Abstract
X-linked cone dysfunction disorders such as Blue Cone Monochromacy and X-linked Cone Dystrophy are characterized by complete loss (of) or reduced L- and M- cone function due to defects in the OPN1LW/OPN1MW gene cluster. Here we investigated 24 affected males from 16 families with either a structurally intact gene cluster or at least one intact single (hybrid) gene but harbouring rare combinations of common SNPs in exon 3 in single or multiple OPN1LW and OPN1MW gene copies. We assessed twelve different OPN1LW/MW exon 3 haplotypes by semi-quantitative minigene splicing assay. Nine haplotypes resulted in aberrant splicing of ≥20% of transcripts including the known pathogenic haplotypes (i.e. ‘LIAVA’, ‘LVAVA’) with absent or minute amounts of correctly spliced transcripts, respectively. De novo formation of the ‘LIAVA’ haplotype derived from an ancestral less deleterious ‘LIAVS’ haplotype was observed in one family with strikingly different phenotypes among affected family members. We could establish intrachromosomal gene conversion in the male germline as underlying mechanism. Gene conversion in the OPN1LW/OPN1MW genes has been postulated, however, we are first to demonstrate a de novo gene conversion within the lineage of a pedigree.
Similar content being viewed by others
Introduction
The apo-proteins of the human long-wavelength and middle-wavelength sensitive cone photoreceptor pigments are encoded by the OPN1LW (LW; OMIM 300822) and OPN1MW genes (MW; OMIM 300821), respectively. Arranged in a head-to-tail tandem array on the long arm of the X-chromosome1, these duplicated genes share 98% sequence identity2. The prototypic gene array structure consists of a LW followed by a MW gene, however, considerable variability has been observed in gene copy number in the human LW/MW gene array3,4. Yet, studies on the human retina showed that only the two most proximal genes in the array are expressed at appreciable levels5. Duplicated genes are prone to unequal homologous recombination and gene conversion, the non-reciprocal transfer of genetic information in which a recipient sequence is replaced by a donor sequence that remains unaltered6. Concurrently, gene conversion and recombination themselves, enhance the interchange of DNA variants, thus reducing the divergence between duplicates while expediting a considerably high haplotype diversity7. In accordance with gene variability at the population level, both mechanisms are presumed to occur at the LW/MW gene cluster; nonetheless, they have never been directly observed.
Blue Cone Monochromacy (BCM, OMIM 303700) is an X-linked inherited cone dysfunction disorder. Affected subjects present with colour vision abnormalities and reduced visual acuity, but nystagmus and photophobia are also common8. In this condition S-cone function is retained, while the function of both L- and/or M-cones is absent. In the allelic condition, X-linked Cone Dystrophy (XLCD), cone function is strongly and progressively impaired. Such patients regularly present with myopia and astigmatism and dichromatic or monochromatic colour vision. The two most common genetic causes of BCM are large deletions encompassing the Locus Control Region (LCR) preventing the expression of the LW/MW genes9,10 or the presence of either a single LW/MW hybrid opsin gene or multiple LW/MW opsin genes inactivated by the p.C203R missense mutation9,11. Very few additional disease-causing point mutations or small internal deletions in the LW/MW array have been reported12,13,14,15.
A particular combination of common polymorphisms in LW exon 3 segregating in a single BCM family was first noted by Nathans and colleagues but not considered as disease-causing at that time9. This haplotype, referred to as ‘LIAVA’ in the literature, comprises the following SNPs: c.(453A > G; 457A > C; 465C > G; 511G > A; 513G > T; 521C; 532A > G; 538T > G) and a thereof deduced cone pigment variant with a certain combination of amino acid exchanges: p.[(=); M153L; (=); V171I; A174A; I178V; S180A]. Neither the spectral properties of this variant cone pigment nor its membrane trafficking is altered16. Still the ‘LIAVA’ haplotype has never been reported in individuals with normal colour vision but in patients with incomplete achromatopsia or XLCD17,18 and is associated with widespread alterations of the retinal morphology and disorganized cone structure, different from findings in BCM patients carrying the p.C203R mutation19. In a recent study involving subjects with protan colour vision defects, the ‘LIAVA’ haplotype and other rare exon 3 haplotypes in LW were found to induce exon 3 skipping in vitro20. These findings have lately been corroborated and extended to patients with severe cone disorders including BCM and the identification of two further haplotypes, ‘LVAVA’ and ‘MIAVA’, that impair splicing21.
Hitherto, the interplay of cis-regulatory elements and their contribution to exon 3 splicing is still unclear. To better ascertain to what extent different exon 3 haplotypes lead to splicing aberrations and thereby contribute to severe cone dysfunction disorders, we pursued a semi-quantitative assessment of transcripts from minigene assays performed on a comprehensive set of rare exon 3 haplotypes observed in a total of sixteen families diagnosed with BCM or XLCD. Gene conversion has been proposed as one mechanism underlying the formation of rare exon 3 haplotyopes. However, little is known about the specific features of such gene conversion in the human cone opsin genes. Taking advantage of a family with strikingly different ocular phenotype between the grandfather and grandson, we were able to identify an intrachromosomal de novo gene conversion event in the male germline which results in the replacement of a permissive haplotype by the strongly deleterious ‘LIAVA’ haplotype in the LW gene and explains the severe BCM phenotype in the grandson.
Materials and Methods
Patient recruitment and clinical evaluation
The study was performed in compliance with the tenets of the WMA Declaration of Helsinki. Study participants were recruited ad hoc at different centers specialized in inherited retinal diseases during routine clinical diagnostics. All participants gave written informed consent – approved by the respective local research and ethical review boards - for participation in the study for which blood or DNA samples were sent to Tuebingen for genetic analysis. Procedures of the genetic analysis were approved by the Ethics Committee of the Medical Faculty, Eberhard-Karls University Tuebingen. Patients underwent basic ophthalmologic examination according to the standards of the recruiting centers (for details see Supplementary information).
Genotyping of the LW/MW gene cluster
Genomic DNA was isolated from blood samples according to standard procedures. We analyzed the basic structure and integrity of the LW/MW gene cluster and the absence of the common p.C203R mutation by means of an established PCR and PCR/RFLP protocol. For those subjects having a structurally intact array, LW or MW specific long distance PCRs (LD-PCRs) were performed and LD-PCR products sequenced. For subjects with multiple gene copies, the total number of LW/MW opsin genes was determined by means of real-time quantitative PCR (for details see Supplementary information).
Minigene preparation
The prototype LW opsin minigene construct was kindly provided by Dr. Ueyama (Shiga University, Japan) and used to generate minigene constructs with LW/MW gene variants in exon 3, flanked by its native intronic sequences and the remaining human LW cDNA sequence (for details see Supplementary information).
Transfection and RNA extraction
HEK293 cells at 80–90% confluency were transfected with 4 μg DNA of the minigene construct using 20 μl Lipofectamine 2000 (ThermoFischer GmbH, Dreieich, Germany) per well. 24 h post-transfection, total RNA was extracted (for details see Supplementary information).
RT-PCR and relative quantification
First strand cDNA synthesis was performed using 2 μg of total RNA and random hexamer primers. Subsequent PCR was performed with a 5′ FAM (6-carboxyfluorescein) labeled forward primer and using the QIAGEN Multiplex PCR Kit (Qiagen, Hilden, Germany). FAM-labeled RT-PCR products were diluted 1:10 in water; mixed with 1 μl of GeneScan ROX500 size standard (Life Technologies, Darmstadt, Germany) and 8 μl of Hi-Di Formamide (Life Technologies) in a total volume of 10 μl. Mixes were separated by capillary electrophoresis on an ABI 3130XL Genetic Analyzer instrument (Life Technologies). The area-under-the-curve (AUC) was calculated with GeneMapper 5 (Life Technologies) software. Ratios of splicing products were determined as the AUC for individual peaks divided by the sum of AUC of all differentially spliced products.
Microsatellite analysis
Microsatellite markers locating either centromeric (DXS8011, DXS8103, DXS1356, DXS8087) or telomeric (L441TA, L441CA, AF277A, AF277B and DXS1073) to the LW/MW cluster were genotyped in the three BCM72 family members. Markers for which the mother BCM72-II:1 was heterozygous were used to reconstruct haplotypes.
Mapping of the gene conversion event
LD-PCRs were performed for all three members of BCM72 for the proximal gene copy and for the distal copies. Upon cloning of digested LD-PCR products, we selected two independent clones of the following gene copies for further analysis: (i) LW-derived clones bearing the ‘LIAVA’ haplotype from subjects BCM72-II:1 and BCM72-III:1, (ii) MW-derived clones bearing the ‘LIAVA’ haplotype from subject BCM72-I:1, and (iii) LW-derived clones bearing the ‘LIAVS’ haplotype from subject BCM72-I:1. We sequenced the clones by primer walking (for details see Supplementary information).
Bioinformatic predictions and reference sequences
Results
LW/MW genotypes of subjects with X-linked cone dysfunction disorders
We genotyped the LW/MW gene cluster in twenty-four affected subjects from sixteen families diagnosed with BCM or XLCD not carrying mutations commonly reported in BCM. Eleven families had structurally intact arrays with a proximal LW gene followed by one (n = 6) or multiple copies of the MW gene (n = 5). The remaining five families harboured either a single LW (n = 3) or a single LW/MW hybrid gene (n = 2) (Fig. 1a).
(a) Overview of the various LW/MW gene arrays observed in the study cohort. The LW/MW gene cluster comprises the LCR, represented by a black circle, LW exons depicted as red and MW exons as green boxes, respectively. We observed single LW or LW/MW hybrid arrays and multiple gene arrays with one LW copy and one up to three MW copies. (b) The twelve different exon 3 haplotypes found in this cohort comprise eight common SNPs, depicted by the coding nucleotide position (left) and the corresponding variant amino acid residues (right) and a novel missense variant c.556C > T/p.P186S as part of haplotype 6.
Sequencing of LW/MW exons revealed a high proportion of different (n=12) rare exon 3 haplotypes (Fig. 1b). Table 1 depicts the LW/MW gene array composition and the exon 3 haplotypes for each subject. For consistency we designate haplotypes according to the amino acid residues they encode (i.e. ‘LIAVA’ for p.[L153-I171-A174-V178-A180]). Exon 3 haplotypes comprise two synonymous variants, c.453A > G and c.465C > G. While the c.453A allele was in strict linkage disequilibrium with c.457A/p.V153, we observed both alleles of the variant c.465C > G in cis with c.457A/p.V152. For ease, we distinguish these alleles hereafter with a superscript add-on where appropriate (e.g. MIAVAc.465C versus MIAVAc.465G). Except for a missense mutation c.556C > T/p.P186S found in the LW gene of subject BCM142–21958, all other variants comprised in the haplotypes are per se common in the population of colour normal subjects (minor allele frequency; MAF > 0.05; 22, 23). A large fraction of these haplotypes is rare: seven out of the twelve haplotypes observed here (Table 2) were not present in the population sample investigated by Winderickx and colleagues22. In all our sixteen families the opsin gene harbouring such a rare haplotype occupies the most proximal position with respect to the LCR. In subjects ZD379–19194, BCM101–19818, and BCM126–20616 also the distal gene copies bear rare exon 3 haplotypes.
Minigene assay for rare exon 3 haplotypes
We analyzed exon 3 transcript processing for a total of twelve different exon 3 haplotypes observed in our patient cohort and the ‘MVAIS’ control haplotype by an established minigene assay20. RT-PCR showed three differently spliced products with relative quantities depending on the actual haplotype (Fig. 2a). The 450 bp product is derived from the correctly processed transcript, whereas the two smaller products are derived from aberrantly spliced transcripts either lacking exon 3 (281 bp) or lacking exon 3 and 72 bp of the 3′ end of exon 2 (214 bp), respectively (Fig. 2a). Both aberrant transcripts cause a frame shift leading to a premature termination codon (PTC) in exon 4. To quantify the splicing defect more precisely we performed RT-PCR with a FAM-labeled forward primer and separated the PCR products on a capillary electrophoresis instrument. From the AUC of fluorescence intensity for each fragment we calculated the relative abundance of individual fragments and correctly spliced RT-PCR products (Fig. 2b, Table 2). The control haplotype ‘MVAIS’ yielded no aberrantly spliced products while the ‘LIAVA’ construct resulted in the complete absence of correctly spliced products (Fig. 2b). Three haplotypes, ‘LVAVA’, ‘MIAVAc.465C’ and ‘MIAVAc.465G’ were found to yield only minor amounts (below ∼10%) of correctly spliced products and were classified as strongly deleterious (+++, Table 2). An intermediate fraction (20–50%) of correctly spliced products was obtained with three further haplotypes, ‘LIAIA’, ‘LIAVS’ and ‘MVAVA’ and were considered intermediate (++) in terms of the magnitude of the splicing defect. The remaining five haplotypes, ‘LVAIA’, ‘LVAISS’, ‘MVAIA’, ‘MVVVAc.465G’ and ‘MVVVAc.465C’ yielded predominantly (>75%) correctly spliced products. (Fig. 2b, Table 2) and therefore were considered minor defective (+).
(a) Agarose gel electrophoresis of RT-PCR products obtained with RNA from HEK293 cells transfected with minigene constructs bearing various exon 3 haplotypes. The tested haplotype is given above the corresponding gel lane. A 100 bp ladder size standard was loaded in the leftmost lane. Both lanes ‘MVAIS’ and ‘MVAIS*’ refer to minigenes carrying the control haplotype. ‘MVAIS*’ has a modified Multiple Cloning Site from the prototype construct ‘MVAIS’ (see Supplementary Materials and Methods). NTC: non-template negative control. A scheme on the composition of the RT-PCR products is given on the right. Full length gel picture is presented in Supplementary Fig. S3. (b) Examples of capillary electrophoresis and quantitative analysis of fluorescent labeled RT-PCR products of the minigene assay for four different haplotypes (‘MVAIS’, ‘LVAVA’, ‘LIAVA’ and ‘MIAVAc.465G’). The fragment size scale is given on the x-axis and fluorescence intensity (in arbitrary units) on the y-axis. Relative amounts of each fragment are given for the corresponding peak as determined by Gene Mapper. The three different sized products correspond to correctly spliced transcripts (450 bp), aberrantly spliced transcripts lacking exon 3 (281 bp), and a minor species of aberrantly spliced products lacking exon 3 and further 72 bp of exon 2 (214 bp).
Correlation of LW/MW arrays and splicing defects with clinical presentation
LW/MW gene cluster genotypes were greatly diverse in our cohort of families with clinical diagnosis of BCM or XLCD. One family had an LW/MW gene array made up of a total of four LW/MW gene copies, four families had three copies, six families had two copies and five families carried a single LW, or LW/MW hybrid gene.
All families with single gene arrays carried strongly deleterious haplotypes: ‘LIAVA’ in BCM73 and BCM93, and ‘LVAVA’ in another three families (BCM66, BCM112, and BCM194). Clinical data were available for nine subjects from these families (Supplementary Table S1). Except for one very young subject (BCM194–25474), all these subjects had a best corrected visual acuity (BCVA) of ≤0.3, absent or strongly reduced photopic and flicker ERG responses, and impaired colour vision. Seven out of nine subjects had nystagmus and two experienced mild photophobia. Funduscopy revealed minor macular pigmentary irregularities but were dominated by myopia-related alterations such as optic disc crescents and peripapillary atrophy. All but two subjects in this group were highly myopic. Within this group we noted a tendency that subjects with the ‘LIAVA’ haplotype are more severely affected than those with the ‘LVAVA’ haplotype.
The families with multiple LW/MW opsin genes were more heterogeneous in terms of genotypes and clinical presentation. Clinical data were available for fifteen subjects from eleven families. Current genotyping technologies cannot determine the actual order of multiple distal MW gene copies. Therefore, for ordering of multiple MW gene copies we took into account impaired MW cone function in the patients along with the positional bias of the opsin gene expression5. Three families had identical MW gene copies with respect to exon 3 haplotypes whereas BCM142 and BCM72 had different MW gene copies (BCM142–21958, BCM72–17075 and BCM72–16874, Table 1). In the latter we assume that the MW gene copy harbouring the exon 3 haplotype with the most severe splicing defect occupies the second position in the array. Four families (ZD379, BCM101, BCM126 and BCM72–17075 [BCM72-III:1]) had highly deleterious (+++) haplotypes in all gene copies either being completely (BCM126) or partially isogenic (i.e. all copies with identical haplotypes or two of three gene copies sharing an identical haplotype, respectively). While all subjects in this group were clinically typical for BCM, myopia was rather mild and one was even slightly hyperopic.
The second sub-category of subjects with multiple copies are characterized by a LW gene comprising a strongly deleterious exon 3 haplotype and a single or multiple MW genes carrying exon 3 haplotypes causing intermediate or minor splicing defects. Although the clinical findings of the seven subjects from five families (BCM51, BCM133, BCM160, ZD314, and ZD547) were rather variable (namely visual acuity, photophobia and nystagmus), we consistently noted myopia, impaired colour vision and reduced but never absent cone or flicker ERG responses in these subjects. Consistent with some preserved central, cone-mediated vision there was at least for the better eye a BCVA of ≥0.5 in four of them.
The remaining subjects are less likely to be explained simply by the splicing defect due to exon 3 haplotypes: BCM98–19713 comprises a ‘LIAIA’ haplotype in the LW gene associated with a moderate splicing defect and a common haplotype with minimally compromised splicing in the MW gene. BCM142–21958 only had exon 3 haplotypes that result in minor amounts of aberrant transcripts, but carries a novel missense variant, c.556C > T/p.P186S, in exon 3 of the LW opsin gene which does not compromise splicing, but may impair folding and/or function of the derived photopigment. Notably, an analogous proline to serine substitution at amino acid position 187 has been reported previously in the LW opsin gene of a deuteranope subject24.
Gene conversion results in the deleterious haplotype ‘LIAVA’
We observed a family, BCM72, with rather discordant phenotypes (Fig. 3) between the index subject (BCM72-III:1, Fig. 4a) diagnosed with BCM, and his grandfather (BCM72-I:1, Fig. 4a) who developed macular dystrophy at the age of 40 and presented with a deutan defect. A crucial difference is documented in the cone-derived ERG recordings: Responses in BCM72-I:1 were not impaired, whereas in BCM72-III:1 photopic single-flash and 30 Hz-flicker-responses were extinguished as shown by a flat light-adapted (LA) ERG (Fig. 3a). Rod-derived responses were normal in both cases (Fig. 3a). Subject BCM72-I:1 performed as a deuteranomalous in the Farnsworth Panel D-15 desaturated test while BCM72-III:1 had a majority of confusions along the protan axis (Fig. 3b). A protan-like arrangement of colour discs in the Panel D-15 is frequently observed in BCM subjects8,25. The index subject’s mother (BCM72-II:2) had normal visual acuity and colour vision but borderline reduced photopic ERG responses. The macular dystrophy of subject BCM72-I:1 is evident in the fundus autofluorescence image of his left eye (Fig. 3c, upper image). OCT of the left eye of both subjects is shown in Fig. 3d. Retinal pigment epithelium and mostly the outer cone photoreceptor layers, namely the cone outer segment termination and the inner segment ellipsoid layers, were severely damaged in the macular area in subject BCM72-I:1, but the retinal thickness is also diminished in subject BCM72-III:1.
(a) Fullfield-ERG with nearly normal responses in BCM72-I:I and not detectable photopic responses in BCM72-III:1. DA: dark-adapted, LA: light-adapted, stimulus strengths: 0.01 or 3.0 cd.s.m−2. (b) Panel D-15 desaturated with protan defects in BCM72-I:I, and deutan defects in BCM72-III:I. (c) Autofluorescence and (d) infrared picture (left panel) and OCT (right panel) with macular dystrophy in BCM72-I:I and normal retinal architecture with thinned photoreceptor layer in BCM72-III:I.
(a) Pedigree of family BCM72. Subject BCM72-I:1 presented with macular dystrophy and deuteranopia, subject BCM72-II:2 is an asymptomatic female carrier and subject BCM72-III:1 was diagnosed with BCM. Reconstructed haplotypes based on microsatellite markers encompassing the LW/MW gene cluster on Xq28 revealed inheritance of the X-chromosome from the grandfather to the grandson with no evidence for recombination. (b) Scheme of the structure of the LW/MW gene array and the LW and MW exon 3 haplotypes in crucial members of the BCM72 family. The LW/MW gene array on the transmitted chromosome comprises the locus control region (LCR), a single LW gene and three MW gene copies. Red and green coloured and numbered boxes represent LW exons and MW exons, respectively. Exon 3 haplotypes are indicated below the respective exon boxes. For multiple distinct MW copies, their most likely order with respect to exon 3 haplotypes is depicted. Note that female subject BCM72-II:2 is heterozygous for LW exon 3 haplotypes ‘LIAVA’ and ‘LIAIS’, the latter inherited from her mother. The findings support an intrachromosomal gene conversion event transforming the more ancestral haplotype ‘LIAVS’ into the ‘LIAVA’ haplotype in the germline of subject BCM72-I:1.
Genotyping of the grandson BCM72-III:1 revealed an intact LW/MW gene array with a proximal LW gene and three copies of the MW gene (Fig. 4b). Sequence analysis showed a ‘LIAVA’ exon 3 haplotype for the LW gene and ‘LIAVA’ and ‘MVVVAc.465C’ for the MW gene copies. No further mutations were found in other exons (including exon 6, which is not covered by the LD-PCRs) of the opsin genes of the affected subjects. Considering the expression bias of the opsin gene cluster, and the BCM phenotype of the subject, we reasoned that the ‘LIAVA’-bearing MW gene copy occupies the first position downstream of the LW gene. Surprisingly, the same downstream MW haplotypes but a distinct LW exon 3 haplotype ‘LIAVS’ were found in the grandfather BCM72-I:1. Applying the minigene splicing assay we found that the ‘LIAVS’ haplotype resulted in an intermediate splicing defect with still approximately 20% correctly spliced transcripts (Fig. 5b).
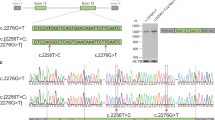
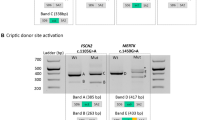
(a) The gene conversion occurred in a ‘telomeric to centromeric’ direction in the X-chromosome of subject BCM72-I:1 with the ‘LIAVA’ bearing MW gene serving as donor and the ancestral ‘LIAVS’ bearing LW gene serving as recipient. Below, the X-chromosome of subject BCM72-III:1 presents the product of the gene conversion event with the ‘LIAVA’ haplotype present on both LW and the most proximal MW as demonstrated by corresponding sequences traces for the c.538T > G variant. Sequencing of cloned LW and MW gene fragments revealed a maximal converted region of 1,297 bp (c.409 + 950_c.578 + 90conNM_000513.2: c.409 + 950_c.578 + 90) covering exon 3 and flanking intron sequences (see also Supplementary Fig. S2). (b) Direct comparison of RT-PCR products from the minigene splicing assays shows a substantial amount of correctly spliced transcripts (450 bp) for the ‘LIAVS’ exon 3 haplotype, whereas such products are undetectable for the ‘LIAVA’ haplotype. Full length gel picture is presented in Supplementary Fig. S4.
Segregation analysis with microsatellite markers flanking the LW/MW gene cluster supported the transmission of the Xq28 segment from grandfather to grandson without evidence for recombination (Fig. 4 and Supplementary Fig. S1), suggesting a de novo gene conversion in the LW/MW gene cluster in this family. We further explored whether the gene conversion event has occurred in trans between X-chromosomes (in the mother’s germline), or in cis between LW and MW gene copies in either the grandfather’s or the mother’s germline. Genotyping of the mother BCM72-II:2 revealed a common ‘LIAIS’ exon 3 haplotype in one of her proximal LW genes and the ‘LIAVA’-bearing haplotype in the other LW gene copy (Fig. 4b). This finding strongly supports transmission of the ‘LIAVA’-bearing LW gene from her father (BCM72-I:1) and from BCM72-II:2 to her offspring and thereby asserts the occurrence of the gene conversion in the grandfather’s germline as a de novo mutation. To define the extent of this intrachromosomal event we sequenced exon 3 and the flanking introns of the putative gene conversion donor (‘LIAVA’-bearing MW gene copy) and recipient gene copy (‘LIAVS’-bearing LW gene copy) in BCM72-I:1 and its product (‘LIAVA’-bearing LW gene copy) in both BCM72-II:2 and BCM72-III:1. By comparative sequencing we narrowed down the size of the maximal converted sequence in the recipient LW to a region of 1,297 bp (c.409 + 950_578 + 90conNM_000513.2:c.409 + 950_578 + 90), which is delimited by the discriminative SNP rs3788802 (c.409 + 949 G > A) in intron 2 and rs369018729 (c.578 + 91 G > A) in intron 3, and includes the discriminating variant c.538 T > G/p.S180A/rs949431 in exon 3 (Fig. 4a and Supplementary Fig. S2).
Discussion
Splicing defects caused by rare LW/MW exon 3 haplotypes
The presence of rare combinations (foremost ‘LIAVA’) of otherwise common coding variants in exon 3 of the LW/MW genes in subjects with X-linked colour vision or cone dysfunction disorders has been noted in several publications9,18,24. Ueyama and colleagues were the first who showed that such rare combinations of variants induce a splicing defect and hence, explain the protan defect in probands with otherwise normal LW gene sequence20. Lately – and during the course of our study – Gardner et al. reported LW/MW gene splicing defects as underlying disease mechanism in nine BCM/XLCD families carrying exon 3 “interchange haplotypes”21. These patients had only a single LW/MW gene copy carrying an exon 3 haplotype inducing a splicing defect or multiple opsin gene copies, in which at least the two most proximal carry such haplotypes. In this study we could corroborate their findings to a broader extent by including a large series of subjects with BCM or XLCD (24 affected subjects from 16 families). We observed an extensive diversity in terms of structure and composition of the LW/MW gene cluster and exon 3 haplotypes. Besides, we explored the functional consequences of twelve exon 3 haplotyes on RNA processing by means of heterologous minigene assays and took advantage of fluorescence-based capillary electrophoresis of RT-PCR products for accurate relative quantification of the proportions of differentially spliced transcripts. Most of the investigated haplotypes resulted in a substantial fraction of mis-spliced transcript with two main aberrant mRNA species that result in a PTC in exon 4. Such transcripts presumably elicit nonsense-mediated mRNA decay26. Accurate quantification of the relative amounts of the different species disclosed clearly distinct ratios of correctly and aberrantly spliced transcripts depending on the haplotype. ‘LIAVA’ as the extreme led to complete exon skipping. The two related haplotypes ‘MIAVAc.465C’ and ‘MIAVAc.465G’, as well as ‘LVAVA’, yielded high levels of aberrantly spliced transcripts but still detectable amounts of correctly spliced transcripts. Notably, we observed the latter haplotype being associated with a less severe clinical phenotype in comparison with patients harbouring ‘LIAVA’. Three further haplotypes, ‘LIAVS’, ‘LIAIA’ and ‘MVAVA’ have an intermediate effect in terms of aberrant splicing (20–50% of correctly spliced opsin), whereas the remaining six haplotypes confer only minor splicing defects. The (patho-)physiological consequences of such intermediate or minor defects on disease expression or colour in general still needs to be explored.
The consequences of genotypes that comprise highly deleterious exon 3 haplotypes in the relevant, most proximal gene copies may approximate the situation of a full null mutation like patients with larger deletions at the LW/MW gene cluster9,12,13,14. Less straightforward are cases with multiple gene copies combining copies with highly deleterious exon 3 haplotypes and copies with haplotypes causing intermediate splicing defects (e.g. LW: ‘LVAVA’+MW: ‘MVAVA’ as in ZD547). The diversity of genotypes and quantitative difference in the magnitude of the splicing defect caused by different haplotypes is reflected by the variability in clinical presentations ranging from a BCM typical phenotype to cone disorders with considerably preserved cone function and rather good visual acuity. This clinical variability is in line with previous reports on subjects with BCM, “atypical BCM”, or XLCD who carry rare LW/MW exon 3 haplotypes9,16,17,18.
Since accurate quantification of the different spliced forms is lacking in prior studies, we could not directly compare their results at the quantitative level. However, a qualitative comparison points out some differences. While in this study as well as in the study of Gardner and colleagues21 the three distinct transcript species were observed, the correctly spliced form, an exon 3-skipped transcript and an additional aberrant transcript, the nature of the latter one seems to differ. This third minor species is produced for instance from the ‘LVAVA’ and ‘MIAVA’ haplotypes (and from ‘LIAVA’ in our study) which in both studies is a result of internal exon 2 splicing but involves different acceptor sites in intron 3 (in the study of Gardner and colleagues) and at the correct intron 3/exon 4 junction (in this study), respectively. Microheterogeneity in terms of minor aberrant splicing products have not only been observed in many heterologous minigene assays but also in native tissue of subjects carrying splicing-inducing mutations27 and may be influenced by technical parameters.
SROOGLE28 predicted several Splicing Regulatory Sequences (SRSs) to be disrupted and/or created when exon 3 SNPs from deleterious haplotypes were interchanged. In fact, every other SNP within the haplotype disrupts or creates at least one SRS. Half of the overall putative Exonic Enhancers of Splicing (EESs) overlapping the eight SNPs were predicted to be disrupted by ‘LIAVA’. For instance, the c.521C > T substitution is foreseen to create a novel EES, which is consistent with the increased correctly spliced transcripts abundance in vitro (Table 2, haplotype 9–10). Comparison of closely related haplotypes indicates that the c.532A/p.178Ile allele confers “protection”: for instance ‘LVAVA’ yielded about 7% of correctly spliced transcripts whereas ‘LVAIA’ about 80% (Table 2, haplotype 4–5). These patterns may suggest the presence of motifs that exert combined enhancer and silencer effects to a certain extent as described in SMN1 and CFTR genes29.
Gene conversion as a mechanism leading to a pathogenic haplotype in the cone opsin array
We here report a de novo gene conversion event in the LW/MW gene cluster which we proved in the lineage of a single family (BCM72). This rare instance further enabled us (i) to discriminate between inter- and intrachromosomal event, (ii) to differentiate between its occurrence in either the male versus the female germline, (iii) to define the directionality of the event (from LW to MW or vice versa), (iv) to determine the actual sequence homology between the donor and recipient prior to the gene conversion event and finally (v) to refine the extent of the converted sequence. These findings are novel since no other de novo gene conversion events in a parent-offspring transmission have been reported for the LW/MW gene cluster so far.
Our study is in congruency with the population-based evidence for gene conversion in the opsin genes as described by Verrelli and Tishkoff23. Gene conversion together with selective forces are proposed to have an influence on the observed variation in LW/MW genes, as mainly advantageous polymorphisms have been spread6,23. However, beyond the evolutionary perspective, gene conversion in the LW/MW gene cluster may also lead to the occurrence of deleterious genotypes that are associated with colour vision deficiencies, XLCD or BCM.
Signatures of gene conversion in the LW/MW gene cluster at the level of individual chromosomes have so far been proposed specifically in the context of BCM subjects with rare point mutations present in all copies of the LW/MW gene cluster13,30, yet the gene conversion event could not be directly pinpointed, hindering a full characterization. For instance, Gardner and colleagues reported a BCM family in which the c. 529T > C/p. W177R missense mutation in exon 3 was observed in both LW and MW genes13. Albeit a gene conversion event was deduced based on the observation that both genes carry this unique mutation and share a block of SNPs in exon 3, occurrence of the gene conversion event (i.e. a family member with the ancestral genotype prior to the event) could not be observed in this family.
Different from these previous reports, we herein were able to study for the first time a gene conversion event at the LW/MW gene cluster within the lineage of a single pedigree (i.e. the genotypes prior to and following the gene conversion event) and, importantly, to associate these two different genotypes to their cognate phenotypes. Furthermore, we could correlate these differences at the phenotypic level prior and post-gene conversion with a quantitative difference in the in vitro levels of correctly spliced LW gene transcripts that results from the distinct exon 3 haplotypes prior and post gene conversion. While ‘LIAVA’ has been previously assessed for splicing20, the ancestral prior gene conversion ‘LIAVS’ haplotype has been reported in the literature18 but has – to our knowledge – never been tested for splicing before.
With respect to directionality of the gene conversion, we could demonstrate in our family that the MW copy acted as donor and LW copy served as recipient gene copy. The same ‘telomeric to centromeric’ direction has been proposed for the “spreading” mutation cases13,30. Seemingly, the position of a recombination hotspot has an influence over the directionality of gene conversion31. The donor MW copy carries the haplotype ‘LIAVA’, which most likely occupies the first position downstream of the LW gene, explaining the deutan defect in BCM72-I:1. The LW gene of this subject carries the exon 3 haplotype ‘LIAVS’, which only differs from ‘LIAVA’ by a single SNP (c.538G > T/p.S180A). Compared to the latter haplotype which results in a fully penetrant splicing defect, the ‘LIAVS’ haplotype produces 20% of correctly spliced transcripts which is perfectly compatible with the much milder phenotype of the grandfather (BCM72-I:1). In contrast, the grandson (BCM72-III:1) presenting with BCM has the two first LW/MW copies carrying the fully deleterious haplotype ‘LIAVA’, consistent with his BCM phenotype. The ERG responses were the most informative clinical feature that documents the phenotypic differences between BCM72-I:1 and BCM72-III:1. The ‘LIAVS’ haplotype has been reported before in affected males of two families with X-linked cone-dominated phenotypes18, who compared to the ‘LIAVA’-carrying individuals from an independent family seemed to have a better visual performance. One of the ‘LIAVS’-carrying individuals presented with evidence of maculopathy, similarly to BCM72-I:1 here, however, BCM72-I:1 developed this maculopathy later in life.
Given the sequence identity between LW and MW, we could only define the outermost borders of the gene conversion event in intron 2 and intron 3 (Fig. 5a) setting a maximal size of about 1,300 bp which is beyond the average size estimates for converted tracts in humans32. Significantly reduced linkage disequilibrium of variants in a region of about 400 bp covering LW exon 3 and flanking intronic sequences supported the presence of a hotspot for gene conversion at this region23 centering around a Chi-like sequence element22. Although gene conversion at a certain site is expected to be a rare event, recent single cell analysis technologies may allow to actually determining the frequency and pattern of such events at the LM/MW locus in human sperm cells33.
While formally a de novo point mutation could be an alternative explanation, we reason that a de novo gene conversion is the most likely event since (i) the very same c.538T> G variant is present in the downstream MW gene copy, (ii) the c.538T > G is a common variant frequently observed in LW genes22, (iii) population studies have revealed strong evidence for frequent gene conversion between LW and MW genes23,34, and (iv) it has been estimated that the de novo rate of gene conversion between paralogous loci in the human genome is approximately 100-fold higher than the de novo point mutation rate35,36,37,38.
The current study, not only supports the predictions of population genetics studies and “spreading” mutation cases on gene conversion in the opsin genes, but also, adds direct evidence on how gene conversion events may arise spontaneously in the germline having an important impact on human disease. Moreover, this type of mutations might be overlooked by employing next-generation sequencing technologies which complicates its identification as well as its prediction.
In conclusion, our study confirms that LW/MW gene array genotypes bearing certain rare exon 3 haplotypes cause BCM and XLCD in a large patient cohort and explains the deleterious functional effect of these haplotypes. Reliable quantitative analysis of splice products has been implemented and revealed that the relative proportions of correctly and aberrantly spliced transcripts obtained for the twelve different haplotypes cluster in three main categories. Lastly, we traced back the origin of the strongly deleterious ‘LIAVA’ haplotype in a BCM subject. This haplotype was inherited from his mother and, in all likelihood arose as a result of a de novo intrachromosomal gene conversion event from an ancestral ‘LIAVS’ in the grandfather’s germline and explains the phenotypic transformation from deuteranopia in the grandfather to BCM in his grandson.
Additional Information
How to cite this article: Buena-Atienza, E. et al. De novo intrachromosomal gene conversion from OPN1MW to OPN1LW in the male germline results in Blue Cone Monochromacy. Sci. Rep. 6, 28253; doi: 10.1038/srep28253 (2016).
References
Vollrath, D., Nathans, J. & Davis, R. W. Tandem array of human visual pigment genes at Xq28. Science 240, 1669–1672 (1988).
Nathans, J., Piantanida, T. P., Eddy, R. L., Shows, T. B. & Hogness, D. S. Molecular genetics of inherited variation in human color vision. Science 232, 203–210 (1986).
Macke, J. P. & Nathans, J. Individual variation in size of the human red and green visual pigment gene array. Invest Ophthalmol Vis Sci 38, 1040–1043 (1997).
Wolf, S. et al. Direct visual resolution of gene copy number in the human photopigment gene array. Invest Ophthalmol Vis Sci 40, 1585–1589 (1999).
Winderickx, J., Battisti, L., Motulsky, A. G. & Deeb, S. S. Selective expression of human X chromosome-linked green opsin genes. Proc Natl Acad Sci USA 89, 9710–9714 (1992).
Chen, J. M., Cooper, D. N., Chuzhanova, N., Ferec, C. & Patrinos, G. P. Gene conversion: Mechanisms, evolution and human disease. Nat Rev Genet 8, 762–775 (2007).
Innan, H. & Kondrashov, F. The evolution of gene duplications: Classifying and distinguishing between models. Nat Rev Genet 11, 97–108 (2010).
Michaelides, M. et al. Blue cone monochromatism: A phenotype and genotype assessment with evidence of progressive loss of cone function in older individuals. Eye (Lond.) 19, 2–10 (2005).
Nathans, J. et al. Molecular genetics of human blue cone monochromacy. Science 245, 831–838 (1989).
Smallwood, P. M., Wang, Y. & Nathans, J. Role of a locus control region in the mutually exclusive expression of human red and green cone pigment genes. Proc Natl Acad Sci USA 99, 1008–1011 (2002).
Kazmi, M. A., Sakmar, T. P. & Ostrer, H. Mutation of a conserved cysteine in the X-linked cone opsins causes color vision deficiencies by disrupting protein folding and stability. Invest Ophthalmol Vis Sci 38, 1074–1081(1997).
Nathans, J. et al. Genetic heterogeneity among blue-cone monochromats. Am J Hum Genet 53, 987–1000 (1993).
Gardner, J. C. et al. X-linked cone dystrophy caused by mutation of the red and green cone opsins. Am J Hum Genet 87, 26–39 (2010).
Ladekjaer-Mikkelsen, A. S., Rosenberg, T. & Jorgensen, A. L. A new mechanism in blue cone monochromatism. Hum Genet 98, 403–408 (1996).
Kellner, U. et al. Blue cone monochromatism: Clinical findings in patients with mutations in the red/green opsin gene cluster. Graefes Arch Clin Exp Ophthalmol 242, 729–735 (2004).
McClements, M. et al. Variations in opsin coding sequences cause x-linked cone dysfunction syndrome with myopia and dichromacy. Invest Ophthalmol Vis Sci 54, 1361–1369 (2013).
Crognale, M. A. et al. Characterization of a novel form of X-linked incomplete achromatopsia. Vis Neurosci 21, 197–203 (2004).
Mizrahi-Meissonnier, L., Merin, S., Banin, E. & Sharon, D. Variable retinal phenotypes caused by mutations in the X-linked photopigment gene array. Invest Ophthalmol Vis Sci 51, 3884–3892 (2010).
Carroll, J. et al. The effect of cone opsin mutations on retinal structure and the integrity of the photoreceptor mosaic. Invest Ophthalmol Vis Sci 53, 8006–8015 (2012).
Ueyama, H. et al. Unique haplotype in exon 3 of cone opsin mRNA affects splicing of its precursor, leading to congenital color vision defect. Biochem Biophys Res Commun 424, 152–157 (2012).
Gardner, J. C. et al. Three Different Cone Opsin Gene Array Mutational Mechanisms with Genotype–Phenotype Correlation and Functional Investigation of Cone Opsin Variants. Hum Mutat 35, 1354–1362 (2014).
Winderickx, J., Battisti, L., Hibiya, Y., Motulsky, A. G. & Deeb, S. S. Haplotype diversity in the human red and green opsin genes: Evidence for frequent sequence exchange in exon 3. Hum Mol Genet 2, 1413–1421 (1993).
Verrelli, B. C. & Tishkoff, S. A. Signatures of selection and gene conversion associated with human color vision variation. Am J Hum Genet 75, 363–375 (2004).
Neitz, M. et al. Variety of genotypes in males diagnosed as dichromatic on a conventional clinical anomaloscope. Vis Neurosci 21, 205–216 (2004).
Weiss, A. H. & Biersdorf W. R. Blue cone monochromatism. J Pediatr Ophthalmol Strabismus 26, 218−223 (1989).
Maquat, L. E. Nonsense-mediated mRNA decay: splicing, translation and mRNP dynamics. Nat Rew Mol Cell Biol 5, 89–99 (2004).
Steffensen, A. Y. et al. Functional characterization of BRCA1 gene variants by mini-gene splicing assay. Eur J Hum Genet 22, 1362–1368 (2014).
Schwartz, S., Hall, E. & Ast, G. SROOGLE: Webserver for integrative, user-friendly visualization of splicing signals. Nucleic Acids Res 37, 189–192 (2009).
Pagani, F. & Baralle, F. E. Genomic variants in exons and introns: Identifying the splicing spoilers. Nat Rev Genet 5, 389–396 (2004).
Reyniers, E. et al. Gene conversion between red and defective green opsin gene in blue cone monochromacy. Genomics 29, 323–328 (1995).
Hallast, P., Nagirnaja, L., Margus, T. & Laan, M. Segmental duplications and gene conversion: Human luteinizing hormone/chorionic gonadotropin β gene cluster. Genome Res 15, 1535–1546 (2005).
Reiter, L. T. et al. Human meiotic recombination products revealed by sequencing a hotspot for homologous strand exchange in multiple HNPP deletion patients. Am J Hum Genet 62, 1023–1033 (1998).
Bhang, H. E. et al. Studying clonal dynamics in response to cancer therapy using high-complexity barcoding. Nat Med 21, 440–448 (2015).
Zhou, Y. H. & Li, W. H. Gene conversion and natural selection in the evolution of X-linked color vision genes in higher primates. Mol Biol Evol 13, 780–783 (1996).
Kong, A. et al. Rate of de novo mutations and the importance of father’s age to disease risk. Nature 488, 471–475 (2012).
Campbell, D. et al. Estimating human mutation rate using autozygosity in a founder population. Nat Genet 44, 1277–1281 (2012).
Dumont, B. L. & Eichler, E. E. Signals of historical interlocus gene conversion in human segmental duplications. PLoS One 8, e75949 (2013).
Dumont, B. L. Interlocus gene conversion explains at least 2.7% of single nucleotide variants in human segmental duplications. BMC Genomics 16, 456 (2015).
Acknowledgements
The authors like to thank Dr. Ueyama (Shiga University, Japan) for providing the minigene construct. This work was supported by the European Union’s Seventh Framework Programme for research, technological development and demonstration [grant agreement no 317472], and by Dr. Renata Sarno (BCM Families Foundation). E.D.B. and B.P.L. are senior clinical investigators of the Research Foundation Flanders (FWO), and are further supported by FWO grant number OZP 3G0C6715. We further acknowledge support by the Deutsche Forschungsgemeinschaft and the Open Access Publishing Fund of the University of Tübingen.
Author information
Authors and Affiliations
Contributions
E.B.-A. and B.B. performed the experiments. K.R., R.B., D.B., E.D.B., H.D., M.T.G., P.G., C.P.H., J.R.H., B.P.L., A.S.P., J.W.R.P., K.R., T.R., Z.S., J.B.G.M.V., R.W. and D.Z. recruited the families and collected the clinical data. N.W., S.K. and B.W. supervised the study and reviewed the data. E.B.-A., S.K. and B.W. drafted the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Buena-Atienza, E., Rüther, K., Baumann, B. et al. De novo intrachromosomal gene conversion from OPN1MW to OPN1LW in the male germline results in Blue Cone Monochromacy. Sci Rep 6, 28253 (2016). https://doi.org/10.1038/srep28253
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep28253
This article is cited by
-
The causal mutation in ARR3 gene for high myopia and progressive color vision defect
Scientific Reports (2023)
-
Diagnostic analysis of the highly complex OPN1LW/OPN1MW gene cluster using long-read sequencing and MLPA
npj Genomic Medicine (2022)
-
Achromatopsia: Genetics and Gene Therapy
Molecular Diagnosis & Therapy (2022)
-
A 73,128 bp de novo deletion encompassing the OPN1LW/OPN1MW gene cluster in sporadic Blue Cone Monochromacy: a case report
BMC Medical Genetics (2018)
-
Genotype determination of the OPN1LW/OPN1MW genes: novel disease-causing mechanisms in Japanese patients with blue cone monochromacy
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.