.Y. critically revised the complete structure and organization of the manuscripts. All the authors have read and approved the final version of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Senthil Kumar, R., Shen, CH., Wu, PY. et al. Nitric oxide participates in plant flowering repression by ascorbate. Sci Rep 6, 35246 (2016). https://doi.org/10.1038/srep35246
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep35246
This article is cited by
-
Influence of heat treatment on the optoelectronic performance of electrodeposited CdSe thin films
Journal of Materials Science: Materials in Electronics (2022)
-
Florigen governs shoot regeneration
Scientific Reports (2021)
-
Isolation, purification and characterization of an ascorbate peroxidase from celery and overexpression of the AgAPX1 gene enhanced ascorbate content and drought tolerance in Arabidopsis
BMC Plant Biology (2019)
-
Cyclic nucleotide-gated ion channel 6 mediates thermotolerance in Arabidopsis seedlings by regulating nitric oxide production via cytosolic calcium ions
BMC Plant Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.