Abstract
The emergence and spread of antimicrobial resistance (AMR) mechanisms in bacterial pathogens, coupled with the dwindling number of effective antibiotics, has created a global health crisis. Being able to identify the genetic mechanisms of AMR and predict the resistance phenotypes of bacterial pathogens prior to culturing could inform clinical decision-making and improve reaction time. At PATRIC (http://patricbrc.org/), we have been collecting bacterial genomes with AMR metadata for several years. In order to advance phenotype prediction and the identification of genomic regions relating to AMR, we have updated the PATRIC FTP server to enable access to genomes that are binned by their AMR phenotypes, as well as metadata including minimum inhibitory concentrations. Using this infrastructure, we custom built AdaBoost (adaptive boosting) machine learning classifiers for identifying carbapenem resistance in Acinetobacter baumannii, methicillin resistance in Staphylococcus aureus, and beta-lactam and co-trimoxazole resistance in Streptococcus pneumoniae with accuracies ranging from 88–99%. We also did this for isoniazid, kanamycin, ofloxacin, rifampicin, and streptomycin resistance in Mycobacterium tuberculosis, achieving accuracies ranging from 71–88%. This set of classifiers has been used to provide an initial framework for species-specific AMR phenotype and genomic feature prediction in the RAST and PATRIC annotation services.
Similar content being viewed by others
Introduction
Over the last 30 years, many antibiotics have become ineffective due to the rise antimicrobial resistance (AMR) mechanisms in bacterial pathogens1,2. The World Health Organization and the United States Centers for Disease Control and Prevention estimate that that approximately 2 million people are infected by AMR pathogens annually in the U.S., at an estimated annual health care cost of $20–35 billion3,4. Meanwhile, the identification and development of novel antimicrobial compounds for clinical use has remained tedious and major breakthroughs are infrequent5.
One contributing factor to the epidemic of AMR pathogens is the overuse and administration of ineffective antibiotics in the clinical setting2,6,7. Clinicians are often faced with the decision to treat a patient immediately using their best judgment, or wait for laboratory results in order to determine the most efficacious treatment. The traditional culture-based detection of AMR phenotypes has drawbacks because many organisms are fastidious growers or may be unculturable8,9. Molecular approaches, including the high-throughput detection of biomarkers, has improved dramatically over the years10,11,12 but is often limited by the number of markers that can be detected and is constrained by our current knowledge of AMR resistance mechanisms, which are evolving rapidly1,13,14,15.
As the cost of genome sequencing is decreasing it is becoming feasible to sequence clinical isolates and detect their AMR phenotypes directly from the sequence data. This approach is advantageous because phenotype prediction is not reliant on bacterial growth, pure cultures or predefined sets of marker genes. To detect AMR phenotypes from sequence data, it is first necessary to gain an understanding of the mutations that exist among strains and the impact that these variations have on the phenotype. One way to do this is through traditional genome-wide association studies (GWAS). In a GWAS study, reads for many closely related strains are aligned against a reference strain in order to determine the significant mutations16,17,18,19. A drawback of using GWAS on bacterial genomes is that strain-to-strain variations can be extensive, and the knowledge base of variations is being established relative to a single reference genome. To narrow the scope of this problem, several recent studies have sequenced hundreds of genomes for a given species and extensively characterized the variations in known AMR genes that confer the resistance phenotype. Using this knowledge base of variants, the researchers were able to build accurate phenotype identification algorithms for Escherichia, Klebsiella, Streptococcus, Staphylococcus and Mycobacterium20,21,22,23,24. This is a successful approach because in many cases AMR can be the result of a single gene or a small set of variations13,25. However, like the detection protocols that are currently in use, the main drawback is that by focusing on a small set of genes with known phenotypes any potentially novel AMR determinants are missed.
Machine learning algorithms may provide an alternative approach to traditional GWAS studies. In general, machine learning algorithms work by finding the relevant features in a complex data set that enable the ability to make a strong prediction26. These algorithms are currently being used extensively in genomics, particularly in cancer research, in order to elucidate clinically important information such as, patient genotypes, gene expression related phenotypes and patient outcomes26,27,28,29. Recently, two studies have used machine learning algorithms to find relevant genomic regions associated with AMR30,31. These studies used a machine learning algorithm to select the oligonucleotide k-mers that are relevant to antibiotic resistance. These relevant k-mers were then used as a phenotype “classifier” for unknown genomes and as a source for identifying important genomic regions.
One of the reasons that machine learning classifiers are not in widespread use for AMR phenotype detection is the difficulty in obtaining large numbers of bacterial genomes with AMR metadata. PATRIC (Pathosystems Resource Integration Center, patricbrc.org) is a NIH supported bioinformatics resource center that has been built to enable comparative genomic analysis of bacterial pathogens32,33. For the last two years, we have been gathering, assembling and maintaining a database containing genomes with AMR panel data from the literature, collaborators and public repositories like GenBank and the SRA34,35. In this study, we describe an infrastructure that we have built and deployed in PATRIC that enables users easy access to sets of genomes that are binned by their AMR phenotype so that they can do research on AMR detection. We then describe how we have used this infrastructure to build several machine learning classifiers to detect AMR phenotypes in Acinetobacter Mycobacterium, Staphylococcus and Streptococcus. These classifiers have been deployed in the RAST36,37 and PATRIC annotation services in order to provide an initial prediction of AMR-related genomic features and phenotype.
Methods
Building AdaBoost classifiers
We start by computing the k-mer occurrences for the contigs of each genome by using the k-mer counting program KMC38. We use the contigs as the starting point because we are maintaining the contigs for each genome on the PATRIC FTP site, although in principle sequence reads could also be used after preprocessing with a suitable error correction or filtering method. The files of the k-mer counts are then merged to form a matrix where the rows represent each individual k-mer and the columns depict presence “1” or absence “0” of each k-mer in a given genome. We chose to abstract the k-mer occurrences to presence versus absence in order to avoid potential complications resulting from repeat regions and low complexity sequence. Since we are seeking k-mers that are signatures of resistance, the most informative k-mers will be the ones that are present in the resistant strains and absent in susceptible strains. For simplicity, we invert the values for the columns corresponding to susceptible genomes so that the most informative k-mer will be the row with the fewest zeros. In other words, if a row in the matrix has a cell with a value of zero, it can then be thought of as an error in that k-mer’s ability to accurately classify the data set. This matrix is then passed to the AdaBoost algorithm39. For clarity, we describe our use of AdaBoost in this context below using equations adapted from39. For a more in-depth description of AdaBoost please refer to refs 39,40.
The first part of the AdaBoost algorithm seeks the most informative k-mers for classification. During this part of the algorithm, two arrays of data are maintained. The first array is the row of values from the matrix corresponding to the given k-mer (we call this the matrix array). The second array contains probability values assigned to each column in the matrix (we call this the probability array). In the first round (m), each element in the probability array (pm) is set to 1/n, where n is the number of columns in the matrix, which amounts to the number of genomes.
Then for each k-mer, a weighted error value equal to the sum of the probabilities for the missed elements in the matrix array is assigned to each k-mer. After iterating over all of the k-mers (rows in the matrix), the “best” k-mer with the lowest weighted error is retained. A final weight for the best k-mer is then computed as shown in equation 1, where α is the final weight and ε is the weighted error for the missed elements in the matrix array for that given round (t).

Each element in the probability array is then updated based on the corresponding matrix array index for the best k-mer. If the corresponding index position was a miss, the unnormalized probability (pu) for that element is updated as shown in equation 2.

If the corresponding index position was a match, the unnormalized probability is updated as shown in equation 3.

The unnormalized probability values (pu) are normalized at each index position by dividing by the sum of unnormalized probabilites (Z) for each element in the probability array as shown in equation 4.

These normalized probabilities now become the elements of the updated probability array.
Thus, the elements in the matrix that were matched by the best k-mer result in lower probability values, while positions that were missed result in higher probability values. In this way, through iterative rounds of “boosting”, the algorithm seeks the set of k-mers that are most representative of the data by tiling over the elements in the matrix that were missed in the previous round. This process is repeated until the α-value reaches a global minimum. For the classifiers that we have built to date, the α-value typically reaches a minimum by the tenth round of boosting (Supplementary Fig. S1).
Classifying a genome
The second part of the AdaBoost algorithm is the classification of an unknown genome. To do this we use the set of “best” k-mers found from successive rounds of boosting and their α values. Unless otherwise indicated, classification results in this study are presented using the k-mer set found in the first 10 rounds of boosting. For each k-mer chosen by AdaBoost, we search for the corresponding k-mer in the target genome. If the k-mer exists, it votes by contributing a positive weighted vote corresponding to the α value for the k-mer. If it is absent, it votes by contributing a negative α value. In practice, there can be multiple k-mers with identical matching patterns that are the best in a given round. When this happens we issue a single weighted vote corresponding to the presence of one, or the absence of all members. If the sum of all weighted votes is negative the genome is predicted to be susceptible to the antibiotic; if the sum is positive, the genome is predicted to be resistant to the antibiotic. The magnitude of the resulting weighted vote is a measure of the confidence in the prediction.
Displaying AMR-related regions
Using the set of distinguishing k-mers found by AdaBoost, the genomic regions corresponding to AMR are displayed by performing a BLASTN41 search using the k-mers as the query and the contigs or genes from the target genome as the subject. High scoring segment pairs (HSPs) where the k-mers match with 100% coverage and 100% identity are presented as AMR features. If k-mers are found to be overlapping, or if they exist in a proximity less than or equal to one k-mer length, the region is presented as a single genomic feature relating to AMR.
Selection of Mycobacterium tuberculosis genome sets
The worldwide health threat of multidrug resistant tuberculosis has resulted in several studies that have generated a large amount of AMR data for M. tuberculosis strains23,42,43. At PATRIC, M. tuberculosis is currently the species with the largest amount of AMR metadata (Table 1). However, many M. tuberculosis genomes are resistant to multiple antibiotics, making an unambiguous classification of individual antibiotics challenging (Supplementary Table S1). For example, this makes it difficult to build a classifier to detect isoniazid-related k-mers that is not also biased by rifampicin-related k-mers and vice versa. As a result, we chose to build classifiers for each antibiotic by selecting subsets of genomes with AMR profiles that are less correlated between antibiotics (Supplementary Table S2).
We converted the AMR SIR (Susceptible, Intermediate, or Resistant) data for each genome to numeric values: +1 for resistant, 0 for intermediate or unknown and –1 for susceptible. Then using these values for all of the genomes, we computed the correlation between each antibiotic. We systematically removed each genome from the set and then recomputed the correlation summing the difference in the correlation across all antibiotics for each genome. After doing this for all genomes, we chose the set of genomes for each antibiotic that minimizes the magnitude of the correlation with the other antibiotics. These less correlated genome sets were chosen for building the M. tuberculosis classifiers (Supplementary Table S2). After performing this analysis, we built classifiers for genome sets built from 100, 150, 200, 250 and 300 genomes respectively. We chose to use classifiers built from the 250 genomes because they offer an adequate number of genomes for classification, while still minimizing correlations. This can be seen in Supplementary Tables S2 and 3, where the top k-mer for the rifampicin classifier is located within the RNA polymerase beta-subunit gene (the expected target) in the 150-, 200- and 250-genome sets (Pearson correlation coefficients (PCC) equal to 0.370, 0.524 and 0.617 respectively), but then becomes located within the katG gene (the expected target of isoniazid resistance) in the 300-genome set (with a PCC equal to 0.746).
Results
Accessing AMR genomes and metadata in PATRIC
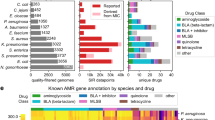
PATRIC is a bioinformatic resource that is designed to enable comparative genomic analysis of bacterial pathogens. At the time of writing, the database at the core of PATRIC contains ~33,000 bacterial genomes, and drives the user interface on the website as well as the service environment (which includes assembly, annotation and RNA-seq analysis). Each time there is a data release, newly obtained bacterial genomes that have been obtained are annotated and released on the website. These genomes are also pushed to the PATRIC FTP site (ftp://ftp.patricbrc.org/patric2/patric3/genomes/) where users can download the original source data and the PATRIC-specific analyses including the RASTtk annotation results36, biochemical pathways44,45, and BLAST matches to the ARDB (antibiotic resistance genes database) and CARD (the comprehensive antibiotic resistance database) compendia of hand-curated AMR-related genes41,46,47.
Until recently, it has been difficult to publicly deposit or obtain AMR data for specific strains48. At PATRIC we have been collecting AMR data for strains that have been given to us by collaborators and that have been published in the literature. These AMR metadata are typically represented as minimum inhibitory concentrations (MIC) for commonly used antibiotics, which are generated using standard laboratory techniques49. They can also be in the form of susceptible, intermediate or resistant (SIR) determinations that have been made by the researcher. In the case of MIC data, the laboratory tests can vary depending on the growth requirements of the organism. SIR data are usually a researcher’s conclusion based on a MIC test, but sometimes they can be an assertion that is based on the MIC result for a similar antibiotic. To date, we have not actively incorporated AMR determinations resulting from purely genomic inference17,49. The PATRIC FTP site also contains AMR metadata that have been reported for an entire family of antibiotics, such as beta-lactam or carbapenem resistance.
We maintain a list of genomes with AMR data on the FTP site (ftp://ftp.patricbrc.org/BRC_Mirrors/AMR/) and report both MIC and SIR data when they are available. For species with at least 100 susceptible and resistant genomes for a given antibiotic, we also maintain directories of the susceptible and resistant genomes (ftp://ftp.patricbrc.org/patric2/current_release/AMR_genome_sets/). The goal of this organized distribution of AMR data is to enable more rapid improvements in the detection and annotation of AMR-related genomic features. Bacterial species on the PATRIC FTP site that have over 100 susceptible and resistant genomes for a given antibiotic are presented in Table 1.
Using the AdaBoost algorithm for whole-genome AMR classification
Given a framework for distributing sets of genomes binned according to their AMR phenotype, we built a set of classifiers for predicting the AMR phenotypes for the genomes that are being submitted to our annotation services in RAST and PATRIC. Since we want to be able to detect phenotypic determinants found in intergenic regions as well as within genes, we built our classifiers from whole genome data using 31 base pair nucleotide k-mers. We decided to use 31-mers because they offer considerable specificity and a manageable memory footprint. In general, k-mers ranging in size from 24–31 nucleotides (and perhaps longer) work well for similar classification problems50,51. In order to make an accurate phenotype prediction, it is necessary to find the set of k-mers that distinguishes resistant genomes from susceptible genomes. In this initial implementation, we have chosen to find these distinguishing k-mers by using the AdaBoost (Adaptive Boosting) algorithm39,52,53. AdaBoost works by taking a weak ensemble of classifiers—in this case the occurrence of our 31-mers across a set of genomes—and ranking them through a process of iterative refinement (i.e., “boosting” or “learning”). This set of distinguishing k-mers and their associated weights becomes the classifier that is used to predict the phenotype of an incoming genome (Materials and Methods). The overall workflow is described in Fig. 1. We chose AdaBoost for its simplicity and ease of implementation, although in principle other machine learning algorithms, such as random forests, support vector machines and the set covering machine might also work well in this context, and may ultimately be used to improve upon this initial set of classifiers30,31,54,55,56.
Genomes for a given species are binned according to whether they are resistant or susceptible to an antibiotic and the k-mer counts are computed for each genome. The k-mer counts are then merged to form a matrix. A machine learning algorithm searches this matrix to find the k-mers that distinguish the resistant and susceptible genomes. These distinguishing k-mers are then used as a “classifier” to predict the phenotype for a new genome.
AMR classification for Acinetobacter Staphylococcus and Streptococcus
We started by building AdaBoost classifiers to detect carbapenem resistance in Acinetobacter baumannii, methicillin resistance in Staphylococcus aureus and beta-lactam and co-trimoxazole resistance Streptococcus pneumoniae. We chose these four examples because we have a sufficient number of sequenced resistant and susceptible strains in the PATRIC database. In our experience with AdaBoost, the genomes in each set should be balanced, so in all cases we present classifiers that have been built with an equal number of susceptible and resistant genomes. Unless otherwise indicated, in this study we describe classifiers learned with at least 100 genomes because we have observed lower accuracies when we try to build classifiers using less than 100 genomes (Supplementary Fig. S2). However, the ease of classification can depend on the AMR mechanism with single nucleotide polymorphism (SNP) based resistance being more difficult to classify than resistance resulting from the presence of entire genes or mobile elements.
We tested the ability to accurately classify the AMR phenotype for genomes of the same species by randomly selecting 80% of the available genomes for training and the remaining 20% of the genomes for testing. This was repeated ten times as a cross validation experiment. In all four cases, we plotted the results of the combined test sets as receiver operating characteristic (ROC) curves which depict the true-positive rate versus the false positive rate for each classifier (Fig. 2). In all four cases, the classifiers are very accurate with the area under the ROC curve (AUC) values greater than 94%, and accuracies ranging from 87% for classifying co-trimoxazole resistance in S. pneumoniae to 99% for classifying methicillin resistance S. aureus. The F1 score, which is the harmonic mean of precision and recall and is commonly used to compare different classification methods, is similarly high (Table 2). In order to understand the effect of misclassified genomes adversely influencing the accuracy of the classifiers, we introduced error into the analysis by mixing susceptible and resistant genomes in the training sets. Although the accuracies diminish with mixing, the classifiers retain some ability to classify even when mixing exceeds 20% of the total genomes in the training set (Supplementary Fig. S3).
Data are the results of cross validation on the set of genomes described in Table 2. Equal numbers of susceptible and resistant genomes were used in the experiment.
For each classifier it is important to establish whether the k-mers that are used for classification are indeed related to a known mechanism of AMR. Since we are using AdaBoost to seek a minimum number of k-mers for classification, we will not find all known AMR determinants, but rather those that describe the most variation in the training set genomes. In the case of carbapenem resistance in A. baumannii, the top ranking k-mer is found in a LysR transcriptional regulator gene that is located immediately downstream of a metallo-beta-lactamase gene that may be conferring resistance to carbapenem antibiotics57 (Table 3). In the case of S. aureus, the set of top ranking k-mers relating to methicillin resistance span a 3-gene region containing the mecA gene, which is known for conferring methicillin resistance e.g.,58. The 17 highest ranking k-mers, and the 14 third-highest ranking k-mers for S. pneumoniae β-lactam resistance are found in the peptidoglycan synthase gene (pbp2x), a penicillin binding protein for which SNPs conferring β-lactam resistance have been extensively characterized17. The 5 highest and 2 second-highest ranking k-mers found relating to co-trimoxazole resistance in S. pneumoniae are found in an intergenic region immediately downstream of the dihydropteroate synthase and within the dihydrofolate reductase genes respectively. Co-trimoxazole is a combination drug comprised of sulfamethoxazole and trimethoprim. Both drugs inhibit steps in folate synthesis with sulfamethoxazole inhibiting dihydropterate synthase and trimethoprim inhibiting dihydrofolate reductase59. Overall, in all four cases, the selection of the highest-ranking k-mers by AdaBoost results in k-mers that correspond with known AMR determinants.
In some cases, the second and third highest-ranking k-mers do not appear to have an obvious role in AMR. For instance the 4 second-highest ranking k-mers in A. baumannii carbapenem resistance correspond with a NAD+–asparagine ADP-ribosyltransferase gene and the 3 third-highest ranking k-mers correspond with a gene encoding Dihydrodipicolinate synthase family protein-encoding gene. In the case of Streptococcus β-lactam resistance, the second highest-ranking k-mers occur in an intergenic region between ABC transporter genes. If any of these genes are involved in AMR, their role seems to be unclear from the current annotations. It is also possible that these regions reflect overfitting or variation from linear descent that may ultimately be eliminated as the diversity of the strains used in the training can be expanded.
AMR classification for Mycobacterium tuberculosis
Since antibiotics are often co-prescribed, many sequenced isolates are resistant to multiple antibiotics23,42,43. This is particularly the case for M. tuberculosis (Supplementary Table S1). In order to obtain classifiers for individual antibiotics, we selected subsets of M. tuberculosis genomes that minimize the correlation between antibiotic resistance profiles (see Methods section) (Supplementary Table S2). The AdaBoost classifiers built from subsets of M. tuberculosis genomes perform well with AUC values >0.70 for all antibiotics. Overall, the classifiers built for isoniazid, kanamycin, and rifampicin resistance are the best, with AUC values ≈0.9 and accuracies between 86–88% (Fig. 3, Table 4). The classifier built for ethambutol was the worst with an AUC value of 0.72 and an accuracy of 58%. Since we had a large number of genomes that were resistant to at least six of the seven antibiotics (83 genomes) and susceptible to at least six of the seven antibiotics (139 genomes), we also built a classifier of “pan-resistance” to the available antibiotics (Supplementary Tables S4 and 5). This combined classifier performed well with an AUC value of 0.97 and an accuracy of 93% (Fig. 3, Table 4).
Genome sets and classifier statistics are described in Table 3. Classifiers for individual antibiotics were chosen for minimal correlation between AMR patterns and up to 250 resistant and susceptible genomes were used. Equal numbers of susceptible and resistant genomes were used all experiments. All curves depict cross validation experiments and are for ethambutol (red line with square symbols), ethionamide (orange line with diamond symbols), isoniazid (green line with triangle symbols), kanamycin (light blue line with circle symbols), ofloxacin (dark blue line with square symbols), rifampicin (purple line with diamond symbols) and streptomycin (brown line with triangle symbols). The black line with circle plot points depicts the combined multidrug resistance classifier described in Tables 3 and Supplementary Tables S4–6.
For five of the seven antibiotics listed in Table 5, we observe a top matching k-mer hit in the expected drug target for the antibiotic. The highest ranking set of k-mers for isoniazid resistance match the katG gene60, the highest-ranking k-mer for kanamycin resistance corresponds to a region of the 16S rRNA gene that is known to be involved in kanamycin resistance61, the highest-ranking k-mer for ofloxacin resistance corresponds with the DNA gyrase subunit A gene62, the highest-ranking k-mer for rifampicin resistance corresponds to the RNA polymerase beta-subunit gene63 and the highest ranking k-mer for streptomycin is found in the small subunit ribosomal protein S12 gene64. In the case of ethambutol, the top ranking k-mer does not appear to have an obvious link to an AMR mechanism and instead matches the beta subunit of the RNA polymerase gene, which is a likely correlation with rifampicin (Table 5, Supplementary Table S2). The second-highest ranking k-mers match the embB gene, which is one of the the targets of ethambutol65. If more genomes were available with ethambutol panel data that were also uncorrelated with the other antibiotics, we would expect the embB gene to become the top match. The three highest-ranking sets of k-mers for the ethionamide classifier do not match the mechanistic target for ethionamide, which is an analog of isoniazid. In this case, we would have expected the top k-mers to match ethA, inhA or katG66. It is possible that by attempting to decouple isoniazid and ethionamide resistance profiles that the signal for ethionamide was lost. The combined multidrug resistance classifier identifies the same targets for isoniazid, kanamycin, rifampicin, ofloxacin and ethambutol in the top ten k-mers (Table S6).
Like the classifiers built for A. baumannii, S. aureus, and S. pneumoniae, the classifiers built for M. tuberculosis also contain matches to genomic regions with no known relationship to AMR. In the case of A. baumannii carbapenem resistance and S. aureus methicillin resistance, nearly all of the resistant genomes contain a match to the top ranking k-mers. However, in the case of the other classifiers, additional rounds of boosting are necessary for the entire set of resistant genomes to be represented by the classifier (Figures S4–7). Thus, these lower ranking targets contribute the accuracy of the classifiers, albeit with lower weights than the main AMR target. These data also suggest that in many instances a classifier built from a single target gene, such as rpoB or katG in M. tuberculosis, may be insufficient for accurately classifying the phenotypes of these organisms.
Availability
Antibiotic resistance phenotype prediction and genomic feature identification for the genomes and antibiotics described in this study (excluding ethambutol and ethionamide in M. tuberculosis) has been made available as an option in the RASTtk workflow36, from the RAST website (http://rast.nmpdr.org/), and in the PATRIC annotation service32 (http://patricbrc.org/). The analysis is performed when a user provides a genome with a species name that matches a species for which we have built classifiers. The classification output includes the weighted AdaBoost vote for the genome, which is a measure of confidence in the prediction, as well as information about each classifier and the genomic regions, which are annotated as features and can be browsed in a compare regions context. The classification output is recorded in the genome object, feature file and annotation history for the genome. AMR-related regions predicted by the classifiers for a newly annotated genome can be browsed on both the PATRIC and RAST websites. As the number of genomes with panel data increases in PATRIC, we plan to expand the number of classifiers. We also plan to expand upon the set of machine-learning algorithms that are used to build the classifiers.
Discussion
As the number of effective antibiotics dwindles, it becomes increasingly important to rapidly identify AMR in bacterial genomes. In order to advance the state of detection and AMR classification, we have provided FTP access to the AMR metadata for the genomes that are available at PATRIC. We have also used these data sets to build classifiers of antibiotic resistance in order to provide a reliable AMR classification capability in the RAST and PATRIC genome annotation pipelines. Given the current data sets available at PATRIC, we built AdaBoost classifiers for A. baumannii, S. aureus, S. pneumoniae and M. tuberculosis. In the cases of A. baumannii (carbapenems), S. aureus (methicillin) and S. pneumoniae (beta-lactams and co-trimoxazole), the classifiers were highly accurate and performed classifications based on k-mers that corresponded to regions in the genome that are known to confer AMR. In the case of M. tuberculosis, we built a classifier for pan-resistance to seven antibiotics that was 93% accurate. Since the AMR profiles from the panel data for each antibiotic were similar, we selected subsets of genomes with AMR profiles that were less correlated between antibiotics in order to identify AMR determinants that were specific to each antibiotic. This was an effective strategy, yielding accurate classifiers with top ranking k-mers in known AMR-conferring regions for five of the seven antibiotics (isoniazid, kanamycin, ofloxacin, rifampicin and streptomycin). In the case of the remaining two antibiotics (ethambutol and ethionamide), we predict that better classifiers can be built if we can obtain more susceptible and resistant genomes with AMR profiles that are unrelated to the other antibiotics.
There are caveats to building classifiers and providing predictions that are based upon them. In all cases, the accuracy of a prediction is contingent upon the set of genomes that was used to train the classifier. For instance, if a strain is resistant because of variation that was not originally sampled in the training set genomes, such as novel SNPs or a horizontally acquired element, the genome will receive an incorrect classification. Likewise, having a limited or biased training set can result in high ranking k-mers that are the result of overfitting rather than true AMR-related variation. We were able to construct classifiers that have top-ranking k-mers linked to well known AMR mechanisms; however, it is unclear if the lower ranking k-mers have a role in AMR, are lineage-related variation rather than AMR-related variation, or are noise due to overfitting. Maintaining these lower ranking k-mers represents a tradeoff in our ability to find relevant but poorly understood AMR-related features, and potentially classifying based on noise or lineage-related variation. As the number and diversity of available genomes increases, it may be possible to improve upon this initial set of classifiers by exploring methods for maximizing genetic diversity in the training sets.
Being able to build classifiers with a balanced number of susceptible and resistant genomes is also important for building accurate classifiers, but is currently a major limitation. In most cases, the number of available genomes with AMR panel data is skewed toward genomes that are resistant because these are the genomes that are of clinical importance to hospitals and epidemiologists. For instance, at the time of writing there are currently over 100 resistant genomes for five additional antibiotics for M. tuberculosis, four antibiotics for Pseudomonas aeruginosa and three antibiotics for S. pneumoniae in PATRIC, but an insufficient number of known susceptible genomes to build these classifiers. In our opinion, emphasis should be given to sequencing diverse susceptible genomes in order to advance the ability to predict phenotypes. Despite these limitations, the ability to classify based upon the current data is an important capability that will improve over time.
GWAS studies are able to detect significant regions of variation by comparing genomes that are binned by phenotype against a reference genome. In principle, machine learning approaches could be used to do the same analysis without requiring alignment against a reference genome—a property that would be useful for bacterial comparisons since they can differ extensively across strains. In this study, the k-mers relating beta-lactam resistance in S. pneumoniae that were identified by AdaBoost corresponded with the pbp2x gene that was also identified in a previous GWAS study17. In that study, Chewapreecha and colleagues also found significant variations relating to resistance in the pbp1a and pbp2a penicillin binding proteins, which were not identified in this study using AdaBoost. This could be the result of AdaBoost selecting the most highly ranked k-mers, or it could be the result of the all-versus-all comparison that was performed by AdaBoost rather than mapping against a single reference genome. Either way, this study suggests that machine learning algorithms can complement a traditional GWAS study, but potentially using them as a replacement for GWAS is an area that requires more research.
The RAST and PATRIC annotation systems currently project functions from the annotated proteins in the SEED database37. They also offer BLAST matches to the AMR-related proteins that have been curated at ARDB (the Antibiotic Resistance genes Database) and CARD (Comprehensive Antibiotic Resistance Database)36,46,47. In addition to being the first phenotype assertion offered by these services, the use of classifiers offers a complementary approach to identifying AMR-related features within the genome. This is advantageous because the classifiers can identify entire regions, such as the mecA containing region in S. aureus, as well as SNP-level variations, such as the kanamycin resistance in M. tuberculosis that was the result of mutations in the 16S rRNA gene. The ability to catalog variants, particularly those found in intergenic regions and within genes, provides a more detailed level of resolution for these annotation systems.
Additional Information
How to cite this article: Davis, J. J. et al. Antimicrobial Resistance Prediction in PATRIC and RAST. Sci. Rep. 6, 27930; doi: 10.1038/srep27930 (2016).
References
Blair, J. M., Webber, M. A., Baylay, A. J., Ogbolu, D. O. & Piddock, L. J. Molecular mechanisms of antibiotic resistance. Nature Reviews Microbiology 13, 42–51 (2015).
Alanis, A. J. Resistance to antibiotics: are we in the post-antibiotic era? Archives of medical research 36, 697–705 (2005).
World Heath Organization, Antimicrobial resistance: global report on surveillance. http://www.who.int/drugresistance/documents/surveillancereport/en/ (2014) (Date of access: 11/04/2016).
United States Department of Health and Human Services. Antibiotic resistance threats in the United States, 2013. Centers for Disease Control and Prevention, Atlanta, GA:http://www.cdc.gov/drugresistance/threat-report-2013/ (2013) (access:04/01/2016).
Ling, L. L. et al. A new antibiotic kills pathogens without detectable resistance. Nature 517, 455–459 (2015).
Hicks, L. A. & Blaser, M. J. Variability in Antibiotic Prescribing: An Inconvenient Truth. Journal of the Pediatric Infectious Diseases Society, piu106 (2014). doi: 10.1093/jpids/piu106.
Gerber, J. S. et al. Variation in antibiotic prescribing across a pediatric primary care network. Journal of the Pediatric Infectious Diseases Society, piu086, doi: 10.1093/jpids/piu086 (2014).
Tang, Y.-W., Procop, G. W. & Persing, D. H. Molecular diagnostics of infectious diseases. Clinical Chemistry 43, 2021–2038 (1997).
Wade, W. Unculturable bacteria—the uncharacterized organisms that cause oral infections. Journal of the Royal Society of Medicine 95, 81–83 (2002).
Taitt, C. R. et al. Antimicrobial resistance determinants in Acinetobacter baumannii isolates taken from military treatment facilities. Antimicrobial agents and chemotherapy, AAC. 01897–01813 (2013).
Leski, T. A. et al. Molecular characterization of multidrug resistant hospital isolates using the Antimicrobial Resistance Determinant Microarray, doi: 10.1371/journal.pone.0069507 (2013).
Inouye, M. et al. SRST2: Rapid genomic surveillance for public health and hospital microbiology labs. Genome Med 6, 90 (2014).
Kumarasamy, K. K. et al. Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: a molecular, biological, and epidemiological study. The Lancet infectious diseases 10, 597–602 (2010).
Dijkshoorn, L., Nemec, A. & Seifert, H. An increasing threat in hospitals: multidrug-resistant Acinetobacter baumannii. Nature Reviews Microbiology 5, 939–951 (2007).
Liu, Y.-Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. The Lancet Infectious Diseases, doi: 10.1016/S1473-3099(15)00424-7 (2015).
Chewapreecha, C. et al. Dense genomic sampling identifies highways of pneumococcal recombination. Nature genetics 46, 305–309 (2014).
Chewapreecha, C. et al. Comprehensive identification of single nucleotide polymorphisms associated with beta-lactam resistance within pneumococcal mosaic genes, doi: 10.1371/journal.pgen.1004547 (2014).
Croucher, N. J. et al. Population genomics of post-vaccine changes in pneumococcal epidemiology. Nature genetics 45, 656–663 (2013).
Read, T. D. & Massey, R. C. Characterizing the genetic basis of bacterial phenotypes using genome-wide association studies: a new direction for bacteriology. Genome Med 6, 109 (2014).
Stoesser, N. et al. Predicting antimicrobial susceptibilities for Escherichia coli and Klebsiella pneumoniae isolates using whole genomic sequence data. Journal of Antimicrobial Chemotherapy 68, 2234–2244 (2013).
Gordon, N. et al. Prediction of Staphylococcus aureus antimicrobial resistance by whole-genome sequencing. Journal of clinical microbiology 52, 1182–1191 (2014).
Bradley, P. et al. Rapid antibiotic resistance predictions from genome sequence data for S. aureus and M. tuberculosis. bioRxiv, 018564, doi: 10.1038/ncomms10063 (2015).
Walker, T. M. et al. Whole-genome sequencing for prediction of Mycobacterium tuberculosis drug susceptibility and resistance: A retrospective cohort study. The Lancet Infectious Diseases 15, 1193–1202 (2015).
Bradley, P. et al. Rapid antibiotic-resistance predictions from genome sequence data for Staphylococcus aureus and Mycobacterium tuberculosis. Nature communications 6, doi: 10.1038/ncomms10063 (2015).
Hiramatsu, K., Cui, L., Kuroda, M. & Ito, T. The emergence and evolution of methicillin-resistant Staphylococcus aureus. Trends in microbiology 9, 486–493 (2001).
Libbrecht, M. W. & Noble, W. S. Machine learning applications in genetics and genomics. Nature Reviews Genetics (2015), doi: 10.1038/nrg3920.
Whitney, D. H. et al. Derivation of a bronchial genomic classifier for lung cancer in a prospective study of patients undergoing diagnostic bronchoscopy. BMC medical genomics 8, 18 (2015).
Griffith, O. L. et al. A robust prognostic signature for hormone-positive node-negative breast cancer. Genome Med 5, 92 (2013).
Shipp, M. A. et al. Diffuse large B-cell lymphoma outcome prediction by gene-expression profiling and supervised machine learning. Nature medicine 8, 68–74 (2002).
Drouin, A. et al. Learning interpretable models of phenotypes from whole genome sequences with the Set Covering Machine. arXiv preprint arXiv:1412.1074 (2014).
Santerre, J., Boisvert, S., Davis, J., Xia, F. & Stevens, R. Gene identification and strain classification using Random Forests. Great Lakes Bioinformatics Conference 2015. Purdue University, West Lafayette, Indiana, USA (2015).
Wattam, A. R. et al. PATRIC, the bacterial bioinformatics database and analysis resource. Nucleic acids research 42, D581–D591 (2014).
Gillespie, J. J. et al. PATRIC: the comprehensive bacterial bioinformatics resource with a focus on human pathogenic species. Infection and immunity 79, 4286–4298 (2011).
Benson, D. A. et al. GenBank. Nucleic acids research 43, D30 (2015).
Kodama, Y., Shumway, M. & Leinonen, R. The Sequence Read Archive: explosive growth of sequencing data. Nucleic acids research 40, D54–D56 (2012).
Brettin, T. et al. RASTtk: A modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Scientific reports 5, doi: 10.1038/srep08365 (2015).
Overbeek, R. et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic acids research 42, D206–D214 (2014).
Deorowicz, S., Kokot, M., Grabowski, S. & Debudaj-Grabysz, A. KMC 2: Fast and resource-frugal k-mer counting. Bioinformatics 31, 1569–1576 (2015).
Freund, Y. & Schapire, R. E. A decision-theoretic generalization of on-line learning and an application to boosting. Journal of computer and system sciences 55, 119–139 (1997).
Zhu, J., Zou, H., Rosset, S. & Hastie, T. Multi-class adaboost. Statistics and its Interface 2, 349–360 (2009).
Camacho, C. et al. BLAST+: architecture and applications. BMC bioinformatics 10, 421 (2009).
TB-ARC MRC SA initiative, Broad Institute.https://olive.broadinstitute.org/projects/tb_mrc_sa. (2016) (Date of access:11/04/2016).
Merker, M. et al. Evolutionary history and global spread of the Mycobacterium tuberculosis Beijing lineage. Nature genetics, doi: 10.1038/ng.3195 (2015).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic acids research 28, 27–30 (2000).
Kanehisa, M. et al. Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic acids research 42, D199–D205 (2014).
Liu, B. & Pop, M. ARDB—antibiotic resistance genes database. Nucleic acids research 37, D443–D447 (2009).
McArthur, A. G. et al. The comprehensive antibiotic resistance database. Antimicrobial agents and chemotherapy 57, 3348–3357 (2013).
Barrett, T. et al. BioProject and BioSample databases at NCBI: facilitating capture and organization of metadata. Nucleic acids research 40, D57–D63 (2012).
Cockerill, Franklin R., and Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing: twenty-second informational supplement; [… provides updated tables for… M02-A11 and M07-A9]. National Committee for Clinical Laboratory Standards. (2012).
Edwards, R. A. et al. Real Time Metagenomics: Using k-mers to annotate metagenomes. Bioinformatics 28, 3316–3317 (2012).
Wood, D. E. & Salzberg, S. L. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome biology 15, R46 (2014).
Rojas, R. AdaBoost and the super bowl of classifiers a tutorial introduction to adaptive boosting. Computer Science Department, Freie Universitat, Berlin. http://www.inf.fu-berlin.de/inst/ag-ki/adaboost4.pdf (2009) (Date of access: 11/04/2016).
Schapire, R. E. Explaining AdaBoost. In Empirical inference. pp. 37–52. Springer, doi: 10.1007/978-3-642-41136-6_5 (2013).
Breiman, L. Random forests. Machine learning 45, 5–32 (2001).
Cortes, C. & Vapnik, V. Support-vector networks. Machine learning 20, 273–297 (1995).
Marchand, M. & Taylor, J. S. The set covering machine. The Journal of Machine Learning Research 3, 723–746 (2003).
Poirel, L. & Nordmann, P. Carbapenem resistance in Acinetobacter baumannii: mechanisms and epidemiology. Clinical Microbiology and Infection 12, 826–836 (2006).
Wielders, C., Fluit, A., Brisse, S., Verhoef, J. & Schmitz, F. mecA gene is widely disseminated in Staphylococcus aureus population. Journal of clinical microbiology 40, 3970–3975 (2002).
Huovinen, P., Sundström, L., Swedberg, G. & Sköld, O. Trimethoprim and sulfonamide resistance. Antimicrobial Agents and Chemotherapy 39, 279 (1995).
Heym, B., Alzari, P. M., Honore, N. & Cole, S. T. Missense mutations in the catalase‐peroxidase gene, katG, are associated with isoniazid resistance in Mycobacterium tuberculosis. Molecular microbiology 15, 235–245 (1995).
Suzuki, Y. et al. Detection of Kanamycin-ResistantMycobacterium tuberculosis by Identifying Mutations in the 16S rRNA Gene. Journal of clinical microbiology 36, 1220–1225 (1998).
Shi, R., Zhang, J., Li, C., Kazumi, Y. & Sugawara, I. Emergence of ofloxacin resistance in Mycobacterium tuberculosis clinical isolates from China as determined by gyrA mutation analysis using denaturing high-pressure liquid chromatography and DNA sequencing. Journal of clinical microbiology 44, 4566–4568 (2006).
Telenti, A. et al. Detection of rifampicin-resistance mutations in Mycobacterium tuberculosis. The Lancet 341, 647–651 (1993).
Honore, N. & Cole, S. T. Streptomycin resistance in mycobacteria. Antimicrobial agents and chemotherapy 38, 238–242 (1994).
Sreevatsan, S. et al. Ethambutol resistance in Mycobacterium tuberculosis: critical role of embB mutations. Antimicrobial Agents and Chemotherapy 41, 1677–1681 (1997).
Morlock, G. P., Metchock, B., Sikes, D., Crawford, J. T. & Cooksey, R. C. ethA, inhA, and katG loci of ethionamide-resistant clinical Mycobacterium tuberculosis isolates. Antimicrobial agents and chemotherapy 47, 3799–3805 (2003).
National Institute of Allergy and Infectious Diseases, Genomic Centers for Infectious Diseases, https://www.niaid.nih.gov/labsandresources/resources/dmid/gsc/pages/default.aspx (2016) (Date of access:01/04/2016)
Acknowledgements
This work was supported by the United States National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Service [Contract No. HHSN272201400027C]. We thank Emily Dietrich for her careful editing.
Author information
Authors and Affiliations
Contributions
J.J.D. Experimental design, data collection, software design, wrote and prepared manuscript. S.B., R.O.v., J.S., F.X. and R.S. Experimental design. T.B., R.W.K., C.M. and A.R.W. AMR data curation. R.O.l. Software engineering and design. M.S. FTP site management and data curation. R.W. AMR data curation and deployment of AMR metadata
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Davis, J., Boisvert, S., Brettin, T. et al. Antimicrobial Resistance Prediction in PATRIC and RAST. Sci Rep 6, 27930 (2016). https://doi.org/10.1038/srep27930
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep27930
This article is cited by
-
Application of machine learning in the prevention of periprosthetic joint infection following total knee arthroplasty: a systematic review
Arthroplasty (2023)
-
Global pathogenomic analysis identifies known and candidate genetic antimicrobial resistance determinants in twelve species
Nature Communications (2023)
-
Exploring antibiotic resistance genes, mobile gene elements, and virulence gene factors in an urban freshwater samples using metagenomic analysis
Environmental Science and Pollution Research (2023)
-
Enhancing predictions of antimicrobial resistance of pathogens by expanding the potential resistance gene repertoire using a pan-genome-based feature selection approach
BMC Bioinformatics (2022)
-
Accelerating antibiotic discovery through artificial intelligence
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.