Abstract
Transcriptional regulation of capsule expression is critical for pneumococcal transition from carriage to infection, yet the underlying mechanism remains incompletely understood. Here, we describe the regulation of capsular polysaccharide, one of the most important pneumococcal virulence factor by a GntR family regulator, CpsR. Electrophoretic mobility-shift assays have shown the direct interaction between CpsR and the cps promoter (cpsp), and their interaction could be competitively interfered by glucose. DNase I footprinting assays localized the binding site to a region −146 to −114 base pairs relative to the transcriptional start site of the cps locus in S. pneumoniae D39. We found that CpsR negatively controlled the transcription of the cps locus and hence CPS production, which was confirmed by fine-tuning expression of CpsR in a ΔcpsR complemented strain. Increased expression of CpsR in complemented strain led to a decreased resistance to the whole-blood-mediated killing, suggesting a protective role for CpsR-cpsp interaction in the establishment of invasive infection. Finally, animal experiments showed that CpsR-cpsp interaction was necessary for both pneumococcal colonization and invasive infection. Taken together, our results provide a thorough insight into the regulation of capsule production mediated by CpsR and its important roles in pneumococcal pathogenesis.
Similar content being viewed by others
Introduction
Streptococcus pneumoniae (pneumococcus) is known as an important human bacterial pathogen, which usually resides in the upper respiratory tract and has the capacity to cause pneumonia, otitis media, meningitis and sepsis1. It has been estimated in 2010 that at least 400,000 children under 6 years of age die each year from invasive pneumococcal disease worldwide2.
An infection generally occurs when host factors are in favor of bacterial growth3 and allow a pathogen to modulate the expression of virulence factors to survive a host’s defense mechanism4,5. The pneumococcus harbors a number of virulence factors, including capsular polysaccharide, pneumolysin, pneumococcal surface antigen A, choline binding proteins6. Of them, the capsular polysaccharide (capsule, CPS) is recognized as the most important one contributing to the disease process6. CPS has several distinguishing features7,8,9,10, one of the most striking being the role of its protection against host opsonophagocytic killing11. Despite CPS contributes to both pneumococcal colonization and invasive disease12, different lines of evidence indicate that CPS is differentially produced between carriage pneumococci and blood-grown planktonic pneumococci13,14,15,16; nevertheless, the underlying mechanism is not yet well understood.
It’s accepted that transcript levels of the cps locus are positively correlated with the amount of CPS17, and cps2A gene has been normally used to reflect the transcription of the cps locus in S. pneumoniae18. Although different S. pneumoniae serotypes varied in the ability to colonize the nasopharynx or cause invasive infection, estimation of the level of cps2A transcript produced by 14 different serotypes showed that strains with high carriage potential (6A, 6B, 9V, 15, 18C, 19F, 23F, and 33) expressed an average of two-fold more cps2A than the invasive serotypes (1, 4, 5, 7F, 8 and 14)19. In addition, a detailed comparison of the strain cultured in vitro or isolated from the blood has revealed that transcript level of the cps2A gene is linked to the survival of pneumococci in the blood20. Thus, transcriptional regulation of CPS production in response to surrounding stimuli is probably a principal mechanism responsible for pneumococcal pathogenesis.
With the exception of serotype 37, the biosynthesis of CPS is closely dependent on the cps locus which is located between the dexB and aliA genes in S. pneumoniae21,22. Sequences of the cps locus promoter (cpsp) and the first four genes are highly conserved among pneumococcal strains23,24. In contrast, cps2D downstream genes are serotype-specific, which are involved in either polymerization or export of capsular polysaccharide24. By searching the cps promoter-proximal region investigators have revealed some motifs with homology to the binding sites of transcription regulators including RegM, CcpA and GlnR; however, there is no experimental evidence indicating the direct regulation between the regulators and the cps locus25. In addition, Spd0729 was reported to transcriptionally regulate the expression of the cps locus, but the underlying mechanism was not investigated in their work18. Two recent studies have demonstrated the essential roles for the spacing sequence and the core promoter region in transcriptional regulation of the cps locus; specifically, unmarked deletion of spacing sequence or core promoter region in the promoter region led to a significantly reduced cps transcription and hence reduced CPS production17,26. Despite of these observations, transcription factors that can directly bind the cpsp and control CPS production remain uncharacterized, and the role of transcription regulators in pneumococcal pathogenesis remains incompletely understood.
To gain insights into the transcriptional regulation of the cps locus, 5′-biotin labeled cpsp DNA was used to fish out potential transcriptional regulators using DNA affinity chromatography, which led to identification of six potential regulators. In this study, we have performed in-depth molecular analyses of SPD_0064 (referred to as CpsR, the cps locus repressor), and show that CpsR binds directly to the cpsp and hence negatively regulates CPS production. More importantly, we found that the interaction of CpsR with cpsp is further controlled by glucose, which may be one of the mechanisms accounting for the transition from carriage to invasive disease in S. pneumoniae. Fine-tuning expression of CpsR in a ΔcpsR-C strain was used to reveal the impact of CpsR-mediated CPS production on resistance to whole-blood-mediated killing. Finally, we provide evidence that CpsR is required for the pneumococcal nasopharyngeal colonization and systemic infection in mice.
Results
Discovery of the transcriptional regulators that bound to the upstream region of the cps locus
It’s known that the transcription of cps genes is closely dependent on the activity of the cps promoter (cpsp). Wen et al. has recently updated the information regarding the transcription start site (TSS) of the cps locus in S. pneumoniae strain D39, which is located 25 bp upstream from the first nucleotide of the cps2A translation initiation codon17. By searching the promoter-proximal sequence, a previous bioinformatic analysis suggested the existence of several operator sequences similar to the binding sites including of CCPA, CodY, GlnR, and RitR25. However, whether these regulators are involved in transcriptional activation of the cps locus is still lack of experimental evidence. Previous studies demonstrated that the core promoter region, spacing sequence (SS) and repeat sequence of pneumococcus (RUP) are required for the transcriptional regulation of the cps locus17,26. Based on these findings, a 5′-end biotin-labeled cpsp DNA (218 bp in length, from −177 to +41 nt relative to the transcriptional start site) was used to search for the potential regulators of the cps locus using DNA affinity chromatography. DNA-binding proteins were visualized by SDS-PAGE and identified by MALDI-TOF mass spectrometry after tryptic digestion. This screening led to the discovery of six candidate regulators including CcpA, a GntR family transcriptional regulator SPD_0064, a MarR family transcriptional regulator SPD_0379, DNA binding protein HU and DNA gyrase subunit A (Fig. 1). Of these proteins, SPD_0064 was of particular interest because it belongs to the GntR family transcription regulator (Fig. S1) and may have the capacity to bind glucose27. Examination of the localization of SPD_0064 gene revealed that it is located approximately 25,000 nt apart from the cps locus in S. pneumoniae D39. Because SPD_0064 negatively controls the transcription of the cps locus, it was then designated as CpsR (for the cps locus repressor, and hence the cpsR gene). The sequence of CpsR consists of 247 amino acids, and the protein has a calculated mass weight of 28.9 kDa. Like other GntR family regulators, CpsR has two typical domains, including a highly conserved N-terminal helix-turn-helix DNA-binding domain (amino acid residues 13 to 78, Fig. S1A) and a C-terminal regulatory ligand-binding domain (amino acid residues 97 to 237, Fig. S1B). In the following sections, the role and underlying mechanisms of CpsR in cps transcription and pneumococcal pathogenesis are described.
Binding of GntR to the cps locus promoter region
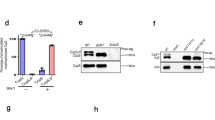
Because CpsR was obtained from the DNA affinity chromatography screening, we hypothesized that CpsR could bind directly to the cpsp DNA fragment. To this end, electrophoretic mobility-shift assays (EMSAs) were carried out. 6His-CpsR protein was produced in E. coli BL21 (DE3) and purified by Ni-NTA affinity chromatography (Fig. 2A). Next, purified 6His-CpsR protein was incubated with the 5′-biotin labeled cpsp probe used in the DNA affinity chromatography screening. Gel shift assays demonstrated that purified 6His-CpsR could bind to the cpsp, showing a dose-dependent pattern (Fig. 2B). Shifted bands were observed when over 1.0 μg of CpsR was added to the reaction mixture. Super-shifted bands were observed at higher CpsR concentrations, which may reflect the formation of dimmers or tetramers of the protein. The incubation of the cpsp DNA fragment with 10 μg of nonspecific protein LytR or nonspecific DNA with CpsR did not result in band shifting (Fig. S2). The mobility shift could be inhibited by coincubation of biotin-labeled cpsp with CpsR and a 400-fold excess of unlabeled cpsp DNA (Fig. 2B), suggesting a specific shift for CpsR. These data provide strong evidence for the binding of CpsR to the cpsp.
(A) The purity of 6His-CpsR protein analyzed with SDS-PAGE. The molecular weight of 6His-CpsR is about 29 kDa. M, mass makers; lane 1, purified 6His-CpsR protein. (B) EMSA of 6His-CpsR protein. The biotin-labeled 218-bp cpsp probe (8 fmol) was incubated with the indicated amounts of 6His-CpsR protein. For the competition analysis, the same but unlabeled (UL) DNA probe was included as 400-fold concentration relative to the biotin-labeled probe. (C) DNase I footprinting protection assay of CpsR. The addition of 3.2 μg 6His-CpsR protected a 33-bp fragment from digestion by DNase I. (D) Structural organization of the cps promoter-proximal region. The CpsR-protected 33-bp DNA sequence is underlined and named binding box, the putative −10 and −35 elements, transcriptional start site (TSS), start codon are in bold. (E) Mutated sequences are either underlined (M7) or in italic (M6). EMSA of 6His-CpsR with wild-type probe (WT) or mutants of the probe (M6 and M7), different amounts of 6His-CpsR protein were incubated with 0.25 μM of the wild-type or mutated probes.
To investigate further the mechanism of CpsR in regulation of the cps locus, we searched for potential CpsR-binding sites in the promoter region of the cps locus using a dye-based DNase I footprinting analysis. As shown in Fig. 2C, in the presence of 6His-CpsR, an apparent region of 33 nt of the cps promoter, 5′-TGTCATGTTCTTATTTCATTTTACTATATTTTT-3′, was protected against DNase I digestion, spanning from nucleotide −146 to −114 relative to the TSS of the cps locus in S. pneumoniae D39 (Fig. 2D). To confirm binding of CpsR to the binding site obtained from DNase I footprinting assays, we mutated the CpsR-protected region with the indicated nucleotides. Wild-type and mutated probes were synthesized and 5′-biotin labeled for EMSA with purified 6His-tagged CpsR. As shown in Fig. 2E, wild-type probe was obviously retarded; whereas the mutated fragments showed less interaction, particularly with the M7 probe. These findings indicate that the binding site is crucial for the specific interaction between CpsR and the promoter of the cps locus.
Effect of cpsR gene deletion on transcription of the cps locus and CPS production
To gain insight into the function of cpsR, we constructed D39ΔcpsR mutant using insertion-deletion mutagenesis strategy. The cpsR gene was deleted from the chromosome of S. pneumoniae by erm cassette replacement, which was a nonpolar event verified by reverse transcription (RT)-PCR. An ectopically complemented strain (ΔcpsR-C) was constructed by integrating the pJWV25-cpsR plasmid via double cross-over into the bgaA locus in D39ΔcpsR strain. This resulted in GFP + -CpsR expression under the control of the Zn2+. Sequencing of the respective genome regions confirmed that the mutations or insertions generated as expected. Immunoblots with murine polyclonal CpsR antiserum revealed the production of CpsR in wild-type D39 strain, and the absence of CpsR in D39ΔcpsR (Fig. 3A). As expected, the expression of CpsR could be well fine-tuned by the addition of Zn2+ in growth medium of the ΔcpsR-C strain (Fig. 3A).
(A) Detection of CPS production in D39 and its derivatives by dot blotting. A representative dot blot image is shown. WT: wild-type D39 strain; M: D39ΔcpsR; Compl: complemented strain. HI: complemented cells were grown in medium supplemented with 0.15 μM ZnCl2; LO: complemented cells were grown in normal C+Y medium. Proteins in lysate of D39, ΔcpsR, and Compl were probed with either polyclonal murine anti-CpsR or anti-GADPH serum. CPS was probed with rabbit anti-serotype 2 serum. The numbers on the left side indicate the dilutions of the sample. (B) Quantitative CPS levels were determined by ELISA. Bacteria were lysed by 0.5% DOC solution and then 1: 20 diluted by PBS before being subjected to ELISA assays. Results are mean (SD) from three independent experiments. **P < 0.01. (C) Representative TEM images of wild-type D39 and ΔcpsR mutant. The bar (magnification is 120,000) indicates 100 nm. (D) Capsule thickness (nm) measured with ImageJ software. The data are expressed as means ± S.D. **P < 0.01. Statistical difference was determined by unpaired two-tailed Student’s t test. (E) Quantitative ELISA analysis of CPS levels in the supernatants of cultures of bacteria. Results are mean (SD) from three independent experiments. **P < 0.01. Statistical difference was determined by unpaired two-tailed Student’s t test.
Immnoblotting and ELISA assays were performed to determine the production of CPS in lysates of D39, D39ΔcpsR, and ΔcpsR-C. Wild-type D39 and D39ΔcpsR strains were grown in normal C+Y medium. For ΔcpsR-C strain, cells were grown in either normal C+Y medium (8 μM ZnCl2) to have a phenotype of CpsRLOWCPSHIGH (ΔcpsR-C-LO) or C+Y medium supplemented with high concentration of ZnCl2 (0.15 mM) to have a phenotype of CpsRHIGHCPSLOW (ΔcpsR-C-HI). Strains were grown to identical culture densities of OD600 = 0.4. Bacteria were harvested, re-suspended in lysis buffer, and then subjected to immunoblotting assays using rabbit anti-serotype 2 serum and Western blotting using polyclonal mouse anti-CpsR sera. GADPH was used as an internal control. Dot blot results showed an increased production of CPS in D39ΔcpsR mutant compared to the wild-type D39 strain (Fig. 3A). Compared to ΔcpsR-C cells grown in normal C+Y medium, ΔcpsR-C cells grown in C+Y medium supplemented with 0.15 mM ZnCl2 produced an increased expression of CpsR but a reduced amount of CPS (Fig. 3A). To confirm the immunoblot results, we directly quantified the amount of CPS of the strains by ELISA. We observed a significantly increased CPS production for D39ΔcpsR mutant than the wild-type D39 strain (Fig. 3B). Also, ΔcpsR-C-LO cells produced significantly more CPS than ΔcpsR-C-HI cells, indicating a reverse relationship between CPS production and CpsR expression.
We further examined the capsule thickness of wild-type D39 strain and ΔcpsR mutant using transmission electron microscopy. The result demonstrated that the capsules of the ∆cpsR mutant were thicker than those of wild-type strain (Fig. 3C). The mean capsule thickness for wild-type D39 strain and D39∆cpsR mutant were 20.5 nm and 33.75 nm respectively (Fig. 3D). The statistical result showed that the average capsule thickness of the ∆cpsR strain was significantly increased compared with that of the wild-type strain (Fig. 3E).
To explore whether the effect of CpsR on cps gene expression was influenced by host strain, cpsR deletion mutant was constructed in a serotype 19F strain (CMCC(B)31693). The 19FΔcpsR mutant also exhibited an increased CPS expression relative to the wild-type 19F strain, as judged by immunoblotting and ELISA assays (Fig. S3). To rule out the possibility that the reduced CPS production is caused by an impaired anchorage of CPS to the cell surface, the amount of CPS in the culture supernatants were tested. The result showed a similar increased CPS in the culture supernatant of the D39ΔcpsR strain (Fig. 3E), confirming that CpsR could negatively control CPS production.
To determine whether the expression of the cps locus is transcriptionally regulated by CpsR, qRT-PCR was performed to determine the relative levels of cps2A-C mRNA in cultures of wild-type D39, D39ΔcpsR grown in C+Y to an OD600 of 0.4. The 16S rRNA levels were used as an internal control. Real time RT-PCR was performed in duplicate on three independently isolated mRNA samples for each strain. In D39ΔcpsR, the cps2A/B/C mRNA levels were up-regulated approximate 6- to 8-fold relative to those in wild-type D39 (P < 0.001, Fig. 4A).
(A) Real time RT-PCR analysis of cps2A-C mRNA. mRNA levels are expressed relative to 16sRNA. Data represents the mean (SD) from three independent experiments each determined in duplicate. ***P < 0.001, analyzed by unpaired two-tailed Student’s t test. (B) The promoter activities of wild-type and mutated cps promoters in S. pneumoniae D39 background. The expression of GFP, driven by the wild-type cpsp (lane 1) or mutated promoters, cpspM6 (lane 2) or cpspM7 (lane 3), was determined by Western blot with monoclonal anti-GFP antibody, and the expression of GADPH was served as loading control.
To address whether the binding site was also important for the transcriptional activity, we constructed transcriptional reporter plasmids by fusing the wild-type promoter (cpspWT) and the mutated promoters (cpspM6 and cpspM7) to the reporter gene gfp respectively. Plasmids were integrated into the cps locus of S. pneumoniae D39 strain via single crossover recombination. Western blotting analyses of GFP were conducted to reflect the activities of the promoters. Immunoblots with polyclonal GFP antibody showed that the expression of GFP was enhanced under the mutated promoters than the wild-type promoter, particularly with the cpspM7 (Fig. 4B). These results indicate that CpsR could directly bind to the promoter region of the cps locus and the 33 nt sequence is essential for the binding of CpsR with the cpsp.
Together these data indicate that the transcription of the cps locus is negatively and directly regulated by the CpsR in S. pneumoniae.
CpsR is a glucose sensing regulator
To test whether glucose could be a signal trigger in controlling cps transcription, wild-type D39 strain was grown in C+Y medium supplemented with different concentrations of glucose. The result showed that CPS production of the wild-type D39 strain was increased with the increasing of glucose concentrations. In contrast, alternation in CPS production was not observed for D39ΔcpsR mutant grown in media supplemented with concentrations of glucose tested in this study (Fig. 5A). As can be seen in Fig. 5A, the addition of glucose has a relatively little influence on the expression of CpsR. To further explore molecular mechanisms associated with glucose in regulation of CPS expression, we tested whether the addition of glucose influences the binding of CpsR with its operator using gel shift assays. A serially 10-fold-increased concentration of glucose was added into the EMSA reaction mixture. Gel shift assays showed evident liberation of the cpsp DNA from the CpsR-cpsp complex at glucose concentrations above 0.5 mM (Fig. 5B). These in vitro data indicate that the glucose can control CPS production, mainly by interfering with the binding of CpsR with the cpsp.
(A) wild-type D39 strain and D39ΔcpsR mutant were grown in C+Y media supplemented with the indicated concentrations of glucose. Bacteria were collected for Western blots of CpsR and dot blots of CPS production. CpsR and GADPH were probed with murine anti-CpsR or anti-GADPH serum, respectively. CPS was probed with rabbit anti-serotype 2 serum. (B) EMSA of CpsR-cpsp complex in the presence of the indicated concentrations of glucose. Unbound DNA is indicated with an arrow.
Role of CpsR in resistance to the whole-blood killing
Given the apparent role of CpsR in regulation of CPS production, we next determined the contribution of CpsR to pneumococcal survival with mouse whole blood. For comparative purposes, the experiment was undertaken with ΔcpsR-C cells whose CPS production could be controlled by Zn2+ in the culture medium. ΔcpsR-C cells were grown in normal C+Y medium (referred to as ΔcpsR-C-LO, a CpsRLOW-CPSHIGH phenotype) or C+Y medium supplemented with 0.15 mM ZnCl2 (referred to as ΔcpsR-C-HI, a CpsRHIGH-CPSLOW phenotype) to generate cells with apparent different expression of CpsR and CPS in the same genetic background. Then, bacterial cells (~120 CFU) in 10 μl were incubated with 90 μl mouse heparinized whole blood and survival rates of the strains were determined. As shown in Fig. 6, 41.8%, 37.1% and 29.1% of ΔcpsR-C-LO cells survived from killing by mouse blood following 1-, 2- and 3-h incubation, respectively. On the contrary, only 15.6%, 10.6% and 13.1% of ΔcpsR-C-HI cells survived from the killing following 1-, 2- and 3-h incubation, respectively. Thus, overall, there was approximately 2- to 3-fold reduction in survival of the ΔcpsR-C-HI variant compared to the survival of the ΔcpsR-C-LO variant. These results indicate that down-regulation of CpsR increases the pneumococcal resistance to whole-blood-mediated killing.
Survival for ΔcpsR-C-LO cells (a CpsRLOW-CPSHIGH phenotype) or ΔcpsR-C-HI cells (a CpsRHIGH-CPSLOW phenotype) following incubation with mouse whole blood. Bacteria (~120 CFU) in 10 μl were added to mouse heparinized whole blood (90 μl), then mixed and incubated for 1, 2, or 3 h at 37 °C. Following incubation, the number of CFU was determined. Data represents the mean (SD) (n = 5). *P < 0.05. Statistical difference was determined by unpaired two-tailed Student’s t test.
The role of CpsR in systemic virulence and nasopharyngeal colonization
As mentioned above, the down-regulation of cpsR resulted in increased CPS production which confers resistance to whole-blood-mediated killing. Therefore we sought to assess whether CpsR expression has impact on pneumococcal colonization and virulence. To determine whether CpsR is required by S. pneumoniae to colonization, groups of 10 Balb/c mice were inoculated intranasally with wild-type D39, D39ΔcpsR, or ΔcpsR-C-HI (a CpsRHIGH-CPSLOW phenotype) at a dose of 106 CFU. Five mice from each group were killed after 16 and 24 h, and the numbers of bacteria recovered from nasopharynx were determined and compared. Compared to wild-type D39 strain, ΔcpsR mutant showed a significant decrease in numbers of pneumococci colonizing the nasopharynx at 16 h (two-tailed Student’s t test, P < 0.01; Fig. 7A). One day after the inoculation, ΔcpsR mutant was also reduced in colonization compared to wild-type strain, despite not showing any statistically significant difference. Furthermore, a significant decrease in bacterial load of the nasopharynx was observed at both 16 h and 24 h postinoculation with the ΔcpsR-C-HI variant compared with the wild-type D39 strain. These results suggest that CpsR-regulated encapsulation is important for pneumococcal colonization of the nasopharynx in mice.
(A) Mice were challenged with 106 CFU and pneumococci were recovered from the nasopharynx 16 or 24 h after intranasal infection of Balb/c mice. The graphs show log10 CFUs (SD) recovered from nasopharynx of five Balb/c mice for D39, ΔcpsR or ΔcpsR-C-HI (a CpsRHIGH-CPSLOW phenotype). Data are the mean (SD). **P < 0.01; *P < 0.05; ns, not significant. Statistical difference was determined by unpaired two-tailed Student’s t test. Balb/c mice were infected intraperitoneally with 103 cells of S. pneumoniae D39 or their derivates (B) or 107 cells of a serotype 19F strain or the 19FΔcpsR mutant (C), respectively. Survival times were recorded. D39 WT vs ΔcpsR, P = 0.029; D39 WT vs ΔcpsR-C-HI (a CpsRHIGH-CPSLOW phenotype), P = 0.0037; ΔcpsR vs ΔcpsR-C-HI (a CpsRHIGH-CPSLOW phenotype), P < 0.0001. 19F WT vs 19FΔcpsR, P = 0.0057. Statistical difference was analyzed by the log-rank (Mantel-Cox) test.
To investigate the role of CpsR in systemic infection, groups of mice were challenged intraperitoneally with wild-type D39, D39ΔcpsR, or ΔcpsR-C-HI (a CpsRHIGH-CPSLOW phenotype) at a dose of 103 CFU. As shown in Fig. 7B, although mice were eventually die from infection, mice infected with wild-type D39 strain survived much longer than those infected with D39ΔcpsR mutant (Log-rank test, P = 0.029). ΔcpsR-C-HI was less virulent compared to the wild-type D39 strain (Log-rank test, P = 0.0037). In addition, virulence potential was evaluated using a serotype 19F strain (CMCC(B)31693) and 19FΔcpsR mutant. As expected, 19FΔcpsR mutant was much more virulent than wild-type 19F strain in invasive disease (Fig. 7C; Log-rank test, P = 0.0057). Overall, consistent with the results obtained from whole-blood-mediated killing assays, the results demonstrate that the capsule increases pneumococcal virulence in the mouse sepsis model.
Discussion
Capsular polysaccharide plays important roles in S. pneumoniae disease processes. Transcriptional regulation of the cps locus is recognized to be an efficient way to modulate pneumococcal CPS production17,26. However, regulators involved in the direct regulation of cps transcription have not been identified. In the present study, we report for the first time that cps transcription is negatively and directly regulated by a GntR family regulator, CpsR. Furthermore, our results show that glucose may act as a signal trigger from the extracellular milieu to activate cps gene expression, which may be a crucial aspect of invasive pneumococcal disease. Finally, we provide evidence that CpsR-regulated CPS production is important for both pneumococcal colonization and invasive disease.
In the present study, DNA affinity chromatography and MALDI-TOF mass spectrometry were used to search for and identify possible transcriptional regulators of the cps locus. This is a well established method to fish for regulators that potentially involve in regulation of the target genes27,28. The present work further confirms the efficiency of this strategy by showing the identification of six possible regulators for the cps locus.
The GntR family has a large number of transcriptional regulators that contain a conserved N-terminal helix-turn-helix DNA-binding domain and a C-terminal ligand regulatory domain. For example, V. cholerae FadR, a GntR family member, has typical N-terminal DNA binding domain and a C-terminal ligand regulatory domain29. Like V. cholerae FadR, S. pneumoniae CpsR has the similar general domains. By sequence alignment using Clustal W method, we observed that amino acid sequences of CpsR spanning residues Y17 to F76 showed 33.3% identity to the region of V. cholerae FadR (Y15 to F74), suggesting that S. pneumoniae CpsR may also bind to the specific binding site as a homodimer. This notion was supported by gel shift results showing the existence of super shift bands (Fig. 2B). A typical mirror repeat, 5′-TTATTTCATT TTACTATATT-3′, was found in the binding site, which may provide binding sequence basis for CpsR to be a homodimer after binding to its operator. Nevertheless, structural study of CpsR and its complexes with the operator need to be further investigated to address this hypothesis.
The upstream sequence of the cps2A start codon (promoter-proximal region) is generally composed of some insertion elements, a repeat unit of pneumococcus (RUP), the spacing sequence (SS) and the core promoter (−10 and −35 boxes) in the majority of the pneumococcal strains17. Work by Shainheit et al. has revealed the essential role of the core promoter of the cps operon (between the −35 box and the transcription start site of the cps operon) in transcriptional regulation of capsule level26. Wen et al. confirmed their results and revealed further the requirement of the SS sequence in transcription regulation of capsule locus17. In this work, we also show the importance of the region spanning RUP and SS sequences in transcription regulation of the cps locus in S. pneumoniae, extending our knowledge on the importance of the upstream sequence in regulation of capsule biosynthesis.
It is clear that transcriptional regulation has a major influence on adaption of S. pneumoniae to changing host environments during infection. Several studies have demonstrated the changes in the expression of virulence factors following the invasive disease14,16,30. The molecular mechanisms underlying S. pneumoniae carriage and virulence remain largely incomplete. A better understanding of how this pathogen controls its gene expression, particularly those having critical roles in invasive disease, is essential to design more effective treatment and prophylaxis strategies against pneumococcal infection. Previous studies have demonstrated the contribution of transcriptional regulators in virulence but were partially characterized5,31. For example, S. pneumoniae two-component signal transduction systems have been reported to play a role in the ability of S. pneumoniae to adapt to changing environments5. The current study extends our understanding on the regulation mechanism of pneumococcal pathogenesis showing that CpsR is required for the transcription of the cps locus and CPS production.
Like GntR family transcription factor PckR, S. pneumoniae CpsR also has an UbiC transcription regulator-associated domain in its C-terminal region, and this domain was proposed to have the capacity to bind small molecules including glucose27. It has been reported in V. cholerae that, in the presence of long chain fatty acyl-CoA, FadR will activate the plsB transcription by detaching itself from its operator sites29,32. Thus, we hypothesized that glucose may work in the same way. To this end, we incubated CpsR with the cpsp in the presence of glucose and gel shift assays were used to show the role of glucose on CpsR-cpsp binding. We found that the cpsp-binding activity of CpsR was influenced by glucose in a dose dependent pattern, suggesting that glucose may interact with CpsR to control its repressor activity.
Recent clinical and epidemiological data suggest respiratory virus infections are important risk factors for acquisition of pneumococcal disease14. Host immune modulation following the viral infection has been shown to contribute to secondary pneumococcal infection33,34, but investigation was seldom performed from the perspective of pathogen itself. Recently, work by Siegel and coworkers has shown that influenza-induced release of sialic acid contributes to pneumococcal growth and colonization, underscoring the importance of nutrient substances in the following pneumococcal infection35. Glucose is another important nutrient factor, which is not normally present in airways secretions but appears in patients with hyperglycemia or having viral infection who were at high risk of developing pneumonia36,37,38,39. It has been reported that glucose can augment bacterial growth by acting as an energy source in the respiratory tracts3,40; however, this alone seems not enough for colonizing pneumococci to cause invasive infection, since Sanchez et al. demonstrated that pneumococci in biofilm were not able to cause invasive disease15. We therefore hypothesized that glucose may serve as a host signal to trigger pneumococci modulating virulence gene expression and hence adapting to different microniches. In the present study, we show that the amount of CPS was increased with increasing of glucose concentrations in wild-type D39 strain (Fig. 5A), suggesting glucose is a positive regulator for CPS biosynthesis in S. pneumoniae. This was confirmed by band shift assays showing that the interaction of CpsR-cpsp could be interfered with the addition of glucose (Fig. 5B). These findings implicate the important carbon source, glucose, in regulating the pneumococcal virulence by controlling CpsR-mediated cps gene expression. In addition, our findings may be used to explain why S. pneumoniae cps gene can be differentially expressed in changing in vitro and in vivo environments.
It is of note that despite we named ΔcpsR-C as the complemented strain, as shown in Fig. 3A, the phenotype of ΔcpsR-C variant was determined by the Zn2+ concentration in growth conditions. ΔcpsR-C-HI variant produced less capsule than the wild-type D39 strain before challenge of mice and the concentrations of Zn2+ may not be optimal to sustain cps expression after inoculation, thus, the phenotype of the complemented strain (ΔcpsR-C-HI) does not fully reflect that of the wild-type D39 strain (Fig. 7A,B). In the present study, ΔcpsR-C-HI variant colonized the nasopharynx at a significant lower level than the wild-type D39 strain. This might be unexpected because a reduced capsule should facilitate interaction between pneumococci and host cells, and therefore increased numbers should be present in the nasopharynx of mice. However, this observation may be at least partially explained by the impairment of ΔcpsR-C-HI cells in resistance to host killing (Fig. 6). In addition, our results show that ΔcpsR mutants were reduced in their ability to colonize the nasopharynx in comparison with the wild-type parents. One possible explanation could be that thick capsule may preclude the interaction between pneumococci and cells41. In consistent with the in vitro result that increased CPS production is linked to a greater resistance to whole-blood-mediated killing (Fig. 6), ΔcpsR mutants are increased in virulence in systemic infection models (Fig. 7B,C). In this sense, upregulation of CPS production seems to be advantageous for invasive pneumococcal infection because of its protective effect against host killing. Taken together, our data suggest the importance of CpsR-regulated encapsulation in both colonization and invasive infection.
In conclusion, our results provide strong evidence that pneumococci could modulate cps gene expression to adapt to changing host environments through CpsR-cpsp interaction (Fig. 8). Given the importance of CPS in capsule production and invasive pneumococcal disease, it is rational that the development of small molecular inhibitors targeting the CpsR-mediated encapsulation might be a promising approach against pneumococcal infection.
In the absence of glucose, the cps transcription was inhibited by the CpsR. In the presence of glucose, glucose may lead to conformation changes of CpsR and hence facilitate the transcription of the cps locus. IE: insertion element; RUP: repeat sequence of pneumococcus; SS: spacing sequence; Core: core promoter; SC: start codon.
Methods
Bacterial strains and growth conditions
All strains and plasmids are listed in Table S1. Unless stated otherwise, S. pneumoniae D39 strain (serotype 2), CMCC31693 strain (serotype 19F) and their derivatives were grown in semi-synthetic casein hydrolysate medium supplemented with 5% yeast extract (C+Y, pH 7.0) medium or blood agar plates at 37 °C under 5% CO2. Escherichia coli strains were grown in Luria-Bertani (LB) broth with shaking or on LB agar plates at 37 °C. Selective antibiotics were added when necessary.
DNA affinity purification
Primers were synthesized by Takara Bio, Dalian and listed in Table S2. The forward primer (Pulldown-cps-F) was 5′- labeled with biotin. DNA affinity purification was carried out essentially as described elsewhere28. Briefly, the 218-bp biotinylated cps promoter fragment (extending from 204 bp upstream of 14 bp downstream of the cps locus start codon) was amplified from chromosomal DNA of the Streptococcus pneumoniae D39 strain using primers Pulldown-cps-F/R2. The resulting 218-bp DNA fragment was bound to streptavidin-coated magnetic beads (DynabeadsM-280 Streptavidin; Invitrogen, Darmstadt, Germany), followed by incubation with crude extracts obtained from S. pneumoniae D39 WT cells grown in the C+Y medium. The eluted proteins were separated by SDS-PAGE, stained with Coomassie, and identified by mass fingerprinting. Nonspecifically bound proteins were removed by several low-salt washing steps (0.1 M NaCl) with subsequent magnetic separation. Specifically bound proteins were eluted with buffers containing 0.3 M, 0.5 M NaCl and subjected to SDS-PAGE and Coomassie staining.
Electrophoretic mobility-shift assays (EMSA)
The DNA fragments used in electrophoretic mobility shift assays (EMSAs) were generated either by PCR using the corresponding oligonucleotides listed in Table S2 in the supplemental material or by annealing two complementary oligonucleotides by incubating the min an equimolar ratio in annealing buffer (10 mM Tris-HCl [pH 8.0], 50 mM NaCl, 1 mM EDTA) for 5 min at 95 °C, followed by cooling at room temperature. LightShift Chemiluminescent EMSA Kits were purchased from Thermo Scientific. EMSAs were performed according to the manufacturer’s instructions and as described elsewhere with modification42. Briefly, EMSA reactions were performed in 20 μl total volumes, containing indicated concentrations of biotinylated DNA probe in buffer containing 50 ng/μl poly (dI–dC). Reaction mixes were incubated for 20 min at room temperature. After incubation, the reaction solution was mixed with 5 μl of the sample loading dye and 20 μl of the solution was subsequently loaded onto pre-cast 6% acrylamide gels (Invitrogen) and electrophoresed at 100 V for 90 min. DNAs were transferred to nylon membranes (Pierce), and cross-linked by exposure to ultraviolet (UV) light. Labeled DNAs were detected using chemiluminescent nucleic acid detection kits (Pierce).
The binding fragment of the 33-mer DNA oligonucleotides was mutated to the indicated sequences, labeled as M6 and M7 (Table S2). Double-stranded DNA were prepared as described elsewhere by annealing complementary oligonucleotides (100 mM each) in 10 mM Tris-HCl, pH 8.0, 20 mM NaCl, heating the reaction to 95 °C for 5 min and allowing it to cool to 25 °C43. Wild-type probe or mutated probes (0.25 μM) were incubated with different amounts of purified 6His-CpsR in a 20 μl buffer as that used above for EMSA. In the assays for effects of glucose on the interaction of CpsR and DNA, wild-type probe of 8 fmol was preincubated with 8.0 μg CpsR at room temperature for 15 min before mixing with different concentrations of glucose, followed by the same procedure as in the other assays.
DNase I footprinting assay
DNase I footprinting assays were performed as described elsewhere by Wang et al44. Briefly, the cps promoter-proximal PCR product was cloned into the HincII digested pUC18H vector (Tolo Biotech). The obtained plasmid was verified via DNA sequencing and used as the template for further preparation of fluorescent FAM labeled probes with primers M13F-47(FAM) and M13R-48. The FAM-labeled probes were purified by the Wizard SV Gel and PCR Clean-Up System (Promega, USA) and were quantified with NanoDrop 2000C (Thermo, USA). For each assay, 400 ng probes were incubated with different amounts of CpsR in a total volume of 40 μl. After incubation for 30 min at 25 °C, 10 μl solution containing about 0.015 unit DNase I (Promega) and 100 nmol freshly prepared CaCl2 was added and further incubate for 1 min at 25 °C. The reaction was stopped by adding 140 μl DNase I stop solution (200 mM unbuffered sodium acetate, 30 mM EDTA and 0.15% SDS). Samples were firstly extracted with phenol/chloroform, then precipitated with ethanol and the pellets were dissolved in 30 μl MiniQ water. The preparation of the DNA ladder, electrophoresis and data analysis were the same as described before44, except that the GeneScan-LIZ500 size standard (Applied Biosystems) was used.
Construction of the mutant and complemented (C) Strains
Primers and sequences are listed in Table S2. cpsR (SPD_0064) gene was deleted from the chromosome of S. pneumoniae by insertion-deletion mutagenesis which was performed essentially according to our established protocol45. Recombinants were selected on blood agar plates containing erythromycin. Successful replacement by the erm cassette was confirmed by sequencing chromosomal DNA in the deleted region.
The full length cpsR was amplified from genome DNA of S. pneumoniae D39 with primers cpsR-SpeI-F/cpsR-NotI-R (Table S2), respectively. The resulting fragment was cloned into plasmid pJWV25 after digestion with restriction enzymes Not I and Spe I. A high-fidelity PrimeSTAR DNA polymerase (TaKaRa, Dalian, China) was used to minimize sequence errors during PCR amplification in this study. All PCR amplified sequences were verified by DNA sequencing. The recombinant plasmids were used to transform to obtain the desired strains. All mutations were confirmed by sequencing of chromosomal DNA in the targeted regions.
For pAE03-plasmid based complementation, the fragment including approximately 700 bp upstream of cps2A gene was cloned into plasmid pAE03 with EcoR I and Not I as restriction sites to form cpspWT-gfp transcriptional fusion. For construction of cpspM6-gfp and cpspM7-gfp reporter fusions, cpspWT-gfp fusion was used as template. All primers were listed in Table S2. PCR products generated with Pfu polymerase were digested with 1 U Dpn I at 37 °C overnight. The resulting products were transformed into E. coli DH5α competent cells. Viable mutants were selected on TSA-BA plates supplemented with appropriate antibiotics. Plasmids were used to transform the S. pneumoniae D39. The cloned promoter sequences were confirmed by DNA sequencing.
Overproduction and purification of CpsR protein
The CpsR protein was overproduced in E. coli BL21 (DE3) using the expression plasmid pW28-cpsR and purified by nickel chelate affinity chromatography according to the established protocol46. Expression was induced at an OD600 of 2.0 by addition of 1 mM isopropyl-D-thiogalactoside (IPTG) at 20 °C. Eight hours after induction, cells were harvested by centrifugation, washed once with the appropriate disruption buffer, and disrupted by ultrasonication. The recombinant protein was purified by nickel-nitrilotriacetic acid (Ni-NTA) affinity chromatography (GE Healthcare). Protein concentrations were determined as described above, and the purified proteins were stored at −80 °C until use.
Immunization of Mice
Mice were immunized as described elsewhere with modification47. Briefly, 6His-CpsR antigen was formulated in aluminum hydroxide adjuvant (Alum) to a final ratio of 100 μg of antigen to 1 mg of Alum adjuvant. Balb/c mice were then immunized i.p. with three doses of 10 μg at 14-day intervals. Polyclonal mouse serum was collected by cardiac puncture 7 days after the final immunization and stored at 4 °C until use. Mouse antiserum against GADPH was prepared previously in the laboratory by using methods similar to those described for CpsR antigen.
RNA Extraction and Real-Time RT-PCR
The liquid culture was grown at 37 °C to an OD600 = 0.4 determined using a cell density meter (Ultrospec 10, GE Healthcare formerly Amersham Biosciences). Total RNA was immediately stabilized with RNAprotect bacteria reagent (QIAGEN, Valencia, CA) and then extracted from S. pneumoniae strains by using a Qiagen RNeasy kit according to the manufacturer’s instructions. RNA was resuspended in nuclease-free water in the presence of RNasin RNase inhibitor (Promega). Contaminating DNA was digested with DNase I (Roche Diagnostics). Real-time RT-PCR was performed on a Rotorgene RG-2000 (Corbett Research, Mortlake NSW, Australia) and as described elsewhere5. Briefly, the cDNA (20 ng/reaction) was used for real-time amplification using the primers listed in Table S2. The mRNA level of 16S rRNA was used as an internal control in order to normalize all data. The amounts of transcripts were expressed as the n-fold difference relative to the control gene (2−ΔCT, where ΔCT represents the difference in threshold cycle between the target and control genes).
SDS-PAGE and Western Blotting
Samples were subjected to SDS-PAGE, and separated samples were then electroblotted onto nitrocellulose. After transfer, the membrane was probed with polyclonal murine antisera or antibodies at a dilution of 1:3,000 and then reacted with blotting grade goat anti-mouse IgG alkaline phosphatase conjugate (BioRad). Labeled bands were visualized by Western ECL reagent from Millipore according to the instructions of the manufactures (ChemiDOC XRS+, BIO-RAD).
Transmission electron microscopy (TEM)
Cells were grown in C+Y medium at 37 °C to the mid-exponential phase. Bacteria were harvested by centrifugation at 4 °C for 10 min at 10,000 × g. Samples were fixed in 2% glutaraldehyde in sodium cacodylate buffer (pH 7.4) for 12 h and then processed by the Electron Microscopy Research Service of Chongqing Medical University. For visualization, cells were sectioned and imaged with a Hitachi H-7500 transmission electron microscope. Image J software was used to determine capsule thickness of 20 randomly chosen bacteria.
Immunoblotting and ELISA
Pneumococcal CPS was assayed by immunoblotting as described elsewhere with modification17. Briefly, one ml of bacterial cultures of the same optical density of 0.4 were pelleted by centrifugation and resuspended in 0.5 ml of DOC solution for 10 min before being used for CPS immunoblotting. The type-2 CPS was detected with a rabbit anti-serotype 2 serum (States Serum Institut, Denmark) and peroxidase-conjugated goat anti-rabbit IgG (H+L) at dilutions of 1: 10000. Binding of the CPS-antibody was visualized by Western ECL reagent from Millipore according to the instructions of the manufactures (ChemiDOC XRS+, BIO-RAD). The results of representative experiments are presented.
Whole-blood killing assays
Whole-blood-mediated killing assays were carried out as described previously with modification18. Bacterial cultures (OD600 = 0.4) of the indicated strains were washed twice and resuspended in PBS to generate a suspension of about 1 × 105 CFU/ml. Whole blood from adult Balb/c mice was collected and heparinized. Ten μl bacterial suspensions were mixed with 90 μl heparinized mouse blood and incubated at 37 °C with shaking (180 rpm) for 1, 2, and 3 h, at which time samples were plated on a blood agar plate for enumeration of surviving CFU.
Animal studies
All animals were purchased from the Laboratory Animal Center of Chongqing Medical University (certificate number: SYXK (yu) 2007-0001). All experimental protocols were approved by the Ethics Committee of Chongqing Medical University. The research was carried out in accordance with the Declaration of Helsinki and with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. To investigate the role of CpsR-mediated CPS production in nasopharyngeal colonization, 6- to 8- week old female Balb/c mice were randomly divided into designed groups. Bacteria were grown in C+Y medium to an OD600 of approximately 0.4. Mice were anesthetized with pentobarbital sodium (1.5%) prior to challenge by intranasal inoculation with 106 CFU. Bacteria were recovered from mouse nasopharynx at 16 and 24 h, and plated on blood agar plate after appropriate dilutions. For systemic virulence evaluation, 6- to 8- week old female Balb/c mice were intraperitoneally infected with 103 cells of S. pneumoniae D39 or their derivates. To evaluate whether the role of CpsR in systemic virulence was influenced by host strain, groups of 10 Balb/c mice were intraperitoneally infected with 107 cells of 31693 strain (serotype 19F) or the ΔcpsR mutant, respectively. Mice were monitored at approximately 4-h intervals, and the survival time of each mouse was recorded.
Statistical analysis
Statistical difference between groups were compared with Student’s t test, the non-parametric Mann-Whitney or Wilcoxon test using GraphPad Prism 5 (GraphPad Software, San Diego, CA). Survival data were analyzed using the log rank (Mantel-Cox) test. Statistical significance was defined as P < 0.05.
Additional Information
How to cite this article: Wu, K. et al. CpsR, a GntR family regulator, transcriptionally regulates capsular polysaccharide biosynthesis and governs bacterial virulence in Streptococcus pneumoniae. Sci. Rep. 6, 29255; doi: 10.1038/srep29255 (2016).
References
Bogaert, D., de Groot, R. & Hermans, P. W. M. Streptococcus pneumoniae colonisation: the key to pneumococcal disease. Lancet Infect Dis. 4, 144–154, doi: 10.1016/s1473-3099(04)00938-7 (2004).
Walker, C. L. F. et al. Global burden of childhood pneumonia and diarrhoea. The Lancet 381, 1405–1416, doi: 10.1016/s0140-6736(13)60222-6 (2013).
Brennan, A. L. et al. Airway glucose concentrations and effect on growth of respiratory pathogens in cystic fibrosis. J Cyst Fibros. 6, 101–109, doi: 10.1016/j.jcf.2006.03.009 (2007).
Hava, D. L., LeMieux, J. & Camilli, A. From nose to lung: the regulation behind Streptococcus pneumoniae virulence factors. Mol Microbiol. 50, 1103–1110 (2003).
Standish, A. J., Stroeher, U. H. & Paton, J. C. The two-component signal transduction system RR06/HK06 regulates expression of cbpA in Streptococcus pneumoniae . Proc Natl Acad Sci USA 102, 7701–7706, doi: 10.1073/pnas.0409377102 (2005).
Kadioglu, A., Weiser, J. N., Paton, J. C. & Andrew, P. W. The role of Streptococcus pneumoniae virulence factors in host respiratory colonization and disease. Nat Rev Microbiol. 6, 288–301, doi: 10.1038/nrmicro1871 (2008).
Nelson, A. L. et al. Capsule enhances pneumococcal colonization by limiting mucus-mediated clearance. Infect Immun. 75, 83–90. Epub 2006 Nov 2006 (2007).
Hyams, C. et al. Streptococcus pneumoniae resistance to complement-mediated immunity is dependent on the capsular serotype. Infect Immun. 78, 716–725, doi: 10.1128/IAI.01056-09 (2010).
Trappetti, C., Ogunniyi, A. D., Oggioni, M. R. & Paton, J. C. Extracellular matrix formation enhances the ability of Streptococcus pneumoniae to cause invasive disease. PLoS One. 6, e19844, doi: 10.1371/journal.pone.0019844 (2011).
Wartha, F. et al. Capsule and D-alanylated lipoteichoic acids protect Streptococcus pneumoniae against neutrophil extracellular traps. Cell Microbiol. 9, 1162–1171. Epub 2007 Jan 1169 (2007).
Hyams, C., Camberlein, E., Cohen, J. M., Bax, K. & Brown, J. S. The Streptococcus pneumoniae capsule inhibits complement activity and neutrophil phagocytosis by multiple mechanisms. Infect Immun. 78, 704–715, doi: 10.1128/IAI.00881-09 (2010).
Magee, A. D. & Yother, J. Requirement for capsule in colonization by Streptococcus pneumoniae . Infect Immun. 69, 3755–3761, doi: 10.1128/IAI.69.6.3755-3761.2001 (2001).
Hammerschmidt, S. et al. Illustration of pneumococcal polysaccharide capsule during adherence and invasion of epithelial cells. Infect Immun. 73, 4653–4667, doi: 10.1128/IAI.73.8.4653-4667 (2005).
Marks, L. R., Davidson, B. A., Knight, P. R. & Hakansson, A. P. Interkingdom signaling induces Streptococcus pneumoniae biofilm dispersion and transition from asymptomatic colonization to disease. mBio 4, doi: 10.1128/mBio.00438-13 (2013).
Sanchez, C. J. et al. Streptococcus pneumoniae in biofilms are unable to cause invasive disease due to altered virulence determinant production. PLoS One. 6, e28738, doi: 10.1371/journal.pone.0028738 (2011).
Kim, J. O. & Weiser, J. N. Association of intrastrain phase variation in quantity of capsular polysaccharide and teichoic acid with the virulence of Streptococcus pneumoniae . J Infect Dis. 177, 368–377 (1997).
Wen, Z. et al. Sequence elements upstream of the core promoter are necessary for full transcription of the capsule gene operon in Streptococcus pneumoniae strain D39. Infect Immun. 83, 1957–1972, doi: 10.1128/IAI.02944-14 (2015).
Yamaguchi, M. et al. Nrc of Streptococcus pneumoniae suppresses capsule expression and enhances anti-phagocytosis. Biochem Biophys Res Commun. 390, 155–160, doi: 10.1016/j.bbrc.2009.09.099 (2009).
Hathaway, L. J., Battig, P. & Muhlemann, K. In vitro expression of the first capsule gene of Streptococcus pneumoniae, cpsA, is associated with serotype-specific colonization prevalence and invasiveness. Microbiol. 153, 2465–2471, doi: 10.1099/mic.0.2006/005066-0 (2007).
Ogunniyi, A. D., Giammarinaro, P. & Paton, J. C. The genes encoding virulence-associated proteins and the capsule of Streptococcus pneumoniae are upregulated and differentially expressed in vivo . Microbiol. 148, 2045–2053 (2002).
Bentley, S. D. et al. Genetic analysis of the capsular biosynthetic locus from all 90 pneumococcal serotypes. PLoS Genet. 2, e31, doi: 10.1371/journal.pgen.0020031 (2006).
Garcia, E., Llull, D., Munoz, R., Mollerach, M. & Lopez, R. Current trends in capsular polysaccharide biosynthesis of Streptococcus pneumoniae . Res Microbiol. 151, 429–435 (2000).
Morona, J. K., Miller, D. C., Morona, R. & Paton, J. C. The effect that mutations in the conserved capsular polysaccharide biosynthesis genes cpsA, cpsB, and cpsD have on virulence of Streptococcus pneumoniae. J Infect Dis. 189, 1905–1913. Epub 2004 Apr 1927 (2004).
Aanensen, D. M., Mavroidi, A., Bentley, S. D., Reeves, P. R. & Spratt, B. G. Predicted functions and linkage specificities of the products of the Streptococcus pneumoniae capsular biosynthetic loci. J Bacteriol. 189, 7856–7876. Epub 2007 Aug 7831 (2007).
Moscoso, M. & Garcia, E. Transcriptional regulation of the capsular polysaccharide biosynthesis locus of Streptococcus pneumoniae: a bioinformatic analysis. DNA Res. 16, 177–186, doi: 10.1093/dnares/dsp007 (2009).
Shainheit, M. G., Mule, M. & Camilli, A. The core promoter of the capsule operon of Streptococcus pneumoniae is necessary for colonization and invasive disease. Infect Immun. 82, 694–705 (2014).
Hyeon, J. E., Kang, D. H., Kim, Y. I., You, S. K. & Han, S. O. GntR-type transcriptional regulator PckR negatively regulates the expression of phosphoenolpyruvate carboxykinase in Corynebacterium glutamicum. J Bacteriol. 194, 2181–2188, doi: 10.1128/JB.06562-11 (2012).
Jutras, B. L., Verma, A. & Stevenson, B. Identification of novel DNA-binding proteins using DNA-affinity chromatography/pull down. Curr Protoc Microbiol. Chapter, Unit1F.1. doi: 10.1002/9780471729259.mc9780471729201f9780471729201s9780471729224 (2012).
Feng, Y. & Cronan, J. E. The Vibrio cholerae fatty acid regulatory protein, FadR, represses transcription of plsB, the gene encoding the first enzyme of membrane phospholipid biosynthesis. Mol Microbiol. 81, 1020–1033, doi: 10.1111/j.1365-2958.2011.07748.x (2011).
Li-Korotky, H. S., Lo, C. Y. & Banks, J. M. Interaction of pneumococcal phase variation, host and pressure/gas composition: virulence expression of NanA, HylA, PspA and CbpA in simulated otitis media. Microb Pathog. 49, 204–210, doi: 10.1016/j.micpath.2010.05.014 (2010).
Giammarinaro, P. & Paton, J. C. Role of RegM, a homologue of the catabolite repressor protein CcpA, in the virulence of Streptococcus pneumoniae . Infect Immun. 70, 5454–5461, doi: 10.1128/iai.70.10.5454-5461.2002 (2002).
van Aalten, D. M., DiRusso, C. C. & Knudsen, J. The structural basis of acyl coenzyme A-dependent regulation of the transcription factor FadR. Embo J. 20, 2041–2050 (2001).
van der Sluijs, K. F. et al. IL-10 Is an important mediator of the enhanced susceptibility to pneumococcal pneumonia after influenza infection. J Immunol. 172, 7603–7609, doi: 10.4049/jimmunol.172.12.7603 (2004).
Cao, J. et al. Activation of IL-27 signalling promotes development of postinfluenza pneumococcal pneumonia. EMBO Mol Med. 6, 120–140, doi: 110.1002/emmm.201302890 (2014).
Siegel, S. J., Roche, A. M. & Weiser, J. N. Influenza promotes pneumococcal growth during coinfection by providing host sialylated substrates as a nutrient source. Cell host & microbe 16, 55–67, doi: 10.1016/j.chom.2014.06.005 (2014).
Wolter, N. et al. High nasopharyngeal pneumococcal density, increased by viral coinfection, is associated with invasive pneumococcal pneumonia. J Infect Dis. 210, 1649–1657, doi: 1610.1093/infdis/jiu1326. Epub 2014 Jun 1646 (2014).
Ehrlich, S. F., Quesenberry, C. P. Jr., Van Den Eeden, S. K., Shan, J. & Ferrara, A. Patients diagnosed with diabetes are at increased risk for asthma, chronic obstructive pulmonary disease, pulmonary fibrosis, and pneumonia but not lung cancer. Diabetes Care. 33, 55–60, doi: 10.2337/dc09-0880 (2010).
Kornum, J. B. et al. Diabetes, glycemic control, and risk of hospitalization with pneumonia: a population-based case-control study. Diabetes Care. 31, 1541–1545, doi: 10.2337/dc08-0138 (2008).
Zhou, H. et al. Invasive pneumococcal pneumonia and respiratory virus co-infections. Emerg Infect Dis. 18, 294–297, doi: 210.3201/eid1802.102025 (2012).
Philips, B. J., Meguer, J. X., Redman, J. & Baker, E. H. Factors determining the appearance of glucose in upper and lower respiratory tract secretions. Intens Care Med. 29, 2204–2210, doi: 10.1007/s00134-003-1961-2 (2003).
Hammerschmidt, S. et al. Illustration of pneumococcal polysaccharide capsule during adherence and invasion of epithelial cells. Infect Immun. 73, 4653–4667, doi: 10.1128/IAI.73.8.4653-4667 (2005).
Hartmann, T. et al. The catabolite control protein E (CcpE) affects virulence determinant production and pathogenesis of Staphylococcus aureus . J Biol Chem. 289, 29701–29711, doi: 10.1074/jbc.M114.584979 (2014).
Davis, J. R., Brown, B. L., Page, R. & Sello, J. K. Study of PcaV from Streptomyces coelicolor yields new insights into ligand-responsive MarR family transcription factors. Nucleic Acids Res. 41, 3888–3900, doi: 10.1093/nar/gkt009 (2013).
Wang, Y., Cen, X. F., Zhao, G. P. & Wang, J. Characterization of a New GlnR binding box in the promoter of amtB in Streptomyces coelicolor inferred a PhoP/GlnR competitive binding mechanism for transcriptional regulation of amtB. J Bacteriol. 194, 5237–5244, doi: 10.1128/jb.00989-12 (2012).
Meng, J. P. et al. Identification of Streptococcus pneumoniae genes specifically induced in mouse lung tissues. Can J Microbiol. 54, 58–65, doi: 10.1139/w1107-1117 (2008).
Liu, Y. et al. Mucosal immunization with recombinant fusion protein DnaJ-DeltaA146Ply enhances cross-protective immunity against Streptococcus pneumoniae infection in mice via interleukin 17A. Infect Immun. 82, 1666–1675, doi: 10.1128/IAI.01391-13 (2014).
Wu, K. et al. Immunization with a combination of three pneumococcal proteins confers additive and broad protection against Streptococcus pneumoniae infections in mice. Infect Immun. 78, 1276–1283, doi: 10.1128/IAI.00473-09 (2010).
Acknowledgements
We thank Jan-Willem Veening for providing plasmids pAE03 and pJWV25 and Donald A. Morrison for providing pEVP3 and S. pneumoniae CPM8. We thank Hong Wang, Jingchuan Fan, and Xiaogang Liao for electron microscopy work. This study was supported by Projects of the National Natural Science Foundation of China (81560324 and 81171532). This work was also supported by Research Fund for the Doctoral Program of Higher Education of China (No. 20135503110004) and scientific research fund of Guizhou province (Qian Ke He J (2015) No. 2151).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: K.W., X.Z. and Y.Y. Performed the experiments: K.W., H.X., Y.Z. and L.W. Analyzed the data: K.W., H.X. and X.Z. Contributed reagents/materials/analysis tools: K.W., H.X., Y.Z. and L.W. Wrote the paper: K.W. and Y.Y. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wu, K., Xu, H., Zheng, Y. et al. CpsR, a GntR family regulator, transcriptionally regulates capsular polysaccharide biosynthesis and governs bacterial virulence in Streptococcus pneumoniae. Sci Rep 6, 29255 (2016). https://doi.org/10.1038/srep29255
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep29255
This article is cited by
-
Synthetic genetic oscillators demonstrate the functional importance of phenotypic variation in pneumococcal-host interactions
Nature Communications (2023)
-
Global transcriptomic analysis of functional oligosaccharide metabolism in Pediococcus pentosaceus
Applied Microbiology and Biotechnology (2021)
-
Identification of a GntR family regulator BusRTha and its regulatory mechanism in the glycine betaine ABC transport system of Tetragenococcus halophilus
Extremophiles (2019)
-
Naturally occurring a loss of a giant plasmid from Mycobacterium ulcerans subsp. shinshuense makes it non-pathogenic
Scientific Reports (2018)
-
Integrated proteomic and metabolomic analysis reveals that rhodomyrtone reduces the capsule in Streptococcus pneumoniae
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.