Abstract
The abundance and diversity of bacteria in two types of ponds were investigated by quantitative PCR and Illumina MiSeq sequencing. The results revealed that the abundance of bacterial 16S rRNA genes in D ponds (with grass carp fed sudan grass) was significantly lower than that in E ponds (with grass carp fed commercial feed). The microbial communities were dominated by Proteobacteria, Cyanobacteria, Bacteroidetes, and Actinobacteria in both E and D ponds, while the abundance of some genera was significantly different between the two types of ponds. Specifically, some potential pathogens such as Acinetobacter and Aeromonas were found to be significantly decreased, while some probiotics such as Comamonadaceae unclassified and Bacillales unclassified were significantly increased in D ponds. In addition, water quality of D ponds was better than that of E ponds. Temperature, dissolved oxygen and nutrients had significant influence on bacterial communities. The differences in bacterial community compositions between the two types of ponds could be partially explained by the different water conditions.
Similar content being viewed by others
Introduction
Aquaculture is currently among the fastest-growing sector of food production in the world1. Global aquaculture production has been growing at an average rate of 6.2% per year during 2000–2012. It was reported that aquaculture production might account for 62% of the total amount of fish consumed by human beings in 20302. Although the high-density, high-yield aquaculture is rapidly growing in recent years, the development of intensive aquaculture was restricted due to increasing pollution of aquatic environments. One major problem in intensive aquaculture is the accumulation of toxic ammonia and nitrite in water column, which not only reduce water quality but also cause disease or even death of the farmed animals3.
In recent years, numerous studies have been reported that the growth of heterotrophic microorganisms will be stimulated and nitrogen is utilized to generate microbial proteins if organic carbon sources are added to the aquaculture systems4. With additional organic carbon, an accumulation of microorganism, protozoa and algae will develop in the aquaculture systems5. The resulting intensive microbial community can improve water quality through removal of toxic nitrogen species such as ammonia and nitrite, whilst the microbial protein produced can be used as feed6. Given the great advantages of adding carbon sources into aquaculture systems, numerous studies have investigated effects of carbon sources addition on water quality control7,8, protein nutritional contribution7,9, and growth-promoting for the feed animals10. However, there is a lack of information on the effect of adding carbon sources on the overall diversity and composition of bacterial communities in aquaculture ponds. An in-depth analysis of microbial communities will be useful for controlling the bacterial community composition for both optimal water quality and optimal feeding of cultured animal, ultimately for animal’s health.
Traditionally, conventional microbiological techniques, such as cultivation methods, 16S rRNA clone libraries, polymerase chain reaction-denaturing gradient gel electrophoresis (PCR-DGGE) and terminal restriction fragment length polymorphism (T-RFLP) have been used for investigation of bacterial communities in aquaculture11,12,13,14,15. However, these techniques are often laborious, time-consuming and poorly present the diversity of rare bacterial groups. Recently, high-throughput sequencing has been brought in as a new generation sequencing technology, which enables more detailed profiling of microbial populations with high throughput and low cost16,17,18. Several previous studies have been reported the microbial diversity and abundance in different ecosystems by high-throughput sequencing, such as in marine water19, natural wetland20, soil21, drinking water22, activated sludge23 and hot spring24. Therefore, these recent advances in DNA sequencing technologies allow us to develop a broader understanding of complex microbial communities affected by adding carbon sources into aquaculture system.
In this study, we hypothesized that adding carbon sources into aquaculture system will not only improve water quality but also change microbial communities in water column. To validate this hypothesis, we compared three typical ponds using commercial feed (control) with two ponds using sudan grass as a carbon source in the Gong’an county of Jingzhou, Hubei Province, China. Water samples were collected every month from June to October 2014 to explore the microbial communities using MiSeq Illumina sequencing technique. The findings of this study will help to understand the impact of the additional carbon sources on microbial communities in aquaculture ponds, and evaluated the impact of adding carbon sources on culture environment.
Results
Environmental description
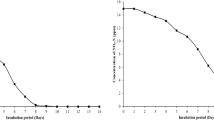
The characteristics of the surface and bottom water samples from two different feeding types of ponds within five months are summarized in Table 1. As observed, there were no significant differences in the pH values and DO concentrations no matter between surface and bottom or between two types of ponds (p > 0.05). The NH4+-N and NO2−-N concentration were also found no significant differences between the surface and bottom from the two type of ponds (p > 0.05). However, both NH4+-N and NO2−-N concentrations were found significantly lower in the ponds D than that in the ponds E (p < 0.05). Variations of water temperature, and the NH4+-N, NO2−-N, NO3−-N, TP, and TOC concentrations of the two types of ponds through five months are shown in Fig. 1. Temperature of the pond water ranged from16.9 ± 0.25 to 32.4 ± 0.06 °C, with the highest record in August. Concentrations of ammonia and nitrite exhibited the same variations in the grass carp fed with sudan grass (D) and those fed with commercial feeds (E). Ammonia and nitrite concentrations were higher in Ponds E than those in Ponds D.
Means ± standard deviations of the of water temperature, NH4+-N, NO2−-N, NO3−-N, TP, and TOC.
Concentrations in water layers of two types of ponds over five months (The capital letters D and E represent the ponds in which grass carp were fed with sudan grass and commercial feed, respectively. The lower case s and b represent the surface and bottom water layer, respectively). Error bars represent standard deviations.
Abundance of bacterial 16S rRNA genes
The abundance of bacterial 16S rRNA genes is shown in Fig. 2 as determined by qPCR from two types of ponds. The abundance of bacterial 16S rRNA genes in all samples had the same changing trend: it showed a decrease first, followed by an increase, a decrease and then an increase again (Fig. 2). The bacterial abundance ranged from3.53 ± 0.62 × 109 to 8.63 ± 1.01 × 1010 copies/mL and from2.81 ± 0.64 × 109 to 7.42 ± 0.87 × 1010 copies/mL for D ponds and E ponds, respectively.
Abundance of bacterial 16S rRNA genes in the surface and bottom water (The capital letters D and E represent the ponds in which grass carp were fed with sudan grass and commercial feed, respectively.
The lower case s and b represent the surface and bottom water layer, respectively). Error bars represent standard deviations.
In D ponds, the abundance of bacterial 16S rRNA genes was significantly higher in the surface water layer than in the bottom layer in August (p < 0.05), while no significant differences were observed between them in other months (p > 0.05). Similar results were obtained for E ponds.
The grass carp were fed with sudan grass in July and August in D ponds, and there were no significant differences in the abundance of bacterial 16S rRNA genes between D ponds and E ponds in these two months (p > 0.05). However, in September when the feed was changed to commercial feed again, the abundance of bacterial 16S rRNA genes in D ponds was significantly lower than that of E ponds (p < 0.05), and no differences were observed between the two types of ponds in October with further supplement of commercial feed (p > 0.05).
Illumina MiSeq sequencing results
After the removal of unqualified reads, the Illumina MiSeq sequencing analysis of fifty water samples yielded 745,337 sequences, with an average length of about 396 bp in the V4–V5 hypervariable regions of the 16S rRNA gene. As summarized in Table 2, the number of operational taxonomy units (OTUs) in the E ponds was larger than that in D ponds no matter in the surface or bottom. By contrast, the Shannon indices showed higher bacterial diversity in D ponds than those in E ponds both on the surface and bottom. It was also noted that bacterial diversity in D ponds was greater than that in E ponds at the bottom layer in August. The rarefaction curves of the twenty samples at the 3% distance cutoff level revealed that the bacterial phylotype richness of sample 7Es was considerably higher than that of other samples (Fig. 3).
Rarefaction curves for a dissimilarity of 3% from twenty samples.
(A) The surface water samples of the ponds in which grass carp were fed with sudan grass. (B) The bottom water samples of the ponds in which grass carp were fed with sudan grass. (C) The surface water samples of the ponds in which grass carp were fed with commercial feed. (D) The bottom water samples of the ponds in which grass carp were fed with commercial feed.
Bacterial community composition
RDP Classifier was used to assign the effective sequence tags into different phylogenetic bacterial taxa. Figure 4 shows the relative abundance of bacterial community at the level of phylum. Proteobacteria, Cyanobacteria, Bacteroidetes and Actinobacteria were four phyla abundant in all the samples from the two types of ponds. Except for July, Proteobacteria were the most abundant phylum in all experimental months, accounting for 34.95% of the total effective bacterial sequences, followed by Cyanobacteria (averaging at 18.8%), Bacteroidetes (averaging at 15.8%) and Actinobacteria (averaging at 14.7%). Other major phyla (average abundance > 1%) of the bacterial communities of the twenty samples included Firmicutes (4.6%), Planctomycetes (3.5%), Chloroflexi (1.9%) and Chlorobi (1.2%). Proteobacteria, Bacteroidetes, and Actinobacteria had the same varying tendency during the five months at the four sampling sites. However, in the surface water layer of D ponds, Cyanobacteria were first increased and then decreased during August to October, while in other three sampling sites it had the opposite varying tendency.
Relative abundance of bacterial community composition in twenty samples at the phylum level over five months (The number before the sample name represents the sampling month).
The capital letters D and E represent the ponds in which grass carp were fed with sudan grass and commercial feed, respectively. The lower case s and bin the sample name represent the surface and bottom water layer, respectively). Taxa represented occurred at >1% abundance in at least one sample.
To further understand the differences in dominant phyla between D and E ponds, the mean relative abundance of the four dominant OTUs at each sampling site in the five months was analyzed (Supplementary Fig. S1). Proteobacteria were found to be the most dominant phylum in all samples (except for the samples in July) followed by Actinobacteria, Bacteroidetes and Cyanobacteria. As observed, the relative abundance of Proteobacteria and Actinobacteria in D ponds were both higher than those in E ponds. However, the relative abundance of Bacteroidetes in D ponds was lower than that in E ponds. The relative abundance of Cyanobacteria fluctuated in D and E ponds, and showed no particular trends. For the rest, Firmicutes and Chlorobi were more abundant in E ponds than that in D ponds (4.0% and 1.1%, respectively), whereas Chloroflexi and Planctomycetes were less abundant.
To conduct a more detailed analysis of the composition of the communities in the water layers, all the reads had been assigned to a phylum into classes (Fig. 5). As observed, Betaproteobacteria were the dominant class among Proteobacteria in all water samples, accounting for 15.7%, followed by Alphaproteobacteria, Gammaproteobacteria and Deltaproteobacteria. Among these classes, Gammaproteobacteria were more abundant in D ponds (9.4% in surface layer, 8.5% in bottom layer) than in E ponds (5.9% in surface layer, 7.6% in bottom layer). However, the mean relative abundance of the five subdivisions within Proteobacteria in the two types of ponds was not significantly different over the five months (p > 0.05). Interestingly, in D ponds, Epsilonproteobacteria were only detected in June and July, averaging at 0.006% in total effective sequences, whereas in E ponds, they were detected in June, July and August, averaging at 0.003%. In addition, Deferribacteres and Thermotogae were only detected in the surface water of E ponds in June.
Hierarchically clustered heatmap showed the similarities and differences of these twenty bacterial communities at genus level (Fig. 6). Cluster analysis classified the samples into five clusters. Except for the samples from September, other samples from the same types of the pond were grouped together first, and then the samples in the same month from different types of the pond were clustered together.
At genus level, some interesting differences between the two types of ponds during the five months were observed. Some probiotics, such as Comamonadaceae unclassified and Bacillales unclassified were found to be significantly higher in D ponds than those in E ponds (p < 0.05) in July and August when the feed was changed to sudan grass feed. On the other hand, some potential pathogens such as Acinetobacter and Aeromonas were found to be significantly lower in D ponds than those in E ponds (p < 0.05) in August.
Similarity analysis of the twenty water samples
The weighted UniFrac clustering method was used to calculate the similarity or dissimilarity of the obtained sequences among different samples25. As shown in Fig. 7, based on abundances of orders, the bacterial communities in the twenty samples could be clustered into five groups, which included all four samples from one of the five months respectively: Group I (September), Group II (August), Group III (October), Group IV (July), Group V (June). Except for those from the samples in September, the communities from the same type of ponds were clustered together.
PCoA was used to estimate the similarities among different water samples using three different approaches: RDP Classifier taxa, OTUs and UniFrac, and PCoA (Fig. 8). The first principal coordinate of the weighted analysis accounted for 47.19% of the variation in the data. It clearly separated the samples in June from those in other four months. PC2 accounted for 20.79% of the variance in the bacterial communities.
Microbial community composition in relation to environmental variables
CCA was used to establish the relationships between the environmental factors and the bacterial community (Fig. 9). CCA plot was carried out using OUTs data together with environmental data (ammonia, nitrite, nitrate, total phosphorus, total organic carbon, temperature, pH, and dissolved oxygen). According to Monte Carlo permutation test (499 permutations), the significant relationships between environmental variables and canonical axes were analyzed by using Canoco program.
Canonical correspondence analysis (CCA) ordination diagram of bacterial communities associated with environmental variables of ammonia (NH4+-N), nitrite (NO2−-N), nitrate (NO3−-N), total phosphorus (TP), total organic carbon (TOC), temperature (T), pH, and dissolved oxygen (DO) (Eigenvalues: Axis 1: 0.385, Axis 2: 0.198, Axis 3: 0.168, Axis 4: 0.091).
Environmental variables were indicated as arrows.
Based on the 5% level in a partial Monte Carlo permutation test, the bacterial community and structure were significantly linked (p < 0.05) to the water environment factors. As shown in Fig. 9, the water samples were clearly clustered according to sampling time rather than sampling water layer. CCA results explained 36.8% and 18.9% of the variation in the first two axes, respectively (Fig. 9). Dissolved oxygen, temperature and total organic carbon were the most important environmental factors to influence the water community composition, and were positively correlated with Axis 1 (p < 0.05) (Table 3). Axis 2 had a positive correlation with temperature, nitrite and nitrate (p < 0.05), but was negatively correlated with ammonia (p < 0.01) (Table 3), suggesting that Axis 2 had a gradient in temperature, nitrite, nitrate and ammonia. By contrast, other nutrient factors (pH and total phosphorus) had no significant correlation to bacterial community (p > 0.05).
Discussion
In this study, we showed the vertical stratification of the 16S rRNA bacterial abundance in the water layers of two different types of aquaculture ponds. The bacterial 16S rRNA gene abundance was significantly higher in August than in other four months in both types of ponds. And the highest temperature was observed in August. These results indicate that temperature may play an important role in the seasonal dynamics of bacterial abundance. Similar phenomenon was observed in other aquaculture ponds, where total bacterial abundance varied with temperature26.
After changing the types of feeds, the bacterial abundance in E ponds was higher than that in D ponds (with sudan grass supplied in July and August) in September (p < 0.05) and October (p < 0.05), respectively. But there was no significant difference in bacterial abundance between D and E ponds in July and August. These results reveal that different types of feeds can influence the bacterial abundance, and possibly the effect of supplying sudan grass on the bacterial abundance was slightly delayed. The higher bacterial abundance in E ponds in September and October may be due to the trophic state of the ponds, as previous studies have demonstrated that the total number of bacterial cells observed in three different sediments may be influenced by the reservoir trophic state27.
The number of OTUs in each sample was estimated with the largest 16S rRNA libraries sequenced to date. However, a lot of “Unclassified” sequences were also detected in the samples, suggesting that there might be a high abundance of unknown microbial lineages in the aquaculture pond environment, which should be studied in detail in the future. At the 3% distance cutoff level, the OTUs ranged from 699 to 1153 in this study. The Shannon diversity values ranged from 3.97 to 5.31 and the highest was observed in 8Ds. These values are in accordance with those reported for some other aquaculture systems28,29. However, according to the study of the microbial diversity of seawater in the East China Sea, the Shannon diversity values were found in the range of 3.42 to 5.6530. Another study showed low diversity values, ranged from 2.4 to 3.4 when studying the microbial community of a warm monomictic tropical freshwater lake31. These results indicate that aquaculture system may harbor higher microbial species richness than other aquatic systems.
Seasonal pattern of bacterioplankton community composition has been reported for aquaculture systems32, river33, lake34, coastal water35 and the open ocean36. However, these previous studies were mostly based on fingerprinting methods, which only include the most abundant members of the community. The present study is the first attempt to apply Illumina MiSeq sequencing to study the seasonal dynamics, including both abundant and rare populations of the observed OTUs in two different types of aquaculture ponds. We showed the vertical stratification of the microbial communities in the pond water layers over five months, and two water layers corresponding to the surface and bottom water bodies were investigated. We hypothesized that there would be significant differences among the water samples from different months. It was shown that the studied microbial community was featured by strong temporal shifts and seasonal clustering. As demonstrated by the clustering, PCoA, and CCA analyses, the water samples from the same sampling month were fairly similar, while the samples from different months were different. This difference may be due to the difference in water conditions in different months, which determine the microbial community distribution. Previous studies have demonstrated that different temperatures and oxygen concentrations significantly alter the microbial community composition in freshwater sediment37,38. One potential driver of seasonal change in community composition is temperature39, and this can also be the case for pond water, as the growth of microbes in this study was at least seasonally affected by temperature. This result is consistent with the results of studies on phytoplankton communities, which showed that pelagic bacterial communities experienced climate-driven seasonal environmental changes40. The CCA result in our study also demonstrates that the temperature is positively correlated with the water community composition. Thus, temperature seems to be an important factor that affects microbial communities.
Many previous studies on the composition of microbial communities in the water have relied heavily on clone library analysis, which only sequences rather few16S rRNA gene fragments11,41,42. A previous study analyzed the bacterial diversity in the influent from a municipal waste water treatment plant using high-throughput sequencing method43. Their results showed that Proteobacteria were the most abundant phylum in the influent sample, which is consistent with our study. However, the clustering of some phyla in the present study is consistent with the findings of the study of the surface water samples in sewage treatment plants44, which revealed that the main phyla were Proteobacteria, Actinobacteria, and Bacteroidetes based on an examination of bacterial community structures by 454 sequencing, but the relative abundance differed from that in the present study. Their results showed that the dominant phyla were Proteobacteria (20.28–67.89%), Bacteroidetes (3.85–16.14%), Acidobacteria (19.78–53.59%), and Cyanobacteria (0.68 to 2.6%) in surface water samples, while our results revealed that Proteobacteria were the most abundant phylum in all twenty water samples and accounted for 24.28–48.22%, followed by Cyanobacteria (5.9–37.61%), Bacteroidetes (5.4–34.82%), and Actinobacteria (8.2–22.37%). Another study reported the bacterial community composition in the Red Sea based on 454 pyrosequencing, and revealed the vertical stratification of the microbial communities in the water layers above the Atlantis II and Discovery Deeps. Their study showed that the classified bacterial reads from the upper layers (20 and 50 m) were dominated by Cyanobacteria, whereas in the deeper layers (200 and 1500 m), the largest group was Proteobacteria19.
The microbial compositions at different depths seemed to be very similar in both types of ponds, and no significant differences of the four dominant phyla (Proteobacteria, Cyanobacteria, Bacteroidetes, and Actinobacteria) were observed between the surface and bottom layers. These results are similar to those of a study using 16S rRNA pyrosequencing in seawater19, which showed that the microbial composition in the surface (20 and 50 m) seems to be very similar. This phenomenon may be due to the similarity of water conditions in the surface and bottom water layers. Freshwater pond is a shallow water body, in which the nutrient concentrations showed no significant difference between the surface and bottom water layers as shown in Table 1. Similar results were also reported in intensive GIFT tilapia (Oreochromisniloticus) ponds45, which demonstrated that the microbial compositions of four different water layers were similar.
There were no significant differences in the four dominant phyla (Proteobacteria, Cyanobacteria, Bacteroidetes and Actinobacteria) between the two types of ponds over the five months. Similar phenomenon was also observed in two rivers, where two clone libraries of 16S rDNA constructed with summer samples from each river were not significantly different and contained typical freshwater bacterioplankton of Betaproteobacteria, Bacteroidetes, and Actinobacteria33. However, as demonstrated by the clustering and PCoA analyses, the communities from the same type of ponds were clustered together in this study, except for the samples in September. Besides, except for in the water samples in June, Proteobacteria were found to be the dominant phylum in the samples from other months, which is similar to the analytical results of bacterial communities in drinking water46. The classified bacterial reads of the samples from D ponds were dominated by Proteobacteria, which were decreased in E ponds in each depth (Fig. 5). Proteobacteria contain a very high level of bacterial metabolic diversity related to global carbon, nitrogen and sulfur cycling47. The effects of different types of feeds on the distributions of the proteobacterial subdivisions in the ponds were also studied. Here, Betaproteobacteria were the most abundant class in the Proteobacteria phylum. This is different from the results of a study using 454 pyrosequencing19, which showed that Gammaproteobacteria were the most abundant Proteobacteria in sea water. However, our finding is similar to the analytical results of bacterial communities in soil48 and activated sludge18, which demonstrated that Betaproteobacteria were the most abundant in Proteobacteria. The classified bacterial reads of the samples from D ponds were dominated by Gammaproteobacteria, which were decreased in samples from E ponds (Fig. 5). A previous study showed the shift of bacterial community structure to a relatively higher abundance of Gammaproteobacteria due to the addition of organic carbon substances49. In our study, the TOC concentration (Table 1) in D ponds was slightly higher than that in E ponds, which might result from the supply of sudan grass for D ponds in July and August. A decrease of Actinobacteria in the samples of E ponds was observed compared within the samples of D ponds. Here, both the nitrate and ammonium concentrations were higher in E ponds than in D ponds, which is supported by a study which demonstrated that both the nitrate and ammonium concentrations negatively affect the abundance of Actinobacteria50. Bacteroidetes were enriched in water samples of E ponds. This might be explained by the reason that the ammonia concentration was higher in E ponds than in D ponds, which is supported by a study showing that Bacteroidetes are positively correlated with ammonia concentration in a hot spring51.
Comamonadaceae unclassifiedand and Bacillales_unclassified were significantly enriched in D ponds compared with in E ponds when the feed was changed to sudan grass feed. Comamonadaceae unclassified was are likely to be nitrate-reducing bacteria52 and could decrease organic carbon in the ponds. Their higher abundance in D ponds might be related to the higher TOC concentration in D ponds resulted from the addition of available organic carbon for denitrification. Some species of Bacillus are used as probiotics, such as Bacillus subtilis, Bacillus licheniformis, in the aquaculture system. Acinetobacter and Aeromonas, many species of which are pathogenic bacteria in aquaculture ponds53,54, were significantly abundant in E ponds in August. This result indicates that sudan grass is effective to decrease Acinetobacter and Aeromonas. The reasons for the higher abundance of Bacillus, Acinetobacter, and Aeromonas in E ponds still need further study.
CCA in this study shows that DO, temperature and nutrients (TOC, nitrate, ammonium and nitrite) significantly influence the composition of bacterial communities in pond water. A previous study showed that DO, temperature, pH and nutrients (total nitrogen, total phosphorus) are the key factors influencing the bacterioplankton diversity of Lake Taihu55. Another study showed that ammonium, chemical oxygen demand and total nitrogen have effects on the composition of bacterial communities in Litopeneaus vannamei aquaculture water56. CCA plots revealed that samples collected in the same month were almost clustered together. This result suggests that the bacterial communities might demonstrate a seasonal pattern, which is consistent with the previous studies57,58.
In conclusion, the community structure differed between the ponds supplied with different types of feeds and the ponds supplied with sudan grass had better water conditions. The pathogenic bacteria genera Acinetobacter and Aeromonas were significantly decreased in the ponds supplied with sudan grass. In addition, the microbial communities in the ponds had a seasonal pattern in our study. These results suggest that the modulation of diets might influence the community structure and thus might effectively change water quality. Our findings could provide a promising direction for the healthy aquaculture of grass carp.
Materials and Methods
All experiments were approved by and carried out in compliance with the guidelines of the Institutional Animal Care and Use Committees (IACUC) of Huazhong Agricultural University, Wuhan, China
Fish culture and water sampling
Water samples were collected from an aquaculture farm (29°54′6.61″ N, 112°15′56.41″ E), Gong’an, Hubei, China. Five aquaculture ponds were selected for investigation. The ponds had a water surface of 2800 m2 (70 m × 40 m), with an average depth of 1.6 m and maximum depth of 1.8–2.0 m, where grass carp (Ctenopharyngodon idellus) were raised as the major species. At the time of sampling, the ponds had been used for 3 years with stable production under intensive cultural management. Five ponds differed mainly on the supply of fish feeds: three of them were provided with commercial feed from June to October 2014, and the other two were provided with commercial feed in June, September, October 2014, and sudan grass (Sorghum sudanense) in July and August 2014, respectively.
The sampling was conducted in 2014 on June 16th, July 16th, August 28th, September 21st, and October 14th. Samples were taken from the surface and bottom water column, which corresponded to 0.2 and 1.6 m of depth, respectively. Five samples (200 mL) were taken from five evenly distributed points within each pond then mixed for each depth. These water samples were collected, homogenized and sub-sampled for further analysis. One part of these water samples were used to measure the physicochemical factors immediately. The other part samples were placed in an incubator box with ice packs until further processing in the laboratory and these water samples were filtered through a 0.22-μm nucleopore filter (diameter: 47 mm) within 12 h after water collection, then kept at −80 °C until DNA extraction. The ponds supplied with sudan grass or commercial feed were designated as D ponds and E ponds, respectively. Thus, the water samples were designated as: Ds and Db (the surface and bottom water of D ponds, respectively), Es and Eb (the surface and bottom water of E ponds, respectively). The number before the sample name represents the sampling month (for example, 6Ds refers to the surface water sample of D ponds collected in June).
Physicochemical analysis
Temperature (T), pH, and dissolved oxygen (DO) concentration were measured using an HQ 30d multi-parameter water quality analyzer (HACH, Loveland, CO). Total phosphorus (TP) was determined with ammonium molybdate spectrophotometric method. Total organic carbon (TOC) was determined on a Total Organic Carbon Analyzer (Elementar, Hanau, Germany). Ammonia (NH4+-N), nitrite (NO2−-N), and nitrate (NO3−-N) were determined by the methods described by Lu et al.55.
Quantitative PCR (qPCR)
Total genomic DNA of each sample was extracted as previously described59. The concentration of the extracted DNA was quantified with a ND-2000 UV-vis spectrophotometer (USA). Quantitative PCR was carried out using a Qiagen Q thermo cycler (Qiagen, Hilden, Germany). The reaction mixture (20 μL) contained 10 μL SYBR Premix Ex Taq II, 0.2 μM of each primer and 1 μL template DNA (10 ng). The specific primers for the amplification of bacterial 16SrRNA genes were 515F (5′-GTGCCAGCMGCCGCGG-3′) and 907R (5′-CCGTCAATTCMTTTRAGTTT-3′) as previously described60. The qPCR reactions were performed as follows: 95 °C for 30 s; 35 cycles of 95 °C for 5 s, 55 °C for 30 s, followed by 72 °C for 1 min. The specificity of the qPCR amplification was determined by melting curve and gel electrophoresis.
The standard curve was constructed using plasmid DNA as the standard sample. The plasmid DNA was extracted from the positive colony, which was confirmed by that bacterial 16S rRNA genes were successfully ligated into the pMD18-T Vector (Takara, Dalian, China). The concentration of the plasmid DNA was determined with a ND-2000 UV-vis Spectrophotometer and bacterial 16S rRNA gene copy numbers of plasmid DNA were calculated using the detected concentration. The standard samples were produced by ten-fold serial dilution of plasmid DNA. The standard curve was observed to have a correlation coefficient (R2) of 0.995 and an efficiency of 93%.
Illumina MiSeq sequencing
In this study, we collected samples for five consecutive months (from June to October) to conduct Illumina MiSeq sequencing. All PCR products of 16S rRNA genes were performed in triplicate using Illumina MiSeq Sequencer (Illumina) by Majorbio Bio-pharm Biotechnology Co. Ltd. (Shanghai, China) as described previously61. PCR products were pooled and purified using an Axy Prep™ DNA Gel Extraction Kit (Axygen). All the sequences used in this study are available from the NCBI Sequence Read Archive (SRA) under accession number SRA3016262.
Bioinformatics and statistical analysis
Sequencing data were processed using the Quantitative Insights Into Microbial Ecology (QIIME) pipeline (http://qiime.sourceforge.net/)62. The sequences with the same barcode were assigned to the same sample, and then the barcode and primer sequences were removed. Denoised sequences with one mismatched base in the barcode, overlapped shorter than 10 bp, containing ambiguous characters, with more than two mismatched bases in the primers, or with a sequence length shorter than 50 bp were eliminated. The chimeric sequences were removed from aligned sequences using the UCHIME method63. And the valid reads obtained from Illumina MiSeq sequencing were normalized to 20000 for comparison of community diversity. The reads were then clustered into operational taxonomic units (OTUs, 97% similarity)64. The Greengenes data base was used to determine the taxonomic identity of each phylotype65. The diversity indices ACE and Chao166 were estimated using Mothur67 to indicate the community richness. Simpson and Shannon indices were estimated to indicate the community diversity. Coverage, which represents the sequencing depth, was calculated using custom R scripts. The differences in overall bacterial community structure between each pair of water samples were detected using the UniFrac metric25. Heatmap was constructed with R software. PCoA (principal coordinate analysis) was used to characterize the changes of the community composition by using R software.
All data are presented as the means ± standard deviation (SD); n refers to the number of samples. Statistical analysis was performed using the SPSS 21.0 software package. The differences in the abundance of bacterial 16S rRNA genes and in the four dominant phyla among the samples from the two types of ponds were evaluated by one-way ANOVA. The differences between the relative abundance of genus from the two types of ponds were also evaluated by one-way ANOVA. CCA (Canonical correspondence analysis) plot and Monte Carlo Permutation test were used to analyze the relationship between the pond water community composition and the environmental factors using CANOCO 4.5 program.
Additional Information
How to cite this article: Qin, Y. et al. Bacterial abundance and diversity in pond water supplied with different feeds. Sci. Rep. 6, 35232; doi: 10.1038/srep35232 (2016).
References
Klinger, D. & Naylor, R. Searching for solutions in aquaculture: charting a sustainable course. Annual Review of Environment and Resources 37, 247–276 (2012).
FAO The State of World Fisheries and Aquaculture: Opportunities and Challenges. FAO, Rome, Italy (2014).
Boyd, C. E. & Tucker, C. S. Pond aquaculture water quality management. (Springer Science & Business Media, 2012).
Avnimelech, Y. Carbon/nitrogen ratio as a control element in aquaculture systems. Aquaculture 176, 227–235 (1999).
Crab, R., Defoirdt, T., Bossier, P. & Verstraete, W. Biofloc technology in aquaculture: beneficial effects and future challenges. Aquaculture 356, 351–356 (2012).
Cardona, E. et al. Bacterial community characterization of water and intestine of the shrimp Litopenaeus stylirostris in a biofloc system. BMC Microbiology 16, 1 (2016).
Azim, M. E. & Little, D. C. The biofloc technology (BFT) in indoor tanks: water quality, biofloc composition, and growth and welfare of Nile tilapia (Oreochromis niloticus). Aquaculture 283, 29–35 (2008).
Hari, B., Kurup, B. M., Varghese, J. T., Schrama, J. & Verdegem, M. The effect of carbohydrate addition on water quality and the nitrogen budget in extensive shrimp culture systems. Aquaculture 252, 248–263 (2006).
Asaduzzaman, M. et al. C/N ratio control and substrate addition for periphyton development jointly enhance freshwater prawn Macrobrachium rosenbergii production in ponds. Aquaculture 280, 117–123 (2008).
Hari, B., Kurup, B. M., Varghese, J. T., Schrama, J. & Verdegem, M. Effects of carbohydrate addition on production in extensive shrimp culture systems. Aquaculture 241, 179–194 (2004).
Cottrell, M. T. & Kirchman, D. L. Community composition of marine bacterioplankton determined by 16S rRNA gene clone libraries and fluorescence in situ hybridization. Applied and Environmental Microbiology 66, 5116–5122 (2000).
Revetta, R. P., Matlib, R. S. & Santo Domingo, J. W. 16S rRNA gene sequence analysis of drinking water using RNA and DNA extracts as targets for clone library development. Current microbiology 63, 50–59 (2011).
Zhang, R. et al. Diversity of bacterioplankton in contrasting Tibetan lakes revealed by high-density microarray and clone library analysis. FEMS microbiology ecology 86, 277–287 (2013).
Bekaert, K., Devriese, L., Maes, S. & Robbens, J. Characterization of the dominant bacterial communities during storage of Norway lobster and Norway lobster tails (Nephrops norvegicus) based on 16S rDNA analysis by PCR-DGGE. Food microbiology 46, 132–138 (2015).
Zhong, F. et al. Bacterial community analysis by PCR-DGGE and 454-pyrosequencing of horizontal subsurface flow constructed wetlands with front aeration. Applied microbiology and biotechnology 99, 1499–1512 (2015).
Portune, K. J., Pérez, M. C., Álvarez-Hornos, F. J. & Gabaldón, C. Investigating bacterial populations in styrene-degrading biofilters by 16S rDNA tag pyrosequencing. Applied microbiology and biotechnology 99, 3–18 (2015).
Wang, X., Hu, M., Xia, Y., Wen, X. & Ding, K. Pyrosequencing analysis of bacterial diversity in 14 wastewater treatment systems in China. Applied and environmental microbiology 78, 7042–7047 (2012).
Zhang, T., Shao, M.-F. & Ye, L. 454 Pyrosequencing reveals bacterial diversity of activated sludge from 14 sewage treatment plants. The ISME journal 6, 1137–1147 (2012).
Qian, P.-Y. et al. Vertical stratification of microbial communities in the Red Sea revealed by 16S rDNA pyrosequencing. The ISME journal 5, 507–518 (2011).
Peralta, R. M., Ahn, C. & Gillevet, P. M. Characterization of soil bacterial community structure and physicochemical properties in created and natural wetlands. Science of the Total Environment 443, 725–732 (2013).
Eilers, K. G., Debenport, S., Anderson, S. & Fierer, N. Digging deeper to find unique microbial communities: the strong effect of depth on the structure of bacterial and archaeal communities in soil. Soil Biology and Biochemistry 50, 58–65 (2012).
Hong, P.-Y. et al. Pyrosequencing analysis of bacterial biofilm communities in water meters of a drinking water distribution system. Applied and Environmental Microbiology 76, 5631–5635 (2010).
Sánchez, O., Ferrera, I., González, J. M. & Mas, J. Assessing bacterial diversity in a seawater‐processing wastewater treatment plant by 454‐pyrosequencing of the 16S rRNA and amoA genes. Microbial biotechnology 6, 435–442 (2013).
Hou, W. et al. A comprehensive census of microbial diversity in hot springs of Tengchong, Yunnan Province China using 16S rRNA gene pyrosequencing. PloS one 8, e53350 (2013).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Applied and environmental microbiology 71, 8228–8235 (2005).
Fan, L. M. et al. Bacterioplankton community analysis in tilapia ponds by Illumina high-throughput sequencing. World J Microb Biot 32 (2016).
Zhang, H.-h. et al. Abundance and diversity of bacteria in oxygen minimum drinking water reservoir sediments studied by quantitative PCR and pyrosequencing. Microbial ecology 69, 618–629 (2015).
Zhang, H. et al. Dynamic changes of microbial communities in Litopenaeus vannamei cultures and the effects of environmental factors. Aquaculture 455, 97–108 (2016).
Li, L., Yan, B., Li, S., Xu, J. & An, X. A comparison of bacterial community structure in seawater pond with shrimp, crab, and shellfish cultures and in non-cultured pond in Ganyu, Eastern China. Annals of Microbiology 66, 317–328 (2016).
Dong, Y. et al. Bacterial diversity and community structure in the East China Sea by 454 sequencing of the 16S rRNA gene. Chinese Journal of Oceanology and Limnology 32, 527–541 (2014).
Santos, A. B. et al. Environmental diversity of bacteria in a warm monomictic tropical freshwater lake. Annals of Microbiology, 1–10 (2015).
Li, Q. et al. Phylogenetic analysis of bacterial communities in the shrimp and sea cucumber aquaculture environment in northern China by culturing and PCR–DGGE. Aquaculture international 18, 977–990 (2010).
Crump, B. C. & Hobbie, J. E. Synchrony and seasonality in bacterioplankton communities of two temperate rivers. Limnology and Oceanography 50, 1718–1729 (2005).
Shade, A. et al. Interannual dynamics and phenology of bacterial communities in a eutrophic lake. Limnology and Oceanography 52, 487–494 (2007).
Fuhrman, J. A. et al. Annually reoccurring bacterial communities are predictable from ocean conditions. Proceedings of the National Academy of Sciences 103, 13104–13109 (2006).
Morris, R. M. et al. Temporal and spatial response of bacterioplankton lineages to annual convective overturn at the Bermuda Atlantic Time‐Series Study Site. Limnology and Oceanography 50, 1687–1696 (2005).
Wu, Y. et al. Autotrophic growth of bacterial and archaeal ammonia oxidizers in freshwater sediment microcosms incubated at different temperatures. Applied and environmental microbiology 79, 3076–3084 (2013).
Bryant, L. D., Little, J. C. & Bürgmann, H. Response of sediment microbial community structure in a freshwater reservoir to manipulations in oxygen availability. FEMS microbiology ecology 80, 248–263 (2012).
Shish, F. K. & Ducklow, H. W. Temperature regulation of heterotrophic bacterioplankton abundance, production, and specific growth rate in Chesapeake Bay. Limnology and Oceanography 39, 1243–1258 (1994).
Reynolds, C. ecology of freshwater phytoplankton (1984).
Sekiguchi, H., Watanabe, M., Nakahara, T., Xu, B. & Uchiyama, H. Succession of bacterial community structure along the Changjiang River determined by denaturing gradient gel electrophoresis and clone library analysis. Applied and Environmental Microbiology 68, 5142–5150 (2002).
Schmidt, T. M., DeLong, E. & Pace, N. Analysis of a marine picoplankton community by 16S rRNA gene cloning and sequencing. Journal of bacteriology 173, 4371–4378 (1991).
McLellan, S., Huse, S., Mueller-Spitz, S., Andreishcheva, E. & Sogin, M. Diversity and population structure of sewage‐derived microorganisms in wastewater treatment plant influent. Environmental microbiology 12, 378–392 (2010).
Wei, Y.-M. et al. Bacterial communities of Beijing surface waters as revealed by 454 pyrosequencing of the 16S rRNA gene. Environmental Science and Pollution Research, 1–10 (2015).
Fan, L. et al. Characterization of microbial communities in intensive GIFT tilapia (Oreochromis niloticus) pond systems during the peak period of breeding. Aquaculture Research (2015).
Pinto, A. J., Xi, C. & Raskin, L. Bacterial community structure in the drinking water microbiome is governed by filtration processes. Environmental science & technology 46, 8851–8859 (2012).
Kersters, K. et al. Introduction to the Proteobacteria. The Prokaryotes: Volume 5: Proteobacteria: Alpha and Beta Subclasses, 3–37 (2006).
Roesch, L. F. et al. Pyrosequencing enumerates and contrasts soil microbial diversity. The ISME journal 1, 283–290 (2007).
Broszat, M. et al. Wastewater irrigation increases the abundance of potentially harmful Gammaproteobacteria in soils in Mezquital Valley, Mexico. Applied and environmental microbiology 80, 5282–5291 (2014).
Philippot, L. et al. Spatial patterns of bacterial taxa in nature reflect ecological traits of deep branches of the 16S rRNA bacterial tree. Environmental Microbiology 11, 3096–3104 (2009).
Jiang, Z. et al. Microbial Community Structure and Arsenic Biogeochemistry in an Acid Vapor-Formed Spring in Tengchong Geothermal Area, China. PloS one 11 (2016).
Sadaie, T. et al. Reducing sludge production and the domination of Comamonadaceae by reducing the oxygen supply in the wastewater treatment procedure of a food-processing factory. Bioscience, biotechnology, and biochemistry 71, 791–799 (2007).
Xia, L. et al. Recovery of Acinetobacter baumannii from diseased channel catfish (Ictalurus punctatus) in China. Aquaculture 284, 285–288 (2008).
Wu, S. et al. Composition, diversity, and origin of the bacterial community in grass carp intestine. PloS one 7, e30440–e30440 (2012).
Tian, C. et al. Spatiotemporal transition of bacterioplankton diversity in a large shallow hypertrophic freshwater lake, as determined by denaturing gradient gel electrophoresis. Journal of Plankton Research, fbp028 (2009).
Zeng, D.-N. et al. Analysis of the bacterial communities associated with different drinking water treatment processes. World Journal of Microbiology and Biotechnology 29, 1573–1584 (2013).
Bosshard, P. P., Stettler, R. & Bachofen, R. Seasonal and spatial community dynamics in the meromictic Lake Cadagno. Archives of microbiology 174, 168–174 (2000).
Yannarell, A., Kent, A., Lauster, G., Kratz, T. & Triplett, E. Temporal patterns in bacterial communities in three temperate lakes of different trophic status. Microbial ecology 46, 391–405 (2003).
Lu, S. et al. Seasonal dynamics of ammonia-oxidizing microorganisms in freshwater aquaculture ponds. Annals of Microbiology, 1–7 (2014).
Großkopf, R., Janssen, P. H. & Liesack, W. Diversity and structure of the methanogenic community in anoxic rice paddy soil microcosms as examined by cultivation and direct 16S rRNA gene sequence retrieval. Applied and environmental microbiology 64, 960–969 (1998).
Xiong, J. et al. Geographic distance and pH drive bacterial distribution in alkaline lake sediments across Tibetan Plateau. Environmental microbiology 14, 2457–2466 (2012).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature methods 7, 335–336 (2010).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Applied and environmental microbiology 72, 5069–5072 (2006).
Chao, A. & Bunge, J. Estimating the number of species in a stochastic abundance model. Biometrics 58, 531–539 (2002).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Applied and environmental microbiology 75, 7537–7541 (2009).
Acknowledgements
The study was funded by the State Science and Technology Support Program of China (2012BAD25B06), Special Fund for Agro-Scientific Research in the Public Interest Project (No. 201203083), the Earmarked Fund for China Agriculture Research System (CARS-46), the Fundamental Research Funds for Innovation Center of Hubei Province (2016ZXPY02) and the Fundamental Research Funds for the Central Universities (No. 2662015PY119).
Author information
Authors and Affiliations
Contributions
X.H., J.H. and Y.Q. conceived and designed the work. Y.Q. performed the experiments. Samples were collected by Y.Q., M.D., Q.L., C.W. and Y.J. Y.Q. drafted the manuscript. J.H. and X.H. modified the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Qin, Y., Hou, J., Deng, M. et al. Bacterial abundance and diversity in pond water supplied with different feeds. Sci Rep 6, 35232 (2016). https://doi.org/10.1038/srep35232
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep35232
This article is cited by
-
Bacterial community structure and bacterial isolates having antimicrobial potential in shrimp pond aquaculture
AMB Express (2022)
-
Mobile resistome of microbial communities and antimicrobial residues from drinking water supply systems in Rio de Janeiro, Brazil
Scientific Reports (2022)
-
The Effect of Two Dietary Protein Sources on Water Quality and the Aquatic Microbial Communities in Marron (Cherax cainii) Culture
Microbial Ecology (2021)
-
Interaction of environmental eukaryotic microorganisms and fungi in the pond-cultured carps: new insights into the potential pathogenic fungi in the freshwater aquaculture
Environmental Science and Pollution Research (2021)
-
Evaluation of the Physiological Bacterial Groups in a Tropical Biosecured, Zero-Exchange System Growing Whiteleg Shrimp, Litopenaeus vannamei
Microbial Ecology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.