Abstract
A fluent method for gene targeting to establish protein tagged and ligand inducible conditional loss-of-function alleles is described. We couple new recombineering applications for one-step cloning of gRNA oligonucleotides and rapid generation of short-arm (~1 kb) targeting constructs with the power of Cas9-assisted targeting to establish protein tagged alleles in embryonic stem cells at high efficiency. RAC (Recombineering And Cas9)-tagging with Venus, BirM, APEX2 and the auxin degron is facilitated by a recombineering-ready plasmid series that permits the reuse of gene-specific reagents to insert different tags. Here we focus on protein tagging with the auxin degron because it is a ligand-regulated loss-of-function strategy that is rapid and reversible. Furthermore it includes the additional challenge of biallelic targeting. Despite high frequencies of monoallelic RAC-targeting, we found that simultaneous biallelic targeting benefits from long-arm (>4 kb) targeting constructs. Consequently an updated recombineering pipeline for fluent generation of long arm targeting constructs is also presented.
Similar content being viewed by others
Introduction
Gene targeting utilizing endogenous homologous recombination has been central to the emergence of the repertoire of genetic technologies including conditional mutagenesis, ligand-inducible loss-of-function and endogenous protein tagging1. However these technologies have been largely restricted to the few model systems that support efficient gene targeting, notably yeast and mouse.
Nuclease-assisted mutagenesis, pioneered by zinc-finger/Fok1 nucleases2 and now exemplified by the RNA-guided CRISPR-Cas9 nuclease3,4, is fundamentally changing genetic engineering. Nuclease-assisted mutagenesis, in particular CRISPR/Cas9, unlocks the advantages of directed mutagenesis for many genomes5,6,7. It also presents new potential for improvements to existing genetic technologies such as gene targeting to establish protein-tagged or conditional alleles. In contrast to the extraordinarily rapid progress with Cas9 nuclease-assisted mutagenesis in many applications, the application of Cas9 to gene targeting has been less straightforward. In contrast to nuclease-assisted mutagenesis, nuclease-assisted targeting requires the assembly of gene specific targeting constructs. Before the advent of recombineering8,9 this was often tedious and laborious. Recombineering greatly simplified the methods for rapid assembly of long homology arm (>4 kb each arm) targeting constructs10 and has been adapted for high-throughput production, resulting in more than 16,000 targeted mouse genes11.
Particularly since the advent of green fluorescent protein (GFP)12,13 protein tagging has empowered cell biology and functional genomics with fluent access to protein localization, live cell imaging and generic affinity purification for proteomics and chromatin immunoprecipitation amongst other applications. Although knock-in targeting into the endogenous gene is conceptually the best way to tag a chosen protein, to date the obstacles involved in achieving this optimum outcome have been circumvented by alternative approaches such as the use of fosmid and BAC transgenes14,15,16.
Because it is highly efficient, nuclease-assisted targeting not only enables gene targeting in many previously inaccessible genomes but could also expedite simultaneous homologous recombination on both alleles in diploid hosts17. Thereby, conditional loss-of-function strategies could be implemented in one step of simultaneous targeting.
Here our first aim was to develop simple and rapid methods for Cas9-assisted targeting for protein tagging. Our second aim was to evaluate whether Cas9 could be used to routinely achieve simultaneous biallelic targeting. Because biallelic targeting would open a variety of options, we explored the possibility that it could be achieved using CRISPR/Cas9 and applied to protein tagging and conditional mutagenesis.
Results
One-step gRNA expression plasmid construction
We applied a recent recombineering advance based on annealing recombination mediated by full-length RecE/RecT18 to establish a one-step method to generate guide RNA (gRNA) expression plasmids (Fig. 1a). An established U6 expression plasmid6 was modified by improving the tracrRNA sequence19 and incorporating the ccdB toxin gene20 at the gRNA cloning site. A batch of this plasmid was digested with BstZ171 and NheI, column purified and 200 ng aliquots mixed with 50 pmol oligonucleotides encoding 20 nucleotide gRNAs. The chosen 20nt gRNA sequence is flanked by homology arms (i.e. identical sequences) to sequences in the plasmid either side of the restriction cleavages (Supplementary Table 1). The gRNA sequences are incorporated into the pBR322-U6 expression plasmid upon co-electroporation into E. coli induced to express RecE/RecT. Due to its toxicity, the ccdB gene ensures no carryover of the parent plasmid so that virtually all ampicillin resistant colonies carry the gRNA insert. For economy, we tested different homology arm lengths to find that 19 bp each is sufficient, so that the single stranded oligonucleotides need only be 60mers. Also, they do not need to be HPLC purified (Fig. 1b). In a recent application, we generated 304 gRNA expression plasmids out of 312 attempts at the first go using 80 mer oligonucleotides and 96 well boxes in a simplified recombineering pipleine21. All 8 initial failures were obtained on the second go using newly synthesized oligonucleotides, indicating that oligonucleotide quality was the cause of the failure (M. Sarov, AFS; unpublished observations). Hence this one-step method to build gRNA expression plasmids is simple fast and cost-effective.
One-step generation of gRNA expression plasmids.
(a) Scheme of the cloning strategy showing pBR322-U6-ccdB-cm-tracrRNA, which is digested with BstZ171 and NheI to release the ccdB-cm cassette and then co-electroporated with a single stranded oligonucleotide into GB05 after arabinose induction of pSC101-BAD-ETγA. The oligonucleotides contained the 20nt gRNA sequence flanked by variable lengths of sequence identity (i.e. homology boxes) to the U6 promoter and tracrRNA regions. (b) Ampicillin resistant colonies after co-electroporation of single stranded oligonucleotides each containing the same 20nt gRNA flanked by homology boxes of different lengths, either 14nt (50mer), 19nt (60mer) or 31nt (84mer). In addition the oligonucleotide included the GG dinucleotide required for the U6 promoter between the 5′ homology arm and the gRNA. Error bars, s.d. (n = 3). RP1 and HPLC-reverse phase or high performance liquid chromatography purifications of the oligonucleotides by the manufacturer. Six clones from each experiment were sequenced. From left to right, the correct cloned product was observed 0/6; 0/6; 3/6; 5/6; 6/6; 6/6 times respectively.
One-step short-arm targeting construct
Potentially, an advantage of nuclease-assisted targeting lies with the employment of short-arm (~1 kb) targeting constructs because genotyping by junction PCR can be employed. This practical advantage is not available for conventional targeting in mouse ESCs because longer arms are required to achieve workable targeting efficiencies1. Because nuclease-assisted targeting is mechanistically different than conventional targeting, the advantages of short-arm targeting constructs are worth re-examination.
Furthermore using full-length RecE/RecT18, short arm targeting constructs can be built rapidly in one recombination step using a 4-way reaction (Fig. 2a). To complement this new application, we built a recombineering-ready plasmid series for C-terminal protein tagging. Coding regions for the auxin degron (AID22), the fluorescent protein Venus23 and the enzymes for biotin proximity labeling BirM24 and APEX225,26 have been cloned into the same context in a plasmid series based on the R6K origin. The context includes on the 5′ side a flexible protein linker based on two copies of the Ty1 peptide, which serves two additional purposes. It is an epitope for the useful monoclonal antibody, BB227 and its coding region serves as one of the standard 50 bp homology boxes for the recombineering reactions (coloured purple in Fig. 2; Supplementary Table 1). On the 3′ side the other standard 50 bp homology box is coloured green. In between the homology boxes reside the coding regions for the various protein tags and the neomycin (neo) antibiotic resistance gene for selection in one of two conformations. If the gene is expressed in the targeted cells, then the polycistronic strategy for selection using 2A peptides28 can be employed. If the gene is not expressed sufficiently for polycistronic selection, then the PGK promoter-driven strategy can be employed. In both cases, the neomycin gene includes an embedded prokaryotic promoter for expression of kanamycin resistance in E. coli10 and is flanked by rox sites for later removal by Dre recombinase29 if required. To utilize the tagging cassettes in recombineering, the plasmids are digested with NheI, which cuts at the outside ends of the homology arms. The plasmids are based on the R6K origin, which will not replicate in standard E. coli strains because they lack its cognate replication A protein, Pir30. Consequently the major source of unwanted recombineering background, which is carry-over of the template plasmid, is eliminated and the tagging cassette does not need to be gel purified after restriction digestion31,21.
Two different recombineering methods to generate targeting constructs for protein tagging.
(a) The 4-way recombination strategy mediated by full-length RecET to generate short arm targeting constructs is illustrated. The C-terminal tagging cassette consisting of the Ty1 linker, tag coding region (either AID, Venus, BirM or APEX2), T2A peptide and promoterless neomycin gene (neo) in R6K-Ty1-Tag-T2A-neo is released from its vector by NdeI digestion. The p15A origin and chloramphenicol (cm) cassette in p15A-cm-ccdB-amp is released by NdeI restriction digestion and mixed with 5′ and 3′ homology arms (HA) that were generated by PCR from genomic or BAC DNA. At their 5′ ends the PCR primers include 50nts of sequence homology to the respective ends of the cassettes indicated by the four colours (Supplementary Table 1). The mixture is electroporated into E. coli induced with arabinose to express RecE/RecT/Redγ from either the genome (strain GB05-dir) or an expression plasmid (pSC101-BAD-ETγA). Double selection for chloramphenicol and kanamycin resistance identifies the targeting vector. Neither of the parental plasmids will be selected because the R6K origin will not replicate in GB05 and ccdB counterselection blocks p15A-cm-ccdB-amp propagation. Brown triangles denote the 34 bp rox sites, red circles denote stop codons and pA is a polyadenylation signal. (b) Recombination efficiencies promoted by full-length RecE/RecT. The 4-way mixture illustrated in (a) based on Ash2l using the T2A version was electroporated into either arabinose induced GB05 containing pSC101-BAD-ETγA plasmid or GB05-dir, in which the PBAD-ETγA operon has been integrated into the genome. The homology arms were amplified from either genomic DNA (gDNA) or a BAC carrying the target gene (BAC). Correct recombinants were retrieved by selection for chloramphenicol (cm) and kanamycin (kan) resistance, followed by restriction mapping. In all four cases, at least 23/24 clones were correct. (c) Diagram of the Red recombineering method to generate long arm targeting constructs for C-terminal tagging. A BAC clone of the gene to be targeted is obtained from a genome provider and transformed with pSC101Redγβα. Oligonucleotides including 50nt homology either side of the chosen C-terminus in the BAC are attached to the rpsL-gentamycin cassette by PCR. After arabinose induction of Red operon expression, the PCR product is electroporated into the E. coli host. Selection for gentamycin resistance identifies the candidate recombinants. The rpsL-gentamycin cassette is flanked by the same standard homology arms illustrated in purple and green in (a). In the next step, PCR is used to attach 50nt homology boxes, which define the ~9 kb region to be copied from the BAC into the p15A-ampicillin subcloning vector. Selection for gentamycin and ampicillin resistance identifies the candidate subcloned intermediate targeting constructs. Protein tags from the R6K series can be readily introduced into these intermediate TCs using the standard purple and green homology boxes with selection for kanamycin and streptomycin resistance.
To build the short-arm targeting constructs, the ~1 kb homology arms are amplified by PCR using either genomic DNA from the target cells, which is therefore isogenic, or BAC DNA carrying the target gene as the template. For PCR, the 60mer oligonucleotides include not only gene specific PCR primers at their 3′ ends but also the four standard 50 bp homology boxes to the tagging cassette (Fig. 2a; purple, green) and the subcloning vector (Fig. 2a; brown, blue). The subcloning vector is also prepared by restriction digest of p15A-cm-ccdB-amp. The presence of the ccdB toxin gene in the unwanted part of this plasmid ensures that only the p15A origin-chloramphenicol (cm) part will be propagated. The two restriction digests are together mixed with the two PCR products and co-electroporated into E. coli after arabinose induction of full-length RecE, RecT and Redγ expression from the BAD-ETγ operon either integrated into the E. coli chromosome (GB05-dir) or in pSC101-BAD-ETγA. Whether amplified from genomic or BAC DNA, or whether RecETγ was expressed from the E. coli chromosome or plasmid, virtually all colonies contained the intended 4-way recombination product upon double selection for chloramphenicol and kanamycin (Fig. 2b; Supplementary Figure 1a). Applying the same principles, a one-step, 5-way recombination reaction can be used to generate targeting constructs for loxP conditional alleles (Supplementary Figures 1b and 2). In our experience, failure to obtain the intended targeting construct is rare and almost always due to errors in oligonucleotide synthesis or PCR amplification.
Recombineering for long-arm targeting constructs
Figure 2c illustrates the recombineering scheme for building long arm (~4 to 5 kb) C-terminal targeting constructs. In this application, the Red operon expression plasmid, pSC101Redγβα is introduced into an E. coli host carrying the selected BAC, which is then modified using a selectable/counterselectable cassette based on gentamycin and rpsL flanked by the same purple and green homology boxes employed in Fig. 2a. After recombineering into the chosen C-terminus, this region is then subcloned by recombineering32 to establish the intermediate targeting construct that can now be modified with any cassette from the R6K plasmid series.
Implementing the auxin degron
Amongst the variety of possible examples, we present work with the auxin degron for two reasons. First, the auxin degron is a ligand-regulated conditional loss-of function strategy that requires the introduction of an effector protein, rice Tir1, which is an F-box protein that binds auxin and then conveys the auxin degron, AID (Auxin Inducible Degron; also known as Auxin/Indole 3-acetic acid or Aux/IAA repressor), to the SCF (Skp1, Cullin, F-box protein) E3 ubiquitin ligase complex for polyubiquitinylation and subsequent degradation by the proteasome22.
Second, for autosomal genes, both alleles need to be tagged (or tagged and mutagenized) to achieve inducible loss-of-function. Hence targeted tagging with the auxin degron requires more work to establish than tagging for fluorescent studies or proximity mapping. Our initial applications with rice Tir1 showed only partial depletion of nuclear proteins (data not shown). Hence we codon optimized Tir1 for mammalian expression, reduced the number of CpG dinucleotides, added a nuclear localization signal (nls; Supplementary Figure 3a) and targeted it to the Rosa26 locus with expression driven by the strong CAG promoter (Supplementary Figure 4). The AID coding region was similarly codon optimized (Supplementary Figure 3b).
Comparing short- and long-arm Cas9 assisted targeting
Short-arm (~1 kb each arm) and long-arm (~4.5 kb each arm) targeting constructs were generated for targeting AID to the C-terminae of two genes, Ash2l and Wdr82, in the Rosa26-CAG-nlsTir1 ESCs. To ensure that Cas9 did not also cut the targeting construct, small gaps were introduced between the homology arms to omit parts of the gRNA target site. The strategy for Ash2l, which included the testing of two different gRNAs, is presented in Fig. 3. Targeting efficiency was scored for monoallelic and biallelic targeting (Table 1). High frequencies of monoallelic targeting were achieved with all four targeting constructs. As expected, most monoallelically targeted ESCs showed Cas9-induced damage on the other allele (10/14; Supplementary Figure 5). Consequently the status of the second allele must be evaluated. With this precaution, the advantages of short-arm targeting constructs can be readily applied to monoallelic applications such as tagging with a fluorescent protein, targeting the sex chromosomes in male cells or heterozygous targeting to generate a mouse. In addition to Ash2l and Wdr82, to date we have successfully monoalleically tagged 11 (of 11) genes in ESCs (AG, MO, Davi Coe Torres, KA, AFS; unpublished observations).
Cas9-assisted targeting at the C-terminus of Ash2l.
(a) Ash2l sequence near its stop codon (highlighted in red) is shown with two chosen gRNAs and their corresponding Cas9 cleavage sites (red triangles, PAM sequences in blue). The Ash2l AID targeting constructs are also illustrated indicating the cassette fusion junctions and the 29 bp deletion to prevent Cas9 loaded with either gRNA 1 or 2 from cleaving the targeting construct. Rox sites (brown triangles), which do not interrupt the open reading frame, flank the neomycin selection cassette for optional later removal by Dre recombinase. In the subcloning step to establish the intermediate targeting construct, the p15A origin of replication and antibiotic resistance gene are flanked by unique restriction sites (R.S.) for digestion before targeting in ESCs. (b) Diagram of the Ash2l locus, Southern strategy and Southern analysis of DNA prepared from resistant Ash2l-AID-T2A-neo ESC colonies, digested with KpnI (K) and hybridized with the 3′ probe. Twenty eight different ESC candidates plus a wt control are shown in the two Southern panels.
For simultaneous biallelic targeting, a different outcome was found. Whereas both Ash2l and Wdr82 were biallelically targeted using long-arm targeting constructs, thereby demonstrating that C-terminal tagging of both proteins with AID was not lethal in ESCs, neither gene was biallelically targeted with the short-arm constructs (Table 1). Notably, the frequencies of monoallelic targeting with short-arm constructs, although high, were also lower than the very high frequencies achieved with long-arm constructs. Although the convenience of short-arm targeting constructs, both from their ease of construction and ease of PCR genotyping, can be productively applied for monoallelic targeting, we conclude that longer homology arms deliver an important increase in efficiency for applications that require biallelic targeting. To explore this issue further, we generated a targeting construct with intermediate length homology arms (1419 and 1509 bps) by digestion of the Ash2l long-arm targeting construct with AseI and BglI. Again monoallelic targeting frequencies were high (Table 1; 69%) and notably a few biallelic events were found (3/70; 4%). These intermediate length homology arms therefore delivered an outcome intermediate between the short- and long-arm targeting constructs.
Functional evaluation of auxin degron
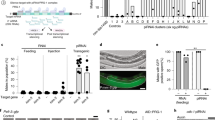
As evaluated using both an Ash2l antibody and the Ty1 monoclonal antibody, administration of auxin (indole-3-acetic acid) in the rosa26 tir1/+, ash2l-AID/ash2l-AID biallelically targeted ESCs resulted in rapid loss of the Ash2l-AID fusion protein, which slowly recovered after auxin removal (Fig. 4) thereby functionally validating the targeting and the auxin degron strategy in ESCs.
Protein degradation promoted by auxin.
Western blot of Ash2l-AID protein (68 kDa + 33 kDa = 101 kDa) using an Ash2l antibody (top panel) and the BB2 monoclonal antibody against Ty1 (middle panel). Cells expressing nlsTir1 and biallelically targeted Ash2l-AID were treated with 500 μM indole-3-acetic acid (IAA) for the indicated times. For protein recovery, cells were washed after 60 minutes of IAA treatment and cultured in ES medium without auxin for 8 and 12 hours. Untargeted E14Tg2a cells expressing Tir1 was the negative control (left hand column) and β-actin as loading control (lower panel). The Ash2l antibody detects a non-specific band marked with an asterisk.
Discussion
We developed RAC-tagging to apply the potential of CRISPR/Cas9 to protein tagging. For monoallelic tagging, RAC-tagging couples the genotyping advantage of junction PCR for short-arm targeting with a very rapid method to build the targeting constructs. For biallelic targeting, our results indicate that long-arm targeting constructs promote the successful outcome. Whether using short- or long-arm constructs, multiple reuse of a gene-specific reagent to introduce a different tag from the R6K series is straightforward because the RAC-tagging system is modular by design. To this end, we are expanding the tagging repertoire by introducing more protein tags into the R6K series.
Recently a generic, homology-independent strategy for monoallelic Cas9-assisted gene tagging was reported, which needs only the gene-specific gRNA and circumvents the need to build gene-specific targeting constructs33. This method is based on ligation of the tagging cassette into the Cas9 genomic cleavage and consequently the intended event is one possibility amongst a variety of different events promoted by the double strand break. The correct clones therefore have to be identified amongst a majority of unwanted events, which include imperfectly ligated junctions. We show here that the gene-specific targeting constructs can be rapidly built and bring the advantage that the majority of clones are correctly targeted.
Amongst the various tags available, we present our experience with the auxin degron because it is relatively unexploited and has great potential for functional studies. As illustrated before22,34 and in Fig. 4, target protein removal can be remarkably rapid. In vertebrate systems, this speed opens new opportunities for real-time functional studies that have been previously inaccessible using currently available loss-of-function strategies such as RNAi knock-down or Cre-mediated site-specific recombination.
Like the widely used gene switch based on ligand-regulated site-specific recombination35, the auxin degron benefits from the advantages of ligand administration to trigger ablation of the target gene product. Both these ligand-inducible strategies require the expression of an effector protein, either rice Tir1 or an SSR-LBD (site specific recombinase/ligand binding domain fusion protein), usually CreERT236. However the auxin degron offers two advantages. First, as opposed to the genetic deletion effected by site-specific recombination, protein ablation by the auxin degron is reversible. Second, protein ablation by auxin is significantly faster than other loss-of-function methods. These advantages suggest that the auxin degron can bring greater accuracy and flexibility to functional studies. However the AID protein tag used here and before22,34 is the full-length Aux/IAA repressor, which can homodimerize37. Reduction of AID to the core 46 amino acids needed for binding to auxin-bound Tir1 without the dimer interface has been reported to work as well as full-length AID in yeast38. However this is not the case in ESCs where the core 46 amino acids does convey auxin-dependent degradation but not as efficiently as full-length AID (Supplementary Figure 6). This indicates the need for sequences N-terminal to the core 46 amino acids, which contribute to the auxin response in plants and include the N-terminal part of the bipartite AID nuclear localization sequence39. Although successful application to cytoplasmic proteins has been described34, some consideration of the potential complications arising from unwanted dimerization and/or nuclear localization appears advisable at this stage.
Methods
DNA constructs
For the 4-way and 5-way recombinations, R6K-Ty1-Tag-neo and p15A-cm-ccdB-amp plasmids were restricted with NdeI and purified using Invisorb Fragment CleanUp Kit (Stratec). The R6K plasmids were grown in EC100D-pir116 (Epicenter). The 5′ and 3′ homology arms were amplified from either genomic or BAC DNA using High-Fidelity polymerase and primer pairs that contained 50 bases of sequence homology to the respective ends of the cassettes (Supplementary Table 1). All four or five DNA fragments (each 500 ng) were co-electroporated into cells induced with L-arabinose to express RecE/RecT/Redγ18 and then plated on LB plates containing chloramphenicol (15 μg/ml) and kanamycin (10 μg/ml). E14Tg2a genomic DNA or bacterial artificial chromosomes (BACs) containing Ash2l (bMQ-333I23) or Wdr82 (CH29-51445) were used to provide the homology arms.
The p15A-cm-ccdB-amp and pBR322-U6-ccdB-cm-tracrRNA plasmids were grown in a ccdB-resistant E. coli strain40. After restriction digestion with BstZ171 and NheI to release the ccdB-cm cassette, pBR322-U6-ccdB-cm-tracrRNA was purified. Linearized plasmid (200 ng) and 50 pmol of single stranded oligonucleotide were co-electroporated into RecET bacteria as described above and plated on LB plates containing ampicillin (100 μg/ml). Only cells that carry the plasmid with oligonucleotide without ccdB expression survive. For the mouse Ash2l and Wdr82 genes, target sites close to endogenous stop codon with the sequence 5′-N20NGG were selected.
For recombineering methods to generate long-arm targeting constructs, please see15,41.
Arabidopsis IAA17 degron (AID) and rice Tir1 were codon-optimized for mammalian expression by Genescript GmbH after manual elimination of CpG dinucleotides. An N-terminal nuclear localization signal (nls) was added to Tir1. Nls-TIR1 and AID were recombineered into a Rosa26-CAG-IRES-puro targeting vector and pR6K-Ty1-T2A-neo respectively. Nls-FLAG-linker-Cas9, optimized for nuclear import of Cas942,43, was obtained from pST1374 plasmid (Addgene; #13426) and cloned into pCAG-IRES-puro. gRNA-AAV1-T2 (Addgene; #41818) was modified to create pBR322-U6-ccdB-cm-tracrRNA by recombineering to insert a ccdB-cm cassette flanked by BstZ171 and NheI restriction sites in place of the 20 bp target sequence. Plasmid sequence files can be obtained from our website http://www.biotec.tu-dresden.de/research/stewart/group-page/plasmids.html.
Cell culture
E14Tg2a embryonic stem cells were maintained in Dulbecco’s modified Eagle’s Medium (DMEM) supplemented with 15% fetal calf serum (FCS), 2 mM L-glutamine, 1 mM sodium pyruvate, 1% penicillin/streptomycin, 100 μM non-essential amino acids (all from Invitrogen), 100 μM β-mercaptoethanol (Sigma) containing leukemia inhibitory factor (LIF) in the presence of 1 μM PD-0325901 and 3 μM CT-99021 (“2i”) at 37 °C with 5% CO2. ESCs (5 × 106) were electroporated with 40 μg linearized targeting vector (Rosa26-CAG-NLS-TIR1-IRES-puro) or co-electroporated with 20 μg linearized targeting vector (Ash2l-, Wdr82-AID), 15 μg gRNA plasmid and 5 μg NLS-FLAG-linker-Cas9 expression vector using a Bio-Rad electroporator (250 V, 500 μF) and selected for 7 days with 1 μg/ml puromycin (Sigma) or with 200 μg/ml G418 (Invitrogen), respectively. Double targeted ES cells were expanded under double selection (puromycin + G418) and induced with 500 μM indole-3-acetic acid (IAA, Sigma).
Western blot
ES cells were sonicated in 20 mM HEPES pH 8.0; 150 mM NaCl; 1.5 mM MgCl2; 10% glycerol; 2 mM EDTA pH 8.0; 0.05% Tween-20) with 1 mM DTT and 1xprotease inhibitor cocktail (Roche) using a BioRaptor waterbath (Diagenode). Whole cell extracts were separated by SDS-PAGE (10% Tris-glycine) transferred to PVDF membranes and probed with primary antibodies: rabbit anti-Ash2l IgG (1:2000; A300-112A, Bethyl), mouse anti-Ty1 IgG (1:5000; home made), mouse anti-β-actin IgG (1:5000; Sigma) overnight at 4 °C, then incubated with goat anti-rabbit (1:30,000)/mouse (1:5000; Sigma) horseradish-peroxidase conjugated secondary antibody for 1 h at RT and visualized with Supersignal West Pico substrate kit by Luminescent Image Analyzer LAS-4000 (Fujifilm Life Science).
Genotyping
The frequencies of long-arm targeting reported in Table 1 were obtained from Southern blots as illustrated in Fig. 3 using standard procedures. The frequencies of the other targetings reported in Table 1 were achieved by PCR using standard procedures. The primers used for PCR genotyping and to generate the Southern probes are listed in Supplementary Table 1.
Additional Information
How to cite this article: Baker, O. et al. RAC-tagging: Recombineering And Cas9-assisted targeting for protein tagging and conditional analyses. Sci. Rep. 6, 25529; doi: 10.1038/srep25529 (2016).
References
Glaser, S., Anastassiadis, K. & Stewart, A. F. Current issues in mouse genome engineering. Nat. Genet. 37, 1187–1193 (2005).
Carroll, D. Genome engineering with targetable nucleases. Annu. Rev. Biochem. 83, 409–439 (2014).
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proceedings of the National Academy of Sciences 109, E2579–86 (2012).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Doudna, J. A. & Charpentier, E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Muyrers, J. P., Zhang, Y., Testa, G. & Stewart, A. F. Rapid modification of bacterial artificial chromosomes by ET-recombination. Nucleic Acids Res 27, 1555–1557 (1999).
Zhang, Y., Buchholz, F., Muyrers, J. P. & Stewart, A. F. A new logic for DNA engineering using recombination in Escherichia coli. Nat. Genet. 20, 123–128 (1998).
Angrand, P. O., Daigle, N., van der Hoeven, F., Schöler, H. R. & Stewart, A. F. Simplified generation of targeting constructs using ET recombination. Nucleic Acids Res 27, e16 (1999).
Skarnes, W. C. et al. A conditional knockout resource for the genome-wide study of mouse gene function. Nature 474, 337–342 (2011).
Chalfie, M., Tu, Y., Euskirchen, G., Ward, W. W. & Prasher, D. C. Green fluorescent protein as a marker for gene expression. Science 263, 802–805 (1994).
Inouye, S. & Tsuji, F. I. Aequorea green fluorescent protein. Expression of the gene and fluorescence characteristics of the recombinant protein. FEBS Letters 341, 277–280 (1994).
Poser, I. et al. BAC TransgeneOmics: a high-throughput method for exploration of protein function in mammals. Nat. Methods 5, 409–415 (2008).
Hofemeister, H. et al. Recombineering, transfection, Western, IP and ChIP methods for protein tagging via gene targeting or BAC transgenesis. Methods 53, 437–452 (2011).
Sarov, M. et al. A Genome-Scale Resource for In Vivo Tag-Based Protein Function Exploration in C. elegans. Cell 150, 855–866 (2012).
Zhang, Y., Vanoli, F., LaRocque, J. R., Krawczyk, P. M. & Jasin, M. Biallelic targeting of expressed genes in mouse embryonic stem cells using the Cas9 system. Methods 69, 171–178 (2014).
Fu, J. et al. Full-length RecE enhances linear-linear homologous recombination and facilitates direct cloning for bioprospecting. Nat. Biotechnol. 30, 440–446 (2012).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Couturier, M., Bahassi, el-M. & Van Melderen, L. Bacterial death by DNA gyrase poisoning. Trends Microbiol. 6, 269–275 (1998).
Sarov, M. et al. A recombineering pipeline for functional genomics applied to Caenorhabditis elegans. Nat. Methods 3, 839–844 (2006).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 20, 87–90 (2002).
Roux, K. J., Kim, D. I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Rhee, H. W. et al. Proteomic Mapping of Mitochondria in Living Cells via Spatially Restricted Enzymatic Tagging. Science 339, 1328–1331 (2013).
Lee, J. et al. An enhanced ascorbate peroxidase 2/antibody-binding domain fusion protein (APEX2-ABD) as a recombinant target-specific signal amplifier. Chem. Commun. (Camb.) 51, 10945–10948 (2015).
Brookman, J. L. et al. An immunological analysis of Ty1 virus-like particle structure. Virology 207, 59–67 (1995).
Szymczak, A. L. et al. Correction of multi-gene deficiency in vivo using a single ‘self-cleaving’ 2A peptide–based retroviral vector. Nat. Biotechnol. 22, 589–594 (2004).
Anastassiadis, K. et al. Dre recombinase, like Cre, is a highly efficient site-specific recombinase in E. coli, mammalian cells and mice. Dis Model Mech 2, 508–515 (2009).
Penfold, R. J. & Pemberton, J. M. An improved suicide vector for construction of chromosomal insertion mutations in bacteria. Gene 118, 145–146 (1992).
Wang, J. et al. An improved recombineering approach by adding RecA to lambda Red recombination. Mol. Biotechnol. 32, 43–53 (2006).
Zhang, Y., Muyrers, J. P., Testa, G. & Stewart, A. F. DNA cloning by homologous recombination in Escherichia coli. Nat. Biotechnol. 18, 1314–1317 (2000).
Lackner, D. H. et al. A generic strategy for CRISPR-Cas9-mediated gene tagging. Nat Commun 6, 10237 (2015).
Holland, A. J., Fachinetti, D., Han, J. S. & Cleveland, D. W. Inducible, reversible system for the rapid and complete degradation of proteins in mammalian cells. Proceedings of the National Academy of Sciences 109, E3350–7 (2012).
Logie, C. & Stewart, A. F. Ligand-regulated site-specific recombination. Proc. Natl. Acad. Sci. USA 92, 5940–5944 (1995).
Feil, R., Wagner, J., Metzger, D. & Chambon, P. Regulation of Cre recombinase activity by mutated estrogen receptor ligand-binding domains. Biochemical and Biophysical Research Communications 237, 752–757 (1997).
Korasick, D. A., Jez, J. M. & Strader, L. C. Refining the nuclear auxin response pathway through structural biology. Curr. Opin. Plant Biol. 27, 22–28 (2015).
Morawska, M. & Ulrich, H. D. An expanded tool kit for the auxin-inducible degron system in budding yeast. Yeast 30, 341–351 (2013).
Moss, B. L. et al. Rate Motifs Tune Auxin/Indole-3-Acetic Acid Degradation Dynamics. Plant Physiology 169, 803–813 (2015).
Wang, H. et al. Improved seamless mutagenesis by recombineering using ccdB for counterselection. Nucleic Acids Res 42, e37–e37 (2014).
Fu, J., Teucher, M., Anastassiadis, K., Skarnes, W. & Stewart, A. F. A recombineering pipeline to make conditional targeting constructs. Meth. Enzymol. 477, 125–144 (2010).
Shen, B. et al. Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Research 23, 720–723 (2013).
Testa, G. et al. A reliable lacZ expression reporter cassette for multipurpose, knockout-first alleles. Genesis 38, 151–158 (2004).
Acknowledgements
We thank Madeleine Walker and Jing Tang for technical assistance. Thanks to I.P. Weber for critical reading of the manuscript. This work was supported by funding to KA from the EU 7th Framework project, PluriMes and to AFS from the EU 7th Framework projects, EUCOMMTools and SyBoSS.
Author information
Authors and Affiliations
Contributions
O.B., Y.Z., K.A., J.F. and A.F.S. planned the experiments. O.B. performed the experiments and generated the reagents except for the 4- and 5-way recombination reactions, which was done by J.F. A.G. helped with the generation of AID constructs. M.O. performed Southern and northern blot analyses. O.B. and A.F.S. wrote the manuscript.
Ethics declarations
Competing interests
Y.Z. and A.F.S. are shareholders in Gene Bridges GmbH, which holds exclusive rights to the commercial applications of Red and RecET recombineering.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Baker, O., Gupta, A., Obst, M. et al. RAC-tagging: Recombineering And Cas9-assisted targeting for protein tagging and conditional analyses. Sci Rep 6, 25529 (2016). https://doi.org/10.1038/srep25529
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep25529
This article is cited by
-
Conditional control of fluorescent protein degradation by an auxin-dependent nanobody
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.