Abstract
The larval zebrafish (Danio rerio) is an excellent vertebrate model for in vivo imaging of biological phenomena at subcellular, cellular and systems levels. However, the optical accessibility of highly pigmented tissues, like the eyes, is limited even in this animal model. Typical strategies to improve the transparency of zebrafish larvae require the use of either highly toxic chemical compounds (e.g. 1-phenyl-2-thiourea, PTU) or pigmentation mutant strains (e.g. casper mutant). To date none of these strategies produce normally behaving larvae that are transparent in both the body and the eyes. Here we present crystal, an optically clear zebrafish mutant obtained by combining different viable mutations affecting skin pigmentation. Compared to the previously described combinatorial mutant casper, the crystal mutant lacks pigmentation also in the retinal pigment epithelium, therefore enabling optical access to the eyes. Unlike PTU-treated animals, crystal larvae are able to perform visually guided behaviours, such as the optomotor response, as efficiently as wild type larvae. To validate the in vivo application of crystal larvae, we performed whole-brain light-sheet imaging and two-photon calcium imaging of neural activity in the retina. In conclusion, this novel combinatorial pigmentation mutant represents an ideal vertebrate tool for completely unobstructed structural and functional in vivo investigations of biological processes, particularly when imaging tissues inside or between the eyes.
Similar content being viewed by others
Introduction
In order to understand complex biological phenomena, structural and functional information has to be extracted from intact animal systems at different spatial scales. Optical transparency of animals is a highly desirable feature to study these biological processes in vivo. Pigment molecules, such as melanin, haemoglobin and myoglobin, are the main sources of visible light absorption in biological tissues1,2. Lipids and collagen, on the other hand, constitute the primary molecules responsible for light scattering. Recently, several tissue clearing techniques have been developed to reduce light scattering in fixed biological tissues by selectively removing lipids in a non-destructive way3. However, the only molecules that can be removed from living systems without dramatically impairing their viability are pigment molecules localised in defined tissues, such as melanin present in the skin. Here, we used a combinatorial genetic approach to generate a viable, fully transparent zebrafish mutant, which we name crystal, lacking the vast majority of skin pigmentation. Compared to the previously described combinatorial pigmentation mutant casper that has pigmented eyes4, the crystal mutant constitutes a significant improvement for in vivo imaging of tissues inside or between the eyes.
In zebrafish, multiple populations of cells produce pigments that ultimately restrict the optical accessibility of tissues5,6. The three main kinds of pigment cells (or chromatophores) are the melanophores (black appearance), iridophores (silvery or blue) and xanthophores (yellow) (Fig. 1a). In addition to these three populations, which in zebrafish larvae derive from neural crest cells, there is another population of pigment cells forming the retinal pigment epithelium (RPE), which originates from optic lobe neuroepithelial cells (Fig. 1a)7. Common strategies to reduce zebrafish skin pigmentation can be grouped in two categories: 1) use of chemical compounds interfering with the synthesis of melanin, the most popular one being 1-phenyl-2-thiourea (PTU)8; 2) use of pigmentation mutants in which genes controlling either pigment formation, chromatophore formation or interactions between chromatophores have been inactivated4,6,9. The first strategy, despite being straightforward (i.e., embryos can be simply raised in medium containing 200 μM PTU), is associated with toxic side effects that impair morphogenesis, behaviour and survival. For example, PTU has been shown to interfere not only with tyrosinase (the enzyme that catalyses the production of melanin)10,11, but also with other enzymes, such as thyroid peroxidase12 and dopamine beta-hydroxylase13. Therefore, the poor selectivity of this drug results in severe consequences including reduced thyroid hormone synthesis14, decreased eye size12, abnormal cranial neural crest and extraocular muscle development15, impaired retinal light adaptation16, as well as anxiety17 and reduced mobility, hatching and survival8 (see also Results and Discussion). In contrast, the second strategy is considerably less disruptive since it takes advantage of viable mutations affecting the function of genes expressed in specific subsets of cells where they are involved in defined processes of pigment production5. Since the formation of each pigment type is controlled independently of the others, the combination of different mutations is required to produce fully transparent zebrafish. This strategy has been previously implemented to generate the double mutant casper, which lacks all melanophores and iridophores4. Here, we further develop this strategy to generate a fully optically clear combinatorial mutant (crystal) that not only lacks all melanophores and iridophores, but also has a non-pigmented RPE. This particular feature makes crystal larvae especially suited for imaging tissues inside or between the eyes while avoiding the use of chemical pigmentation blockers. Importantly, crystal mutants show no behavioural or viability deficits as compared to wild type animals. We also validate the in vivo application of this novel mutant by performing whole-brain light-sheet imaging and two-photon functional recordings of neural activity in the retinae of crystal larvae.
(a) Left: schematic diagram of the main populations of pigment cells in wild type (WT) zebrafish. Centre: cell lineages generating the different populations of pigment cells. Right: mutations affecting genes controlling either pigment cell formation (nacrew2/w2 and roya9/a9) or melanin production (albb4/b4) used to generate the crystal mutant. RPE, retinal pigment epithelium. (b–d) Pigmentation phenotypes of wild type, single mutant, casper and crystal zebrafish at adult (>3 month old, (b)), embryonic (3 dpf, (c)) and larval (7 dpf, (d)) stages. Red dashed boxes indicate crystal mutants. Insets on the right display eye pigmentation phenotypes. 3 dpf and 7 dpf zebrafish treated with 200 μM PTU are shown at the bottom of (c,d). Note that the optical transparency of crystal fish is higher than that of wild type, single mutant, casper and PTU-treated fish.
Results and Discussion
Generation of crystal: a fully transparent zebrafish for in vivo imaging
To generate optically clear zebrafish lacking the vast majority of skin pigmentation without using chemical compounds inhibiting pigment formation, we undertook a combinatorial genetic approach. Three previously described viable mutations affecting different populations of chromatophores (Fig. 1a) were selected and combined through crossbreeding (Fig. 1b): 1) the nacrew2/w2 mutant lacks all melanophores due to a mutation in the mitfa gene9. Since the nacre mutation does not affect the population of cells forming the RPE, this structure is still pigmented in this single mutant. 2) The albino (albb4/b4) mutant is characterised by a general deficiency in the production of melanin due to a mutation in the slc45a2 gene18. 3) The roy orbison (roya9/a9) mutant shows a complete lack of iridophores4. The gene responsible for this mutant phenotype is currently unknown. The resulting combinatorial triple mutant (nacrew2/w2;albb4/b4;roya9/a9; red boxes in Fig. 1b–d), which we name crystal, lacks the vast majority of dark and reflective pigments normally present at the cutaneous level and, as a consequence, appears optically clear. The internal organs are clearly visible in adult (>3 months old) crystal mutants as opposed to wild type and single mutant fish (Fig. 1b). Compared to the previously described double mutant casper (nacrew2/w2;roya9/a9)4, where internal organs are also visible, the triple mutant crystal (nacrew2/w2;albb4/b4;roya9/a9) lacks melanin in the RPE (Fig. 1b). Consequently, the eyes of crystal fish are considerably less pigmented than wild type, single mutant and casper fish, therefore resulting easily accessible to optical investigation (see right insets in Fig. 1b–d). The pigmentation phenotype of crystal fish is already evident at embryonic (e.g., 3 dpf, Fig. 1c) and larval (e.g., 7 dpf, Fig. 1d) stages. Importantly, the optical clarity of crystal larvae is even superior to that of larvae treated with 200 μM PTU (Fig. 1c,d), since PTU inhibits melanin production but does not interfere with iridophore function8. Moreover, unlike PTU-treated animals, adult crystal mutants are viable and produce normal numbers of offspring. Heterozygous fish (nacrew2/+;albb4/+;roya9/+) do not exhibit any visible phenotype (data not shown). Overall, the crystal mutant constitutes a significant improvement in the optical accessibility of both larval and adult zebrafish, even superior to the previously described combinatorial mutant casper4, which is characterised by black-pigmented eyes.
Crystal zebrafish larvae exhibit normal visual behaviour
An ideal system for in vivo imaging has to be characterised not only by optical clarity but also by normal functional and behavioural viability. The use of PTU to quickly obtain transparent zebrafish larvae is widespread across the research community. However, numerous studies reported severe morphological and behavioural side effects caused by the toxicity of PTU treatment (see Introduction). We thus wanted to compare the behavioural viability of crystal larvae to wild type larvae and to larvae raised in medium containing 200 μM PTU. As a representative visually guided behaviour we tested the ability of 5 dpf larvae to perform the optomotor response. During optomotor response assays, freely swimming zebrafish larvae respond to whole-field moving stimuli (e.g., dark and light bars) by swimming in the same direction of stimulus motion (Fig. 2a)16. They do so to compensate for the optic flow-induced perception of apparent involuntary displacement, and therefore regain the desired course of locomotion. Individual larvae were tested five times in total and scored according to the number of trials they responded to (i.e., fish turns and swims in the direction of the moving stimulus). Notably, crystal larvae exhibited a response rate equivalent to wild type larvae as opposed to PTU-treated larvae, which instead showed a dramatic behavioural impairment (Fig. 2b).
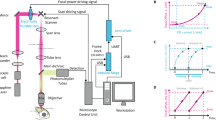
(a) Schematic diagram illustrating the optomotor response behavioural assay. Individual 5 dpf larvae were positioned in a petri dish containing Danieau solution. An LCD screen controlled by a computer was used to display black and white square-wave gratings moving in 4 directions (red arrows) at the bottom of the petri dish. Each larva was tested 5 times in total (each trial lasted 6 seconds) and scored according to the trials it responded to (i.e., fish turns and swims in the direction of the moving gratings). (b) Quantification of the optomotor response assays for 5 dpf wild type (WT, black), crystal (blue) and PTU-treated (red) larvae (n = 25 larvae in each group). The frequency distribution (left), cumulative frequency distribution (centre) and mean ± SEM (right) of number of responsive trials per larva are reported. Note that PTU-treated larvae show a significant decrease in the number of trials they respond to. ns, non-significant; ***p < 0.001; one-way ANOVA with post-hoc Tukey’s HSD test. (c) Functional calcium imaging of tectal cells and retinal ganglion cell axons expressing GCaMP5G (green) in 4 dpf Tg(elavl3:GCaMP5G) larvae. Distance of right eye from projection screen is 3 cm. Recordings are performed from 3 Z-planes (approximately 20 μm total volume thickness) in the contralateral optic tectum at 4 Hz image acquisition rate. Dashed box shows the angles of moving bars relative to zebrafish larva orientation. Mean ΔF/F0 images of calcium recordings in untreated (control) and PTU-treated larvae followed by mapping of direction-selective (DS) and orientation-selective (OS) voxels are displayed. Np, neuropil; SPV, stratum periventriculare; DSI, direction selectivity index; OSI, orientation selectivity index. Scale bar is 40 μm. (d) Average number (top) and relative frequency (bottom) of DS, OS, visually responsive and non-DS/non-OS voxels per Z-plane in control and PTU-treated 4 dpf larvae (n = 12 larvae in each group). Criteria used to identify DS and OS voxels are reported at the top. Note the dramatic reduction in visually responsive, DS and OS voxels following 200 μM PTU treatment. Error bars are ± SEM. ***p < 0.001, unpaired two-tailed Student’s t test.
Given the observed behavioural consequences of PTU treatment, we further investigated whether PTU has any effect on zebrafish visual system function. We did so by recording visually induced neural responses in the optic tectum, which in zebrafish is the main retinorecipient brain area. The zebrafish optic tectum receives inputs from all functional types of retinal ganglion cells19, the sole output neurons of the retina, including ganglion cells tuned to stimulus direction of motion (direction-selective cells) or stimulus orientation (orientation-selective cells)20. Using a transgenic fish line where the genetically encoded calcium indicator GCaMP5G is expressed pan-neuronally21, we analysed visual responses to moving dark and light bars in both the tectal neuropil and periventricular neurons through in vivo calcium imaging, as previously described (Fig. 2c and Supplementary Video 1)22,23. Compared to untreated Tg(elavl3:GCaMP5G) larvae, 4 dpf PTU-treated larvae (treatment from 1 dpf to 3 dpf) showed a large decrease in visual responses with direction- and orientation-selective neural responses being absent or severely impaired, respectively (Fig. 2c). From these data, we conclude that PTU can cause deleterious consequences on both zebrafish behaviour and neural function, and should be therefore avoided whenever these two biological processes are under investigation. Since PTU is a highly non-selective drug (see Introduction) and is generally applied at the whole-animal level, it is difficult to attribute any of the detrimental effects we observed to a specific biochemical pathway.
Whole-brain light-sheet imaging of crystal larvae
We next aimed to validate the in vivo application of crystal larvae by performing whole-brain light-sheet imaging24. Light-sheet microscopy has recently experienced a series of significant technological advancements and is generally used to study nervous system activity and development with cellular resolution at the whole-animal level21,25,26,27. In typical zebrafish light-sheet preparations, the excitation light is provided laterally and fluorophore emission light is collected through an objective positioned orthogonally to the illumination plane. However, due to the strong pigmentation of the eyes, imaging of areas inside or between these structures is normally problematic. We therefore compared the optical accessibility of the nervous system between crystal, nacre and casper larvae at 4 dpf using the pan-neuronal Tg(elavl3:GCaMP6f) line28 (Fig. 3a and Supplementary Video 2). In our setup, the sheet of laser light (488 nm) is generated by two objectives positioned on the lateral sides of the larva. In crystal larvae, the excitation light can easily reach deep (i.e., ventral) regions of the brain (Fig. 3b) as well as the retinae (Fig. 3c) without being absorbed or reflected by pigments normally present on the surface of the eyes, like in nacre or casper larvae (note the dark regions in the left and middle panels of Fig. 3b,c). Therefore, crystal allows a fully unobstructed optical access of the larval zebrafish brain in its entirety (i.e., including the eyes), as opposed to nacre and casper mutants where a substantial portion of the nervous system (~18%) is not accessible through standard light-sheet imaging (Fig. 3a,d,e; mean volume ± SEM, crystal 3.04 ± 0.12 × 107 μm3, n = 4; casper 2.51 ± 0.07 × 107 μm3, n = 4; nacre 2.47 ± 0.10 × 107 μm3, n = 5; p = 0.0040, F2,10 = 10.09, one-way ANOVA with post-hoc Tukey’s HSD test). Even though the optical accessibility of crystal larvae through light-sheet imaging is significantly higher than nacre and casper mutants, it is noteworthy that the spatial definition of brain regions between the eyes appears lower than other regions of the brain, likely due to scattering of excitation light by the lenses. Despite this fact, the light-sheet signal detected from these regions in crystal mutants is still many-fold higher than in the other animal groups (Fig. 3a,b).
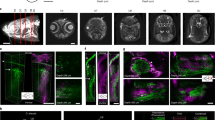
(a) Volumetric imaging of the larval zebrafish brain with light-sheet microscopy in 4 dpf nacre (top), casper (middle) and crystal (bottom) Tg(elavl3:GCaMP6f) larvae (n = 4–5 larvae in each group). Six different volume sections per larva out of 450 (total xyz volume of 798 × 623 × 283 μm3) are displayed. L, left; A, anterior. Scale bar is 200 μm. (b) Single volume sections of the brain (100 μm Z plane depth) in nacre (left), casper (middle) and crystal (right) larvae. Note the dark region between the eyes of nacre and casper larvae due to the excitation light being absorbed or reflected by pigments present on the surface of the eyes. L, left; A, anterior. Scale bar is 100 μm. (c) Insets showing the labelling of amacrine and ganglion cells in the left retina of the crystal larva (right) compared to nacre (left) and casper (middle) larvae, where no GCaMP6f fluorescence is detected in the eyes. T, temporal; N, nasal. Scale bar is 50 μm. (d) 3D reconstructions of the brain (lateral view) of nacre (left), casper (middle) and crystal (right) Tg(elavl3:GCaMP6f) larvae shown in (a). Note the improved optical accessibility (~18% of brain volume) allowed by crystal. P, posterior; V, ventral. Scale bar is 100 μm. (e) Average imaged brain volume in nacre, casper and crystal 4 dpf Tg(elavl3:GCaMP6f) larvae. Error bars are ± SEM. ns, non-significant; *p < 0.05; **p < 0.01; one-way ANOVA with post-hoc Tukey’s HSD test.
Two-photon calcium imaging in the intact retina using crystal
To further assess the in vivo application of crystal larvae we performed two-photon functional imaging of neural activity in the retina (Fig. 4a), a brain region that is not optically accessible in wild type, single pigmentation mutant or casper larvae (Fig. 1) unless embryos are raised in medium containing PTU29,30. The retina is a sensory neural circuit formed by multiple classes of excitatory cells (photoreceptors, bipolar and ganglion cells) and inhibitory cells (horizontal and amacrine cells)31. Its primary function is to detect and process light stimuli and, subsequently, send the processed visual information to higher brain areas through different types of functionally specialised ganglion cells32. To record visual responses in the larval retina through calcium imaging, 4 dpf Tg(elavl3:GCaMP6f) crystal larvae were immobilised in 2% low melting point agarose with one eye facing an LCD screen where square-wave gratings moving in 12 different directions were displayed (n = 8 larvae). Visually evoked calcium transients were recorded from amacrine and ganglion cells at 4 Hz using near-infrared (930 nm) two-photon laser excitation (Supplementary Video 3). Voxel-wise analysis was then used to identify visually responsive voxels (Fig. 4b) and quantify direction and orientation selectivity of visual responses at subcellular resolution (0.248 × 0.248 μm voxel XY size, Fig. 4c) as previously described22,33. ΔF/F0 calcium traces from 6 selected regions of interest (ROIs, Fig. 4b) are displayed as examples (Fig. 4d). Interestingly, not only we could record stimulus-locked responses, but also we observed the presence of direction-selective (direction selectivity index (DSI) > 0.5, ROI# 4 and 5) and orientation-selective (orientation selectivity index (OSI) > 0.5, ROI# 1–3) responses (Fig. 4c,d), indicating that the retina is functional in crystal larvae. A previous study has reported that adult zebrafish mutants with hypopigmented eyes show deficits in performing visual escape assays under defined luminance conditions34. Thus, even though our visual behaviour assays demonstrated that crystal larvae perform the optomotor response as well as wild type larvae (Fig. 2a,b), we cannot exclude that crystal mutants might show abnormal retinal responses under certain luminance conditions. However, given the deleterious effects caused by PTU treatment (Fig. 2c,d; see Introduction), the crystal mutant represents, to the best of our knowledge, the most viable strategy to perform in vivo functional imaging in the intact retina of larval zebrafish.
(a) Two-photon functional calcium imaging of amacrine cells and ganglion cell dendrites expressing GCaMP6f (green) in 4 dpf crystal Tg(elavl3:GCaMP6f) larvae (n = 8 larvae). Distance of the eye from LCD screen is 2 cm. Recordings are performed from 2–4 Z-planes (approximately 20 μm total volume thickness) at 4 Hz image acquisition rate. Dashed box shows the angles of moving gratings relative to zebrafish larva orientation. INL, inner nuclear layer; GCL, ganglion cell layer; IPL, inner plexiform layer. Scale bar is 20 μm. (b,c) Mean ΔF/F0 image of a representative calcium recording (b) followed by voxel-wise analysis of direction and orientation selectivity of visual responses (c). DSI, direction selectivity index; OSI, orientation selectivity index. (d) ΔF/F0 calcium traces during a representative tuning experiment from the 6 selected regions of interest (ROIs) shown in (b). Polar plots showing the tuning profiles (i.e., integral ΔF/F0 responses to different stimuli) of individual ROIs are reported on the right. Stimulus epochs are shown in grey. Dark arrows indicate the different directions of gratings motion. The blank-screen null condition is indicated by a ‘-’ sign.
Conclusions
In this study, we generated a viable, optically transparent combinatorial pigmentation mutant zebrafish, named crystal, which constitutes an ideal tool for completely unobstructed in vivo imaging of biological phenomena. More specifically, compared to casper mutants4, crystal fish are superior in terms of optical transparency when imaging inside or between the eyes. We validated the viability and in vivo application of crystal larvae through the optomotor response assay, whole-brain light-sheet imaging and two-photon functional imaging of neural activity in the intact retina. Importantly, crystal larvae show a higher viability than larvae treated with the chemical pigmentation blocker PTU. We envisage that crystal will be an invaluable tool for other in vivo applications, such as one-photon21 or two-photon27 volumetric calcium imaging of neural activity across the entire brain in semi-restrained behaving35 or paralysed fictively swimming36 zebrafish larvae, as well as to study the wiring, function and plasticity of neural circuits in normally highly pigmented, optically inaccessible structures like the eyes30,37,38.
Methods
Animals
Zebrafish were maintained at 28.5 °C on a 14 hr ON/10 hr OFF light cycle in Danieau solution [58 mM NaCl, 0.7 mM KCl, 0.4 mM MgSO4, 0.6 mM Ca(NO3)2 5.0 mM HEPES, pH 7.6]. The combinatorial crystal mutant (albb4/b4;nacrew2/w2;roya9/a9) was generated by sequentially crossbreeding three different single mutant strains, namely nacrew2/w2 (ref. 9), roya9/a9 (ref. 4) and albb4/b4 (ref. 18) mutants. Importantly, both larvae and adult crystal zebrafish are viable and do not display visible morphological, functional or behavioural abnormalities other than the pigmentation phenotype. The casper mutant was obtained by crossing nacrew2/w2 and roya9/a9 mutants, as previously described4. The AB line was used to obtain wild type zebrafish. Transgenic lines used in this study include Tg(elavl3:GCaMP5G)21 and Tg(elavl3:GCaMP6f)28. Confocal functional imaging experiments were performed in the nacre mutant. Light-sheet imaging experiments were performed in nacre, casper and crystal mutants. Two-photon functional imaging experiments were performed in crystal larvae. To treat zebrafish larvae with PTU we followed standard procedures8, specifically larvae were raised in 200 μM PTU (Sigma) in Danieau solution from 24 hours post fertilisation (hpf). Tg(elavl3:GCaMP5G) larvae used for confocal functional imaging of visually evoked neural activity in the optic tectum were treated with 200 μM PTU from 24 hpf to 3 dpf and imaged at 4 dpf. This work was approved by the local Animal Welfare and Ethical Review Body (King’s College London), and was carried out in accordance with the Animals (Scientific Procedures) Act 1986, under license from the United Kingdom Home Office (PPL70/8057).
Imaging
Whole-animal in vivo microscopy
Whole-animal images of adult zebrafish were taken with a Nikon D7000 digital SLR camera equipped with a Sigma 150 mm macro lens. Adult zebrafish were anesthetized with 0.2% tricaine (MS222, Sigma) in fish facility water and placed in a 90 mm petri dish containing fish facility water. Imaging of larvae was performed using a ZEISS Axioskop microscope connected to EXi Blue CCD cameras (Retiga) and Volocity acquisition software (PerkinElmer). Larval zebrafish were anesthetized with 0.02% tricaine in Danieau solution and immobilized in 1% low melting point agarose (Sigma) on glass slides.
Confocal calcium imaging
Imaging was performed using a ZEISS LSM 710 confocal microscope equipped with a spectral detection scan head and a 20X/1.0 NA water-immersion objective (Carl Zeiss). Functional time-series of visually evoked calcium responses in retinal ganglion cells (RGCs) were acquired at a rate of 4.1 Hz and 0.415 × 0.415 μm resolution (256 × 256 pixels), and 1 AU pinhole aperture. Excitation light was provided by a 488 nm multi-line laser. Non-anaesthetised Tg(elavl3:GCaMP5G) larvae were immobilized in 2% low melting point agarose (Sigma) prepared in Danieau solution and mounted dorsal side up on a raised glass platform that was placed in a custom-made Danieau-filled chamber. The agarose was sufficient to restrain the larvae so that anaesthesia was not required. Imaging was performed in the afternoon (1–8 pm).
Light-sheet imaging
Whole-brain light-sheet imaging was performed using a ZEISS Lightsheet Z.1 microscope equipped with two 10X/0.2 NA illumination objectives and one 20X/1.0 NA water-immersion detection objective (Carl Zeiss). 488 nm laser excitation light was used to elicit GCaMP6f fluorescence and a 505–545 BP filter was used for emitted light detection. The pivot scanner (Carl Zeiss) was used to deliver homogeneous illumination and, therefore, avoid shadows along the illumination axis. The thickness of the light sheet was 5.39 μm at the centre and 10.8 μm at the edges of the field of view. Exposure time was 29.97 ms. The size of volumetric images was 623 × 798 × 283 μm3 (1500 × 1920 × 490 pixels) with a resolution of 0.415 × 0.415 × 0.631 μm. 4 dpf nacre, casper and crystal Tg(elavl3:GCaMP6f) larvae were first paralysed for 10–15 minutes in α-bungarotoxin (1 mg/ml; Biotium) prepared in Danieau solution. Subsequently, larvae were immobilized in 2% low melting point agarose (Sigma) and placed inside a glass capillary (20 μl volume, 701904; Brand). We subsequently extruded the section of the agarose cylinder containing the head of the larva from the capillary, and oriented the larvae so that the dorsal side of the head was facing the detection objective and the eyes were facing the two illumination objectives. Whole-brain light-sheet imaging of casper mutant larvae was performed using a custom-made light-sheet microscope built by Dr Martin Meyer (King’s College London) and equipped with a 20X/1.0 NA water-immersion XLUMPlanFLN detection objective (Olympus).
Two-photon calcium imaging
Two-photon functional imaging in the retina was performed using a Nikon A1R MP microscope equipped with a 4-channel GaAsP NDD and an Apochromat 25X/1.1 NA water-immersion objective (Nikon). Excitation was provided by a Chameleon Ultra II Mode-locked titanium-sapphire laser (Coherent) tuned to 930 nm. Time-series of visually evoked calcium responses were acquired at a rate of 4 Hz and 0.248 × 0.248 μm resolution (512 × 256 pixels). Following activation of the laser scanning, we waited 60 seconds before starting the visual stimulation to ensure the retina adapted to the background light level caused by the multi-photon laser. 4 dpf crystal Tg(elavl3:GCaMP6f) larvae were first paralysed for 10–15 minutes in α-bungarotoxin (1 mg/ml; Biotium) prepared in Danieau solution. Subsequently, larvae were immobilized in 2% low melting point agarose (Sigma) and mounted on a raised custom-made glass platform with the dorsal side up (45° angle tilt) and one eye facing an LCD screen (see Visual stimulation) that was placed underneath a custom-made Danieau-filled chamber. Imaging was performed in the afternoon (1–8 pm).
Visual stimulation
Moving bars in confocal preparation
Moving bars stimuli were generated as previously described22,33. A diffusion filter (3026, Rosco) was bonded to one side of the chamber to serve as a projection screen. The agarose in front of the eye facing the projection screen was removed, allowing an unobstructed view of the projected image on the side of the chamber. Larvae were positioned 3 cm away from the screen and the projected image filled a visual field of ~97° × 63°. Visual stimuli consisted of light (56 cd/m2) or dark bars (8 cd/m2) (175% and 25% of mean luminance, respectively) on a mean grey background (32 cd/m2). As no qualitative differences between light and dark bars were noted, data obtained using the two stimuli were combined. Each bar was 10° in width moving at a speed of 20°/s and separated from the preceding bar by 30°, enabling more than one bar on the screen at any one time. The long axes of the bars were orthogonal to the direction of motion. Each of the 12 directions of motion was presented once (3 seconds) in a pseudo-random order unique to each slice in every animal imaged. Each inter-epoch interval was 10 seconds to enable GCaMP5G signals to return to baseline. A blank-screen null condition of 2 seconds was also interleaved. Visual experiments were generated and controlled using custom-written Labview and MATLAB code (MathWorks), implemented on a ViSaGe stimulus presenter (Cambridge Research Systems) and delivered via a DLP Pico Projector (Optoma).
Moving gratings in two-photon preparation
Moving gratings stimuli in the two-photon preparation were generated and controlled using PsychoPy39, and delivered through an LCD screen (SKD5VA-3, GoodWell Technology) positioned underneath a custom-made Perspex chamber. A long-pass red glass filter (FGL610, Thorlabs) was positioned between the LCD screen and the chamber to allow for simultaneous imaging and visual stimulation. Larvae were positioned 2 cm away from the screen and the image on the LCD screen filled a visual field of ~140° × 100° (mean background luminance 30.4 cd/m2). Visual stimuli consisted of square-wave gratings (100% contrast, spatial frequency 1.66 cycles/cm, temporal frequency 1 cycles/s). Each grating bar was 8.5° in width and the long axes of the bars were orthogonal to the direction of motion. Each of the 12 directions of motion was presented once (6 seconds) with and inter-epoch interval of 10 seconds to enable GCaMP6f signals to return to baseline. A blank-screen null condition of 6 seconds was also interleaved. TTL triggers (0-5-0 Volts) to record epoch time events where generated through a LabJack USB DAQ device (U3-LV, LabJack Corporation). Following activation of the laser scanning, we waited 60 seconds before starting the visual stimulation to ensure the retina adapted to the background light level caused by the multi-photon laser.
Optomotor response assay
Individual 5 dpf larvae were positioned in a 35 mm petri dish containing Danieau solution. The LCD screen of an iPhone 5 (Apple) controlled by a MacBook Pro (Apple) through Duet Display (Kairos Technologies) was used to display black and white square-wave gratings (85% contrast, spatial frequency 0.33 cycles/mm, temporal frequency 3.5 cycles/s) moving in 4 directions (90° angular distance) at the bottom of the petri dish. Visual stimuli were generated in Keynote (Apple). Each larva was tested 5 times in total (each trial lasted 6 s followed by 10 s of static gratings) and scored according to the trials it responded to (i.e., fish turns and swims in the direction of the moving gratings). The behaviour of larvae was visually monitored using an M165 FC stereomicroscope (Leica).
Analysis
Functional Analyses
In vivo calcium imaging data were analysed as previously described22,33. In summary, functional time-series were processed before analysis as follows: time-series images from each experiment were corrected for motion with a rigid-body algorithm (SPM12; http://www.fil.ion.ucl.ac.uk/spm/), median filtered with a kernel size of 1 voxel to remove dark and shot noise, and spatially smoothed with a 2D Gaussian kernel = 2 voxels to improve signal-to-noise. A baseline (B) that corrects for low-frequency drifts was determined using a cubic-spline algorithm extrapolating between knots averaged from 5 s of the inter-epoch interval data. Both relative signal intensity changes (ΔF = F-B; where F = raw fluorescence) and normalised signal intensity changes [%ΔF/F0 = (F-B)/B] were calculated at each voxel. ΔF was used for population functional data (voxel-wise analysis), whereas %ΔF/F0 was used for manually defined regions of interest (ROIs). For each voxel or ROI the integral response over the epoch-interval was calculated to provide a single response metric of each presented direction of stimulus motion. The integral within each epoch window is a summary metric more resistant to saturation effects of the calcium probe than maximum signal change. A threshold for each voxel within an acquisition image sequence was determined from the variance of ΔF changes during the inter-epoch intervals and null condition, threshold = 5 × SDs. All voxels that were supra-threshold within at least two visual presentation epochs were regarded as visually responsive and subjected to further characterization.
To analyse the direction and orientation selectivity of visually responsive voxels direction- and orientation-selective indices (DSI and OSI)40, based on fitted von-Mises or Gaussian profiles41, were calculated together with an estimate for their goodness of fit, R2. The DSI was defined as (Rpref−Rnull)/(Rpref + Rnull), where Rpref, the response to the preferred direction, was the integral response over the preferred direction epoch-interval. Rnull was similarly calculated as the integral response evoked by the direction opposite to the preferred direction. The OSI was defined as (Rpref−Rorth)/(Rpref + Rorth), where Rpref, the response to the preferred orientation, was the integral response over the preferred orientation epoch-interval. Rorth was similarly calculated as the integral response evoked by the orthogonal orientation. To minimize cross talk and over-fitting associated with DSI and OSI metrics, a stringent approach was undertaken. For a voxel to be regarded as direction-selective (DS) or orientation-selective (OS), mutually exclusive criteria were used: DS if DSI > 0.5 and OSI < 0.5; and OS if OSI > 0.5 and DSI < 0.5. In both cases, the goodness of fit (R2) for DSI and OSI, respectively, had to be >0.8; thus, the fitted curves explained at least 80% of the integral responses. A single von-Mises distribution was used to fit responses of DS voxels and estimate their preferred direction of motion angle from the centre of the fitted curve. The sum of two von-Mises (180° angular distance apart) was used to fit responses of OS voxels and estimate their preferred orientation of motion angles from the centres of the fitted curves. Circular variance was also calculated for comparison as an alternative metric of orientation selectivity (Circular variance < 0.4)41.
Morphological Analyses
To determine the brain volume imaged in 4 dpf nacre, casper and crystal Tg(elavl3:GCaMP6f) larvae, we calculated the number of GCaMP6f+ voxels in each volumetric image by applying the adjust>threshold function followed by the analyse>histogram>list command in ImageJ42. Subsequently, the obtained values were multiplied by the volume of a single voxel (0.415 × 0.415 × 0.631 μm3 = 1.086 × 10−1 μm3).
Statistical Analyses
Statistical test results are reported in Figures and Figure legends. Statistical analyses and tests were carried out using Prism 6 (GraphPad) or MATLAB R2014b (MathWorks). Before performing statistical tests, descriptive statistics (e.g., normality tests to see whether values come from a Gaussian distribution or F-test to compare variances) were used to choose the appropriate statistical test (reported in Figure legends together with test results). The criterion for statistical significance was set at p < 0.05.
Additional Information
How to cite this article: Antinucci, P. and Hindges, R. A crystal-clear zebrafish for in vivo imaging. Sci. Rep. 6, 29490; doi: 10.1038/srep29490 (2016).
References
Jacques, S. L. Optical properties of biological tissues: a review. Phys Med Biol 58, R37–R61 (2013).
Pawley, J. B. Handbook of biological confocal microscopy. (Springer, 2005).
Richardson, D. S. & Lichtman, J. W. Clarifying Tissue Clearing. Cell 162, 246–257 (2015).
White, R. M. et al. Transparent adult zebrafish as a tool for in vivo transplantation analysis. Cell Stem Cell 2, 183–189 (2008).
Singh, A. P. & Nusslein-Volhard, C. Zebrafish stripes as a model for vertebrate colour pattern formation. Curr Biol 25, R81–R92 (2015).
Kelsh, R. N. et al. Zebrafish pigmentation mutations and the processes of neural crest development. Development 123, 369–389 (1996).
Bharti, K., Nguyen, M. T., Skuntz, S., Bertuzzi, S. & Arnheiter, H. The other pigment cell: specification and development of the pigmented epithelium of the vertebrate eye. Pigment Cell Res 19, 380–394 (2006).
Karlsson, J., von Hofsten, J. & Olsson, P. E. Generating transparent zebrafish: a refined method to improve detection of gene expression during embryonic development. Mar Biotechnol (NY) 3, 522–527 (2001).
Lister, J. A., Robertson, C. P., Lepage, T., Johnson, S. L. & Raible, D. W. Nacre encodes a zebrafish microphthalmia-related protein that regulates neural-crest-derived pigment cell fate. Development 126, 3757–3767 (1999).
Whittaker, J. R. An analysis of melanogenesis in differentiating pigment cells of ascidian embryos. Dev Biol 14, 1–39 (1966).
Ryazanova, A. D., Alekseev, A. A. & Slepneva, I. A. The phenylthiourea is a competitive inhibitor of the enzymatic oxidation of DOPA by phenoloxidase. J Enzyme Inhib Med Chem 27, 78–83 (2012).
Li, Z. et al. Phenylthiourea specifically reduces zebrafish eye size. PLoS One 7, e40132 (2012).
Johnson, G. A., Boukma, S. J. & Kim, E. G. In vivo inhibition of dopamine beta-hydroxylase by 1-phenyl-3-(2-thiazolyl)-2-thiourea (U-14,624). J Pharmacol Exp Ther 171, 80–87 (1970).
Elsalini, O. A. & Rohr, K. B. Phenylthiourea disrupts thyroid function in developing zebrafish. Dev Genes Evol 212, 593–598 (2003).
Bohnsack, B. L., Gallina, D. & Kahana, A. Phenothiourea sensitizes zebrafish cranial neural crest and extraocular muscle development to changes in retinoic acid and IGF signaling. PLoS One 6, e22991 (2011).
Orger, M. B. et al. Behavioral screening assays in zebrafish. Methods Cell Biol 77, 53–68 (2004).
Parker, M. O., Brock, A. J., Millington, M. E. & Brennan, C. H. Behavioural phenotyping of casper mutant and 1-pheny-2-thiourea treated adult zebrafish. Zebrafish 10, 466–471 (2013).
Streisinger, G., Singer, F., Walker, C., Knauber, D. & Dower, N. Segregation analyses and gene-centromere distances in zebrafish. Genetics 112, 311–319 (1986).
Robles, E., Laurell, E. & Baier, H. The retinal projectome reveals brain-area-specific visual representations generated by ganglion cell diversity. Curr Biol 24, 2085–2096 (2014).
Nikolaou, N. et al. Parametric functional maps of visual inputs to the tectum. Neuron 76, 317–324 (2012).
Ahrens, M. B., Orger, M. B., Robson, D. N., Li, J. M. & Keller, P. J. Whole-brain functional imaging at cellular resolution using light-sheet microscopy. Nat Methods 10, 413–420 (2013).
Antinucci, P., Nikolaou, N., Meyer, M. P. & Hindges, R. Teneurin-3 specifies morphological and functional connectivity of retinal ganglion cells in the vertebrate visual system. Cell Rep 5, 582–592 (2013).
Nikolaou, N. & Meyer, M. P. Lamination Speeds the Functional Development of Visual Circuits. Neuron 88, 999–1013 (2015).
Keller, P. J. & Ahrens, M. B. Visualizing whole-brain activity and development at the single-cell level using light-sheet microscopy. Neuron 85, 462–483 (2015).
Tomer, R. et al. SPED Light Sheet Microscopy: Fast Mapping of Biological System Structure and Function. Cell 163, 1796–1806 (2015).
Chhetri, R. K., Amat, F., Wan, Y., Hockendorf, B. & Lemon, W. C. Whole-animal functional and developmental imaging with isotropic spatial resolution. Nat Methods 12, 1171–1178 (2015).
Wolf, S. et al. Whole-brain functional imaging with two-photon light-sheet microscopy. Nat Methods 12, 379–380 (2015).
Dunn, T. W. et al. Brain-wide mapping of neural activity controlling zebrafish exploratory locomotion. Elife 5 (2016).
Odermatt, B., Nikolaev, A. & Lagnado, L. Encoding of luminance and contrast by linear and nonlinear synapses in the retina. Neuron 73, 758–773 (2012).
Nikolaev, A., Leung, K. M., Odermatt, B. & Lagnado, L. Synaptic mechanisms of adaptation and sensitization in the retina. Nat Neurosci 16, 934–941 (2013).
Masland, R. H. The neuronal organization of the retina. Neuron 76, 266–280 (2012).
Baden, T. et al. The functional diversity of retinal ganglion cells in the mouse. Nature 529, 345–350 (2016).
Lowe, A. S., Nikolaou, N., Hunter, P. R., Thompson, I. D. & Meyer, M. P. A systems-based dissection of retinal inputs to the zebrafish tectum reveals different rules for different functional classes during development. J Neurosci 33, 13946–13956 (2013).
Ren, J. Q., McCarthy, W. R., Zhang, H., Adolph, A. R. & Li, L. Behavioral visual responses of wild-type and hypopigmented zebrafish. Vision Res 42, 293–299 (2002).
Bianco, I. H. & Engert, F. Visuomotor transformations underlying hunting behavior in zebrafish. Curr Biol 25, 831–846 (2015).
Vladimirov, N. et al. Light-sheet functional imaging in fictively behaving zebrafish. Nat Methods 11, 883–884 (2014).
Randlett, O. et al. Cellular requirements for building a retinal neuropil. Cell Rep 3, 282–290 (2013).
Antinucci, P., Suleyman, O., Monfries, C. & Hindges, R. Neural mechanisms generating orientation selectivity in the retina. Current Biology, http://dx.doi.org/10.1016/j.cub.2016.05.035 (2016).
Peirce, J. W. Generating Stimuli for Neuroscience Using PsychoPy. Front Neuroinform 2, 10 (2008).
Niell, C. M. & Stryker, M. P. Highly selective receptive fields in mouse visual cortex. J Neurosci 28, 7520–7536 (2008).
Swindale, N. V. Orientation tuning curves: empirical description and estimation of parameters. Biol Cybern 78, 45–56 (1998).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9, 671–675 (2012).
Acknowledgements
We would like to thank Andrew Lowe for analysis tools, Vladimir Grigoriev for providing the Nikon D7000 digital SLR camera, and Martin Meyer for providing zebrafish transgenic lines, the custom-made light-sheet microscope and comments on the manuscript. We are grateful to ZEISS (Ben Ewins) for providing the Lightsheet Z.1 microscope. We thank the Nikon Imaging Centre at King’s College London for additional support and instrumentation. This work was supported by the Biotechnology and Biological Sciences Research Council (BB/M000664/1 to R.H.) and by the Medical Research Council (1413592 to P.A.).
Author information
Authors and Affiliations
Contributions
P.A. and R.H. designed the study. P.A. performed the experiments and analysed the data. P.A. and R.H. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Antinucci, P., Hindges, R. A crystal-clear zebrafish for in vivo imaging. Sci Rep 6, 29490 (2016). https://doi.org/10.1038/srep29490
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep29490
This article is cited by
-
Generation of Albino Phenotype in Ornamental Fish by CRISPR/Cas9-Mediated Genome Editing of slc45a2 Gene
Marine Biotechnology (2023)
-
Disease modeling by efficient genome editing using a near PAM-less base editor in vivo
Nature Communications (2022)
-
Optical transparency and label-free vessel imaging of zebrafish larvae in shortwave infrared range as a tool for prolonged studying of cardiovascular system development
Scientific Reports (2022)
-
Manipulation of the Tyrosinase gene permits improved CRISPR/Cas editing and neural imaging in cichlid fish
Scientific Reports (2021)
-
Overexpressed GATA3 enhances the sensitivity of colorectal cancer cells to oxaliplatin through regulating MiR-29b
Cancer Cell International (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.