Abstract
We have created a transgenic reporter for virus infection and used it to study Nora virus infection in Drosophila melanogaster. The transgenic construct, Munin, expresses the yeast transcription factor Gal4, tethered to a transmembrane anchor via a linker that can be cleaved by a viral protease. In infected cells, liberated Gal4 will then transcribe any gene that is linked to a promoter with a UAS motif, the target for Gal4 transcription. For instance, infected cells will glow red in the offspring of a cross between the Munin stock and flies with a UAS-RFPnls transgene (expressing a red fluorescent protein). In such flies we show that after natural infection, via the faecal-oral route, 5–15% of the midgut cells are infected, but there is little if any infection elsewhere. By contrast, we can detect infection in many other tissues after injection of virus into the body cavity. The same principle could be applied for other viruses and it could also be used to express or suppress any gene of interest in infected cells.
Similar content being viewed by others
Introduction
Nora virus is a currently unclassified picorna-like virus that naturally infects the fruit fly Drosophila melanogaster1. This virus causes persistent infection with no obvious symptoms or pathogenesis. Like many other picorna-like viruses, it is spread by faecal-oral transmission2. A striking characteristic of the Nora virus infection is that the virus titre differs enormously between individual fruit flies that are kept in the same vial and the titres may vary from 102 up to 1010 viral genomes per fly2,3. We study Nora virus as an animal model for persistent virus infection.
Virus detection is crucial in all virus research and several methods have been used for this purpose. Viruses can be detected by their specific pathogenesis in human and animal models, changes in morphology and viability of cultured cells, detection of viral nucleic acid or use of antibodies and other molecular markers4,5,6,7,8,9,10. The Nora virus does not cause obvious symptoms and there are no antibodies available for immunohistochemical labelling. This encouraged us to develop a new molecular technique to facilitate viral studies in the fruit fly. Besides detecting the mere presence of viruses, this technique permits virus-triggered expression of any gene of choice specifically in virus-infected cells. This can be used to express a molecular marker for virus detection or, for example, for targeted expression or suppression of genes involved in the host’s viral immunity.
Several techniques are developed to modify gene expression in Drosophila melanogaster, for example the popular Gal4-UAS system11. Transgenic fruit flies can express the transcription factor Gal4 from the yeast Saccharomyces cerevisiae, which acts on a yeast-specific promoter motif, the upstream activating sequence (UAS). Tens of thousands of such “driver” fly stocks are available for the Gal4-UAS expression system, which in a combinatorial way allow expression of virtually any gene in any tissue. A common application is to express an RNA hairpin that feeds into the fruit fly’s RNA interference (RNAi) pathway, in order to suppress the expression of an endogenous Drosophila gene.
We designed a Gal4-UAS-based expression system that depends on the activity of the Nora virus-encoded protease. For this purpose we constructed a Drosophila driver stock with constitutive and ubiquitous expression of a target to the viral protease. The target is the Gal4 protein fused to a transmembrane anchor, via a linker that has a cleavage site for the virus-encoded protease (Fig. 1A). This anchors Gal4 in the cytoplasmic space, preventing this transcription factor from entering the nucleus and from initiating transcription from UAS promoters. Upon virus infection the linker is cleaved by the virus protease, allowing Gal4 to enter the nucleus and initiate gene expression. Like in the system reported recently by Ren et al.10 this amplifies a signal created by the action of a virus-specific protease. In addition, the system described here will permit expression of any gene of the researcher’s choice specifically in virus-infected cells.
Nora virus replication in flies infected by oral ingestion.
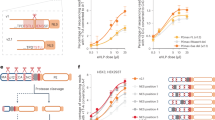
(A, A′) Construction and molecular mechanism for the Munin virus reporter. (A) The transformation vector pUASTattB21 was adapted to constitutively drive expression of the inserted gene and the Munin reporter construct was inserted. (A′) The Munin reporter is ubiquitously expressed in the fly but the transcription factor Gal4 can enter nucleus only when there is an active virus-encoded protease in the cytoplasm. MCS, multiple cloning site. α–tubP, α-tubulin 84B promoter. lb, late bloomer transmembrane region. NV, Nora virus protease target site. (B–D) RFPnls expression in the Nora virus-infected midgut. C and D are post-processed to enhance the red fluorescence. (E) Correlation between RFPnls expression and the amount of Nora virus RNA excreted in the faeces, as determined by quantitative RT-PCR. Each data point represents a single fly. The graph shows the results of 56 flies from three independent experiments. An additional 16 low-titre flies were not scored for fluorescence. ND, not detected. (F,G) No obvious difference between infected and uninfected whole flies when observed from the outside. Infection was confirmed in F by a detected high virus titre in the faeces. Scale bars show 100 μm in (B), 25 μm in (C,D) and 250 μm in (E–G).
We call this virus protease-dependent gene expression system Munin. As an application we used it to drive red fluorescent protein expression for in vivo detection of Nora virus in the fruit fly. This reporter system allowed us to determine which tissues the Nora virus naturally infects.
Results and Discussion
The Munin reporter construct contains a tetraspanin transmembrane anchor, fused to Gal4 via a Nora virus protease cleavage motif (Fig. 1A,A′). The construct was inserted into a Drosophila P element vector and transgenic fly lines were generated. The w; UAS-RFPnls; Munin reporter fly stock also expresses red fluorescent protein tagged with a nuclear localisation signal (RFPnls), under the control of the Munin reporter. To test the Munin reporter we established a persistent Nora virus infection in the fly stock. The flies were thus environmentally exposed to the virus via faeces deposited in the fly vial and they ingested the virus with the food. We dissected adult flies one week after eclosion and examined their abdominal organs for RFPnls expression by fluorescence microscopy. Our previous results suggested that Nora virus is an enteric virus2 and in line with this we observed about 20 cells in the midgut with strong red fluorescence (Fig. 1B). Enhancing the red fluorescence channel by image processing revealed a large number of more weakly expressing cells. Thus, RFPnls expression varies considerably between the infected cells (Fig. 1C), possibly indicating progressive infection of previously uninfected cells. In total we find that 5–15% of the midgut cells expressed RFPnls.
A characteristic of the Nora virus infection is that the virus titres differ several orders of magnitude between the individuals in an infected fly stock. This is reflected by highly variable amounts of viral RNA excreted in the faeces3. To identify high-titre individuals we determined the amount of Nora virus genomic RNA in the faeces by quantitative RT-PCR. Blinded from the viral quantities we also examined the abdominal organs for RFPnls expression. We observed RFPnls expression in the midgut in all flies with a high virus titre and in some flies with an intermediate titre, but no RFPnls was observed in flies with a low virus titre (Fig. 1C) or in uninfected flies (Fig. 1D). The quantification is shown in Fig. 1E. We conclude that the Munin reporter is a reliable reporter of Nora virus infection. The low levels of Nora virus RNA in the low-titre flies probably originate from virus that has been ingested with the food without producing an active infection in the animal. We have previously shown that the virus can be cleared from such low-titre animals by serial transfer of the flies to fresh uncontaminated food2, as expected if no cells are infected. Thus, expression of the Munin reporter is tightly correlated with the presence of a productive Nora virus infection.
To test the general usefulness of the Munin reporter design, we constructed a corresponding reporter fly stock for the Drosophila C virus. When injected with the C virus, the flies showed classical signs of disease by swelled abdomen and death after a few days. C virus-infected flies also showed obvious RFP expression in their abdomen, whereas uninfected flies did not express RFP (Supplementary Fig. S1). Importantly, C virus reporter flies injected with Nora virus did not express RFP two weeks after the viral infection. This further supports virus-specific expression by the Munin reporter and it also suggests that the reporter can be designed to recognise only a certain group of virus.
The precise labelling of virus-infected cells with the Munin reporter allowed us for the first time to investigate in detail which tissues that are infected by Nora virus. We investigated flies up to four weeks after eclosion, but we never observed RFPnls expression in any other tissues besides the midgut. Four different cell types have been described in the Drosophila midgut epithelium. A self-renewing population of intestinal stem cells produces enteroblasts, which differentiate to enterocytes (about 90%) or enteroendocrine cells (about 10%). Enterocytes are morphologically distinguishable from the other cell types by their big nuclei12,13. We only saw RFPnls in cells with a big nucleus, indicating that Nora virus has tropism for enterocytes (Fig. 1C; see also Fig. 2C). Quantitative RT-PCR for Nora virus RNA in different tissues supports the conclusion that Nora virus only infects the alimentary canal (Fig. 3 and ref. 2). Nora virus RNA was mainly found in the gut, whereas viral RNA concentration in the reproductive tract and other parts of the body was at least a 1000-fold lower (Fig. 3 and Supplementary Fig. S2). Such low concentrations are most likely due to contamination from faeces during dissection of the flies.
Nora virus replication in flies infected by injection.
(A–D) All injected animals showed RFPnls expression in epidermis, cardia and midgut. (A) RFPnls was expressed in a spotted pattern in epidermal cells. In addition, stray light from fluorescing cells lights up the whole body, except where it is pigmented. (A′) RFPnls expression in the wing veins. (B) Only faint autofluorescence was seen in uninfected flies, at the same level as in wild-type control flies. (C) Midgut. Only enterocytes (cells with a big nucleus) expressed RFPnls. (D) Cardia. (E–I) In rare cases crop, Malpighian tubules, hindgut, rectum and parts of the reproductive tract expressed RFPnls. (E) Malpighian tubules (Mt), here shown on each side of midgut (mg) and hindgut (hg). (F) Crop. (G) Hindgut and rectum. The posterior section of hindgut (hg), rectal epithelium (r) and rectal papillae (rp). (H) The male reproductive tract. Testis (t), seminal vesicle (sv), accessory gland (ag) and ejaculatory duct (ed). This specimen has strong expression in the proximal end of the testes, just next to the testicular ducts that connect the testes to the seminal vesicles. (I) Parts of the female reproductive tract. Uterus (u), seminal receptacle (sr) and spermathecae (s), with strong expression in cells surrounding the melanized cuticle. Scale bars show 250 μm in A,B, 25 μm in C and 100 μm in (D–I).
Quantitative RT-PCR for Nora virus RNA in different body parts.
Animals infected by oral ingestion have viral RNA essentially only in the gut, whereas animals injected with the virus have a substantial amount of viral RNA in additional tissues. “Body” refers to all tissues except gut and reproductive tract. The flies were four weeks old at the time for the analysis. Error bars show standard deviation. Data are based on two independent experiments involving in total 10 flies in each group. Data for each fly are shown in Supplementary Fig. S2.
As most viral studies performed in Drosophila are based on animals that are infected by injection of the virus into the blood, we also investigated Nora virus replication in flies infected this way. In all injected animals we saw RFPnls expressed in epidermis and cardia in addition to the midgut (Fig. 2A–D). In rare cases we also observed RFPnls expression in the crop, Malpighian tubules, the posterior part of hindgut, rectum or the reproductive tract (Fig. 2E–I). For the reproductive tract, our data do not allow characterisation of which specific cells are infected, but this may include sperm heads in the proximal end of the testes, epithelial cells and secretory cells in the spermathecae (Fig. 2H,I). Quantitative RT-PCR show that injected animals had increased amount of viral RNA in the body and in the reproductive tract (Fig. 3 and Supplementary Fig. S2). Furthermore, all injected flies had a high virus titre and there was no longer a group of animals with low virus titre as seen after oral ingestion (Supplementary Fig. S3). Altogether, we conclude that additional tissues are infected after injection of the virus into the blood. Thus, Nora virus infection by injection causes a radically different outcome compared to infection by oral ingestion.
Although Nora virus has the capacity to infect other tissues in the fruit fly, it remains contained in the midgut after oral ingestion. The gut thus constitutes an efficient barrier that prevents Nora virus spread to other tissues. This could also be the case for other viruses that are naturally transmitted by the faecal-oral route, for example the Drosophila C virus. When infecting by oral ingestion, the C virus is usually non-pathogenic14, but it causes lethality within a few days after injection into the blood15. C virus infection after oral ingestion is surprisingly little studied. The virus replicates in the larval midgut16, but to our knowledge it is not yet described which tissues that are infected in the adult fly. In contrast, the pathogenesis after infection by injection is well characterised. The C virus replicates in many different tissues, including the visceral muscles15,17, causing obstruction of the gut and starvation or dehydration to death18. Thus, systemic infection of the C virus has dramatic consequences to the fly. The gut epithelium might therefore play a crucial role in limiting spread of the Drosophila C virus, as for the Nora virus. However, in studies of insect viral immunity it is currently common practise to inject the virus into the blood, thereby bypassing the normal infection route. The gut is therefore very little studied for its role in viral immunity.
Our results show that the Munin construct is a useful tool to study Nora virus infection and that the technique can be adapted also to other viruses and it can most likely be adapted to other hosts. We had initially hoped to use the Munin reporter to score viral infection directly in living flies, but this was only possible with animals infected by injection. RFPnls expression is too faint to detect gut infections with our instruments without dissection of the abdomen (Fig. 1F,G). Future developments could be to combine Munin with other UAS-dependent reporters, or to use more sensitive imaging equipment.
Material and Methods
Munin reporter, fly strains and fly maintenance
We constructed the Munin reporter gene by inserting the first 216 bp (encoding the first two transmembrane helices) of the Drosophila tetraspanin late bloomer, followed by 180 bp cDNA sequence of the Nora virus genome encoding the VP4B-VP4C junction protease cleavage site19 and the complete 2646 bp Gal4 gene, in a cloning vector. In the corresponding Drosophila C virus reporter we inserted a 180 bp cDNA sequence encoding the VP2-VP3 junction protease cleavage site, as determined for the closely related cricket paralysis virus20. For constitutive and ubiquitous expression we modified the pUASTattB transformation vector21 by replacing the UAS promoter with the α-tubulin 84B promoter22,23 and we inserted the Munin reporter construct in the new transformation vector. We call the final construct pMunin. The complete sequence of the final transformation plasmid is available in the Supplementary Appendix. The Munin reporter was transformed into y1, w1118; PBac (y+−attP-9A) VK00013 flies, which have a predestined landing site at 3L:1919745824. Transformation was performed by BestGene Inc., USA.
A w1118; P{w[+mC] = UAS-RedStinger}4/CyO stock25 was obtained from the Bloomington Drosophila Stock Center. We refer to RedStinger as RFPnls in this work. The flies were reared on standard mashed potato-agar fly food26 and kept in 25 °C and 60% relative humidity. All fly stocks were treated with Tetracycline as described27 to remove potential Wolbachia infections.
Virus and virus infection procedures
Drosophila Nora virus strain Umea 2007 was derived from an infectious cDNA clone19 (accession GQ257737) and had been continuously passaged in w; +; RelishE23 flies28 for more than two years before performing the experiments described in this work.
Flies were infected by environmental exposure to the virus via contamination of the food by the faeces of their virus-infected parents, or by microinjection of a virus-containing suspension into the blood. For the latter, Nora virus suspension was prepared by homogenising approximately 100 infected w; +; RelishE23 flies in 0.5 ml PBS and filtering the suspension through a 0.45 μm filter. Using an Inject + matic microinjector (Inject + matic, Geneva, Switzerland), we injected approximately 100 nl of the virus suspension into the thorax of each fly via the thin cuticle under the wing hinge. Infected this way, the flies show an 100–1000 fold increase of Nora virus RNA within ten days after injection2. For environmental exposure, we established a persistently infected fly stock by injecting the virus into flies and we used their progeny in the experiments.
We used a Drosophila C virus strain of uncertain origin that has been kept in our lab. The region around the protease cleavage site was sequenced (see Supplementary Appendix) and found 97% identical to Drosophila C virus strain EB (accession AF014388). The virus was propagated in Drosophila S2 cells until 50% of the cells had lysed. The suspension was freeze-thawed three times to completely lyse the cells, diluted 105 times in phosphate-buffered saline, pH 7.5 (KCl 0.2 g/l, KH2PO4 0.2 g/l, NaCl 8 g/l, Na2HPO4 anhyd. 1.15 g/l) and filtered through a 0.22 μm filter before injection into flies as described above.
Nora virus quantification
Nora virus titres were determined by one-step quantitative RT-PCR as described2,19. For the amount of Nora virus in the faeces, we kept individual flies in clean fly vials and we exchanged to clean vials once per day. On the third day we transferred the flies to 1.5 ml centrifuge tubes with 10–20 μg fly food and the flies were kept in these tubes for 18 h. The viral RNA deposited with faeces was collected by vortexing the tubes with Total RNA Lysis Solution provided in the Aurum Total RNA Mini Kit (Bio-Rad).
Microscopy
Dissected organs were fixed in 3.5% paraformaldehyde for 10 min at room temperature, permeabilised with 0.1% Triton X-100 for 10 min, incubated in CytoPainter Phalloidin iFlour 488 (Abcam) for 30 min and mounted in ProLong Gold antifade reagent with DAPI (Life Technologies). Images were acquired in Zeiss ApoTome.2 and Zeiss Lumar.V12 microscopes and the contrast was enhanced with the Adobe Photoshop software. Images of control and infected animals were always processed the same way. Contrast was equally enhanced in Figs 1 and 3.
Etymology
In the Norse mythology, Hugin and Munin is a pair of ravens that fly over the worlds every day. They see everything that happens and they bring all news to the god Odin, who is thereby kept informed29.
Additional Information
How to cite this article: Ekström, J.-O. and Hultmark, D. A Novel Strategy for Live Detection of Viral Infection in Drosophila melanogaster. Sci. Rep. 6, 26250; doi: 10.1038/srep26250 (2016).
References
Habayeb, M. S., Ekengren, S. K. & Hultmark, D. Nora virus, a persistent virus in Drosophila, defines a new picorna-like virus family. J. Gen. Virol. 87, 3045–3051 (2006).
Habayeb, M. S. et al. The Drosophila Nora virus is an enteric virus, transmitted via feces. J. Invertebr. Pathol. 101, 29–33 (2009).
Habayeb, M. S., Ekström, J.-O. & Hultmark, D. Nora virus persistent infections are not affected by the RNAi machinery. PLos One 4, e5731 (2009).
Chesebro, B. & Wehrly, K. Development of a sensitive quantitative focal assay for human immunodeficiency virus infectivity. J. Virol. 62, 3779–3788 (1988).
Felber, B. K. & Pavlakis, G. N. A quantitative bioassay for HIV-1 based on trans-activation. Science 239, 184–187 (1988).
Lutz, A., Dyall, J., Olivo, P. D. & Pekosz, A. Virus-inducible reporter genes as a tool for detecting and quantifying influenza A virus replication. J. Virol. Methods 126, 13–20, 10.1016/j.jviromet.2005.01.016 (2005).
Olivo, P. D., Frolov, I. & Schlesinger, S. A cell line that expresses a reporter gene in response to infection by Sindbis virus: a prototype for detection of positive strand RNA viruses. Virology 198, 381–384, 10.1006/viro.1994.1046 (1994).
Sivaraman, D., Biswas, P., Cella, L. N., Yates, M. V. & Chen, W. Detecting RNA viruses in living mammalian cells by fluorescence microscopy. Trends Biotechnol. 29, 307–313, 10.1016/j.tibtech.2011.02.006 (2011).
Hwang, Y. C., Chen, W. & Yates, M. V. Use of fluorescence resonance energy transfer for rapid detection of enteroviral infection in vivo. Appl. Environ. Microbiol. 72, 3710–3715, 10.1128/AEM.72.5.3710-3715.2006 (2006).
Ren, Q. et al. A Dual-reporter system for real-time monitoring and high-throughput CRISPR/Cas9 library screening of the hepatitis C virus. Sci. Rep. 5, 8865, 10.1038/srep08865 (2015).
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Ohlstein, B. & Spradling, A. The adult Drosophila posterior midgut is maintained by pluripotent stem cells. Nature 439, 470–474 (2006).
Micchelli, C. A. & Perrimon, N. Evidence that stem cells reside in the adult Drosophila midgut epithelium. Nature 439, 475–479 (2006).
Jousset, F. X. & Plus, N. Étude de la transmission horizontale et de la transmission verticale des picornavirus de Drosophila melanogaster et de Drosophila immigrans [Study of the vertical transmission and horizontal transmission of Drosophila melanogaster and Drosophila immigrans picornavirus]. Ann. Microbiol. (Paris) 126, 231–249 (1975).
Jousset, F. X., Plus, N., Croizier, G. & Thomas, M. Existence chez Drosophila de deux groupes de Picornavirus de propriétés sérologiques et biologiques différentes. C. R. Acad. Sci. (Paris), Série D 275, 3043–3046 (1972).
Lautié-Harivel, N. Drosophila C virus cycle during the development of two Drosophila melanogaster strains (Charolles and Champetières) after larval contamination by food. Biol. Cell 76, 151–157 (1992).
Lautié-Harivel, N. & Thomas-Orillard, M. Location of Drosophila C virus target organs in Drosophila host population by an immunofluorescence technique. Biol. Cell 69, 35–39 (1990).
Chtarbanova, S. et al. Drosophila C virus systemic infection leads to intestinal obstruction. J. Virol. 88, 14057–14069, 10.1128/JVI.02320-14 (2014).
Ekström, J.-O. et al. Drosophila Nora virus capsid proteins differ from those of other picorna-like viruses. Virus Res. 160, 51–58, 10.1016/j.virusres.2011.05.006 (2011).
Tate, J. et al. The crystal structure of cricket paralysis virus: the first view of a new virus family. Nat. Struct. Biol. 6, 765–774, 10.1038/11543 (1999).
Bischof, J., Maeda, R. K., Hediger, M., Karch, F. & Basler, K. An optimized transgenesis system for Drosophila using germ-line-specific φ C31 integrases. Proc. Natl. Acad. Sci. USA 104, 3312–3317, 10.1073/pnas.0611511104 (2007).
Theurkauf, W. E., Baum, H., Bo, J. & Wensink, P. C. Tissue-specific and constitutive α-tubulin genes of Drosophila melanogaster code for structurally distinct proteins. Proc. Natl. Acad. Sci. USA 83, 8477–8481 (1986).
Thibault, S. T. et al. A complementary transposon tool kit for Drosophila melanogaster using P and piggyBac. Nat. Genet. 36, 283–287, 10.1038/ng1314 (2004).
Venken, K. J., He, Y., Hoskins, R. A. & Bellen, H. J. P[acman]: a BAC transgenic platform for targeted insertion of large DNA fragments in D. melanogaster. Science 314, 1747–1751, 10.1126/science.1134426 (2006).
Barolo, S., Castro, B. & Posakony, J. W. New Drosophila transgenic reporters: insulated P-element vectors expressing fast-maturing RFP. Biotechniques 36, 436–440, 442 (2004).
Yang, H., Kronhamn, J., Ekström, J.-O., Korkut, G. G. & Hultmark, D. JAK/STAT signaling in Drosophila muscles controls the immune response against parasitoid infection. EMBO Rep. 16, 1664–1672, 10.15252/embr.201540277 (2015).
Teixeira, L., Ferreira, A. & Ashburner, M. The bacterial symbiont Wolbachia induces resistance to RNA viral infections in Drosophila melanogaster. PLos Biol. 6, e2, 10.1371/journal.pbio.1000002 (2008).
Hedengren, M. et al. Relish, a central factor in the control of humoral but not cellular immunity in Drosophila. Mol. Cell 4, 827–837 (1999).
Sturluson, S. Edda. Codex Upsaliensis (DG11), Carolina Rediviva, Uppsala University, Sweden. (ca 1220).
Acknowledgements
We thank Sajna Anand Sadanandan for preliminary experiments and for constructive discussions. This research was supported by grants from the Swedish Research Council, the Swedish Cancer Society, the Academy of Finland and the Sigrid Juselius Foundation.
Author information
Authors and Affiliations
Contributions
J.-O.E. and D.H. initiated the project. J.-O.E. designed the experimental approach and performed the research. J.-O.E. and D.H. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ekström, JO., Hultmark, D. A Novel Strategy for Live Detection of Viral Infection in Drosophila melanogaster. Sci Rep 6, 26250 (2016). https://doi.org/10.1038/srep26250
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep26250
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.