Abstract
Constitutive functional HIF-2α was recently identified in cancer and stem cell lines under normoxia. In this study, BEAS-2B, a bronchial epithelial cell line, was shown to constitutively express active HIF-2α under normoxia and exhibit markers of pluripotency including Oct-4, Nanog and sphere formation. Oct-4 expression was reduced after knockdown of HIF-2α under normoxia. Global enrichment analysis of HIF-2α demonstrated the diverse functions of HIF-2α under normoxia. Bioinformatics analysis of the enriched loci revealed an enhancer role of HIF-2α binding sites, involvement of HIF-2α interacting proteins and enriched de novo motifs which suggest the diverse role of HIF-2α in pseudohypoxia. The low ratio of the discovered loci overlapping with those revealed in cancer cell lines 786-O (16.1%) and MCF-7 (15.9%) under hypoxia indicated a prevailing non-canonical mechanism. Hypoxia had positive, marginal or adverse effects on the enrichment of the selected loci in ChIP-PCR assays. Deletion of the N-terminal activation domain (N-TAD) of HIF-2α disrupted the reporting activity of two of the loci annotated to ELN and ANKRD31. Hypoxia incurring abundance variation of HIF-2α may misrepresent the N-TAD functions as canonical hypoxia inducible features via C-TAD activation. Elucidation of the pseudohypoxia functions of constitutive HIF-2α is useful for resolving its role in malignancy and pluripotency.
Similar content being viewed by others
Introduction
The cellular response to limited oxygen tensions is primarily via induction of the hypoxia inducible factors, HIF-1α and HIF-2α which are involved at crucial stages of embryonic development1,2,3, homeostasis4,5 and cancer pathogenesis6,7 such as sensing nutrient availability and intracellular pH, regulation of anaerobic glycolysis, modulation of the tumor microenvironment and cell survival. HIFα protein molecules tend to become hydroxylated under normoxia at an oxygen dependent degradation domain by prolyl hydroxylase (PHD) leading to their ubiquitination and subsequent degradation via the von Hippel-Lindau protein (VHL)-proteasomal route8,9. Asparaginyl hydroxylase (also named factor inhibiting HIF, FIH) hydroxylates the C-terminal activation domain (C-TAD) leading to diminished recruitment of co-activator CBP/P300 and hence much reduced activation function8,10. Both PHD and FIH are members of the dioxygenase family which utilize molecular oxygen as a co-substrate for the hydroxylation of the HIFαs11,12. The former affects the abundance of HIFα and the latter exerts tight regulation on HIFα C-TAD activity with a higher affinity for oxygen than the former13. Thus, it is widely believed that hypoxia activated HIFs primarily through interaction between the C-TAD and CBP/P30014,15. To contrast with the results of the reported oxygen tension dependent inducible features, it is of considerable interest to investigate further some types of cells such as stem cells4, cancer stem cells16,17 and certain cancer cells5,17 that have been shown to exhibit constitutive expression of HIF-2α under normoxia. HIF-2αlikely has a major role in malignant transformation and maintenance of the pluripotent state2,10,17,18. HIF-2α molecules which have frequently been shown functional under normoxia though their C-TAD can still be inhibited by FIH17,19. Its underlying mechanism is very intriguing. Knockdown of PHD2 in HeLa cells intended to stabilize HIFs under normoxia was shown to partially induce a set of genes characterizing hypoxia, whereas combined silencing of both PHD2 and FIH induced a full set of genes characterizing hypoxia20. A conceivable mechanism for this is that interaction of HIF-2α with other transcription factors such as ETS family, NEMO, c-Myc and those cited in literature12,21,22,23 mutually enhances their transcription function, , indirectly resulting in activation of a set of genes under normoxia. Furthermore, it has been postulated that such induction of a partial set of genes under normoxia could be mediated by interaction of other domains, such as the N-TAD rather than the C-TAD, with distinct co-activators which were shown to be different from those of its C-TAD9,24,25. C-TAD truncated HIF-2α was able to activate both HIF enriched elements and artificial promoters9,20. The N-TAD of HIF-2α has major differences in sequence from that of HIF-1α25,26. “Domain-swapping” experiments between HIF-1α and HIF-2α provided clear evidence of gene selectivity which was attributed to the N-TAD of HIF-α25,27. The PAS domain was also reported to recruit specific co-activators and to contribute to the target specificity of HIF-2α28,29. Genes uniquely regulated by HIF-2α but not by HIF-1α include some implicated in malignant transformation, such as PAI-1, CXCR4, TGFα, Flk-1 and some in pluripotency, such as Oct-4, Sox2, Nanog and cyclin D12,18,22. Whether epigenetic cues plus environmental factors can favor the expression of stemness-related HIF-2α downstream genes and/or suppress expression of malignancy-related HIF-2α downstream genes in a normal cell is of considerable interest and may reveal novel gene regulatory networks.
Given the considerable interest in the versatile biological functions of this kind of proteins, many gene profiles30,31,32 and chromatin enrichment analyses of HIFs33,34,35,36 have been performed under hypoxic conditions mostly in neoplastic cells. It is of interest to exploit the broad, hypoxia-inducible and beneficial functions of HIFs in lung. The constitutive expression of HIF-2α but not HIF-1α was found in lung epithelial cells BEAS-2B to maintain viability and to exhibit a protective function against H2O2 insult (unpublished observation). Also, some markers and features characterizing pluripotency were found in BEAS-2B cells under normoxia. Furthermore, a genome wide analysis of HIF-2α chromatin enrichment was conducted in BEAS-2B cells in the absence of HIF-1α competition for enrichment at certain sites; this protocol contrasts with previous investigations under hypoxia conditions33,34,35,36, in which HIF-1α was shown to up-regulate histone demethylase and histone modification could dramatically affect chromatin structure and gene expression37,38. High enrichment of HIF-2α at loci was confirmed by using ChIP-PCR and reporter assays under normoxia. Enrichment of some of the candidate loci were shown less or un-responsive to hypoxia or sometimes even labile to hypoxia. Both N-TAD and C-TAD were found to be involved in specific enrichments with different weightings to each of the loci. In combination with bioinformatics analysis of sequence motifs, the present work provides insights into the pseudohypoxia functions of HIF-2α in BEAS-2B.
Results
Expression of pluripotency-related markers and a sphere culture of BEAS-2B under normoxia
In hypoxic but not normoxic embryonic stem cells, HIF-2α translocates into the nucleus to regulate pluripotency markers such as Oct-4, Sox2 and Nanog39,40. Generally, stem cells start to differentiate following the disappearance of Oct-4 expression. It is not common to detect Oct-4 expression in a differentiated normal cell. However, intriguingly, in our study, both constitutive expression of HIF-2α in the nucleus (Fig. 1a) and Oct-4 (Fig. 1b) were detected in BEAS-2B under normoxia while HIF-1α was not (unpublished observation). Knockdown of HIF-2α led to reduced Oct-4 expression (Fig. 1c) even under normoxia. In addition, expression of Nanog and a culture of BEAS-2B containing floating spheres (Fig. 1d) were also observed under normoxia. These results are similar to reports that HIF-2α expressed in several malignant and stem cells is functional under normoxia8,17,41. However, the possible pseudohypoxia function of HIF-2α has not been delineated. Therefore next, a genome wide analysis of HIF-2α under normoxia was conducted to explore its possible function and features.
Expression of pluripotency related markers and a sphere culture of BEAS-2B under normoxia.
(a) HIF-2α expression was detected in the nucleus of BEAS-2B under normoxia by western blotting. Beta tubulin and TATA binding protein (TBP) were used as protein markers for the cytosol and nucleus fraction, respectively. (b) Western blotting for Oct-4. (c) For knockdown of HIF-2α, BEAS-2B cells were transfected with shRNA targeting HIF-2α and sh-Luc was used as control. (d) Western blotting of Nanog and a culture of BEAS-2B containing floating spheres. Representative result from two or three experiments was presented.
ChIP-Seq and analysis of NGS data
Further investigation of the enrichment of nuclear HIF-2α on chromatin in BEAS-2B under normoxia was performed by ChIP-Seq analysis. After alignment of our ChIP-Seq data to hg19, more than 10 million mapped reads enriched by HIF-2α antibody (10,313,454) and IgG (10,469,717) respectively, were identified (Supplementary Data 1 Table S2). CisGenome software v2.0 was adopted for further analysis of the mapped reads and we selected “Two Sample Peak Calling” to look for binding peaks from IP versus control samples (http://www.biostat.jhsph.edu/~hji/cisgenome/index.htm). The final criteria, as described in Methods, for analysis led to identification of 1,991 binding peaks (Supplementary Data 2) and the top 20 are listed in Table 1.
Among these binding peaks were loci corresponding to 1741 annotated closest genes, including 96 that were annotated as regulated areas for non-protein coding genes and several of them annotated to the same genes. ChIP-Seq analysis of HIF enrichments conducted for a few cancer cells under hypoxic condition has been reported by several groups33,34,35,36. These studies led to the discovery of more potential functional features of HIFs under hypoxia. A small portion of the annotated gene loci in this study coincided with those reported under hypoxic conditions, around 16.1% of those in MCF-7 which expresses both HIF-1α and HIF-2α under hypoxia and 15.9% of those in 786-O, which expresses only HIF-2α under normoxia but not HIF-1α even under hypoxia (Supplementary data 1 Table S3A,B)34,36. These overlapping annotated gene loci were previously attributed as hypoxia inducible features.
Distribution of HIF-2α ChIP-Seq peaks
Similar to the previously reported ChIP-Seq work on HIFs under hypoxia, it was revealed that HIF-2αenrichment occupied fewer promoters (2.26%) but more in introns (44.80%) and more in remote regions far away from the transcription start site (TSS) (Fig. 2a,b). HIF-2α enrichment is inclined to distal regions away from the TSS, implying a role for enhancers in its target regulation34,36,42. The actual proximal interactions between distant enriched loci and regulated genes may be brought about through three dimensional chromatin looping structure with the aid of other proteins and remains intriguing, as what was studied about the regulation of cyclin D in 786-O by HIF-2α using chromatin conformation capture36.
Distribution of HIF-2α ChIP-seq peaks in BEAS-2B under normoxia.
(a) Pie chart illustrating the genomic locations of HIF-2α binding sites. (b) Bar graph showing the percentage of region–gene associations according to genomic regions’ distance to TSS computed by Genomic Regions Enrichment of Annotations Tool (GREAT) (http://bejerano.stanford.edu/great/public/html/).
Differential enrichment of HIF-2α under normoxia and hypoxia
Among representative loci chosen for verification of enrichment by using ChIP-PCR assay, nine of them here were shown highly enriched with HIF-2α under normoxia with percentage input ranging from 2.71% to 10.73% (Fig. 3a). A fragment of β-actin exon was chosen as a negative control34. In comparison to other previously reported data for hypoxic conditions, enrichments of the selected loci under normoxia were relatively high34,35,36,43. Hypoxia was reported to activate HIF transcriptional activity via recruitment of the co-activator P300 to the C-TAD14,15. In the case of BEAS-2B, abundance of HIF-2α was not apparently affected under hypoxia in our study. Thus, the effect of hypoxia on enrichments of HIF-2α and P300 on the representative binding loci was further investigated. HIF-2α enrichments at those loci annotated to IL10, JMJD5 and PPP2R3A respectively were shown apparently enhanced under hypoxia mimetic condition using CoCl2 (Fig. 3b). A concomitant rise of P300 enrichment at these sites was also observed under hypoxia. These data reflect the notion as documented in the literature that cooperation between the N-TAD and C-TAD facilitates some of the HIF transcription activity under hypoxia15,25. On the other hand, hypoxia exerted relatively less effect on enrichment of HIF-2α at loci annotated to ELN, SP3 and ANKRD31. We surmised that the less responsiveness to hypoxia was because there was less involvement of the C-TAD of HIF-2α in enrichment. Less involvement of the C-TAD was also reflected in less responsiveness of the other P300 enrichments to hypoxia except that at the locus annotated to CBY3. The apparent adverse hypoxia effect resulting in diminished enrichment of HIF-2α at those loci annotated to CBY3 (with high error bar), SCG2 and SUDS3 likely implies unknown interactions between HIF-2α and chromatin possibly involving recruitment of repressors4. The obvious inhibitory effect on enrichments of both HIF-2α and P300 could enable gene down-regulation , an area of study that remains largely unexplored. Next, four of the representative loci were selected for luciferase reporter assay to determine their activation function(s) with HIF-2α (Fig. 4a). All of the four loci were shown to be active relative to the backbone vector. Further hypoxia treatment to the reporter assay on these four loci consistently showed a response pattern similar to that of enrichment (Fig. 4b). The loci annotated to IL10 and PPP2R3A, wherein enrichments were increased under hypoxia, were also shown to respond positively to hypoxia in the established reporter assay. The loci annotated to ELN and ANKRD31, at which enrichments were less responsive to hypoxia, resulted in a minor scale of variation of observed reporter activity with a reduction similar to that of HIF-2α but not P300 enrichments under hypoxia. Further reporter assays with N-TAD or C-TAD truncated HIF-2α (dN and dC respectively) with these four loci in HEK293 cells clearly illustrated the relative contribution of HIF-2α domains in their activation function (Fig. 4c–f). A stabilized full-length active HIF-2α (M2) mutant (see details in Materials and Methods) was included for comparison. For both loci annotated to IL10 and PPP2R3A, the dC mutant demonstrated significantly more reduction of activation activity relative to M2, indicating relatively higher involvement of the C-TAD. This was consistent with the observation of their positive response to hypoxia. In the case of the locus for PPP2R3A, truncation of either domain resulted in loss of most of the activation activity indicating that there is a high degree of cooperation required. For both loci annotated to ELN and ANKRD31, dN elicited only minimal or no reporter activity relative to M2, although apparent reduction of activity was also seen for dC mutants. Deletion of the N-TAD disrupted most of the activation activity of HIF-2α in these loci suggesting an absolute requirement for the N-TAD for activation activity. Overall, both the N-TAD and the C-TAD and other domains of HIF-2α seem to be required to cooperate to achieve the observed activation activities of HIF-2α with different weightings at each representative locus selected for investigation.
Chromatin enrichment of HIF-2α analyzed by using ChIP-PCR.
(a) Representative enrichments of HIF-2α provided by ChIP-Seq analysis were further verified using semiquantitative PCR. Data are presented as signal relative to that of input DNA (100%). *P < 0.05 and **P < 0.01 vs. negative control. (b) The confirmed enrichments were further subjected to hypoxia mimetic treatment using CoCl2 (150 μM). Antibody against HIF-2α or P300 was used with the same primer pair. Signal of semiquantitative PCR relative to that of input DNA was used to calculate the relative fold binding between hypoxia and normoxia. All values were estimated by using image J (mean ± SD, n = 3).
Reporter assay on enriched loci annotated to IL10, PPP2R3A, ELN and ANKRD31.
Cypridina luc reporter plasmid loaded with or without sequences of enriched loci annotated to IL10, PPP2R3A, ELN or ANKRD31was used for the assay. Cypridina luciferase activity was normalized to the red firefly luciferase activity. (a) Luciferase activity is presented relative to empty vector in BEAS-2B under normoxia. (b) Relative luciferase activity between hypoxia (treated with 150 μM CoCl2) and normoxia. (c–f) Relative luciferase activity when the reporter plasmid was co-transfected with HIF-2α variants in HEK293 in a separate expression plasmid from Addgene. M2, a stabilized full length HIF-2α mutant, dN or dC, the N-TAD or the C-TAD truncated HIF-2α respectively. Data are shown as mean ± SD, n = 3. *P < 0.05; **P < 0.01 and ***P < 0.001 vs. control.
None of the promoters of pluripotency-related genes which were reported to be bound by HIF-2α in stem cells, such as those of Oct-4, Sox2 and Nanog were found in our ChIP-seq data. ChIP-PCR assay did not show significant enrichment of HIF-2α at the reported Oct-4 promoter under normoxia or hypoxia (data not shown). Other factors for suppressing enrichment of the loci known to be bound by HIF-2α in stem cells could exist. However, further treatment with trichostatin A, a HDAC inhibitor, increased Oct-4 transcript and protein level (Supplementary Data 1 Figure S1 A,B) in BEAS-2B under normoxia, suggesting the existence of epigenetic modification in the regulation of stemness. It is also plausible that there are other as yet unrevealed cells and context dependent loci bound by HIF-2α for its expression.
Analysis of HIF-2α binding motifs
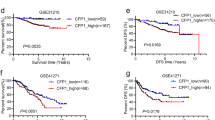
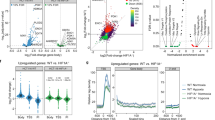
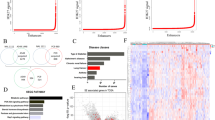
Using RSAT for nucleotide analysis, it was apparent that occurrence of single nucleotide C or G were somewhat less than A or T by around 2% in a single nucleotide composition profile of −100 to 100 bps across the HIF-2α binding peaks (Fig. 5a). In the dinucleotide composition profile, CG was clearly much less than all other combinations within the HIF-2α binding peaks whereas CA and CT dinucleotide compositions were the most frequent (Fig. 5b). When MEME-ChIP44 and MEME45 using a longer oligomer up to 30 mers was applied to motif analysis, three significant motifs (E-value ≤ 0.05) were discovered (Fig. 5c). There were 133, 13 and 6 sequences containing motif 1, motif 2 and motif 3, respectively (Supplementary Data 3). The remainder of the various peaks could have been derived from complex indirect interactions with other proteins as described below. Since the ChIP-Seq data of BEAS-2B was derived from experiments performed in the absence of HIF-1α under normoxia, the observed enrichment pattern of HIF-2α has been determined without any interference from competition of HIF-1α activities which have been previously reported to overlap with that of HIF-2α to a significant extent. Furthermore, HIF-1α is known to enable transcription of histone demethylase known to affect chromatin structure and hence gene expression37,38. It can be concluded that there were a small portion of genes commonly annotated between normoxic BEAS-2B and hypoxic MCF-7 based on HIF-2α enrichment and a significantly higher portion between normoxic BEAS-2B and hypoxic 786-O cells (Supplementary Data 1 Table S3A,B). Cell lineage defining factors could have also played a pivotal role in exerting cell-type specific effects. The renal cancer cell line 786-O fails to express HIF-1α under hypoxia but constitutively expresses HIF-2α even under normoxia. The 611 binding peaks of HIF-2α and its closest annotated genes were obtained by utilizing CisGenome 2.0 for re-analysis of the deposited ChIP-Seq data of 786-O (GSE34871) (Supplementary Data 2). The above three motifs of BEAS-2B and the 611 peaks of 786-O were matched using MAST46. There were approximately 4.4%, 9.2% and 3.6% of 611 sequences of 786-O ChIP-Seq containing motif 1, motif 2 and motif 3 with an E-value less than 10 and a position P-value of less than 0.0001, respectively (Supplementary Data 1 Figure S2A–C). As shown in Fig. 5d, motif 2 and motif 1 possess a similar pattern to the EPAS1 (HIF-2α) and ARI3A binding region in the databank, respectively. PWM Tools47 were applied to scan the HOCOMOCO48 with the motif of EPAS1 and the motif of ARI3A against BEAS-2B and 786-O (Fig. 5d). Results showed that the EPAS1 binding region was more frequent in 786-O (59.7%) than in BEAS-2B (35.3%) likely due to the fact that the experiment described with 786-O was performed under hypoxia which induced the complete activation of HIF-2α. Additionally, binding regions of ARI3A, which encodes a member of the AT-rich interaction domain family of DNA binding proteins and has clear roles in transcriptional regulation and chromatin structure modification, is more abundant in BEAS-2B (30.8%) than in 786-O (20.7%). These results imply that at least some common regulatory mechanisms exist between the hypoxia and normoxia response reactions.
Analysis of HIF-2α binding motifs.
HIF-2α binding profile of single nucleotide (a) or dinucleotide (b) composition generated by using RSAT. (c) The three most significant HIF-2α binding motifs under normoxia produced by MEME. (d) PWM Tools was used to scan the HOCOMOCO for EPAS1 motif 1 and ARI3A motif 2 against BEAS-2B and 786-O. (e) Analysis of HIF-2α potential heterodimers binding motif under normoxia. Results were generated by using dyad-analysis in RSAT. Significance index (sig) was computed by “sig = −log10(E value)” and E value = P value × number of tested character. Higher value means less error. *, de novo binding motifs that did not match those of any of known DNA binding proteins in the databank. (f) Co-enrichment of HIF-2α and SP-1 on Peak ID 151 annotated to PDXK.
In BEAS-2B, a low portion of peaks, i.e., 11.9% comprising the core hypoxia response element (HRE) RCGTG and a low ratio of overlapping content with loci presented via the strategy of hypoxia treatment of 786-O and MCF-7 indicated a prevailing non-canonical mechanism under normoxia43,49. For comparison, the reported canonical HRE enrichment in primary macrophage under hypoxia was 43%43. Hypoxia could have resulted in epigenetic modifications since activity of chromatin histone demethylases is oxygen dependent and some of them are known to be regulated by HIF-1α. Enrichment at loci lacking HRE may represent events where HIF-2α was tethered with other transcription factors (or via proteins indirectly) to enhance the specificity of one other, especially at loci far from TSS. It has been demonstrated that HIF-2α interacts with various protein partners, like ETS, NEMO and MYC, resulting in mutual enhancement of transcription activities at various targets. A de novo search for HIF-2α heterodimer binding motifs was conducted using dyad-analysis of RSAT. Five of the predicted motifs are listed (Fig. 5e). Two of them were annotated to similar binding motifs of several proteins. However, three of them did not match any motif of known DNA binding proteins in the databank. They may be de novo and in fact elicit functions via long range chromatin interaction, which would merit further study. Enriched loci within binding motifs as above could be directly derived from enrichment of HIF-2α or indirectly via its interacting protein partners. The motif analysis provides important clues about the de novo influence of other protein factors to HIF-2α binding to each site of chromatin or the de novo influence of HIF-2α to binding of other proteins to other sites of chromatin and thus their site specific transactivation activity. For exemplification, we detected the co-enrichment of HIF-2α and SP1 on the locus represented by peak ID151 (Fig. 5f).
Gene ontology (GO) and pathway enrichment analysis of annotated genes
Several HIF-2α downstream genes have been reported to be pluripotency-related and actively transcribed in stem cells and some have been successfully used for induction of pluripotent stem cells. For instance in this study, Oct-4 protein was detected though we could not find enrichment of HIF-2α at a locus in the promoter reported to be actively involved in transcription of Oct-4 in embryonic stem cells. It is not impossible that an unknown cell dependent locus exists to fulfill an enhancer role via long range interaction of chromatin. For comparison, a pathway analysis on published ChIP-Seq data indicated implication of HIF-2α in pluripotency in hypoxic MCF-7 but not in hypoxic 786-O cells34,36. Thus, GO analysis was performed on the 1741 annotated genes. HIF-2α was found to be associated with differentiation, regulation of transcription, vasculature development and so forth (Fig. 6a). Interestingly, the truncated N-TAD of HIF-2α was also reported to debase the transcription activation of vascular endothelial growth factor (VEGF), a critical factor in vasculature development, implicating HIF-2α involvement with vasculature development via N-TAD50. Our results echo the concept that HIF-2α drives vascular development under less hypoxia as proposed by Koh and Powis, possibly suggesting less involvement with C-TAD of HIF-2α50,51. Using Ingenuity Pathway Analysis, HIF-2α was implicated in pluripotency of embryonic stem cells involving Oct-4 and Nanog (Fig. 6b)34. It was also shown that HIF-2α is involved in specific cancer-related pathways such as the Wnt signaling pathway. Several reports have actually linked HIF-2α to Wnt-β-catenin pathway in cancer cells49,52,53. Notably, constitutive expression of HIF-2α was also detected in colon cancer cells under normoxia and it was reported to be involved in Wnt signaling under normoxia53. Furthermore, HIF-2α brought more contribution to Wnt signaling under hypoxia, suggesting N-TAD and C-TAD of HIF-2α synergetic involvement in Wnt signaling but with different weightings53. Furthermore, in analysis of “Diseases and Functions” by IPA (Supplementary Data 4), it was determined that HIF-2α in BEAS-2B also plays a significant role in maintaining cell survival, consistent with our data (unpublished observations).
Analysis of GO and Pathway for HIF-2α annotated genes in BEAS-2B.
(a) Analysis of GO was achieved by using DAVID, in which we adopted Gene Ontology/GOTERM_BP_FAT. Fold Enrichment >1.5 and P-Value, Bonferroni and FDR <0.05 were set as criteria for analysis. (b) Analysis of pathway by IPA and top 20 are shown according to the value of −log (P value).
Discussion
Herein, lung bronchial epithelial cells (non-malignant cells) constitutively expressing HIF-2α within the nucleus were shown to exhibit a few markers and weak features of pluripotency. Global analysis of enrichments of HIF-2α was performed to explore the diverse functions of HIF-2α under normoxia to contrast with those described for cancer cell lines 786-O and MCF-7 under hypoxia. This provided de novo loci enriched by HIF-2α under normoxia. Interaction with protein complexes via the N-TAD (or other domains) was found to mediate non-canonical diverse transcription functions of HIF-2α which is different from the well-studied canonical HRE enrichment via interaction of the C-TAD with P300 under hypoxia. Overall the cooperation between domains played a specific role in improving functions. Τargeting specificities under normoxia apparently extend wider range due to the diverse interactions of HIF-2α domains with other important proteins. The targeting mechanism under normoxia could also involve interactions between domains of HIF-2α with different specific co-activators. Furthermore, interactions between the enriched loci and those beyond the closest genes are conceivably likely via a chromatin looping structure54. The study of the hypoxia effect reported in numerous publications may have obscured non-canonical enrichments and resulted in the interpretation of the altered gene expressions as hypoxia inducible features via C-TAD activation. In fact, it may have been due to a change in abundance of HIF-2α and could be via interaction with other domain function. Hypoxia may also affect some of HIF-2α activities by epigenetic regulation apart from competition from HIF-1α, especially those involving oxygen dependent chromatin demethylase. In this study, enrichment of HIF-2α was not detected at the reported promoter core sequence of Oct-4 either under normoxia or hypoxia, in contrast to the hypoxia activated HIF-2α and expression of Oct-4 reported for (cancer) stem cells. Knockdown of HIF-2α reduced Oct-4 expression which was subject to epigenetic regulation but not hypoxia. Thus, a cell dependent unknown locus could be directly or indirectly enhanced by HIF-2α to activate Oct-4 transcription via long range interaction of chromatin. Actually, the reported enrichment levels of HIF-2α on loci within promoters of Oct-4 and other pluripotency-related genes are very low (<0.03%) in stem cells under hypoxia40. It is possible that de novo non-canonical loci which are less or un-responsive to hypoxia could be overlooked when aiming for hypoxia inducible features using an approach such as microarray or RT-PCR analysis. ChIP-Seq analyses have led to an increasing appreciation of the significance of cell and context specific enrichment of HIFs. At the same locus, protein partners co-enriched with HIFs could be context dependent. Thus, the correlation between enrichment of HIF-2α and the expression of regulated genes is not only context and cell dependent but also target dependent. It is of interest whether non-cancer cells like BEAS-2B expressing constitutive HIF-2α could assume epigenetic privilege plus environmental cues to favor pluripotency rather than malignancy. Elucidation of the regulatory mechanism of each pseudohypoxia function of constituent functional HIF-2α should be useful for resolving its role in malignancy or pluripotency in the growing number of cancer/stem cells accommodating constituent HIF-2α. Thereafter, a somatic cell which constitutively expresses HIF-2α would seem a likely candidate for developing induced pluripotent stem cells. Exploitation of the potential pluripotency of BEAS-2B cells could be a valuable model study.
ChIP-seq data is available at http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE81635.
Methods
Cell culture
The BEAS-2B cell line was obtained from ATCC and routinely cultured in RPMI1640 medium (Gibco) with high glucose containing 10% FBS (Gibco) and 2g/L sodium bicarbonate (Wako Pure Chemical Industries). The HEK293 cell line from ATCC was cultured in high glucose DMEM medium (Gibco) containing 10% FBS and 3.7g/L sodium bicarbonate.
Western blotting
The Nuclear/Cytosol Fractionation Kit (BioVision) was used for nuclear fractionation. Extracted proteins were separated by running SDS-PAGE and transferred to polyvinylidene fluoride (PVDF) membrane. After the membrane was blocked in 5% fat-free milk in 1 × TBST and washed three times with 1 × TBST, it was immersed in 5% fat-free milk in 1 × TBST solution containing a primary antibody against HIF-2α (Novus, NB100-122; 1:200), Oct-4 (abcam, ab19857; 1 μg/ml), Nanog (ThermoFisher, PA1-097; 1:2000), Beta-tubulin (Novus, NB600-936; 1:1000), TATA binding protein (Novus, NB500-700; 1:1000), or Beta-actin (Novus, NB600-501; 1:5000), respectively at 4 °C overnight. After the membrane was washed three times with 1 × TBST, a secondary antibody against HRP-conjugated goat polyclonal anti-rabbit IgG (1:10000) or HRP-conjugated rabbit polyclonal anti-mouse IgG (1:10000) were added for 30 minutes at room temperature (RT). Finally, results were photographed by using LAS-3000 (FUJIFILM).
Chromatin immunoprecipitation and polymerase chain reaction
Magna ChIP-Seq™ ChIP and NGS Library Preparation Kit (Millipore) were used following manufacturer’s instructions except for cell lysis buffer and nuclei lysis buffer. Five hundred and fifty microliters of 37% formaldehyde was added to a final concentration of 1% (V/V) for crosslinking of DNA and protein in BEAS-2B cells under normoxia for 10 minutes at RT. To stop this reaction, 0.125 M glycine (final concentration) was added and thoroughly mixed for 5 minutes. Ice-cold 1 × PBS containing protease inhibitor cocktail (Thermo Scientific™) was used to wash cells twice. Cells, (approximately 7.5 × 106), were harvested in 1.5 ml ice-cold PBS and centrifuged at 800 × g for 5 minutes at 4 °C, followed by incubation in 1 ml cell lysis buffer (0.5% NP40, 85 mM KCl, 20 mM Tris-HCl, pH8.0) with protease inhibitors for 30 minutes on ice and vortex-mixed briefly every five minutes. Cells were collected by centrifugation at 800 × g for 5 minutes at 4 °C and re-suspended with 500 μl of nuclei lysis buffer (1% SDS, 10 mM EDTA, 50 mM Tris-HCl, pH8.0) containing protease inhibitors. Chromatins were sheared by using BioRuptor (Diagenode) to yield fragments mostly within 100–300 base pair range. Centrifugation at 10,000 × g was performed for 10 minutes at 4 °C after the ultrasonic shearing. The supernatant was divided into 50 μl aliquots. For immunoprecipitation, each aliquot was added to 450 μl of the dilution buffer, protease inhibitors, HIF-2α antibody (Novus, NB100-122), or p300 antibody (Novus, NB500-161), or SP1 (Millipore, 07–645) or IgG antibody (Millipore, 12–371) , together with 20 μl of protein A/G magnetic beads for subsequent incubation at 4 °C with agitation overnight. PureProteome Magnetic Stand (Millipore) was used to adsorb bead-immune complexes in each tube. These were washed sequentially by low salt-, high salt-, LiCl-immune complex wash buffer and TE buffer. Elution buffer with RNase A was added at 37 °C for 30 minutes and then proteinase K was added at 62 °C for 2 hours to elute the immune complexes from beads. The crosslinks of the immune complexes were disrupted by incubation at 95 °C for 10 minutes. DNA purification was conducted by utilizing commercial spin columns (Millipore) following the manufacturer’s instructions. Lastly, eluted DNA aliquots in TE buffer were stored at −20 °C immediately until needed. For the ChIP-PCR assay, input DNA or antibody-enriched DNA fragments were amplified with primers (Supplementary data 1 Table S1) using PCR with reagent mixture (VELOCITY DNA Polymerase, BIOLINE) according to manufacturer’s instructions. PCR products were analyzed on a 2% agarose gel. The signal strength was estimated and quantified by using ImageJ software. Results are presented as percentage input (=ChIP/input), relative fold binding, or the ratio of individual percentage input under hypoxia divided by that under normoxia.
ChIP-sequencing and data analysis
ChIP-sequencing (ChIP-Seq) and basic bioinformatics analysis were executed by the Center for Genomic Medicine of Taiwan National Cheng Kung University. These tasks included library construction, emulsion PCR and sequencing analysis conducted by using 5500xl SOLiD sequencer (Applied Biosystems). SOLiD Accuracy Enhancement Tool (SAET) was used to analyze raw sequence data, including enhancement of color call accuracy and mapping reads to GRCh37/hg19 by employing LifeScope™ Genomic Analysis Software. CisGenome software v2.055 was used to identify binding peaks from mapped reads set at least 30 reads each window and minimum of 30 percent of total reads was limited for each single direction. Peaks were retained only when the false discovery rate (FDR) was less than 0.05 and the maximal |log2(fold change)| for all 100 bp windows within the peak was more than or equal to 556. Deduction of putative HIF-2α target genes was also accomplished by performing CisGenome software v2.0. In motif analysis, MEME-ChIP and MEME were applied to identify HIF-2α binding motifs44,45. For heterodimer binding motif, dyad analysis of Regulatory Sequence Analysis Tools (RSAT) was utilized to explore potential HIF-2α partners referring to the guide to the software57,58. In addition, MAST was used for match of motifs discovered in BEAS-2B with that in previously published ChIP-seq data of 786-O under hypoxia46. Among the motifs, PWM Tools47 was used to scan the HOCOMOCO48 for EPAS1 motif and ARI3A motif against BEAS-2B and 786-O. In Gene ontology (GO) analysis, “The Database for Annotation, Visualization and Integrated Discovery (DAVID)” was used to explore the biological denotation of putative HIF-2α downstream genes59,60. Ingenuity Pathway Analysis61 was performed for analysis of pathway and “Diseases and Functions”.
Reporter assay
All procedures followed the instructions in the Pierce™ Cypridina-Firefly Luciferase Dual Assay Kit (Thermo Scientific) manual. Briefly, pMCS-Cypridina Luc Vector inserted with or without HIF-2α enriched locus, ID 49 (ELN), 62 (IL10), 201 (PPP2R3A) or 402 (ANKRD31) together with red firefly luc control plasmid were co-transfected into BEAS-2B for 24 hours and either further exposed to normoxia or hypoxia mimicking conditions (150 μM cobalt chloride) for 4 hours. Primers are listed in Supplementary Data 1 Table S1. Backbone vector was used for background control. HIF-2α variants denoted as M2, dN and dC were purchased from Addgene and were co-transfected with each pMCS-Cypridina Luc Vector with or without the above insert and red firefly luc control plasmid into HEK293 under normoxia as described below. Data were normalized to that of red firefly luciferase that was yielded from pTK-Red Firefly Luc Vector. Measurement of luciferase activity was conducted by using FLx800 (BioTek). M2 is a plasmid for expression of full length of active HIF-2α in which the amino acid sequences was mutated at position 405 (P- > A), 531 (P- > A) and at 847 (N- > A). dN is a plasmid for expression of the N-TAD (450–572) deleted variant of HIF-2α and is mutated at 405 (P- > A). dC is a plasmid for expression of the C-TAD (820–870) deleted variant of HIF-2α and is mutated at position 405 (P- > A) and 531 (P- > A)50.
RT-PCR and Knockdown
RNA was extracted by using REzolTM C&T following the manufacturer’s instruction. Next, High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems™) was used to reverse transcribe RNA to cDNA. Master Mix Kit 2.0X (Topbio PCR MIX) was used to conduct PCR and signal intensity was quantified by using ImageJ software. Data was normalized to beta-actin. For Knockdown of HIF-2α, BEAS-2B cells were transfected with shHIF-2α (TRCN000003806) or shLuc (TRCN0000072245) by using lipofectamine 2000 reagent (Invitrogen) according to the manufacturer’s protocol. RNA was extracted after one day of transfection. Primers are listed in Supplementary Data 1 Table S1.
Sphere culture
First, BEAS-2B cells were cultured as a monolayer as above. Then, cells were harvested and around 20,000 cells/well were seeded into Ultra-low Attachment 6-well plates (Corning , Corning, NY) containing DMEM/F12 (Gibco) with 20% KnockOut™ Serum Replacement (Gibco), 1X Non-essential amino acids (Biological Industries), 1 mM L-alanyl-L-glutamine (Biological Industries), 100 μM 2-mercaptoethanol (Sigma) and 10 ng/ml recombinant human fibroblast growth factor-basic (PEPROTECH). Two hundred and fifty microliters of fresh medium was added every three days. Spheres were photographed on day 10.
Statistical analysis
Data are shown as mean ± the standard deviation. T-test was used to determine statistical significance (P < 0.05) manipulated using Excel.
Additional Information
How to cite this article: Lee, M.-C. et al. Genome-wide analysis of HIF-2α chromatin binding sites under normoxia in human bronchial epithelial cells (BEAS-2B) suggests its diverse functions. Sci. Rep. 6, 29311; doi: 10.1038/srep29311 (2016).
References
Wiesener, M. S. et al. Induction of endothelial PAS domain protein-1 by hypoxia: characterization and comparison with hypoxia-inducible factor-1alpha. Blood 92, 2260–2268 (1998).
Covello, K. L. et al. HIF-2alpha regulates Oct-4: effects of hypoxia on stem cell function, embryonic development and tumor growth. Genes Dev 20, 557–570, doi: 10.1101/gad.1399906 (2006).
Tian, H., McKnight, S. L. & Russell, D. W. Endothelial PAS domain protein 1 (EPAS1), a transcription factor selectively expressed in endothelial cells. Genes Dev 11, 72–82 (1997).
Hu, C. J. et al. Differential regulation of the transcriptional activities of hypoxia-inducible factor 1 alpha (HIF-1alpha) and HIF-2alpha in stem cells. Mol Cell Biol 26, 3514–3526, doi: 10.1128/MCB.26.9.3514-3526.2006 (2006).
Holmquist-Mengelbier, L. et al. Recruitment of HIF-1alpha and HIF-2alpha to common target genes is differentially regulated in neuroblastoma: HIF-2alpha promotes an aggressive phenotype. Cancer Cell 10, 413–423, doi: 10.1016/j.ccr.2006.08.026 (2006).
Kaelin, W. G., Jr. The von Hippel-Lindau tumour suppressor protein: O2 sensing and cancer. Nat Rev Cancer 8, 865–873, doi: 10.1038/nrc2502 (2008).
Gordan, J. D. & Simon, M. C. Hypoxia-inducible factors: central regulators of the tumor phenotype. Curr Opin Genet Dev 17, 71–77 (2007).
Qing, G. & Simon, M. C. Hypoxia inducible factor-2alpha: a critical mediator of aggressive tumor phenotypes. Curr Opin Genet Dev 19, 60–66, doi: 10.1016/j.gde.2008.12.001 (2009).
Pawlus, M. R., Wang, L., Ware, K. & Hu, C. J. Upstream stimulatory factor 2 and hypoxia-inducible factor 2alpha (HIF2alpha) cooperatively activate HIF2 target genes during hypoxia. Mol Cell Biol 32, 4595–4610, doi: 10.1128/MCB.00724-12 (2012).
Mimeault, M. & Batra, S. K. Hypoxia-inducing factors as master regulators of stemness properties and altered metabolism of cancer- and metastasis-initiating cells. J Cell Mol Med 17, 30–54, doi: 10.1111/jcmm.12004 (2013).
Kaelin, W. G. Proline hydroxylation and gene expression. Annu Rev Biochem 74, 115–128 (2005).
Elvert, G. et al. Cooperative interaction of hypoxia-inducible factor-2alpha (HIF-2alpha) and Ets-1 in the transcriptional activation of vascular endothelial growth factor receptor-2 (Flk-1). J Biol Chem 278, 7520–7530, doi: 10.1074/jbc.M211298200 (2003).
Koivunen, P., Hirsila, M., Gunzler, V., Kivirikko, K. I. & Myllyharju, J. Catalytic properties of the asparaginyl hydroxylase (FIH) in the oxygen sensing pathway are distinct from those of its prolyl 4-hydroxylases. J Biol Chem 279, 9899–9904, doi: 10.1074/jbc.M312254200 (2004).
Kasper, L. H. et al. Two transactivation mechanisms cooperate for the bulk of HIF-1-responsive gene expression. EMBO J 24, 3846–3858, doi: 10.1038/sj.emboj.7600846 (2005).
Lisy, K. & Peet, D. J. Turn me on: regulating HIF transcriptional activity. Cell Death Differ 15, 642–649, doi: 10.1038/sj.cdd.4402315 (2008).
Seidel, S. et al. A hypoxic niche regulates glioblastoma stem cells through hypoxia inducible factor 2 alpha. Brain 133, 983–995, doi: 10.1093/brain/awq042 (2010).
Pietras, A., Johnsson, A. S. & Pahlman, S. The HIF-2alpha-driven pseudo-hypoxic phenotype in tumor aggressiveness, differentiation and vascularization. Curr Top Microbiol Immunol 345, 1–20, doi: 10.1007/82_2010_72 (2010).
Patel, S. A. & Simon, M. C. Biology of hypoxia-inducible factor-2alpha in development and disease. Cell Death Differ 15, 628–634, doi: 10.1038/cdd.2008.17 (2008).
Franovic, A. & Lee, S. HIF-2alpha: many cancers, one engine? Cell Cycle 9, 859–860 (2010).
Dayan, F., Roux, D., Brahimi-Horn, M. C., Pouyssegur, J. & Mazure, N. M. The oxygen sensor factor-inhibiting hypoxia-inducible factor-1 controls expression of distinct genes through the bifunctional transcriptional character of hypoxia-inducible factor-1alpha. Cancer Res 66, 3688–3698, doi: 10.1158/0008-5472.CAN-05-4564 (2006).
Bracken, C. P., Whitelaw, M. L. & Peet, D. J. Activity of hypoxia-inducible factor 2alpha is regulated by association with the NF-kappaB essential modulator. J Biol Chem 280, 14240–14251, doi: 10.1074/jbc.M409987200 (2005).
Aprelikova, O., Wood, M., Tackett, S., Chandramouli, G. V. & Barrett, J. C. Role of ETS transcription factors in the hypoxia-inducible factor-2 target gene selection. Cancer Res 66, 5641–5647, doi: 10.1158/0008-5472.CAN-05-3345 (2006).
Dang, C. V., Kim, J. W., Gao, P. & Yustein, J. The interplay between MYC and HIF in cancer. Nat Rev Cancer 8, 51–56, doi: 10.1038/nrc2274 (2008).
Pawlus, M. R., Wang, L., Murakami, A., Dai, G. & Hu, C. J. STAT3 or USF2 contributes to HIF target gene specificity. PLoS One 8, e72358, doi: 10.1371/journal.pone.0072358 (2013).
Hu, C. J., Sataur, A., Wang, L., Chen, H. & Simon, M. C. The N-terminal transactivation domain confers target gene specificity of hypoxia-inducible factors HIF-1alpha and HIF-2alpha. Mol Biol Cell 18, 4528–4542, doi: 10.1091/mbc.E06-05-0419 (2007).
O’Rourke, J. F., Tian, Y. M., Ratcliffe, P. J. & Pugh, C. W. Oxygen-regulated and transactivating domains in endothelial PAS protein 1: comparison with hypoxia-inducible factor-1alpha. J Biol Chem 274, 2060–2071 (1999).
Lau, K. W., Tian, Y. M., Raval, R. R., Ratcliffe, P. J. & Pugh, C. W. Target gene selectivity of hypoxia-inducible factor-alpha in renal cancer cells is conveyed by post-DNA-binding mechanisms. Br J Cancer 96, 1284–1292, doi: 10.1038/sj.bjc.6603675 (2007).
Partch, C. L. & Gardner, K. H. Coactivators necessary for transcriptional output of the hypoxia inducible factor, HIF, are directly recruited by ARNT PAS-B. Proc Natl Acad Sci USA 108, 7739–7744, doi: 10.1073/pnas.1101357108 (2011).
Masetti, M., Falchi, F. & Recanatini, M. Protein dynamics of the HIF-2alpha PAS-B domain upon heterodimerization and ligand binding. PLoS One 9, e94986, doi: 10.1371/journal.pone.0094986 (2014).
Malard, V. et al. Global gene expression profiling in human lung cells exposed to cobalt. BMC Genomics 8, 147 (2007).
Zaman, K. et al. Protection from oxidative stress-induced apoptosis in cortical neuronal cultures by iron chelators is associated with enhanced DNA binding of hypoxia-inducible factor-1 and ATF-1/CREB and increased expression of glycolytic enzymes, p21(waf1/cip1) and erythropoietin. J Neurosci 19, 9821–9830 (1999).
Wang, V., Davis, D. A., Haque, M., Huang, L. E. & Yarchoan, R. Differential gene up-regulation by hypoxia-inducible factor-1alpha and hypoxia-inducible factor-2alpha in HEK293T cells. Cancer Res 65, 3299–3306, doi: 10.1158/0008-5472.CAN-04-4130 (2005).
Tanimoto, K. et al. Genome-wide identification and annotation of HIF-1alpha binding sites in two cell lines using massively parallel sequencing. Hugo J 4, 35–48, doi: 10.1007/s11568-011-9150-9 (2010).
Schodel, J. et al. High-resolution genome-wide mapping of HIF-binding sites by ChIP-seq. Blood 117, e207–217, doi: 10.1182/blood-2010-10-314427 (2011).
Mole, D. R. et al. Genome-wide association of hypoxia-inducible factor (HIF)-1alpha and HIF-2alpha DNA binding with expression profiling of hypoxia-inducible transcripts. J Biol Chem 284, 16767–16775, doi: 10.1074/jbc.M901790200 (2009).
Schodel, J. et al. Common genetic variants at the 11q13.3 renal cancer susceptibility locus influence binding of HIF to an enhancer of cyclin D1 expression. Nat Genet 44, 420–425, S421–422, doi: 10.1038/ng.2204 (2012).
Mimura, I., Tanaka, T., Wada, Y., Kodama, T. & Nangaku, M. Pathophysiological response to hypoxia - from the molecular mechanisms of malady to drug discovery: epigenetic regulation of the hypoxic response via hypoxia-inducible factor and histone modifying enzymes. J Pharmacol Sci 115, 453–458 (2011).
Xia, X. et al. Integrative analysis of HIF binding and transactivation reveals its role in maintaining histone methylation homeostasis. Proc Natl Acad Sci USA 106, 4260–4265, doi: 10.1073/pnas.0810067106 (2009).
Forristal, C. E., Wright, K. L., Hanley, N. A., Oreffo, R. O. & Houghton, F. D. Hypoxia inducible factors regulate pluripotency and proliferation in human embryonic stem cells cultured at reduced oxygen tensions. Reproduction 139, 85–97, doi: 10.1530/rep-09-0300 (2010).
Petruzzelli, R., Christensen, D. R., Parry, K. L., Sanchez-Elsner, T. & Houghton, F. D. HIF-2alpha regulates NANOG expression in human embryonic stem cells following hypoxia and reoxygenation through the interaction with an Oct-Sox cis regulatory element. PLoS One 9, e108309, doi: 10.1371/journal.pone.0108309 (2014).
Pietras, A. et al. HIF-2alpha maintains an undifferentiated state in neural crest-like human neuroblastoma tumor-initiating cells. Proc Natl Acad Sci USA 106, 16805–16810, doi: 10.1073/pnas.0904606106 (2009).
Schodel, J., Mole, D. R. & Ratcliffe, P. J. Pan-genomic binding of hypoxia-inducible transcription factors. Biol Chem 394, 507–517, doi: 10.1515/hsz-2012-0351 (2013).
Tausendschon, M. et al. Genome-wide identification of hypoxia-inducible factor-1 and -2 binding sites in hypoxic human macrophages alternatively activated by IL-10. Biochim Biophys Acta 1849, 10–22, doi: 10.1016/j.bbagrm.2014.10.006 (2015).
Machanick, P. & Bailey, T. L. MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics 27, 1696–1697, doi: 10.1093/bioinformatics/btr189 (2011).
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol 2, 28–36 (1994).
Bailey, T. L. & Gribskov, M. Combining evidence using p-values: application to sequence homology searches. Bioinformatics 14, 48–54 (1998).
Orenstein, Y. & Shamir, R. A comparative analysis of transcription factor binding models learned from PBM, HT-SELEX and ChIP data. Nucleic Acids Res 42, e63, doi: 10.1093/nar/gku117 (2014).
Kulakovskiy, I. V. et al. HOCOMOCO: a comprehensive collection of human transcription factor binding sites models. Nucleic Acids Res 41, D195–202, doi: 10.1093/nar/gks1089 (2013).
Stiehl, D. P. et al. Non-canonical HIF-2alpha function drives autonomous breast cancer cell growth via an AREG-EGFR/ErbB4 autocrine loop. Oncogene 31, 2283–2297, doi: 10.1038/onc.2011.417 (2012).
Yan, Q., Bartz, S., Mao, M., Li, L. & Kaelin, W. G. Jr. The hypoxia-inducible factor 2alpha N-terminal and C-terminal transactivation domains cooperate to promote renal tumorigenesis in vivo. Mol Cell Biol 27, 2092–2102, doi: 10.1128/mcb.01514-06 (2007).
Koh, M. Y. & Powis, G. Passing the baton: the HIF switch. Trends in biochemical sciences 37, 364–372, doi: 10.1016/j.tibs.2012.06.004 (2012).
Choi, H., Chun, Y. S., Kim, T. Y. & Park, J. W. HIF-2alpha enhances beta-catenin/TCF-driven transcription by interacting with beta-catenin. Cancer Res 70, 10101–10111, doi: 10.1158/0008-5472.CAN-10-0505 (2010).
Santoyo-Ramos, P., Likhatcheva, M., Garcia-Zepeda, E. A., Castaneda-Patlan, M. C. & Robles-Flores, M. Hypoxia-inducible factors modulate the stemness and malignancy of colon cancer cells by playing opposite roles in canonical Wnt signaling. PLoS One 9, e112580, doi: 10.1371/journal.pone.0112580 (2014).
Fullwood, M. J. & Ruan, Y. ChIP-based methods for the identification of long-range chromatin interactions. J Cell Biochem 107, 30–39, doi: 10.1002/jcb.22116 (2009).
Ji, H. et al. An integrated software system for analyzing ChIP-chip and ChIP-seq data. Nat Biotechnol 26, 1293–1300, doi: 10.1038/nbt.1505 (2008).
Shin, H., Liu, T., Duan, X., Zhang, Y. & Liu, X. S. Computational methodology for ChIP-seq analysis. Quant Biol 1, 54–70, doi: 10.1007/s40484-013-0006-2 (2013).
Thomas-Chollier, M. et al. RSAT peak-motifs: motif analysis in full-size ChIP-seq datasets. Nucleic Acids Res 40, e31, doi: 10.1093/nar/gkr1104 (2012).
Thomas-Chollier, M. et al. A complete workflow for the analysis of full-size ChIP-seq (and similar) data sets using peak-motifs. Nat Protoc 7, 1551–1568, doi: 10.1038/nprot.2012.088 (2012).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4, 44–57, doi: 10.1038/nprot.2008.211 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37, 1–13, doi: 10.1093/nar/gkn923 (2009).
Risbud, M. V., Schipani, E. & Shapiro, I. M. Hypoxic regulation of nucleus pulposus cell survival: from niche to notch. Am J Pathol 176, 1577–1583, doi: 10.2353/ajpath.2010.090734 (2010).
Acknowledgements
We thank team at the Center for Genomic Medicine of Taiwan National Cheng Kung University for providing generous advice. We thank Michael Byford at South Bank University of London and Miranda Loney at Academic Sinica Taipei for their help in editing this manuscript. This work was supported by Cheng Hsin General Hospital [104-58] and the National Defense Medical Center [MAB-103-M029] and [MAB-104-69].
Author information
Authors and Affiliations
Contributions
M.-C.L. conducted most experiments and analyzed the results and drafted the manuscript. H.-J.H. revised the manuscript and shared ideas. T.-H.C. provided bioinformatics expertise and some relevant results. H.-C.H. revised the manuscript and shared ideas. S.-Y.H. conducted experiments of subcloning plasmids for reporter assay. Y.-S.C. initiated sonication work and provided some data from ChIP and western blotting. W.-Y.C. revised the manuscript (Y.-S.C. co-supervisor). C.-H.C. revised the manuscript (Y.-S.C. co-supervisor). C.-H.L. proofread the manuscript (co-supervisor). C.-Y.S. designed this study and completed the writing of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Lee, MC., Huang, HJ., Chang, TH. et al. Genome-wide analysis of HIF-2α chromatin binding sites under normoxia in human bronchial epithelial cells (BEAS-2B) suggests its diverse functions. Sci Rep 6, 29311 (2016). https://doi.org/10.1038/srep29311
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep29311
This article is cited by
-
Metabolic implications of hypoxia and pseudohypoxia in pheochromocytoma and paraganglioma
Cell and Tissue Research (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.