Abstract
The female-limited Batesian mimicry polymorphism in Papilio butterflies is an intriguing system for investigating the mechanism of maintenance of genetic polymorphisms. In Papilio polytes, an autosomal region encompassing the sex-determinant gene doublesex controls female-limited mimicry polymorphism. In the closely related species P. memnon, which also exhibits female-limited Batesian mimicry polymorphism, we identified two allelic sequences of the doublesex gene that corresponded exactly with the mimetic and non-mimetic female phenotypes. Thus, the genetic basis of the mimicry polymorphism in P. memnon is similar to that in P. polytes. However, the mimetic and non-mimetic alleles of the two species were not identical, and the divergence of alleles occurred independently in P. memnon and P. polytes. Different mutation-selection processes may have resulted in the convergent patterns of mimicry polymorphism in these Papilio butterflies.
Similar content being viewed by others
Introduction
Batesian mimicry is the phenomenon in which palatable mimics avoid predation by resembling unpalatable species1,2,3. One of the most intriguing types of Batesian mimicry in butterflies is the female-limited polymorphism in which females display both mimetic and non-mimetic forms4,5,6. Some Papilio butterflies are textbook examples of female-limited Batesian mimicry polymorphism, which is controlled by a supergene locus7,8,9,10,11. Recently, the genetic basis of Batesian mimicry was revealed in two Papilio species, P. polytes and P. dardanus, and the allelic differences associated with mimicry polymorphism have been identified12,13,14. In P. polytes, a single ~130-kb autosomal region including the sex-determinant gene doublesex (dsx), with mimetic (H-type) and non-mimetic (h-type) allelic sequences, controls polymorphism, and these two alleles are protected against recombination by an inversion that covers the entire region of the dsx gene12,14. These findings would facilitate the comparative genomic study of female-limited mimicry polymorphisms in Papilio butterflies to understand the evolutionary diversification of genes underlying the complex adaptive traits involved in mimicry.
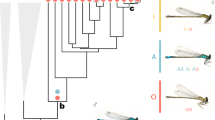
The Great Mormon butterfly, Papilio memnon, is closely related to P. polytes and exhibits similar female-limited Batesian mimicry polymorphism15,16. P. memnon females can exist as either the mimetic or non-mimetic form, whereas P. memnon males exhibit only the non-mimetic form (Fig. 1). Mimetic females have tails in their hindwings and aposematic colours in their hindwings and abdomens (Fig. 1), which allow them to mimic unpalatable species in the genera Pachliopta and Atrophaneura9,11. These phenotypic traits are tightly linked with a putative supergene9,11. Based on the phylogenetic proximity of P. polytes and P. memnon (Fig. 2), we predicted that the genetic basis of female-limited Batesian mimicry in P. memnon would be the same as that in P. polytes and that P. memnon would also have doublesex alleles corresponding to mimetic and non-mimetic forms.
In this study, we first sequenced mRNA from the wing discs of P. memnon pupae to identify dsx ortholog sequences. Next, we identified specific sequences in mimetic (H) and non-mimetic (h) alleles by sequencing cDNA derived from mRNA from reared individuals. Second, we sequenced a portion of dsx exon 1 using genomic DNA from field-collected individuals to confirm that the mimetic phenotype corresponded with the dsx allele type. Finally, we performed high-resolution melting (HRM) analysis to determine dsx genotypes from genomic DNA of males and females collected in the field, and investigated geographic variation in the frequency of the mimetic allele in P. memnon populations of Taiwan and Okinawa, Japan.
Results
Our de novo assembly of 101-bp paired-end reads from P. memnon transcriptomes (DDBJ DRA accession nos.: SAMD00052594-SAMD00052595) yielded 45,887 and 39,035 contigs for one female and one male, respectively (Table S1, Supplementary Information). In a BLAST search of the dsx sequences, two of the female contigs (c1658_g1_i1, c17110_g5_i1) yielded significant matches with the F1 isoform of the dsx H and h alleles of P. polytes, but no male contig yielded a significant match (Table S2, Supplementary Information). The two female contigs were identified as portions of the dsx sequence separated by a 59-bp gap. Upon sequencing of a 567-bp portion of contig c1658_g1_i1, we obtained three heterozygous and three homozygous sequences from 6 female cDNA samples and six heterozygous and five homozygous sequences from 11 male cDNA samples. These sequences were comprised of only two genotypes, probably corresponding to Hh and hh. Next, we discriminated the H and h (hh) sequences by subtracting the h sequence from the Hh sequence (Fig. 3; DDBJ accession: LC155217-LC155218). The 567-bp sequence contained 18 nucleotide substitutions (3.2% of all bp) between H and h, of which 4 were non-synonymous (Fig. 3). In this region, the P. memnon H sequence differed from that of P. polytes by 57 nucleotide substitutions (10.1%), of which 13 were non-synonymous. The P. memnon h sequence differed from that of P. polytes by 33 nucleotide substitutions (5.8%), of which 5 were non-synonymous. None of the non-synonymous substitutions between the H and h sequences were shared between the two species. The P. polytes h sequence contained a 14-bp insertion in the 5′-UTR (untranslated region) that was absent in P. memnon. The reconstructed phylogenetic relationship among the four dsx allelic sequences from P. memnon and P. polytes revealed that the divergence of H and h occurred independently within each species (Fig. 4).
cDNA and amino acid sequences of the Papilio memnon doublesex (dsx) H and h alleles.
The amino acid sequences are shown above the cDNA sequences. Dots indicate sequence identities in the dsx H and h alleles. Single-nucleotide polymorphisms are highlighted in red, and amino acid substitutions are highlighted in blue. The primers used are indicated above the sequences.
Sequencing of the 185-bp portions (containing 10 single-nucleotide polymorphisms) from 134 wild-caught individuals again yielded three sequence types, corresponding to the putative HH, Hh and hh genotypes. In females, the HH and Hh genotypes were obtained from the mimetic form and the hh genotype from the non-mimetic form (Table 1). Both the H and h alleles were found in Taiwan, but not in Okinawa, where no mimetic females occur (Table 1).
HRM analysis using primers targeting the dsx sequences of P. memnon showed that the H and h alleles exhibited unique, non-overlapping melting curves, enabling differentiation of the three dsx genotypes (Fig. S1, Supplementary Information). The sensitivities of the delta Tm values and shape discriminations were 40% and 20%, respectively. HRM analysis of the 134 wild-caught individuals afforded genotyping data consistent with those obtained by direct sequencing in all but one sample (Table 1). This inconsistent sample was genotyped as Hh by direct sequencing but as HH by HRM analysis.
In Taiwan, the proportions of dsx genotypes among males and females did not significantly differ from Hardy-Weinberg equilibrium at all localities (Fisher’s exact probability test; P > 0.05). The estimated frequency of the H allele ranged from 0.36 to 0.55 in males (Table 1). The dsx allele frequencies in males did not significantly differ among nine localities (Fisher’s exact probability test; P = 0.93). The estimations based on females were unreliable because of the small sample sizes; the total number of females collected in Taiwan was only one-third the number of males (Table 1).
Discussion
We found that the mimetic and non-mimetic forms of P. memnon females corresponded exactly with genotypes defined by two distinct allelic sequences in the dsx gene, suggesting that the genetic basis of female-limited Batesian mimicry in P. memnon is similar to that in P. polytes. It is not surprising that the same gene (i.e. dsx) controls mimicry polymorphism in the two closely related species. However, these two Papilio species are not sister species (Fig. 2), and the allelic dsx sequences of P. memnon differ considerably from those of P. polytes. Interestingly, all of the non-synonymous substitutions between the mimetic and non-mimetic alleles in P. memnon differed from those in P. polytes. The phylogenetic tree of the four dsx alleles (Fig. 4) revealed that the mimetic (H) and non-mimetic (h) alleles of P. memnon diverged independently from those of P. polytes. Thus, different mutation-selection processes in the dsx gene region may have resulted in the same pattern of female-limited mimicry polymorphism in these species. In the phylogeny of Papilio, the clade containing P. memnon and P. polytes and three other species includes two species without any sexual differences in terms of mimicry (Fig. 2). Therefore, even if the mimicry polymorphism evolved from their common ancestor, the responsible locus (including dsx) may have been subject to considerable evolutionary changes among the derived species. Furthermore, in P. dardanus, which is distantly related to P. polytes and P. memnon, the female-limited mimicry polymorphism has been found to be controlled by the transcription factor gene engrailed13. Female-limited mimicry evolved repeatedly in the genus Papilio, which originated approximately 55–65 million years ago16, and different mechanisms of controlling female-limited Batesian mimicry polymorphism may have evolved in different lineages. Further genomic studies are needed to reveal the origin of this female-limited Batesian mimicry polymorphism and its relationship with dsx in Papilio butterflies.
Batesian mimicry polymorphism is thought to be maintained by negative frequency-dependent selection6,17,18,19. However, few studies of female-limited Batesian mimicry polymorphism have attempted to estimate the frequencies of model, mimetic and non-mimetic forms6. To the best of our knowledge, no study exists on temporal or spatial variations in the frequencies of mimetic/non-mimetic alleles in natural populations. Our genotyping method using HRM analysis allows quick and precise estimation of allele frequencies in natural populations of P. memnon and facilitates study of the adaptive dynamics of the mimicry polymorphism. A particular merit of our approach is that allele frequencies can be estimated in male populations. Males are collected more frequently than are females, and more importantly, it is assumed that they are not subjected to differential predation mortality with respect to the mimicry locus genotype. Indeed, in this study, far fewer females than males were collected, both in Okinawa and Taiwan (Table 1).
In Taiwan, where the females are polymorphic, the mimetic allele (H) frequency in males was 45% on average and did not differ significantly among localities. In addition, the genotype frequencies at different sites did not depart from Hardy-Weinberg equilibrium, suggesting that mating was random in terms of mimetic alleles in the previous generation, and that the mimetic genotype did not affect juvenile survival. Only the non-mimetic allele was present in Okinawa, although the P. memnon population of Japan may have originated from Taiwan or another Southeast Asian region where both forms occur, and unpalatable papilionids Pachliopta and Atrophaneura occur in Okinawa. The absence of the mimetic allele in Okinawa may be attributed to selection against the mimetic allele, a founder effect, or genetic drift. The ecological and evolutionary factors responsible for maintenance or loss of female-limited Batesian mimicry polymorphism in P. memnon must be examined in both the field and laboratory, and our method of discriminating the mimicry genotypes will be useful in such studies.
Methods
Sequencing dsx mRNA
Two mimetic females of P. memnon were collected at Hualien in eastern Taiwan (23°59′N, 121°32′E). A laboratory colony developed from eggs laid by these females was reared successively (at 25 °C under a 14:10 L:D photoperiod) over three generations. The mimetic phenotype (genotype HH or Hh) is dominant11. One F2 mimetic female (expected genotype Hh) was hand-paired with one F2 male (expected genotype hh). Of their offspring (F3), 12 male and 7 female pupae were fixed in RNAlater (Qiagen) on the seventh day of pupation and stored at −30 °C prior to RNA extraction. Total RNA was extracted from developing wing discs of these pupae using the RNeasy Mini Kit (Qiagen), following the manufacturer’s protocol. Two total RNA samples from one male and one female pupa were sent to Hokkaido System Science Co. Ltd., Sapporo, Japan, for library construction using the Illumina TruSeq method (paired-end, 101 cycles) and for sequencing using the Illumina Hiseq 2000. All raw reads from the two individuals were assembled de novo using Trinity version_r2014071720 running the default settings. Dsx sequences were searched among the contigs using BLAST + 2.2.3021 using the F1 isoform sequences of the dsx H and h alleles of P. polytes14 as query sequences.
After obtaining dsx RNA contigs from P. memnon, the dsx RNAs of 11 F3 males and 6 F3 females (expected genotypes, Hh or hh) were sequenced to identify allele-specific sequences of dsx H and h. Extracted total RNAs were reverse-transcribed into single-stranded cDNAs using a ReverTra Ace® qPCR RT Kit (Toyobo, Osaka, Japan). Based on the dsx mRNA contigs, we designed primers (forward, PmDsxf_F2: 5′-GCCGCCTGTGTGAACCTC-3′; reverse, PmDsxf_R8: 5′-AGTCTGTGACAGTTCTCCACCAAAGATT-3′) amplifying a 567-bp region of dsx. PCR amplification was performed in 8-μL volumes containing 5.72 μL ultrapure water, 0.64 μL 2.5 mM dNTP mix, 0.80 μL Ex-Taq buffer, 0.15 μL each primer (10 μM), 0.04 μL Ex-Taq DNA polymerase (Takara, Shiga, Japan) and 0.5 μL cDNA template. The PCR settings were (1) 94 °C for 3 min, (2) 30 cycles of 98 °C for 10 s and 68 °C for 60 s and (3) 72 °C for 7 min. The targeted region was GC-rich (70.6–71.0%), and the denaturation temperature in step (2) was thus set to a higher than normal temperature. The PCR products were sequenced directly on an ABI 3130xl sequencer (Applied Biosystems, Foster City, CA, USA) using the above primers and a BigDye Terminator Cycle Sequencing FS Ready Reaction Kit version 3.1 (Applied Biosystems). The nucleotide sequences were aligned using MEGA version 6.0622. Heterozygous sites within each individual were manually identified using 4peaks software23. The phylogenetic relationship among the 567-bp sequences of dsx H and h alleles in P. memnon and P. polytes was analysed by the maximum-likelihood method using MEGA version 6.0622. The substitution model used was GTR + G (general-time-reversible model with gamma distribution for rate heterogeneity); a bootstrap analysis with 1000 replications was performed.
Collection of wild butterflies and sequencing
In total 134 P. memnon individuals were collected from nine localities in Taiwan and three localities on the main island of Okinawa in June and July 2013 (Table 1). Both mimetic and non-mimetic females occur in Taiwan, but only non-mimetic females are found in Okinawa. The whole bodies of the collected butterflies were fixed in absolute ethanol after recording the sex and, if applicable, wing type. Total genomic DNA was extracted using a Genomic DNA Purification Kit (Promega, Madison, WI, USA). To genotype the dsx locus, we designed primers for PCR-amplification of a 185-bp region of exon 1 including a part of the open reading frame: forward (alias, PmDsx_Hh_F9), 5′-AACACGGTAGCGCGTCAGCCCGCCA-3′; reverse (alias, PmDsxf_R2), 5′-CACTTCTCGCAGGTGCAGT-3′. The settings for PCR using Ex-Taq were (1) 94 °C for 3 min (2) 30 cycles of 98 °C for 10 s, 66 °C for 30 s and 72 °C for 30 s and (3) 72 °C for 7 min. The PCR products were sequenced directly (using the same primers) on the ABI3130xl sequencer.
High-resolution melting analysis
HRM analysis using a real-time PCR platform is a rapid, economical, and sensitive genotyping tool for the study of natural populations24,25. For HRM analysis, we used the forward primer PmDsx_Hh_F9 described above and a newly designed reverse primer (alias: Pmemnon_HRM_R1), 5′-CGCAGTTGGGCGGCGCSCGCGGCACT-3′, based on the consensus sequences of the dsx H and h loci. These primers were used to amplify a 93-bp portion of exon 1. cDNAs from the 19 laboratory-reared butterflies and genomic DNA from the 156 field-collected butterflies were subjected to pre-amplification and HRM analysis using a LightCycler® 96 System (Roche) with a LightCycler® 480 High Resolution Melting Master (Roche) and the saturating dye ResoLight. Pre-amplification was performed in 10-μL volumes containing 0.3 μL ultrapure water, 5.0 μL 2 × master mix, 0.4 μL each primer (10 μM), 1.4 μL 25 mM MgCl2 and 2.5 μL template DNA. The PCR settings were (1) 95 °C for 10 min, (2) 45 cycles of 95 °C for 10 s, 65 °C for 15 s and 72 °C for 15 s, (3) 95 °C for 1 min and (4) 40 °C for 1 min. HRM analysis was performed from 65 °C to 97 °C, increasing at 0.03 °C/s with 25 readings per degree increase. LightCycler® 96 SW 1.1 software was used to conduct the data analysis. The sensitivities of the delta Tm values and shape discriminations were adjusted manually to enable reliable genotyping.
The proportions of dsx genotypes at each locality in Taiwan were evaluated in terms of Hardy-Weinberg equilibrium using the “HardyWeinberg” package26 in R27. The geographical variations among dsx allele frequencies in Taiwan were analysed using Fisher’s exact probability test.
Additional Information
How to cite this article: Komata, S. et al. Identification of doublesex alleles associated with the female-limited Batesian mimicry polymorphism in Papilio memnon. Sci. Rep. 6, 34782; doi: 10.1038/srep34782 (2016).
References
Bates, H. W. Contributions to an insect fauna of the Amazon Valley (Lepidoptera: Heliconidae). Trans. Linn. Soc. (Lond.) 23, 495–556 (1862).
Mallet, J. & Joron, M. Evolution of diversity in warning color and mimicry: polymorphisms, shifting balance, and speciation. Annu. Rev. Ecol. Syst. 30, 201–233 (1999).
Vane-Wright, R. I. On the definition of mimicry. Biol. J. Linn. Soc. 13, 1–6 (1980).
Wallace, A. R. On the phenomena of variation and geographical distribution as illustrated by the Papilionidae of the Malayan Region. Trans. Linn. Soc. (Lond.) 25, 1–71 (1865).
Ford, E. B. Ecological Genetics, 4th edn. (Chapman and Hall, London, 1975).
Kunte, K. Female-limited mimetic polymorphism: a review of theories and a critique of sexual selection as balancing selection. Anim. Behav. 78, 1029–1036 (2009).
Clarke, C. A. & Sheppard, P. M. Super-genes and mimicry. Heredity 14, 175–185 (1960).
Clarke, C. A. & Sheppard, P. M. Interactions between major genes and polygenes in the determination of the mimetic patterns of Papilio dardanus. Evolution 17, 404–413 (1963).
Clarke, C. A. & Sheppard, P. M. Further studies on the genetics of the mimetic butterfly Papilio memnon L. Phil. Trans. R. Soc. Lond. B 263, 35–70 (1971).
Clarke, C. A. & Sheppard, P. M. The genetics of the mimetic butterfly Papilio polytes L. Phil. Trans. R. Soc. Lond, B 263, 431–458 (1972).
Clarke, C. A., Sheppard, P. M. & Thornton, I. W. B. The genetics of the mimetic butterfly Papilio memnon L. Phil. Trans. R. Soc. Lond, B 254, 37–89 (1968).
Kunte, K. et al. doublesex is a mimicry supergene. Nature, 507, 229–232 (2014).
Timmermans, M. J. T. N. et al. Comparative genomics of the mimicry switch in Papilio dardanus. Proc. R Soc. B Biol. Sci. 281, 20140465–20140465 (2014).
Nishikawa, H. et al. A genetic mechanism for female-limited Batesian mimicry in Papilio butterfly. Nat. genet., 47, 405–409 (2015).
Yagi, T., Sasaki, G. & Takebe, H. Phylogeny of Japanese papilionid butterflies inferred from nucleotide sequences of the mitochondrial ND5 gene. J Mol. Evol. 48, 42–48 (1999).
Zakharov, E. V., Caterino, M. S. & Sperling, F. A. Molecular phylogeny, historical biogeography, and divergence time estimates for swallowtail butterflies of the genus Papilio (Lepidoptera: Papilionidae). Sys. Biol. 53, 193–215 (2004).
Barrett, J. A. The maintenance of non-mimetic forms in a dimorphic Batesian mimic species. Evolution, 30, 82–85 (1976).
Turner, J. R. G. Why male butterflies are non-mimetic: natural selection, sexual selection, group selection, modification and sieving. Biol. J. Linn. Soc. 10, 385–432 (1978).
Ohsaki, N. Preferential predation of female butterflies and the evolution of Batesian mimicry. Nature 378, 173–175 (1995).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Zhang, Z., Schwartz, S., Wagner, L. & Miller, W. A greedy algorithm for aligning DNA sequences. J. Comput. Biol. 7, 203–214 (2000).
Tamura, K. et al. MEGA5: Molecular Evolutionary Genetics Analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Griekspoor, A. & Groothuis, T. 4Peaks: a program that helps molecular biologists to visualize and edit their DNA sequence files. (2005) Available at: http://nucleobytes.com/4peaks/. (Accessed: 13th July 2015).
Smith, B. L., Lu, C. P. & Alvarado Bremer, J. R. High-resolution melting analysis (HRMA): a highly sensitive inexpensive genotyping alternative for population studies. Mol. Ecol. Res. 10, 193–196 (2010).
Kang, D. & Sim, C. Identification of Culex complex species using SNP markers based on high-resolution melting analysis. Mol. Ecol. Res. 13, 369–376 (2013).
Graffelman, J. Exploring Diallelic Genetic Markers: The HardyWeinberg Package. J. Stat. Softw. 64, 1–22 (2015).
R. Core Team. R: A language and environment for statistical computing. (2015) Available at: http://www.R-project.org/. (Accessed: 23th November 2015).
Kunte, K. The diversity and evolution of Batesian mimicry in Papilio swallowtail butterflies. Evolution 63, 2707–2716 (2009).
Acknowledgements
We thank T. Fujisawa for genomic analysis. Supported by JSPS KAKENHI (no. 15K14603) and Fujiwara Natural History Public Interest Incorporated Foundation. Research and collecting permits were obtained from local governments of Taiwan. Import and rearing of butterflies were conducted with permission from Plant Protection Station, the Ministry of Agriculture, Forestry and Fisheries of Japan.
Author information
Authors and Affiliations
Contributions
Conception of the study: S.K., C.P.-L. and T.S. Wrote the paper: S.K. and T.S. Experiment, field study and data analysis: S.K. Interpretation of data: T.I. and H.F.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Komata, S., Lin, CP., Iijima, T. et al. Identification of doublesex alleles associated with the female-limited Batesian mimicry polymorphism in Papilio memnon. Sci Rep 6, 34782 (2016). https://doi.org/10.1038/srep34782
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34782
This article is cited by
-
Rapid parallel adaptation despite gene flow in silent crickets
Nature Communications (2021)
-
A shared genetic basis of mimicry across swallowtail butterflies points to ancestral co-option of doublesex
Nature Communications (2020)
-
Phylogenetic analysis and embryonic expression of panarthropod Dmrt genes
Frontiers in Zoology (2019)
-
Temporal dynamics of the mimetic allele frequency at the doublesex locus, which controls polymorphic Batesian mimicry in Papilio memnon butterflies
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.