Abstract
Most mammalian species have heteromorphic sex chromosomes in males, except for a few enigmatic groups such as the mole voles Ellobius, which do not have the Y chromosome and Sry gene. The Ellobius (XX ♀♂ ) system of sex chromosomes has no analogues among other animals. The structure and meiotic behaviour of the two X chromosomes were investigated for males of the sibling species Ellobius talpinus and Ellobius tancrei. Their sex chromosomes, despite their identical G-structure, demonstrate short synaptic fragments and crossover-associated MLH1 foci in both telomeric regions only. The chromatin undergoes modifications in the meiotic sex chromosomes. SUMO-1 marks a small nucleolus-like body of the meiotic XX. ATR and ubiH2A are localized in the asynaptic area and the histone γH2AFX covers the entire XX bivalent. The distribution of some markers of chromatin inactivation differentiates sex chromosomes of mole voles from those of other mammals. Sex chromosomes of both studied species have identical recombination and meiotic inactivation patterns. In Ellobius, similar chromosome morphology masks the functional heteromorphism of the male sex chromosomes, which can be seen at meiosis.
) system of sex chromosomes has no analogues among other animals. The structure and meiotic behaviour of the two X chromosomes were investigated for males of the sibling species Ellobius talpinus and Ellobius tancrei. Their sex chromosomes, despite their identical G-structure, demonstrate short synaptic fragments and crossover-associated MLH1 foci in both telomeric regions only. The chromatin undergoes modifications in the meiotic sex chromosomes. SUMO-1 marks a small nucleolus-like body of the meiotic XX. ATR and ubiH2A are localized in the asynaptic area and the histone γH2AFX covers the entire XX bivalent. The distribution of some markers of chromatin inactivation differentiates sex chromosomes of mole voles from those of other mammals. Sex chromosomes of both studied species have identical recombination and meiotic inactivation patterns. In Ellobius, similar chromosome morphology masks the functional heteromorphism of the male sex chromosomes, which can be seen at meiosis.
Similar content being viewed by others
Introduction
The existence of two sexes and two types of gametes makes the exchange of genetic information possible, which is the main benefit of sexual reproduction. Meiosis is a key process in sexual reproduction; meiotic checkpoints purify unfit gametes and maintain the genetic integrity of the species.
The evolution of heteromorphic sex chromosomes started with the origin of the sex determination factor on the ancestor proto-Y chromosome1,2,3. In different mammalian groups, it was followed by numerous chromosomal rearrangements and the gradual loss of homologous regions, resulting in damage to the ability of such heteromorphic chromosome pairs to recombine3,4. Meiosis in the heterogametic sex demonstrates specific features, from complete synapsis to the lack of it.
In placental mammals, the gene SRY (Sex-determining Region Y) is a key factor that determines the formation of testes (the Testis Determining Factor, TDF) and is located on the Y chromosome5,6. In most placental mammals, the X and Y chromosomes contain short homologous regions (pseudoautosomal regions, PAR), which limit synapsis between them at the pachytene stage with the formation of short fragments of the synaptonemal complex (SC)7. There are no data for homologous regions of marsupial sex chromosomes8, but the possibility of a metatherian PAR is not excluded.
Sex chromosome systems in placental mammals that differ from the conventional XX/XY could be explained by translocations of autosomes and sex chromosomes or duplications of sex chromosomes. Usually, such atypical sex chromosomes are present in part of a population such as in the lemming Myopus schisticolor9, Mus minutoides10, and Acomys11. Nevertheless, in some species, such as in some howler monkeys, multiple sex chromosomes occur in all specimens12.
Some variants of duplications or nondisjunctions of sex chromosomes have been described for humans, for example XXY, XXXY, XXXXY, and XXYY13,14. Chromosomal changes cause different syndromes, including mental retardation and infertility. For example, de la Chapelle syndrome (46, XX, SRY-positive), which is classified as a testicular disorder of sex development, is diagnosed in 1 in 20,000 newborn boys15. In this syndrome, a fragment of the Y chromosome carrying the SRY gene is translocated to the X chromosome during paternal meiosis16. A similar case has been described for a tortoiseshell cat with 38, XX and the presence of the SRY gene due to Xp;Yp translocation17.
Occasionally, molecular studies have revealed the absence of the SRY gene in humans with male testicular disorder. Only 10% of the patients with XX are SRY-negative, and they usually demonstrate incomplete masculinization18. In nature, the existence of mammalian species with X chromosomes in both sexes and absence of the SRY gene in males seems incredible. Nonetheless, in at least two groups of rodents, that is the case: Tokudaia with X019 and Ellobius with X0 and XX in both sexes20. These males are fertile; moreover, although Tokudaia are a rare species due to their small range of distribution, Ellobius, especially E. talpinus and E. tancrei, have broad distributions and occur in large numbers.
Five species of mole voles Ellobius are very unusual mammals with three types of sex chromosomes21. E. fuscocapillus (2n = 36, fundamental number (NF) or number of chromosomal arms is 60 for female and 59 for male) has a classical system of sex chromosomes, XY♂/XX♀. E. lutescens (2n = 17, NF = 34) has one sex chromosome, X in males and females. X chromosomes have a different morphology in Ellobius. In the E. fuscocapillus and E. lutescens, Xs are submetacentrics with different numbers of G-bands21. In the E. fuscocapillus pachytene XY chromosomes formed the PAR, which is typical for mammalian sex bivalents. In E. lutescens, the X univalent thickened during prophase I, becoming curved and variously shaped22. Three sibling species—the northern mole vole E. talpinus (2n = 54, NF = 54), the eastern mole vole E. tancrei (2n = 54-30, NF = 56) and the alai mole vole E. alaicus (2n = 52, NF = 56)—are characterized by two large acrocentric X chromosomes in females and males, which are equal in length and isomorphic for G- and C-bands21,23. The mechanisms of sex determination in Ellobius species remain enigmatic. There are no published data on whole-genome Ellobius studies; the identity of X chromosomes in males was proven by Zoo-FISH analysis23. Moreover, our previous study of female and male meiosis revealed functional distinctions for their X chromosomes24. We found that SCs that form between XX chromosomes were indistinguishable from autosomal SCs in oocytes—i.e., XX chromosomes underwent complete synapsis over the entire length of the SC bivalent. Male XX bivalent typically formed for the male mammal sex body, which moved to the periphery of the nucleus. Short fragments of SC were formed at telomeric regions of the bivalents only, and a wide zone of asynapsis between X and X chromosomes occurs in the central part of the SC bivalent. In these early studies, we found an electron-dense argyrophilic nucleolus-like body (Nlb) associated with one of the axial elements of the XX bivalent22.
The study of meiosis in males with XX is not a trivial task. The formation of chiasmas in PAR is essential for the proper segregation of the X and Y chromosomes during anaphase at the first meiotic division in the spermatocytes of most mammals. The evolution of heteromorphic sex chromosomes was synchronized with the origin of so-called MSCI (meiotic sex chromosome inactivation). MSCI is a multipurpose process, during which the chromatin of asynaptic regions of sex chromosomes undergoes transcriptional silencing25,26,27. Autosomes undergo silencing as well—this process has been named MSUC (meiotic silencing of unsynapsed chromatin). Upon the completion of autosome synapsis in meiotic prophase I, chromatin inactivation is replaced by its activation. In the case of the absence of synapsis up to the middle pachytene stage, as in heterozygotes with nonreciprocal aberrations for example, such asynaptic chromosomal regions are associated with sex bivalents and activate pachytene arrest, leading to the selection of spermatocytes28. A multistep process of chromatin reorganization underlies MSCI. First, the tumour suppressor protein BRCA1 (breast cancer 1) accumulates at non-synaptic areas of sex chromosomes, which starts the process of phosphokinase ATR (ataxia telangiectasia- and RAD3-related) recruitment29. BRCA1 and ATR proteins participate in further post-translation chromatin modification, including the phosphorylation of H2AFX30, ubiquitination of H2A31, sumoylation32 and others.
Mole voles offer the opportunity to explore these processes in the unique male XX sex chromosome system. We studied the recombination and pachytene chromatin inactivation of the male XX chromosomes, and the enigmatic nucleolus-like body associated with the SC of the sex bivalent in males of E. talpinus and E. tancrei. We also analysed the problem of sex chromosome divergence within the genus Ellobius, whose evolution has remained mysterious for several decades.
Results
Behaviour, synapsis and morphology of mole vole sex chromosomes
We studied the synapsis features of the isomorphic XX chromosome (Fig. 1) by electron microscopy and immunocytochemistry in nuclei spreads of spermatocytes I.
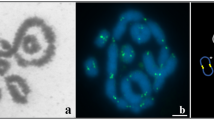
(A) G-band sex chromosomes of E. tancrei (ХХ). For more details concerning G-band E. tancrei/E. talpinus sex chromosomes, see Bakloushinskaya et al.23 (B) C-band sex chromosomes of E. talpinus. (C) C-band sex chromosomes of E. tancrei.
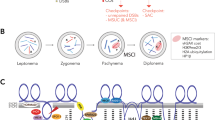
The sex bivalent was located among the autosomes at the zygotene stage. Later, the XX bivalent was gradually displaced to the periphery of the nucleus, forming an electron-dense sex body by the end of the mid pachytene stage as is typical for mammalian males. The XX sex chromosomes of E. talpinus and E. tancrei males underwent a partial synapsis between the short telomeric regions in meiotic prophase I. Thus, their XX sex chromosomes form a closed bivalent with a central asynaptic zone (Figs 2, 3A, 4A,D and S4A,E,I). At synaptic sites (Ss), SYCP1 (synaptonemal complex protein 1) immunostaining revealed a central element, indicating the formation of a typical SC (Fig. 3B,C).
The x-axis shows the average length of sex bivalents in μm; the y-axis shows the relative frequency of MLH1 foci in each 10% of the length of sex bivalent. The arrows point to the central position of MLH1 in Ss2. Axial elements of the XX chromosomes are green, the central element of the SC in areas of synapsis is pink, the centromere is red, and Nlbs are grey. The frequency of MLH1 is presented graphically as blue bars.
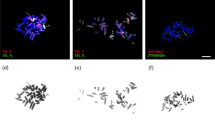
Bar = 10 μm. Localization of SYCP3 (green), SYCP1 (magenta), CREST (red), SUMO-1 (yellow), ubiH2A (cyan), RNA pol II (blue), γH2AFX (violet), MLH1 (grey) and DAPI (grey). (A–H) The same spermatocyte was stained with different antibodies. (I,J) Sex (XX) bivalent (from A–H). MLH1-signals were observed in telomeric synapsis, proteins of inactivation were localized at the asynaptic zone. SUMO-1 was located within the nucleolus-like body (Nlb) (arrow). Intense RNA pol II signal was detected in the main part of spermatocytes; within the sex bivalent, it was less intense. Co-localisation of SUMO-1, ubiH2A, RNA pol II, and γH2AFX is shown in graphs (a–d) (see I’ and J’).
Bar (A–C, H) = 5 μm; Bar (D–G, J–M) = 2 μm. Axial elements were identified using anti-SYCP3 antibodies (green), and anti-CREST for kinetochores (red). (A–C) Spermatocyte at the pachytene stage. The sex bivalent was displaced to the periphery of the nucleus. (D–G) The sex (XX) bivalent (from Figs A–C) has two regions of synapsis (white arrows) and a wide asynaptic area (asterisk) (see D). SUMO-1 (yellow) immunostained Nlb (yellow arrows) in the form of two large round spots (see E). γH2AFX (violet) surrounded the entire sex bivalent (F). DAPI-signal (grey) surrounded the sex bivalent (see C), with the more intense signal in the Nlb (see G). (H) The RNA pol II signal was intense in the main part of the spermatocyte. RNA pol II had a weak staining signal within the sex bivalent. (I) Co-localization of SUMO-1, γH2AFX, and DAPI is shown in graph (a,b) (see E–G). (J) The sex bivalent at the early pachytene stage was intensely γH2AFX stained (violet) (scheme J’). (K) The asynaptic area of the sex bivalent was intensely ATR stained at the mid pachytene stage (scheme K’). (L) The sex bivalent rolled up into a tangle at the late pachytene stage (scheme L’). Axial elements of the XX bivalent had SYCP3 gaps, which were marked with SUMO-1 (yellow). (M) The sex bivalent underwent reorganization at the early diplotene stage (scheme M’). At the same time, the gaps in the axial elements are interspersed with clusters of SYCP3 along the chromosomes, including the area near the centromere, and were surrounded by ubiH2A (cyan).
At the middle pachytene stage, axial elements twisted together and acquired a figure-eight configuration (Figs 3A and 4A). By the end of pachytene, the sex bivalent curled up into a tangle (Figs 4L, S1I and S2). SYCP3, the main protein of the axial and lateral elements of the SC, was gradually removed from the axial elements at early diplotene (Figs 4M and S2), wherein gaps in the axial elements alternated with SYCP3 clusters along the chromosomes and near the centromeres (Fig. 4M). At the late diplotene stage, some nuclei could be observed in which only the sex body was stained with anti-SYCP3 antibodies because autosomal axial elements no longer contained SYCP3.
The average length of E. talpinus sex bivalents was 11.6 ± 2 μm (range: 8.2 to 16.3 μm), while the average length of E. tancrei sex bivalents was 12.6 ± 2.5 μm (range: 8.3 to 17.7 μm). The length of the asynaptic regions of the XX chromosomes in E. tancrei and E. talpinus varied (range: 5.5 to 14.4 μm for E. talpinus and 5.2 to 16.4 μm for E. tancrei). The length of the synaptic regions also varied, in general becoming shorter towards the end of pachytene SC fragments (Fig. S1I).
Centromeres were identified by immunodetection with the CREST antibody. We have designated the pericentromeric SC region between XX chromosomes as Ss1, and the distal part of the SC as Ss2 in both species (Table 1, Figs 2, 3A and 4A). Centromeres were located at 0.2 and 0.3 μm from the telomere in E. talpinus and E. tancrei, respectively (Table 1, Fig. 2). A comparison of the SC fragment lengths (Ss1 and Ss2) revealed no significant differences between these fragments in E. tancrei and E. talpinus (P = 0.5 and P = 0.3, respectively). Thus, the length of the E. tancrei axial elements was greater than those of E. talpinus due to their longer asynaptic regions. It should be emphasized that the distal part of the SC (Ss2) was not detected by immunostaining with antibodies against SYCP3 in the sex bivalent structure at the pachytene stage in 24% of E. talpinus and 27% of E. tancrei nuclei; however in electron micrographs, using AgNO3 for contrast, this SC fragment was usually visible but disappeared later at the diplotene stage (Fig. S2).
In our previous electron microscopy study of sex bivalent structures we revealed one or two nucleolus-like bodies (Nlb) in E. tancrei and E. talpinus28 (Figs 2 and S2). The nature of the Nlbs has not been established. The nucleolar nature of these structures has been proposed based on their high argyrophility, high density and shape. Nlbs vary in size and shape, and the patterns of their structural organization are described below.
MLH1 in mole vole sex (XX) chromosomes structure
Antibodies against the mismatch repair protein MLH1 (mutL homolog 1) detected crossing over sites in 94 E. talpinus and 114 E. tancrei nuclei of pachytene spermatocytes. We could identify the MLH1 signals in XX bivalent structures for 46% of E. talpinus and 65% of E. tancrei nuclei (Table 1, Fig. 2). One or two MLH1 foci were detected at sex bivalents in both species. Simultaneously, one signal for each of the synaptic sites (Ss1 and Ss2) was detected at two E. talpinus and four E. tancrei sex bivalents (Fig. 3J).
The average distance between the telomere and MLH1 focus was 1.3 ± 0.7 in Ss1, and 0.93 ± 0.5 in Ss2 for E. talpinus nuclei; for E. tancrei nuclei, the average distances were 1.04 ± 0.1 in Ss1, and 0.96 ± 0.5 in Ss2 (Table 1, Fig. 2). The average distance between the MLH1 focus and telomeres was 74% of the length of the Ss2 for E. talpinus and 69% for E. tancrei (Fig. 2, highlighted by an arrow). Significant differences in the position of the late recombination nodules in Ss2 for these species were not found (P = 0.08).
However, in E. talpinus nuclei, in the short sections of Ss1 and Ss2 (up to 10% of the length of the zones) flanking the asynaptic zone, MLH1 foci were not identified or detected in only a few cases. Perhaps this difference in the distribution of late recombination nodules in the structure of the XX bivalent between two sibling species arose due to the lower number of identified MLH1 signals in the E. talpinus sex bivalents.
Chromatin remodeling in the mole voles male XX body
At the early pachytene stage, when sex bivalents have not yet become tangled, chromatin was grouped around the axis elements of the XX chromosomes (Fig. 4J). The histone γH2AFX was not detected at telomeric synaptic sites of sex bivalents (Ss1 and Ss2) (Table S1, Fig. 3G,J). In middle pachytene stage spermatocytes of both species, the distribution of histone γH2AFX in the structure of the sex body coincided with XX chromosome chromatin when counterstained with DAPI (Figs 5В and S1M–P). In E. tancrei nuclei, the ATR protein was detected in the pachytene chromatin, which surrounded the sex bivalent, with more intense fluorescence at the asynapsis zone (Fig. 4K and S1A–C).
A pachytene spermatocyte (A) and a sex (XX) bivalent (B) of a mole vole are shown. The sex (XX) chromosomes form a sex body on the periphery of the nucleus. The chromatin of the sex body undergoes reorganization. MSCI markers have different distributions: SUMO-1 (yellow), ATR (black dots), ubiH2A (blue), H2AFX (violet). SUMO-1 marks a small nucleolus-like body of the meiotic XX. ATR and ubiH2A are localized in the asynaptic area of the sex bivalent. γH2AFX covers the entire XX bivalent. The approximate number of autosomal SCs is shown. MLH1 signals are shown only for the sex chromosomes (black balls). The red balls indicate centromeres.
SUMO-1 appears in the structure of sex bivalents of mole voles closer to the middle pachytene. The SUMO-1 signal was detected intensively in the Nlb area of XX bivalents in both species (Figs 3E, 4E and S1E–H). When sex bivalents curled into a tangle at the late pachytene stage, SUMO-1 was often detected in areas of axial elements free of SYCP3 (Fig. 4L).
ubiH2A was detected in the XX asynaptic zone (Fig. 3F). If the sex bivalent rolled into a tangle—i.e., synaptic regions were close to each other, but not on opposite sides (in the post pachytene stages)—ubiH2A was detected within the entire XX bivalent (Figs 4M and S1I–L).
Intensity correlation analysis (ICA) and analysis of the fluorescence intensity profile (FIP) allowed the assessment the degree of MSCI protein co-localization (Figs 3I’-J’, 4I and S3). Thus, the degree of co-localization (m ± SD) is high for γH2AFX and ubiH2A (Pearson correlation coefficients: rp = 0.90 ± 0.10; Overlap correlation coefficients: r = 0.93 ± 0.06 for E. talpinus and rp = 0.82 ± 0.16, r = 0.90 ± 0.09 for E. tancrei; P = 0.45). Regarding the fluorescence-intensity profiles, the ubiH2A-signal path (Fig. 3I’–J’) was similar to the γH2AFX-signal path, but smaller in the width of coverage—as a rule, it covered only the asynaptic zone. γH2AFX has a wide distribution in the XX bivalent. The high degree of ubiH2A and γH2AFX co-localization within the sex body was noted previously31. γH2AFX/ATR and ubiH2A/ATR are highly colocalized (rp = 0.70 ± 0.19, r = 0.88 ± 0.06 and rp = 0.72 ± 0.08, r = 0.89 ± 0.05, respectively; only for E. tancrei) (Fig. S4). A lesser degree of co-localization was revealed for the γH2AFX/SUMO-1 (rp = 0.45 ± 0.10, r = 0.59 ± 0.11 for E. talpinus and rp = 0.48 ± 0.14, r = 0.64 ± 0.13 for E. tancrei; P = 0.17) and ubiH2A/SUMO-1 (rp = 0.63 ± 0.17, r = 0.69 ± 0.14 for E. talpinus and rp = 0.60 ± 0.13, r = 0.73 ± 0.17 for E. tancrei; P = 0.53) combinations (Figs S3 and S4). It should be emphasized that in the case of multiple immunostaining assays, the bright signal of the fluorescent label conjugated with antibodies to SUMO-1 and detected in Nlb did not disappear during photobleaching and thorough washing with PBS, although the “burning” of fluorescence was easily managed for other antibodies. Because anti-SUMO-1 antibodies were used earlier than ubiH2A and γH2AFX, we observed a characteristic peak in the paths of the ubiH2A-, γH2AFX-signal fluorescence-intensity profiles (Fig. 3I’). Thus, it was necessary to duplicate all experiments using one of the MSCI proteins (Fig. S1), eliminating the possibility of the manifestation of signals from earlier rounds of immunostaining.
RNA pol II in mole vole XX bodies
RNA pol II was either not detected in the chromatin of sex bivalents by immunostaining or its fluorescence intensity was significantly lower than that in autosomal chromatin for both species of mole voles (Figs 3H and 4H), a finding that is confirmed in the graph cd (Fig. 3J,J’). The intensity of the RNA pol II signal was analysed via the cd path, drawn from the main part of spermatocytes through the sex bivalent. The signal was less than half as intense in the area surrounding the XX bivalent than in other parts of the pachytene nuclei.
DAPI staining of mole vole meiotic sex chromosomes
The DAPI signal was identified in all mole vole spermatocytes. The most intensive DAPI staining was observed for the Nlb of sex bivalents in E. talpinus and E. tancrei (Figs 3D, 4C,G and S1H,L). When the axis of the sex bivalent formed a tangle at the late pachytene stage, the Nlb was sometimes absent (Fig. S1P).
Discussion
Analysing the distribution of crossover-associated MLH1 signal along mole vole male sex chromosomes, we found a high frequency of foci close to the asynaptic zone and a lower frequency of signals in telomeric areas. At a distance of 65–75% of the length of Ss1 (counting from the telomere-centromere), MLH1 signals were absent in sex bivalents of both species. A decrease or absence of recombination near centromeres has been previously described for different species: muntjac33, elephant shrew34,35, mouse, humans36,37 and others. It is assumed that the suppression of recombination in the centromere area (centromere effect) is due to the presence of centromeric heterochromatin38. Despite the small amount of C-heterochromatin localized in the sex chromosomes of the studied mole voles (Fig. 1B,C), it was still able to influence the distribution of MLH1. The centromeric effect is likely to be significantly linked to the lower rate of recombination in Ss1 than in Ss2 for both species. In Ss2, the recombination peak was identified outside of the telomeric zone. Apparently, that may be due to the mechanical containment of synaptic sites, as if the asynapsis area is severely restricted on both sides.
The low number of sex (XX) bivalents with MLH1 foci (46% E. talpinus, 65% E. tancrei) is likely due to two reasons. First, a heavy and compact protein envelope causing inactivation and covering the sex chromosomes at prophase I could prevent the penetration of anti-MLH1 antibodies. Second, SYCP3 could be removed from Ss2 early in the pachytene stage. In this case, the MLH1 could be detected at the sex bivalent for a short time. The number of sex bivalents with MLH1 varies not only among males of different species but also conspecific males. The percentage of males having sex bivalents with MLH1 was 17,47-45,8% in bulls, 20,56% in barbary sheep39,40, 29–30% in wildebeest and antelopes39,41, 60.6% in muntjacs33, 61–74% in different breeds of pigs42, 73% in humans43, and 80% in horses44. The recombination patterns of the sex chromosomes in both species of mole voles were similar and appeared to be different from those of other mammalian species.
The presence of two recombination peaks, one at each synaptic site of the sex chromosomes suggests that the two ancestral X chromosomes were structurally isomorphic. We hypothesize that later they underwent a secondary heteromorphization and became functionally different in the central region. The evolutionary assumption, based on ancestral male mole voles having the XY system, is that the loss of the Y chromosome was the first event, and doubling of the X chromosome occurred thereafter.
During MSCI, sex chromosomes are evicted to the periphery of the meiotic nucleus and form a so-called sex body25. A sex body is a morphological structure that marks the MSCI. It should be noted that the failure of sex body formation in spermatocytes might correlate with infertility in heterozygotes for chromosomal rearrangements45. Epigenetic chromatin modifications of sex bodies have been conducted using antibodies against proteins involved in the MSCI process.
ATR is involved in the formation of the sex body at prophase I29,46. Normally, ATR starts to be eliminated at middle zygotene upon the development of synapsis between the chromosomes and disappears later; at pachytene, ATR is only detected at asynaptic regions of the sex chromosomes47,48. In E. tancrei, ATR was localized in unsynapsed chromatin of the central part of the pachytene XX bivalent. We observed a similar situation in the spermatocytes of interspecific hybrids of E. tancrei and E. talpinus (unpublished data). Unfortunately, we have not analysed the distribution of ATR foci in E. talpinus, but we expect similar results.
Phosphorylation of H2AFX is an ATR-dependent process, and phosphorylated histone γH2AFX appears twice during prophase I. The first wave of phosphorylation comes at leptotene, when DSB development occurs, and the second wave starts at pachytene coincident with sex body formation30. The phosphorylation of H2AFX is considered to be an important event for the initiation of MSCI. Mouse null mutants of γH2AFX demonstrated deviations in sex body formation49. In mutants of SYCP1 and BRCA1, an aberrant pattern of γH2AFX and ATR localization was observed48,50. In mole vole spermatocytes, we observed γH2AFX in the sex body from early pachytene. At the transition from the early to the middle pachytene stages, γH2AFX was localized only at the XX bivalent asynaptic zone. At the late pachytene stage, γH2AFX appeared throughout the entire sex bivalent. Thus, it is possible that the chromatin in the synaptic sites inside the XX body was also modified.
SUMO (small ubiquitin-related modifier) group proteins are involved in various cellular processes, including DNA replication, recombination, repair and formation of heterochromatin-specific nuclear areas32. SUMO-1 plays an important role in SC and sex body formation, as well as in maintaining the structure of chromatin51,52. In mouse and rat XY bodies, SUMO-1 is located along the asynaptic sites of the sex chromosomes51,53. Therefore, we were surprised to discover that SUMO-1 localized within the Nlb at the XX bivalent. Maximum staining for SUMO-1 at the sex body of E. talpinus and E. tancrei occurred at the middle pachytene stage. During the transition from pachytene to diplotene, when SYCP3 protein is partially eliminated from the axial elements, it is possible to observe SUMO-1 localization. This was shown for E. tancrei only (Fig. 5L). Until now, the nature of the AgNO3-positive Nlb was not clear. Previously, we could not immunodetect nucleolar RNA-binding protein nucleophosmin/B23 in mole vole sex bodies54. In this study, we demonstrate that the Nlbs are SUMO-1- and DAPI-positive sites and seem to have a chromatin nature.
The mono-ubiquitinated histones H2A (ubiH2A) and H2B play an important role in nuclear processes, including transcription, DNA repair and the maintenance of chromatin structure55. In pachytene spermatocytes of mice, rats and humans, ubiH2A was localized within the asynaptic chromatin of the XY bivalent31,53. In mole voles, we observed a similar picture, and an intense signal of ubiH2A was localized in the asynaptic area of the XX bivalent. Assessing the co-localization of ubiH2A with γH2AFX, we showed that ubiH2A was distributed in a smaller area of the XX sex bivalent. As suggested by Baarends et al.31, ubiH2A is not responsible for the formation of silent chromatin but is only involved in maintaining it in its inactive form. Apparently, the same is true for the mole vole sex body.
RNA pol II, a marker of chromatin transcriptional activity, is highly concentrated in autosomal chromatin but much less concentrated in sex body chromatin48,56. There is also a link between low levels of RNA pol II and high levels of γH2AFX and ATR in mouse sex bivalent35. We observed a similar link between the RNA pol II and MSCI proteins in the mole vole sex bivalent.
The Nlb was clearly revealed in the structure of one of the XX bivalent axes early in the zygotene stage. As seen from the above results, asynchronous chromatin inactivation occurs later in asynaptic parts of the XX chromosomes. The XX chromosome isomorphism denotes their homology; additionally, we assume that asynapsis may be caused not by a lack of homology between isomorphic XX but by their early inactivation and specific heterochromatization of some regions of at least one of the X chromosomes. In other words, we assume that early asynchronous epigenetic chromatin changes lead to absence of synapsis between isomorphic XX chromosomes. Obviously, whole-genome sequencing will resolve this issue.
We propose a model of pachytene chromatin remodeling in the sex body for the two mole vole species by the identification of MSCI markers (Fig. 5). Obviously, the processes of MSCI, as well as the patterns of recombination, are similar in E. talpinus and E. tancrei. Thus, isomorphic G-band XX sex chromosomes of mole vole males behave normally in prophase I of meiosis. As in all male mammals, they formed areas of synapsis and asynapsis, shifted to the periphery of pachytene nuclei, and underwent MSCI.
The main characteristic of a Y chromosome is that it carries a testis-determining factor, such as Sry, in placental mammals. The Sry gene encodes a transcription factor that up-regulates the Sox9 gene via binding of the testis-specific enhancer (TESCO)57. Recent studies on Sox9 gene regulation have demonstrated unusual results common between four Ellobius species, with or without Y chromosome and Sry58. All of the studied species carry a 14-bp deletion (Δ14) in the highly conserved module of the Sox9 cis-regulatory element, TESCO. It is possible that the deletion Δ14 led to up-regulation of the Sox9 gene by increased activity of TESCO. It is known that XX transgenic mice over-expressing Sox9 develop as infertile males59, but XX males and females of E. talpinus and E. tancrei are fertile. Moreover, in the Sry-positive species E. fuscocapillus, a point mutation (A to G change) in the TESCO module ECRiii, might abolish Sry binding to its target site R558. In that case, the presence of the Sry gene in the E. fuscocapillus genome is redundant due to loss of its functional target binding site.
In similar model species of Tokudaia, TESCO has lost its enhancer functions for the Sox9 gene in XO species without the Sry gene and in XY species with multiple Sry gene copies60 caused by different mutations than in Ellobius, but with similar effects. Apparently, the loss of the Sry gene triggers an alternative, still unknown gene for sex determination. Such changes in enhancer structure could be crucial for genome stability and provoke the loss of the Sry gene and the Y chromosome in Ellobius and Tokudaia.
The manifestation of typical XY sex bivalent meiotic patterns in case of XX male sex bivalents in E. talpinus and E. tancrei may be a first step in the evolution from isomorphic sex chromosomes into heteromorphic ones. In Ellobius, with XX chromosomes in both sexes, chromosome morphology masked the functional heteromorphism of the sex chromosomes, which can be observed only at meiosis. Ellobius seems to be a natural model for the study of the evolution of sex chromosomes and molecular mechanisms of sex determination in mammals.
Materials and Methods
Animals
We studied two males of E. talpinus, and six males of E. tancrei: one 2n = 54, one 2n = 50 and four 2n = 34. We used animals from a long-term stock, which has been maintained at the Koltzov Institute of Developmental Biology; E. talpinus were originated from the Orenburg District, Russia, and all forms of E. tancrei originated from Pamiro-Alay, Tajikistan. The animals were maintained under comfortable conditions, enabling them to reproduce. All of the specimens were karyotyped. The animals were treated according to established international protocols, such as the Guidelines for Humane Endpoints for Animals Used in Biomedical Research. All of the experimental protocols were approved by the Ethics Committees for Animal Research of the Vavilov Institute of General Genetics and the Koltzov Institute of Developmental Biology in accordance with the Regulations for Laboratory Practice in Russian Federation.
Meiotic spread preparations
The suspension of spermatocytes was placed in a centrifuge tube and washed by centrifugation with 10 ml of Eagle’s medium, 2–3 times for 10 minutes at 1500 rpm. Thereafter, the tube with cells was placed on ice. Synaptonemal complex preparations were made and fixed using the technique previously described22,24.
Electron microscopy study of SCs
Slides for a subsequent electron-microscopic study were covered with Falcon plastic and stained with a 50% AgNO3 solution in a humid chamber for 3 h at 56 °C, then washed 4 times in distilled water and air dried. Stained slides were examined under a light microscope to select suitably spread cells. Once selected, Falcon plastic circles were cut out with a diamond knife, transferred onto grids, and examined at a JEM 100B electron microscope.
Antibody and immunostaining procedure
Poly-L-lysine-coated slides were used for all immunofluorescence studies. The slides were washed with phosphate-buffered saline (PBS) and incubated overnight at 4 °C with primary antibodies diluted in antibody dilution buffer (ADB: 3% bovine serum albumin—BSA, 0.05% Triton X-100 in PBS). The primary antibodies used were rabbit polyclonal anti-SYCP1 (1:500, Abcam, #15090), rabbit polyclonal anti-SYCP3 (1:500–1:1000, Abcam, #15093), mouse monoclonal anti-ATR (1:200, Abcam, #54793), human anticentromere antibody CREST (1:500, Fitzgerald, #90C), mouse monoclonal anti-MLH1 (1:50, Abcam, #59756), mouse monoclonal anti-SUMO-1 (1:250, Zymed, #33-2400), mouse monoclonal anti-ubiquityl histone H2A (1:400, Millipore, #05-678), mouse anti-RNA polymerase II (RNA pol II) (1:1000, Abcam, #5408) and mouse anti-phospho-histone H2AX (1:1000, Abcam, #22551). After washing, we used the corresponding secondary antibodies diluted in PBS: FITC-conjugated bovine anti-rabbit IgG (1:1000, Santa Cruz Biotechnology), goat anti-rabbit Alexa Fluore 488 (1:500, Invitrogen), FITC-conjugated horse anti-mouse IgG (1:500, Vector Lab.), Rodamine-conjugated chicken anti-rabbit IgG (1:400, Santa Cruz Biotechnology), goat anti-human Alexa Fluore 546 (1:500, Invitrogen), and goat anti-mouse Alexa Fluore 546 (1:200, 1:1000, Invitrogen). Immunostaining was carried out sequentially in several rounds. The first round included the simultaneous use of two primary antibodies, SYCP1+MLH1, and then washing in PBS. The slides were subsequently processed with the corresponding secondary antibodies, were washed in PBS, and then were mounted in Vectashield with 4′,6-diamidino-2-phenylindole (DAPI) (Vector Laboratories). The slides were examined using an Axioimager D1 microscope (Carl Zeiss, Jena, Germany) equipped with an Axiocam HRm CCD camera (Carl Zeiss) and image-processing AxioVision Release 4.6.3. software (Carl Zeiss). All of the preparations were counterstained with DAPI. Each cell was photographed three times to document the results of SYCP1, MLH1, DAPI staining. The coordinates of each photographed nucleus were recorded. Next, we “burned” off the fluorochromes that were used during the first round. The slides were kept for several minutes under a halogen lamp for full or partial elimination (“burning”) of fluorochrome fluorescence. The slides were then washed in PBS (5–7 times for 6–8 minutes each). We conducted the second round in the same manner as for the first round, this time using antibodies against SYCP3 + CREST. In the third round we used antibodies against ATR (or SUMO-1). In the fourth round we used antibodies against ubiH2A, in the fifth antibodies against RNA pol II were used. In the sixth round, we used antibodies against γH2AX (also known as γH2AFX). After each round, the slides were washed in PBS and mounted in Vectashield with DAPI (Vector Laboratories). The number of rounds varied. The use of multi-round immunostaining allowed us to identify up to eight antigens in the same nucleus.
Control
As a control for the co-localization of proteins and to avoid errors when using the same host antibodies, control staining was performed in parallel on other slides, which were treated with two or three antibodies (e.g., only antibodies against SYCP3-SUMO-1 or against SYCP3-ubiH2A or against SYCP3-γH2AFX).
Co-localization and intensity correlation analysis (ICA) or the «dye-overlay» method
An assessment of the co-localization of various proteins in the same part of the cell was made by overlaying photographs taken separately using different channels. ICA was performed according to Reitan et al.61. In the analysis of the co-localization of proteins involved in the process of MSCI, we considered several characteristics of the distribution of proteins, using scatter plots, Pearson correlation coefficients (rp), overlap correlation coefficients (r) and individual fluorescence intensities, which were obtained using the plug-Intensity Correlation analysis62, and ImageJ version 1.45o, available in the public domain63,64. ImageJ converts primary (raw) microscopic images in an 8-bit format and analyses the degree of co-localization. rp and r allow us to estimate the correlation between different fluorescence intensities. rp ranged from −1 (in the case of strict negative correlation) to +1 (strictly positive correlation). It is worth noting that various software plugins for co-localization can calculate differing values for rp65; thus it was important to use the same plugin for analogous assessments. In analysing scatter plots, overlaying green and red signal resulted in a yellow signal (see the scatter plots in Fig. S3). The more yellow in the scatter plot was, the higher the level of overlap. The width of the distribution of the yellow signals in scatter plots corresponded to the degree of co-localization of the compared fluorescence signals: the wider the distribution of the signal was, the higher the level of overlap of the two channels in the study area would be. P value was calculated for r (see the Results section).
Fluorescence-intensity profile (FIP)
To assess the degree of co-localization for several proteins along a given line, we made fluorescence-intensity profiles (FIPs) using the RGB profiler function in the ImageJ program according to previously published methods66,67.
Statistical analysis
All of the data are shown as the mean values ± SD. Student’s t-test was performed to determine significant differences in the data. All statistical analyses were conducted using GraphPad Prism Version 5.0 (GraphPad Software, CA, USA). P value < 0.05 was considered to be statistically significant.
Additional Information
How to cite this article: Matveevsky, S. et al. Unique sex chromosome systems in Ellobius: How do male XX chromosomes recombine and undergo pachytene chromatin inactivation? Sci. Rep. 6, 29949; doi: 10.1038/srep29949 (2016).
References
Ohno, S. Sex chromosomes and sex-linked genes. (Springer-Verlag, 1967).
Сharlesworth, D., Charlesworth, B. & Marais, G. Steps in the evolution of heteromorphic sex chromosomes. Heredity 95, 118–128 (2005).
Graves, J. A. M. Sex chromosome specialization and degeneration in mammals. Cell 124, 901–914 (2006).
Rice, W. R. Evolution of the Y sex chromosome in animals. Bioscience 46, 331–343 (1996).
Sinclair, A. H. et al. A gene from the human sex-determining region encodes a protein with homology to a conserved DNA-binding motif. Nature 346, 240–244 (1990).
Koopman, P., Gubbay, J., Vivian, N., Goodfellow, P. & Lovell-Badge, R. Male development of chromosomally female mice transgenic for Sry. Nature 351, 117–121 (1991).
Burgoyne, P. S. Genetic similarity and crossing over in the X and Y chromosomes of humans. Hum. Genet. 61, 85–90 (1982).
Patel, H. R., Delbridge, M. L. & Graves, J. A. M. Organization and evolution of the marsupial X chromosome. In Marsupial Genetics and Genomics (eds Deakin, J. E. et al. ) Ch. 8, 151–171 (Springer, 2010).
Fredga, K., Gropp, A., Winking, H. & Frank, F. Fertile XX- and XY-type females in the wood lemming Myopus schisticolor. Nature 261, 225–227 (1976).
Veyrunes, F. et al. A novel sex determination system in a close relative of the house mouse. Proc Biol Sci 277, 1049–1056 (2010).
Castiglia, R. & Annesi, F. Cytotaxonomic considerations on the sex chromosome variation observed within Acomys ngurui Verheyen et al. 2011 (Rodentia, Muridae). Zootaxa 3493, 35–38 (2012).
Steinberg, E. R., Nieves, M. & Mudry, M. D. Multiple sex chromosome systems in howler monkeys (Platyrrhini, Alouatta). Compar Cytogenet 8, 43–69 (2014).
Matzuk, M. M. & Lamb, D. J. The biology of infertility: research advances and clinical challenges. Nature medicine 14, 1197–1213 (2008).
Maiburg, M., Repping, S. & Giltay, J. The genetic origin of Klinefelter syndrome and its effect on spermatogenesis. Fertil Steril. 98, 253–260 (2012).
Vorona, E., Zitzmann, M., Gromoll, J., Schüring, A. N. & Nieschlag, E. Clinical, endocrinological, and epigenetic features of the 46, XX male syndrome, compared with 47, XXY Klinefelter patients. J Clin Endocrinol Metab. 92, 58–65 (2007).
Visootsak, J. & Graham J. M. Klinefelter syndrome and other sex chromosomal aneuploidies. Orphanet J Rare Dis. 1, 42 (2006).
Szczerbal, I., Stachowiak, M., Dzimira, S., Sliwa, K. & Switonski, M. The first case of 38, XX (SRY-positive) disorder of sex development in a cat. Molec Cytogenet. 8, 22 (2015).
Alves, C., Braid, Z., Coeli, F. B. & Mello, M. P. D. 46, XX male-testicular disorder of sexual differentiation (DSD): hormonal, molecular and cytogenetic studies. Arq Bras Endocrinol Metabol 54, 685–689 (2010).
Kuroiwa, A., Ishiguchi, Y., Yamada, F., Shintaro, A. & Matsuda, Y. The process of a Y-loss event in an XO/XO mammal, the Ryukyu spiny rat. Chromosoma 119, 519–526 (2010).
Just, W. et al. Ellobius lutescens: sex determination and sex chromosome. Sexual Devel. 1, 211–221 (2007).
Vorontsov, N. N., Lyapunova, E. A., Borissov, Y. M. & Dovgal, V. E. Variability of sex chromosomes in mammals. Genetica 52/53, 361–372 (1980).
Kolomiets, O. L., Vorontsov, N. N., Lyapunova, E. A. & Mazurova, T. F. Ultrastructure, meiotic behavior, and evolution of sex chromosomes of the genus Ellobius. Genetica 84, 179–189 (1991).
Bakloushinskaya, I.Yu. et al. A comparative analysis of the mole vole sibling species Ellobius tancrei and E. talpinus (Cricetidae, Rodentia) through chromosome painting and examination of synaptonemal complex structures in hybrids. Cytogenet Genome Res 136, 199–207 (2012).
Kolomiets, O. L., Matveevsky, S. N. & Bakloushinskaya, I. Y. Sexual dimorphism in prophase I of meiosis in the Northern mole vole (Ellobius talpinus Pallas, 1770) with isomorphic (XX) chromosomes in males and females. Compar Cytogenet 4, 55–66 (2010).
Solari, A. J. The behavior of the XY pair in mammals. Int Rev Cytol 38, 273–317 (1974).
Turner, J. M. et al. Meiotic sex chromosome inactivation in male mice with targeted disruptions of Xist. J. Cell Sci. 115, 4097–4105 (2002).
Turner, J. M. Meiotic silencing in mammals. Annu. Rev. Genet. 49, 395–412 (2015).
Jablonka, E. & Lamb, M. J. Meiotic pairing constraints and the activity of sex chromosomes. J Theor Biol 133, 23–36 (1988).
Moens, P. B. et al. The association of ATR protein with mouse meiotic chromosome cores. Chromosoma 108, 95–102 (1999).
Mahadevaiah, S. K. et al. Recombinational DNA double-strand breaks in mice precede synapsis. Nat Genet 27, 271–276 (2001).
Baarends, W. M. et al. Silencing of unpaired chromatin and histone H2A ubiquitination in mammalian meiosis. Mol Cell Biol 25, 1041–1053 (2005).
Gill, G. SUMO and ubiquitin in the nucleus: different functions, similar mechanisms? Genes Dev. 18, 2046–2059 (2004).
Yang, Q. et al. Synapsis and meiotic recombination in male Chinese muntjac (Muntiacus reevesi). PLoS One 6, e19255 (2011).
Waters, P. D., Ruiz-Herrera, A., Dobigny, G., Garcia Caldes, M. & Robinson, T. J. Sex chromosomes of basal placental mammals. Chromosoma 116, 511–518 (2007).
Segura, J. et al. Evolution of recombination in eutherian mammals: insights into mechanisms that affect recombination rates and crossover interference. Proc Biol Sci 280, 2013 1945 (2013).
Anderson, L. K., Reeves, A., Webb, L. M. & Ashley, T. Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1 protein. Genetics 151, 1569–1579 (1999).
Lian, J. et al. Variation in crossover interference levels on individual chromosomes from human males. Hum Mol Genet 17, 2583–2594 (2008).
Froenicke, L., Anderson, L. K., Wienberg, J. & Ashley, T. Male mouse recombination maps for each autosome identified by chromosome painting. Am J Hum Genet 71, 1353–1368 (2002).
Vozdova, M. et al. A comparative study of meiotic recombination in cattle (Bos taurus) and three wildebeest species (Connochaetes gnou, C. taurinus taurinus and C. t. albojubatus). Cytogenet Genome Res 140, 36–45 (2013).
Fröhlich, J. et al. Variation of meiotic recombination rates and MLH1 foci distribution in spermatocytes of cattle, sheep and goats. Cytogenet Genome Res 146, 211–221 (2015).
Vozdova, M. et al. Impact of Robertsonian translocation on meiosis and reproduction: an impala (Aepyceros melampus) model. J Appl Genet 55, 249–258 (2014).
Mary, N. et al. Meiotic recombination analyses of individual chromosomes in male domestic pigs (Sus scrofa domestica). PLoS One 9, e99123 (2014).
Sun, F. et al. Human male recombination maps for individual chromosomes. Am J Hum Genet 74, 521–531 (2004).
Al-Jaru, A., Goodwin, W., Skidmore, J. & Khazanehdari, K. Distribution of MLH1 foci in horse male synaptonemal complex. Cytogenet Genome Res 142, 87–94 (2014).
Homolka, D., Ivanek, R., Capkova, J., Jansa, P. & Forejt, J. Chromosomal rearrangement interferes with meiotic X chromosome inactivation. Genome Res 17, 1431–1437 (2007).
Keegan, K. S. et al. The Atr and Atm protein kinases associate with different sites along meiotically pairing chromosomes. Genes Dev. 10, 2423–2437 (1996).
Turner, J. M. et al. BRCA1, histone H2AX phosphorylation, and male meiotic sex chromosome inactivation. Curr Biol 14, 2135–2142 (2004).
de Vries, F. A. et al. Mouse Sycp1 functions in synaptonemal complex assembly, meiotic recombination, and XY body formation. Genes Dev 19, 1376–1389 (2005).
Fernandez-Capetillo, O. et al. H2AX is required for chromatin remodeling and inactivation of sex chromosomes in male mouse meiosis. Dev Cell 4, 497–508 (2003).
Turner, J. M. et al. Silencing of unsynapsed meiotic chromosomes in the mouse. Nat Genet 37, 41–47 (2005).
Rogers, R. S., Inselman, A., Handel, M. A. & Matunis, M. J. SUMO modified proteins localize to the XY body of pachytene spermatocytes. Chromosoma 113, 233–243 (2004).
La Salle, S., Sun, F., Zhang, X. D., Matunis, M. J. & Handel, M. A. Developmental control of sumoylation pathway proteins in mouse male germ cells. Dev Biol 321, 227–237 (2008).
Manterola, M. et al. A high incidence of meiotic silencing of unsynapsed chromatin is not associated with substantial pachytene loss in heterozygous male mice carrying multiple simple Robertsonian translocations. PLOS Genet 5, 1002009 (2009).
Matveevsky, S. N. Signs of sexual dimorphism in meiosis and karyotype variability of mole vole Ellobius (Rodentia, Mammalia). PhD thesis. NI Vavilov Institute of General Genetics of Russian Academy of Science (Moscow, 2011).
Cao, J. & Yan, Q. Histone ubiquitination and deubiquitination in transcription, DNA damage response, and cancer. Front Oncol 2, 26 (2012).
Richler, C. et al. Splicing components are excluded from the transcriptionally inactive XY body in male meiotic nuclei. Mol Biol Cell 5, 1341–1352 (1994).
Sekido, R. & Lovell-Badge, R. Sex determination involves synergistic action of SRY and SF1 on a specific Sox9 enhancer. Nature 453, 930–934 (2008).
Bagheri-Fam, S. et al. Sox9 gene regulation and the loss of the XY/XX sex-determining mechanism in the mole vole Ellobius lutescens . Chromosome research 20, 191–199 (2012).
Chaboissier, M. C. et al. Functional analysis of Sox8 and Sox9 during sex determination in the mouse. Development 131, 1891–1901 (2004).
Kimura, R., Murata, C., Kuroki, Y. & Kuroiwa, A. Mutations in the testis-specific enhancer of SOX9 in the SRY independent sex-determining mechanism in the genus Tokudaia. PLoS ONE 9, e108779 (2014).
Reitan, N. K., Sporsheim, B., Bjørkøy, A., Strand, S., Davies, C. & de L. Quantitative 3-D colocalization analysis as a tool to study the intracellular trafficking and dissociation of pDNA-chitosan polyplexes. J Biomed Opt 17, 026015 (2012).
Li, Q. et al. A syntaxin 1, Galpha(o), and N-type calcium channel complex at a presynaptic nerve terminal: analysis by quantitative immunocolocalization. J. Neurosci. 24, 4070–4081 (2004).
Rasband, W. S. ImageJ. National U. S. Institutes of Health, Bethesda, Maryland, USA. Available at: http://imagej.nih.gov/ij/ (1997–2016).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9, 671–675 (2012).
French, A. P., Mills, S., Swarup, R., Bennett, M. J. & Pridmore, T. P. Colocalization of fluorescent markers in confocal microscope images of plant cells. Nat Protoc 3, 619–628 (2008).
Barak, B. et al. Tomosyn expression pattern in the mouse hippocampus suggests both presynaptic and postsynaptic functions. Front Neuroanat 4, 149 (2010).
Fargue, S., Lewin, J., Rumsby, G. & Danpure, C. J. Four of the most common mutations in primary hyperoxaluria type 1 unmask the cryptic mitochondrial targeting sequence of alanine:glyoxylate aminotransferase encoded by the polymorphic minor allele. J Biol Chem 288, 2475–2484 (2013).
Acknowledgements
This work was partially supported by research grants of the Russian Foundation for Basic Research № 15-29-02649, 16-04-01447 and by the Presidium of the Russian Academy of Sciences Program “Biodiversity of natural systems”, subprogram “Gene pool of wildlife and conservation”. We thank our colleague Tatyana M. Grishaeva for the bioinformatics analysis of the proteins.
Author information
Authors and Affiliations
Contributions
S.M. prepared the meiotic chromosome samples and did the electron microscopic, immunocytological, statistic and micro photo analysis. I.B. provided and karyotyped animals. O.K. presented electron-microscopic photograph of SCs of two species of Ellobius that have been obtained and analyzed in previous studies. S.M., I.B. and O.K. wrote the main manuscript and the supporting information.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Matveevsky, S., Bakloushinskaya, I. & Kolomiets, O. Unique sex chromosome systems in Ellobius: How do male XX chromosomes recombine and undergo pachytene chromatin inactivation?. Sci Rep 6, 29949 (2016). https://doi.org/10.1038/srep29949
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep29949
This article is cited by
-
Sex differences in the meiotic behavior of an XX sex chromosome pair in males and females of the mole vole Ellobius tancrei: turning an X into a Y chromosome?
Chromosoma (2021)
-
Evolutionary rearrangements of X chromosomes in voles (Arvicolinae, Rodentia)
Scientific Reports (2020)
-
Sex chromosome quadrivalents in oocytes of the African pygmy mouse Mus minutoides that harbors non-conventional sex chromosomes
Chromosoma (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.