Abstract
The FET protein family includes FUS, EWS and TAF15 proteins, all of which have been linked to amyotrophic lateral sclerosis, a fatal neurodegenerative disease affecting motor neurons. Here, we show that a reduction of FET proteins in the nematode Caenorhabditis elegans causes synaptic dysfunction accompanied by impaired motor phenotypes. FET proteins are also involved in the regulation of lifespan and stress resistance, acting partially through the insulin/IGF-signalling pathway. We propose that FET proteins are involved in the maintenance of lifespan, cellular stress resistance and neuronal integrity.
Similar content being viewed by others
Introduction
Mutations in fused-in-sarcoma (FUS) are one of the causes of amyotrophic lateral sclerosis (ALS)1,2,3, a fatal neurodegenerative disease causing loss of motor neurons. Mutations are found in different domains of the protein and cause its cellular mislocalization3. The presence of FUS in the cytoplasm suggests a gain of toxic function mechanism, but the depletion of FUS from the nucleus also points to a loss of normal function being implicated in motor neuron degeneration. Due to its nature as an RNA/DNA binding protein, FUS has been shown to participate in many cellular functions including translation, splicing and RNA transport4. FUS is part of the FET protein family that includes two other RNA binding proteins; Ewing sarcoma breakpoint region 1 (EWSR1 gene encoding the EWS protein) and TBP associated factor 15 (TAF15). These proteins are highly similar and it is thought that they share common functions5,6. Furthermore, EWSR1 and TAF15 mutations have been linked to some sporadic cases of ALS7,8. However, how mutant FET proteins cause neuronal loss is still unclear.
Many proteins associated with ALS are evolutionarily conserved in the nematode Caenorhabditis elegans. C. elegans are transparent nematodes that have been used to make important contributions to the fields of neuroscience and aging. More recently C. elegans has emerged as a useful model to study human diseases, namely conserved aspects of age-dependent neurodegeneration9.
To better understand the function of FET proteins, we characterised the fust-1(tm4439) deletion mutant in worms. C. elegans has a simple, largely non-redundant genome and many highly conserved human genes have a single orthologue in the nematode. Here, fust-1 is the orthologue of FUS, EWSR1 and TAF15. Using a loss of function mutation, we show that fust-1 is a key gene acting to regulate neuronal integrity, lifespan and cellular stress responses. Also, for some of these functions, fust-1 is an active component of the insulin/IGF-like signalling pathway (ISS).
Results
FUST-1 is required for neuronal integrity
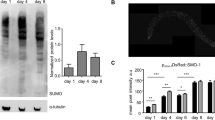
To understand the function and role of the FET proteins, we characterised fust-1, which encodes the C. elegans orthologue of FUS, EWSR1, TAF15. At the protein level, FUST-1 shares 50% identity with the FUS human protein, 32% identity with EWS and 35% identity with TAF15 (Supplementary Material, Fig. S1A). Bioinformatic analyses using NetNes (http://www.cbs.dtu.dk/services/NetNES/) and Prosite (http://prosite.expasy.org/scanprosite/) confirmed the conservation of the main functional domains of FET proteins including the RNA binding domain, the zinc finger motif and the nuclear export signal (Fig. 1A and Supplementary Material, Figs S1 and S2A,B). To investigate the role of FUST-1, we used a C. elegans deletion mutant strain, fust-1(tm4439) which contains a homozygous 240 base pair deletion in the N terminal part of the gene. The tm4439 allele is an in-frame deletion spanning 15 bp of the second intron and 225 bp of exon 3 that is predicted to code for a protein product missing 75 amino acids compared to wild type FUST-1. While fust-1 mRNA level increases during adulthood in wild type animals, fust-1(tm4439) mutants exhibit a similar expression level throughout adulthood which is at least 50% lower than the expression level of wild-type worms at day 1 (Supplementary Material, Fig. S2C). Thus, fust-1(tm4439) is likely a hypomorphic mutation that results in decreased RNA expression and may produce a defective protein product, suggesting that these worms could be used to model a partial loss of function of FUST-1.
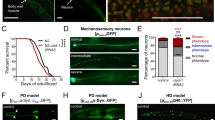
Deletion of fust-1 causes motility impairment and loss of neuronal integrity. (A) C. elegans fust-1 is the ortholog of human FUS, EWSR1 and TAF15 and contains the conserved RNA recognition motif (RRM), nuclear export signal (NES) and zinc finger motif (ZFM). fust-1(tm4439) is a 240 bp deletion. (B–C) Loss of fust-1 expression causes age-dependant paralysis (B) on solid media (p value < 0.0001) and (C) the phenomenon is accelerated in liquid media (p value < 0.0001) (D–F) (D) unc-47 promoter (red) and SNB-1 (green) were used to visualise motor neurons and synapse formation. Shown is the normal expression pattern in wild type animals (i), while decreased expression of fust-1 caused gaps along the motor neurons (ii) and disorganisation of the SNB-1 protein (ii-iii). Quantification of the number of animals at day 1, 5 and 9 with (E) axonal fragmentation in motor neuron (unc-47p::mCherry) (*p value < 0.05, n ≥ 100 for each condition) and (F) synaptic disorganisation (unc-25p::SNB-1::GFP) (*p value < 0.05, n ≥ 100 for each condition). (G) fust-1(tm4439) mutants are more sensitive to aldicarb than N2 and unc-64(e246) but less than the hypersensitive strain unc-47(e307) p < 0.001. (H) Overexpression of fust-1 rescues the age dependant paralysis phenotype observed in fust-1(tm4439) mutants (p value < 0.001).
Previous reports studying the function of the ortholog of FUS in Drosophila, Cabeza, have suggested that a decreased expression of Cabeza in flies induces neuronal dysfunction and defects in neuromuscular junction morphology10,11,12,13. To evaluate if this function was conserved in C. elegans, deletion mutant worms were evaluated for age-dependent paralysis, a motor phenotype that has been shown to be a good predictor of neuronal integrity14,15,16. At day 1 of adulthood, fust-1(tm4439) mutants showed normal motor behaviour when compared to wild-type N2 worms, but as the mutants aged they showed progressive motility defects leading to paralysis, reaching 66% paralysis by day 12 of adulthood compare to the 13% observed for wild-type N2 controls (Fig. 1B and Supplementary Material, Table S1A). We have previously shown that the paralysis phenotype that occurs when worms are grown on solid media after several days can be observed within hours when the worms are grown in liquid culture17,18. The swimming behaviour of C. elegans is an energetically costly activity that actively engages the neuromuscular junction and may be a phenotype relevant to the study of the health of motor neurons. To study the movement of the animals in liquid, worms were placed in a 96-well plate and their movements were evaluated with an automated method that measures locomotion activity based on infrared beam scattering19. With this method, fust-1 mutants initially exhibit normal motility behaviour but show a drastic decrease in movement over time (Fig. 1C and Supplementary Material, Table S1B).
Next we assessed the integrity of GABAergic motor neurons using an unc-47p::mCherry reporter strain20. UNC-47 is a GABA vesicular transporter and is expressed in the 26 GABAergic motor neurons of the worm21. At day 9, fust-1(tm4439) mutants exhibit an increase in the number of gaps or breaks along the ventral cord probably due to axonal fragmentation (Fig. 1D,E) that coincides with the onset of the paralysis phenotype.
To evaluate potential synaptic dysfunction, an unc25p::snb-1::GFP reporter strain was used. SNB-1 is a synaptic vesicle protein and has been used to visualize synapses22. Starting at day 1 of adulthood, fust-1 deletion mutant worms exhibit abnormal organization of SNB-1 protein compared to wild-type worms. SNB-1 abnormal localization affected motor neurons that also showed gaps along their axons (Fig. 1D ii) or could affect neurons prior to breakage of the axons (Fig. 1D iii). The proportion of worms with abnormal SNB-1 localization increased with aging reaching 60% of the worms at day 9 compared to 35% of the wild-type worms (Fig. 1F), however there was no change in the overall intensity of SNB-1 protein between the wild-type and fust-1 mutant worms (Supplementary Material, Fig. S2D).
To evaluate the health of the neuromuscular junction, worms were exposed to aldicarb (2-methyl-2(methylthio) propanal o-[(methylamino)-carbonyl] oxime), an acetylcholine esterase inhibitor that causes the build-up of acetylcholine at the neuromuscular junction leading to paralysis23. Worms with defects in vesicular release at the neuromuscular junction showed hypersensitivity to aldicarb, similar to unc-47 mutants24, or resistance, similar to unc-64/Syntaxin mutants25. fust-1 mutants showed hypersensitivity to aldicarb when compared to wild-type N2, reaching 80% paralysis compared to 40% for the wild-type control after two hours in aldicarb (Fig. 1G). These data suggest an abnormal function of the neuromuscular junction perhaps due to a decrease of GABA release in the fust-1 mutants.
To confirm that the effects observed in our fust-1(tm4439) worms were due to the loss of function of fust-1, we generated a transgenic worm expressing full-length fust-1 linked to GFP under the control of its own promoter (fust-1p::fust-1::GFP). This strain displays GFP expression throughout development and adulthood with expression in the head, pharynx, intestine and tail of the adult worm (Supplementary Material, Fig. S3A). In some cells of the pharynx and the tail of the adult animal, FUST-1 was localized in the nucleus, similar to what is observed in humans (Supplementary Material, Fig. S3B). When crossed to the mutant worms, the fust-1p::fust-1::GFP construct completely rescued the paralysis phenotype of the fust-1(tm4439) worms (Fig. 1H and Supplementary Material, Table S1C) suggesting that indeed the motor phenotype is due to a loss of fust-1. Interestingly, the transgenic worms that overexpress fust-1 do not exhibit motor impairment, suggesting that, in C. elegans, the overexpression of fust-1 under the control of its own promoter is not intrinsically toxic.
Overall, these results suggest that synaptic dysfunction precedes neuronal loss and that aging could promote the development of the motor phenotype observed in the fust-1(tm4439) deletion mutant worms.
FUST-1 is involved in lifespan regulation
Genetic signalling pathways regulating aging have been extensively studied in C. elegans and central to lifespan and stress response mechanisms is the insulin /IGF-like signalling pathway (IIS)26. DAF-2 is the sole insulin/IGF receptor in C. elegans and hypomorphic daf-2 mutants are long-lived and highly resistant to environmental stress26. We constructed a daf-2(e1370); fust-1(tm4439) double mutant strain and observed that the loss of fust-1 completely abolished the extended lifespan phenotype of daf-2(e1370) mutants (Fig. 2A and Supplementary Material, Table S2A). These data suggest that that fust-1 functions within the IIS to regulate longevity.
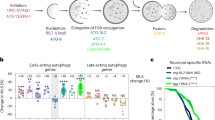
fust-1 functions within the IIS pathway to regulate lifespan.
(A) Decreased expression of fust-1 abolished the long-lived phenotype of daf-2(e1370) (p value < 0.005) but (B) has no effect on daf-16 mutants. Overexpression of fust-1 increased the lifespan (C) of N2 worms (p value < 0.0001) and (D) of daf-2(e1370) mutants (p value < 0.005) but (E) has no effect on daf-16 loss of function (p value 0.4030).
In C. elegans, a crucial downstream effector of the IIS is the forkhead box O (FOXO) transcription factor encoded by daf-16. The long-lived phenotypes of daf-2 mutants is completely dependent on daf-1626. daf-16(mu86) mutants are short-lived but fust-1(tm4439); daf-16(mu86) double mutants have lifespan similar to daf-16(mu86) mutants alone (Fig. 2B and Supplementary Material, Table S2B).
We observed that although fust-1(tm4439) mutants have a normal lifespan at 20 °C and 25 °C (Supplementary Material, Fig. S4A,B and Table S2C,D), the overexpression of fust-1 caused an increased lifespan compared to wild-type worms (Fig. 2C, Supplementary Material Table S2E). Additionally, the overexpression of fust-1 had an additive effect on the lifespan of daf-2 mutants (Fig. 2D, Supplementary Material Table S2F). However, the loss of daf-16 did not affect the long-lived phenotype caused by the overexpression of fust-1 (Fig. 2E). Thus our data suggest that fust-1 is essential regulating lifespan via the IIS and that lifespan-extension is modulated by FUST-1 in a dose-dependent manner.
FUST-1 is involved in resistance to environmental stress
Another important role of the IIS is the regulation of cellular stress responses. Worms were tested against different environmental stresses to evaluate the contribution of fust-1 within the IIS. First, worms were exposed to juglone (5-hydroxy-1,4-naphthoquinone), a natural product that causes the production of intracellular free radical in worms causing an acute oxidative stress27. The fust-1 deletion mutants were more sensitive than wild-type N2 worms (Fig. 3A) and the sensitivity was rescued by the fust-1p::fust-1::GFP transgene in fust-1(tm4439) mutants (Supplementary Material, Fig. S5A). Next we examined daf-2(e1370); fust-1(tm4439) double mutants and observed that these animals were more sensitive to oxidative stress than daf-2(e1370) mutants, but more resistant than fust-1 mutants alone (Fig. 3B). These data suggest that the IIS pathway is only partially reliant on fust-1 in response to oxidative stress and the action of fust-1 in other stress response pathways could explain the intermediate phenotype observed during oxidative stress.
Oxidative and osmotic stress responses require fust-1 (A,B) (A) fust-1 mutants are more sensitive to oxidative stress induced by juglone compared to N2 controls (p value < 0.0005). (B) fust-1; daf-2 mutants are more sensitive to oxidative stress than daf-2 controls (p value < 0.0001. (C,D) (C) fust-1 mutants are sensitive to osmotic stress (p value < 0.001) but the fust-1 mutation does affect the stress sensitivity of (D) daf-2(e1370).
fust-1 deletion mutants were then tested for their resistance to osmotic stress. fust-1 mutants showed sensitivity to a hypertonic environment induced by NaCl (Fig. 3C) and this phenotype was partially rescued by the overexpression of fust-1 (Supplementary Material, Fig S5B). The deletion of fust-1 did not affect the sensitivity of daf-2 mutants to osmotic stress (Fig. 3D) suggesting that fust-1 is completely independent of the IIS for the regulation of osmotic stress.
Finally, worms were exposed to thermal stress. The fust-1 deletion mutants had a normal sensitivity when submitted to 37 °C even after 14 hours (Supplementary Material, Fig. S5C). Also, the deletion of fust-1 in daf-2 mutants did not significantly change their resistance to this stress (Supplementary Material, Fig. S5D) suggesting that fust-1 does not participate in responses to thermal stress.
fust-1 expression is regulated by the IIS
Genes participating in the IIS are known to have their expressions modulated under stress conditions or in IIS mutants28,29. To test if environmental stresses could induce the expression of fust-1, a transcriptional reporter of fust-1 (fust-1p::GFP) (Supplementary Material, Fig. S6) was used to specifically evaluate the gene expression profile of fust-1 and not the protein stability or degradation under these conditions. Worms were submitted to oxidative and osmotic stresses, the two types of stresses where fust-1 seems to be the most involved. Osmotic but not oxidative could induce the expression of fust-1 (Fig. 4A,B). To evaluate if fust-1 expression is regulated by the IIS, expression level was measured in IIS mutant worms. When measured by qRT-PCR the daf-2 mutants exhibit a two-fold higher expression level of fust-1 than wild-type N2 (Fig. 4C). However, daf-16 mutants exhibit a decreased expression level of fust-1 (Fig. 4C) suggesting that fust-1 is overexpressed when the IIS is reduced. Therefore, IIS pathway mutants have an abnormal expression of fust-1 and osmotic stress can induce fust-1 expression independently.
fust-1 expression is induced by osmotic stress and IIS.
(A) Representative, black and white, photo-reversed images of the transgenic fust-1p::GFP reporter strains (control i) showing increased expression in response to osmotic (ii), but not oxidative stress (iii). (B) Relative quantification level of fust-1p::GFP under stress conditions (*p value < 0.0001, n ≥ 30 for each condition). (C) qRT-PCR with ΔΔCT analysis of fust-1 expression showing an increased expression in daf-2(e1370) mutants and decreased expression in daf-16(mu86) mutants.
fust-1 involvement in neuronal integrity is independent of the IIS
The IIS is involved in maintaining neuronal integrity30 and some reports have previously suggested a link between the IIS and neurodegeneration in C. elegans31,32. Therefore the impact of the daf-16 mutants was tested on fust-1 motor phenotypes. The lack of daf-16 neither increased nor decreased the paralysis of the fust-1(tm4439) mutants (Fig. 5A). In previous studies, it was shown that stress sensitivity could cause neurodegeneration14. In order to test this hypothesis, worms were tested against oxidative, osmotic or thermal stresses and neurodegeneration of the GABAergic neurons was evaluated. Oxidative and osmotic stresses cause neurodegeneration in the fust-1 deletion mutants but thermal stress had no significant effect (Fig. 5B), suggesting a link between the role of fust-1 in neuronal integrity and the survival to these stresses. However, the lack of daf-16 did not change the percentage of animals with neurodegeneration (Fig. 5B). Therefore, fust-1 seems to be involved in neuronal integrity independently of the IIS.
Maintenance of neuronal integrity by fust-1 is not regulated by the IIS.
(A) daf-16(mu86); fust-1(tm443) mutants had rates of paralysis similar to fust-1(tm443) mutants alone. (B) Acute osmotic and oxidative stresses induce neurodegeneration in day 1 fust-1 mutants but not thermal stress. daf-16 mutants do not influence the % of animals with neurodegeneration (*p value < 0.05, n ≥ 60 for each conditions).
Discussion
FUS, EWS and TAF15 are RNA binding proteins that form the FET protein family. FET proteins were first identified as being involved in tumorigenesis, resulting from a DNA translocation6. Mutations in FET proteins have recently been linked to ALS3,7,8. In ALS, most mutations affecting these genes are missense mutations and cause their mislocalization from the nucleus to the cytoplasm1,2,7,8 The effect of this mislocalization and how the mutant proteins cause toxicity to motor neurons is unknown. Because of their high sequence and domain similarity, all three proteins have the same orthologue in the nematode C. elegans, encoded by the gene fust-1. Using a deletion mutant strain, we have shown here that a decrease in fust-1 expression can cause impaired stress resistance, lifespan regulation and neuronal integrity.
Using C. elegans, other ALS proteins have been reported to be involved in stress and lifespan regulation. tdp-1, the orthologue of TDP-43 and alfa-1, the orthologue of C9ORF72, have been shown to impair stress response and TDP-1 was shown to be a key regulator of longevity17,33,34,35. In humans, TDP-43 localization was shown to change upon stress where wild-type TDP-43 relocalizes in stress granules36,37. In humans, FUS was also shown to participate in stress granule formation and exhibit a change in localization upon stress induction38,39. More specifically, FUS localization changes after osmotic stress and a reduction of expression of FUS in human cell lines induced a loss of cell viability in a hyperosmolar environment40. EWS and TAF-15 were also shown to translocate to stress granules in human cells and the RNA binding domains of these proteins seem essential for this phenomenon41,42. Together, these results suggest the FET proteins are involved in stress response in C. elegans and humans.
The Drosophila gene Cabeza is the orthologue of FUS, TAF15 and EWSR1 (from http://flybase.org/reports/FBgn0011571.html). Interestingly in Drosophila, Cabeza was also shown to be involved in lifespan, neuronal integrity and synaptic function10,11,13,43. Synaptic dysfunction was also shown to precede neuronal loss recapitulating features observed in C. elegans. In a recent FUS knock-out model, mice with a complete loss of expression of FUS exhibit changes in behaviour but no motor neuron loss44. However, whole transcriptome analysis of spinal cord tissue of these mice has shown an increase in EWSR1 and TAF15 expression44. It is known that FUS can regulate itself though alternative splicing45 and it seems that it can also regulate expression of proteins with similar function such as TAF-15 and EWS-1. Therefore, using simple model organisms where compensatory mechanisms are less frequent we can reveal functions of protein families more easily and suggest here that the FET proteins act together to maintain neuron integrity and lifespan.
Using the fust-1 deletion mutant, we have also shown that the FET protein family is involved in the IIS to maintain longevity and stress resistance. When the IIS pathway is activated, the effect of DAF-2 leads to the phosphorylation of DAF-16 that inhibits its function as a transcription factor26. Our lifespan and expression analysis suggested that fust-1 is downstream of daf-16 and that a reduction in the pathway causes an overexpression of fust-1. In human, IIS exerts its effect through many effectors including its many receptors, the kinases PI3K and AKT as well as FOXO transcription factors. TAF-15, EWS and FUS all contain a RNA binding domain suggestive of a role in RNA expression and metabolism. In a previous study, using UV cross-linking immunoprecipitation followed by whole transcriptome sequencing (CLIP-seq), many RNA targets of the FET protein family have been revealed. Interestingly, members of the AKT and FOXO protein family were identified as targets of wild type TAF-15, EWS and FUS proteins in human cells5. Using different C. elegans mutants, we have shown genetic interactions between the FET proteins and members of the IIS and results from humans supports our hypothesis and suggest direct interactions with members of the IIS.
In conclusion, using C. elegans deletion mutants, several functions of the FET proteins have been revealed. FUST-1 is actively participating and regulated by the IIS for the maintenance of lifespan and for proper resistance to oxidative stress. Independently of the IIS, FUST-1 is involved in neuronal integrity and the osmotic stress response suggesting its participation in other stress response pathways. It is still unclear if mutations of the FET protein family causative of ALS results in a loss of function or a gain of function. However, mutations in essential tremor, another neurodegenerative disorder, were found in FUS and are suggestive of a loss of function mechanism46. Therefore, therapies targeting one of those proteins will have to be highly specific and take into account the impact on the other members of the FET family.
Materials and Methods
Strains and maintenance
Standard methods for culturing and handling the worms were used47. Worms were cultured on standard NGM media streak with OP-50 Escherichia coli strain at 20 °C if not specified otherwise. For a list of strains used see Supplementary Table 4. fust-1p::fust-1::GFP fosmid was obtained by the TransgenOme project48 and confirmed by sequencing and a transgenic strain was created by microinjection in unc-119(ed3) worms.
Bioinformatic analyses
For amino acid alignment, FUST-1 protein sequence was used as query sequence (Wormbase) and compared to FUS (CCDS58454.1), EWS isoform (CCDS54513.1) and TAF15 (CCDS59279.1) as subject sequence and align using BlastP (http://blast.ncbi.nlm.nih.gov/blast/Blast.cgi?PROGRAM=blastp&PAGE_TYPE=BlastSearch&LINK_LOC=blasthome). For prediction of the functional domain of FUST-1 NetNes (http://www.cbs.dtu.dk/services/NetNES/) and Prosite (http://prosite.expasy.org/scanprosite/) were used with FUST-1 coding sequence (Wormbase). Protein alignments were done using Clustal Omega (http://www.ebi.ac.uk/Tools/msa/clustalo/) using the coding sequence mentioned above and followed by processing with BoxShade (http://www.ch.embnet.org/software/BOX_form.html).
Paralysis assay
Worms were transferred to 5 μM FUDR plates one day after L4. Worms were scored daily for movement for 12 days. Worms were counted as paralysed if they failed to move after they were prodded on the nose. Experiments were performed at 20 °C and at least 60 worms were counted per condition. Survival curves and statistics were produced using Log-rank (Mantel-Cox) test. Standard error is shown on graphs.
Liquid culture assay
A synchronised population was obtained using hypochlorite extraction. Worms were grown on solid media up to day 1 of adulthood. At day 1, 30 worms per well were placed in S basal with OP-50 E. coli (optical density 0.5) in a flat-bottom 96-well plate. Measurement was done using Microtracker (Phylumtech), at least 3 wells were done per condition. Standard error is shown on the graph.
Neuronal integrity
To score gaps along the GABAergic neurons, day one, five and nine worms expressing the transgenic unc-47p::mCherry marker were selected for visualisation. To evaluate synaptic integrity, transgenic worms expressing snb-1p::GFP were selected. For neurodegeneration counts during stress tests, adult day one worms were transferred to NGM + 400 mM NaCl at 20 °C (osmotic stress) or normal NGM and put at 37 °C (thermal stress) for six hours or for oxidative test worms were transferred on 240 uM juglone for 30 minutes at 20 °C. For visualization, animals were immobilized in M9 with 5 mM of levamisole and mounted on slides with 2% agarose pads. For all experiments, a minimum of 100 worms was scored for all conditions. The mean and SEM were calculated and two-tailed t-tests were used for statistical analysis.
Lifespan assays
Worms were grown on NGM and transferred on NMG + 5 μM FUDR at day 1 of adulthood. For daf-16 RNAi lifespan RNAi clone from the ORFeome RNAi library (Open Biosystems) was used. RNAi experiments were performed at 20°C. Worms were grown on NGM enriched with 1 mM Isopropyl-b-thiogalacto-pyranoside (IPTG). Worms were counted every two days until their death. At least 100 worms were counted per strain. Survival curves and statistics were produced using Log-rank (Mantel-Cox) test. Standard error is shown on graphs.
Stress sensitivity assay
Worms were grown on NGM until day 1 of adulthood. At day 1, worms were transferred onto 400 mM NaCl plates for osmotic stress, or 240 μM juglone for oxidative stress or onto NGM and put at 37 °C for thermal stress. Worms were counted every two hours for up to 14 hours for osmotic and thermal stress and every 30 minutes for three hours for oxidative stress. Survival curves and statistics were produced using Log-rank (Mantel-Cox) test. Standard error is shown on graphs.
RT-PCR
Worms grown on NGM plates were collected before starvation and froze in Trizol. Total RNA was extracted according to the manufacturer protocol. 1 ug of RNA was used to produce cDNA using Vilo cDNA enzyme. Taqman probes detecting fust-1 (probe Ce02434658_g1 Life technologies) and act-5 (probe Ce02454560_g1 Life technologies) as endogenous control were used. Expression level were calculated by converting the threshold cycle (Ct) values using the 2−ΔΔCt method49
Additional Information
How to cite this article: Therrien, M. et al. FET proteins regulate lifespan and neuronal integrity. Sci. Rep. 6, 25159; doi: 10.1038/srep25159 (2016).
References
Vance, C. et al. Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323, 1208–1211 (2009).
Kwiatkowski, T. J. et al. Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323, 1205–1208 (2009).
Deng, H., Gao, K. & Jankovic, J. The role of FUS gene variants in neurodegenerative diseases. Nat Rev Neurol. doi: 10.1038/nrneurol.2014.78(2014).
Lagier-Tourenne, C., Polymenidou, M. & Cleveland, D. W. TDP-43 and FUS/TLS: emerging roles in RNA processing and neurodegeneration. Hum. Mol. Genet. 19, R46–R64 (2010).
Hoell, J. I. et al. RNA targets of wild-type and mutant FET family proteins. Nature Structural & Molecular Biology 18, 1428–1431 (2011).
Kovar, H. Dr. Jekyll and Mr. Hyde: The Two Faces of the FUS/EWS/TAF15 Protein Family. Sarcoma 2011, 837474–13 (2011).
Couthouis, J. et al. Evaluating the role of the FUS/TLS-related gene EWSR1 in amyotrophic lateral sclerosis. Hum. Mol. Genet. 21, 2899–2911 (2012).
Couthouis, J. et al. A yeast functional screen predicts new candidate ALS disease genes. Proceedings of the National Academy of Sciences 108, 20881–20890 (2011).
Therrien, M. & Parker, J. A. Worming forward: amyotrophic lateral sclerosis toxicity mechanisms and genetic interactions in Caenorhabditis elegans. Front Genet 5, 1–13 (2014).
Xia, R. et al. Motor neuron apoptosis and neuromuscular junction perturbation are prominent features in a Drosophila model of Fus-mediated ALS. Molecular Neurodegeneration 7, 10 (2012).
Sasayama, H. et al. Knockdown of the Drosophila fused in sarcoma (FUS) homologue causes deficient locomotive behavior and shortening of motoneuron terminal branches. PLoS ONE 7, e39483 (2012).
Shahidullah, M. et al. Defects in Synapse Structure and Function Precede Motor Neuron Degeneration in Drosophila Models of FUS-Related ALS. Journal of Neuroscience 33, 19590–19598 (2013).
Frickenhaus, M., Wagner, M., Mallik, M., Catinozzi, M. & Storkebaum, E. Highly efficient cell-type-specific gene inactivation reveals a key function for the Drosophila FUS homolog cabeza in neurons. Sci. Rep. 5, 9107 (2015).
Therrien, M., Rouleau, G. A., Dion, P. A. & Parker, J. A. Deletion of C9ORF72 results in motor neuron degeneration and stress sensitivity in C. elegans. PLoS ONE 8, e83450 (2013).
Vaccaro, A. et al. Mutant TDP-43 and FUS cause age-dependent paralysis and neurodegeneration in C. elegans. PLos ONE 7, e31321 (2012).
Maruyama, I. N. & Brenner, S. A phorbol ester/diacylglycerol-binding protein encoded by the unc-13 gene of Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 88, 5729–5733 (1991).
Therrien, M., Rouleau, G. A., Dion, P. A. & Parker, J. A. Deletion of C9ORF72 Results in Motor Neuron Degeneration and Stress Sensitivity in C. elegans. PLoS ONE 8, e83450 (2013).
Vaccaro, A. et al. Methylene blue protects against TDP-43 and FUS neuronal toxicity in C. elegans and D. rerio. PLoS ONE 7, e42117 (2012).
Simonetta, S. H. & Golombek, D. A. An automated tracking system for Caenorhabditis elegans locomotor behavior and circadian studies application. J. Neurosci. Methods 161, 273–280 (2007).
Barbagallo, B., Prescott, H. A., Boyle, P., Climer, J. & Francis, M. M. A dominant mutation in a neuronal acetylcholine receptor subunit leads to motor neuron degeneration in Caenorhabditis elegans. J. Neurosci. 30, 13932–13942 (2010).
Jorgensen, E. M. GABA. WormBook 1–13. doi: 10.1895/wormbook.1.14.1 (2005).
Jin, Y. Synaptogenesis. WormBook, doi: 10.1895/wormbook.1.44.1 (2005).
Mahoney, T. R., Luo, S. & Nonet, M. L. Analysis of synaptic transmission in Caenorhabditis elegans using an aldicarb-sensitivity assay. Nat Protoc 1, 1772–1777 (2006).
Vashlishan, A. B. et al. An RNAi screen identifies genes that regulate GABA synapses. Neuron 58, 346–361 (2008).
Saifee, O., Wei, L. & Nonet, M. L. The Caenorhabditis elegans unc-64 locus encodes a syntaxin that interacts genetically with synaptobrevin. Mol. Biol. Cell 9, 1235–1252 (1998).
Murphy, C. T. & Hu, P. J. Insulin/insulin-like growth factor signaling in C. elegans. WormBook 1–43, doi: 10.1895/wormbook.1.164.1 (2013).
de Castro, E., Hegi de Castro, S. & Johnson, T. E. Isolation of long-lived mutants in Caenorhabditis elegans using selection for resistance to juglone. Free Radical Biology and Medicine 37, 139–145 (2004).
Murphy, C. T. et al. Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans. Nature 424, 277–283 (2003).
Oliveira, R. P. et al. Condition-adapted stress and longevity gene regulation by Caenorhabditis elegans SKN-1/Nrf. Aging Cell 8, 524–541 (2009).
Broughton, S. & Partridge, L. Insulin/IGF-like signalling, the central nervous system and aging. Biochem. J. 418, 1–12 (2009).
Boccitto, M., Lamitina, T. & Kalb, R. G. Daf-2 signaling modifies mutant SOD1 toxicity in C. elegans. PLoS ONE 7, e33494 (2012).
Teixeira-Castro, A. et al. Neuron-specific proteotoxicity of mutant ataxin-3 in C. elegans: rescue by the DAF-16 and HSF-1 pathways. Hum. Mol. Genet. 20, 2996–3009 (2011).
Vaccaro, A. et al. TDP-1/TDP-43 regulates stress signaling and age-dependent proteotoxicity in Caenorhabditis elegans. PLoS Genet 8, e1002806 (2012).
Zhang, T., Baldie, G., Periz, G. & Wang, J. RNA-Processing Protein TDP-43 Regulates FOXO-Dependent Protein Quality Control in Stress Response. PLoS Genet 10, e1004693 (2014).
Zhang, T., Hwang, H. Y., Hao, H., Talbot, C. & Wang, J. Caenorhabditis elegans RNA-processing Protein TDP-1 Regulates Protein Homeostasis and Life Span. Journal of Biological Chemistry 287, 8371–8382 (2012).
Liu-Yesucevitz, L. et al. Tar DNA binding protein-43 (TDP-43) associates with stress granules: analysis of cultured cells and pathological brain tissue. PLoS ONE 5, e13250 (2010).
Colombrita, C. et al. TDP-43 is recruited to stress granules in conditions of oxidative insult. Journal of Neurochemistry 111, 1051–1061 (2009).
Gal, J. et al. Nuclear localization sequence of FUS and induction of stress granules by ALS mutants. Neurobiology of Aging 32, 2323.e27–40 (2011).
Bosco, D. A. et al. Mutant FUS proteins that cause amyotrophic lateral sclerosis incorporate into stress granules. Hum. Mol. Genet. 19, 4160–4175 (2010).
Sama, R. R. K. et al. FUS/TLS assembles into stress granules and is a prosurvival factor during hyperosmolar stress. J. Cell. Physiol. 228, 2222–2231 (2013).
Andersson, M. K. et al. The multifunctional FUS, EWS and TAF15 proto-oncoproteins show cell type-specific expression patterns and involvement in cell spreading and stress response. BMC Cell Biol. 9, 37 (2008).
Blechingberg, J., Luo, Y., Bolund, L., Damgaard, C. K. & Nielsen, A. L. Gene expression responses to FUS, EWS and TAF15 reduction and stress granule sequestration analyses identifies FET-protein non-redundant functions. PLos ONE 7, e46251 (2012).
Wang, J.-W., Brent, J. R., Tomlinson, A., Shneider, N. A. & McCabe, B. D. The ALS-associated proteins FUS and TDP-43 function together to affect Drosophila locomotion and life span. J. Clin. Invest. 121, 4118–4126 (2011).
Kino, Y. et al. FUS/TLS deficiency causes behavioral and pathological abnormalities distinct from amyotrophic lateral sclerosis. Acta Neuropathol Commun 3, 24 (2015).
Zhou, Y., Liu, S., Liu, G., Oztürk, A. & Hicks, G. G. ALS-associated FUS mutations result in compromised FUS alternative splicing and autoregulation. PLoS Genet 9, e1003895 (2013).
Merner, N. D. et al. Exome sequencing identifies FUS mutations as a cause of essential tremor. Am. J. Hum. Genet. 91, 313–319 (2012).
Stiernagle, T. Maintenance of C. elegans. WormBook, doi: 10.1895/wormbook.1.101.1 (2006).
Sarov, M. et al. A recombineering pipeline for functional genomics applied to Caenorhabditis elegans. Nat Meth 3, 839–844 (2006).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Acknowledgements
Some worm strains were provided by the CGC, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440), the International C. elegans Gene knock-out consortium and the TransgenOme project. We thank Sarah Peyrard for technical support. M.T. is supported by Reseau de médecine généetique appliquée and by the Fonds de recherche du Québec- Santé, J.A.P. holds a career development award from the Fonds de recherche du Québec - Santé. This research was funded by the Canadian Institute of Health Research-CIHR and the ALS Society of Canada.
Author information
Authors and Affiliations
Contributions
M.T. and J.A.P. conceived the study, M.T. performed the experiments, M.T. and J.A.P. wrote the manuscript and G.A.R. and P.A.D. reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Therrien, M., Rouleau, G., Dion, P. et al. FET proteins regulate lifespan and neuronal integrity. Sci Rep 6, 25159 (2016). https://doi.org/10.1038/srep25159
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep25159
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.