Abstract
Transforming growth factor-β1 (TGF-β1) plays an important role on fibrogenesis in heart disease. MicroRNAs have exhibited as crucial regulators of cardiac homeostasis and remodeling in various heart diseases. MiR-19a-3p/19b-3p expresses with low levels in the plasma of heart failure patients. The purpose of our study is to determine the role of MiR-19a-3p/19b-3p in regulating autophagy-mediated fibrosis of human cardiac fibroblasts. We elucidate our hypothesis in clinical samples and human cardiac fibroblasts (HCF) to provide valuable basic information. TGF-β1 promotes collagen I α2 and fibronectin synthesis in HCF and that is paralleled by autophagic activation in these cells. Pharmacological inhibition of autophagy by 3-methyladenine decreases the fibrotic response, while autophagy induction of rapamycin increases the response. BECN1 knockdown and Atg5 over-expression either inhibits or enhances the fibrotic effect of TGF-β1 in experimental HCF. Furthermore, miR-19a-3p/19b-3p mimics inhibit epithelial mesenchymal transition (EMT) and extracellular matrix (ECM) prodution and invasion of HCF. Functional studies suggest that miR-19a-3p/19b-3p inhibits autophagy of HCF through targeting TGF-β R II mRNA. Moreover, enhancement of autophagy rescues inhibition effect of miR-19a-3p/19b-3p on Smad 2 and Akt phosphorylation through TGF-β R II signaling. Our study uncovers a novel mechanism that miR-19a-3p/19b-3p inhibits autophagy-mediated fibrogenesis by targeting TGF-β R II.
Similar content being viewed by others
Introduction
Abnormal expression of cardiomyocyte gene can result in cardiomyocyte hypertrophy and impaired cardiomyocyte viability and contraction, ultimately resulting in heart failure (HF)1,2. The heart function decreases and affects the lungs, liver, and other body systems. HF is considered as the most common ultimate of many cardiovascular disease including dilated cardiomyopathy (DCM)3, myocardial infarction (MI)4,5, diabetic cardiomyopathy6,7, aortic stenosis (AS) and hypertension8,9. Interstitial fibrosis of myocardial cells may initiate with the dysfunctional cardiac remodeling following cardiac injury. Fibrosis is a complex process resulting from activation of some signaling pathways, such as Transforming growth factor (TGF)-β1 signaling10. Indeed, the dynamic mobilization within cardiac extracellular matrix (ECM) is critical during the pathogenesis of ventricular remodeling following DCM, MI, hypertension, and other cardiovascular conditions11. TGF-β1 signaling has broad-ranging effects that may affect cell growth, differentiation and the production of ECM proteins12,13,14.
TGF-β1 is also a known factor in angiotensin II (Ang II)-mediated cardiac fibrosis15. The phenomenon that an abundant latent collagenase system is closely associated with interstitial collagen matrix in heart has been identified for the first time by Montfort and Pérez-Tamayo in 197516. During the harmful remodeling process, cardiac fibroblasts are differentiated into myofibroblasts and the ECM components such as collagen I α2 and fibronectin are accumulated17,18. Moreover, differentiation of fibroblasts into myofibroblasts activates matrix metalloproteinase (MMPs) such as MMP-2 and MMP-9 in the border of remodeling area. The MMPs activation accelerates degradation of adjoining ECM and thus facilitates the highly organized matrix to be replaced with the structureless and thickened matrix19,20. The dysregulation between accumulation and degradation of ECM has been involved in the mobility of ventricular geometry and function and then contributes to the development to heart failure (HF)21.
TGF-β Receptor II is formed with trans-membrane serine/threonine kinase and the TGF-β type II serine/threonine kinase receptor22. TGF-β Receptor II can transduce the TGF-β1, TGF-β2 and TGF-β3 signaling from cell membrane to cytoplasm and then regulate a series of physiological or pathological processes including mesenchymal cell proliferation and differentiation23,24, and ECM production25. Researchers have shown an association between a common TGF-β Receptor II polymorphism and risk of sudden cardiac arrest caused by ventricular arrhythmias in the setting of coronary artery disease26. The formation of the receptor complex composed of TGF-β Receptor I and TGF-β Receptor II molecules symmetrically bound to the cytokine dimer results in the phosphorylation and the activation of TGF-β Receptor I by the constitutively active TGF-β Receptor I27.
Autophagy works as a tightly-regulated process for bulk degradation, through which intracellular components are sequestered into autophagosomes and subsequently degraded by lysosomes28,29,30. Autophagy is critical for the clearance of damaged organelles and protein to maintain cellular homeostasis31,32. Autophagy can communicate with apoptosis as one of the programmed cell death through autodigestive cellular progression, cellular infection with pathogens or extracellular stimulation29,30,33,34. The overall regulation of interstitial fibrosis may contain the complex functioning of various regulatory factors35. It has been reported that the cardiomyocyte-specific deletion of basal autophagy resulting from Atg5 deficiency leads to spontaneous cardiac hypertrophy36,37. TGF-β1 can induce autophagy in other cellular systems, however, there is little evidence supporting the link between autophagy and fibrogenesis in cardiovascular disease.
MicroRNAs (miRNAs) are a series of small non-coding RNAs that regulate gene expression by binding to complementary sequences in un-translated regions (UTR) of target messenger RNAs (mRNAs) bearing fully complementary target sites to trigger either translation repression or mRNA degradation38,39. MiRNAs are involved in a wide spectrum of biological processes including autophagy40, fibrosis41, cell proliferation42, and apoptosis43. An abundant number of evidences indicate that miRNA-mediated gene negative regulation plays important roles in the cardiac homeostasis and pathological remodeling44,45,46,47. William T. Pu’s study has focused on the genome-wide miRNA expression profiling in left ventricular myocardium of 67 patients belonging to four diagnostic groups. The groups comprise ischemic cardiomyopathy (ICM), dilated cardiomyopathy (DCM), aortic stenosis (AS), and nonfailing controls48. They have found that miRNA expression profiles were significantly changed in heart diseases and that the pattern of miRNA expression was distinct in various forms of heart diseases, respectively. Among them, the expresson of miR-19a-3p/19b-3p is low in DCM (p ≦ 0.001), ICM (p < 0.001), and AS (p < 0.001)48. In our present study, we have found that expression of miR-19a-3p/19b-3p in HF patients is lower than in normal. In addition, the over-expression of miR-19a-3p/19b-3p in vitro sufficiently decreases the autophagy and fibrosis induced by TGF-β1 signaling, accompanied by inhibition effect of TGF-β R II. Our further investigation revealed the molecular mechanisms of miR-19a-3p/19b-3p in the regulation of autophagy-related fibrogenesis in human cardiac fibroblasts (which are responsible for the deposition of extracellular matrix).
Method and Materials
Cell culture and treatment
Human Cardiac Fibroblasts (HCF) was purchased from Cell Bank of Tongpai Biotechnology Co., Ltd. (Shanghai, China). The base medium for HCF cell line is formulated Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco Inc.). To make the complete growth medium, we add the following components to the base medium: 4.5 g/L glucose and fetal bovine serum to a final concentration of 10%. Exponentially growing cultures were maintained in a humidified atmosphere of 5% carbon dioxide (CO2) at 37 °C.
Cell proliferation assay
Cell growth was analyzed using the Cell Counting Kit-8 (Dojindo Molecular Technologies, Inc., Japan) according to the manufacturer’s instructions. Briefly, human Cardiac Fibroblasts were seeded at a density of 5 × 103cells/well in 96-well plates and treated with TGF-β1 (10 ng/ml) for 0, 24, 48, 72, and 96 hours at 37 °C with 5% CO2. Then, the medium in each well was substituted with 100 μl of fresh medium containing 10% Cell Counting Kit-8, and the cultures were incubated at 37 °C for 2 hours. The absorbance value (A) was determined using Synergy™ 2 Multi-Mode Microplate Reader (BioTek Instruments, Inc., headquartered in Winooski, VT, USA) at 450 nm. Cell Viability (%) = average absorbance of treated group/average absorbance of control group ×100.
MiR-19a-3p/19b-3p prediction targets
TargetScan 7.0 human, miRDB, miRanda, Ingenuity Pathway Analysis were used to identify predicted miR-19a-3p/19b-3p targets. The mRNAs would be considered as targets if selected miRNA was predicted with high probability to interact with their 3′-UTR. The mRNA targets were then compared to genes and pathways associated with human cellular morphology to evaluate potential miRNA regulation of cell motility.
Dual-luciferase activity assay
The 3′UTR of human TGF-β R II cDNA containing the putative target site for the miR-19a-3p/19b-3p (sequence shown in Supplementary data) was chemically synthesized and inserted at the XbaI site, immediately downstream of the luciferase gene in the pGL3-control vector (Promega, Madison, WI) by Integrated Biotech Solutions Co., Ltd (Shanghai, China) (Supplementary data). Twenty-four hours before transfection, cells were plated at 1.5 × 105 cells/well in 24-well plates. 200 ng of pGL3- TGF-β R II-3′-UTR plus 80 ng pRL-TK (Promega) were transfected in combination with 50 nM of the miR-19a-3p/19b-3p mimics or miRNA mimic control using LipofectamineTM 2000 reagent (Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s protocol, respectively. Luciferase activity was measured 24 hours after transfection using the Dual Luciferase Reporter Assay System (Promega). Firefly luciferase activity was normalized to renilla luciferase activity for each transfected well. Three independent experiments were performed in triplicate.
The sequence of 3′UTR of human TGFBR2 cDNA containing the putative target site for the miR-19a-3p/19b-3p; stark black body stands for the putative target site for miR-19a-3p/19b-3p: CTCTTCTGGGGCAGGCTGGGCCATGTCCAAAGAGGCTGCCCCTCTCACCAAAGAACAGAGGCAGCAGGAAGCTGCCCCTGAACTGATGCTTCCTGGAAAACCAAGGGGGTCACTCCCCTCCCTGTAAGCTGTGGGGATAAGCAGAAACAACAGCAGCAGGGAGTGGGTGACATAGAGCATTCTATGCCTTTGACATTGTCATAGGATAAGCTGTGTTAGCACTTCCTCAGGAAATGAGATTGATTTTTACAATAGCCAATAACATTTGCACTTTATTAATGCCTGTATATAAATATGAATAGCTATGTTTTATATATATATATATATATCTATATATGTCTATAGCTCTATATATATAGCCATACCTTGAAAAGAGACAAGGAAAAACATCAAATATTCC
The use of miR19a/19b mimics
The miR-RiboTM miR19a/19b mimics were chemically synthesized mature double stranded miRNA that could be ready to use (Guangzhou RiboBio Co., Ltd.). The product with lyophilized form was transported at normal temperature. The freeze-dried powder was prepared as a 20 μM stock solution in sterilized ddH2O and stored at −40 °C. For miRNA miR19a/19b mimics transfection, the optimal concentration for human cardiac fibroblasts was 50 nM. For 24-well plate system: 1.25 μl storage solution of miRNA mimic (20 μM) was diluted in 50 μl serum-free medium, mixed gently and incubated at room temperature for 5 minutes. For 6-well plate system: 5 μl storage solution of miRNA mimic (20 μM) was diluted in 250 μl serum-free medium, mixed gently and incubated at room temperature for 5 minutes.
Transient transfections
Cells were transiently transfected with miR-19a-3p/19b-3p mimics, BECN1 siRNA, pCMV-myc-Atg5 plasmid using LipofectamineTM 2000 reagent (Life Technologies, Carlsbad, CA, USA), according to the manufacturer’s instructions49. Short interfering RNA against the human beclin 1 (BECN 1) and the sequence was: sense 5′-AAGAUUGAAGACACAGGAGGC-3′ and antisense 5′-GCCUCCUGUGUCUUCAAUCUU-3′. The universal negative control siRNA were used. The expression vector was transfected 24 hours before treatment with TGF-β130.
IF and confocal fluorescence microscopy
Human cardiac fibroblasts were pro-transfected with miR-19a-3p/19b-3p and then treated with 10 ng/ml TGF-β1. MAP-LC3 (green) was labeled with primary anti-MAP-LC3 polyclonal antibody. Goat anti-rabbit IgG/FITC were used as secondary antibody. Cells were simultaneously imaged in the presence of SNLYSO sensor (an autolysosome fluorescent probe, shown red) to visualize autophagy. The nuclei were stained with 4′, 6-diamidino-2-phenylindole (DAPI, Sigma-Aldrich, St. Louis, MO) for 10 minutes before imaging. An FV10-ASW laser scanning confocal microscope [Ver 2.1] (Olympus Corp, MPE FV1000) was used for co-localization analysis30.
Total RNA extraction, Q-PCR for mRNA and miRNA quantification
Total RNA was extracted using Trizol reagent (Invitrogen, USA). The concentration and purity of the RNA samples were determined spectroscopically. Reverse transcription was performed with M-MLV (Promega, USA) following standard protocols. For the TaqMan-based real-time reverse transcription-polymerase chain reaction (RT-PCR) assays, the ABI 7900 HT Sequence Detection system (Applied Biosystem, Foster City, CA) was used. For quantitative PCR of miRNA, the miR-19a-3p/19b-3p primer and EzOmics SYBR qPCR kit were purchased from Biomics. Amplification procedure was 94 °C for 5 min, followed by 30 cycles at 94 °C for 30 s, 61 °C for 45 s, finally 72 °C for 10 min.
Variations in expression of miR-19a-3p/19b-3p between different RNA samples from cells or plasma were calculated after normalization to U6 or ath-miR-156a, respectively50. The methods were carried out in accordance with the approved guidelines and all experimental protocols were approved by the First Affiliated Hospital of Nanjing Medical University. The ethical code of our study is 2013-SRFA-078 obtained from the licensing committee of Nanjing Medical University. For quantification of TGF-β R II transcripts, Q-PCR was carried out with total RNA samples extracted from human cardiac fibroblasts after miR-19a-3p/19b-3p mimics transfection. TGF-β R II (158 bp) mRNA was amplified at an annealing temperature of 60 °C and using the following primers: TGF-β R II (forword): 5′-CAGAAATCCTGCATGAGC-3′ and TGF-β R II (reverse): 5′-GCAGCATCTTCCAGAATAAAG-3′. TaqMan quantitative assay was performed with the expression level of the corresponding housekeeping gene GAPDH as an internal control. The nucleotide sequences of miR-19a-3p/19b-3p are available in the GenBank database with the accession numbers of LM378760.1, LM378761.1.
Cell synchronization and flow cytometry
Autophagy induction of human cardiac fibroblasts was examined using SNLYSO Autophagy Detection Kit (Catalog Number: E0010, SNPT, Chengdu, China), according to the manufacturer’s instructions. In brief, cells were treated for indicated condition and the medium was gently removed. 250 μl SNLYO Sensor was added to cover the sample and incubated for 20 hours. Finally, cells were treated with trypsin and collected to make the density of 105~106/ml and analyzed by BD FACSCalibur™ Flow Cytometry System (Franklin Lakes) and a computer station running Cell Quest software (BD Biosciences, Franklin Lakes, NJ).
Transwell invasion assay
The invasion behavior of human cardiac fibroblasts was determined using 24-well Millicell Hanging Cell Culture inserts with 8 mm PET membranes (Millipore, Bedford, Massachusetts, USA) as described previously45,51,52,53. Briefly, after the cells were treated for indicated condition, 5.0 × 104 human cardiac fibroblasts in 200 μl serum-free DMEM medium were plated onto BD BioCoatTM MatrigelTM Invasion Chambers (8 μM pore size polycarbonate filters; BD Biosciences), while complete medium containing 10% FBS was added to the lower chamber. After processing the invasion chambers for 48 hours (37 °C, 5% CO2) in accordance with the manufacturer’s protocol, the non-invading cells were removed with a cotton swab; the invading cells were fixed in 100% methanol and then stained with crystal violet solution and counted microscopically. The data are presented as the average number of cells attached to the bottom surface from five randomly chosen fields.
Western blotting
Briefly, after washing twice with PBS, total cellular proteins were extracted with lysis buffer (50 mmol/l Tris [pH 7.4], 150 mmol/l NaCl, 1% Triton X-100, 1% deoxycholic phenylmethylsulfonyl fluoride, 1 mg/ml aprotinin, 5.0 mm sodium pyrophosphate, 1.0 g/ml leupeptin, 0.1 mm phenylmethylsulfonyl fluoride, and 1 mm/l DTT). Protease inhibitors were added immediately before use. The lysates were centrifuged at 13,000× g for 15 min at 4 °C. The concentration of total proteins was measured using the BCA assay method with Varioskan spectrofluorometer and spectrophotometer (Thermo, Waltham, MA) at 562 nm. Protein samples were separated with 12% SDS-PAGE gel and electrophoretically transferred onto the polyvinylidene difluoride (PVDF) membranes (Millipore, Boston, MA). Immune complexes were formed by incubation of proteins with primary antibodies overnight at 4 °C. After incubation with the appropriate secondary antibodies, blots were visualized using the ECL plus Western blotting detection reagents (Bio-Rad) and the ChemiDoc XRS Plus luminescent image analyzer (Bio-Rad, Hercules, CA, USA). Densitometric analysis of band intensity was performed using Image lab software (Bio-Rad, Hercules, CA, USA)30.
Immunocytochemistry
IHC staining of heart valve tissue using MAP-LC3, TGF-β R I, TGF-β R II, Fibronectin, MMP-2, MMP-9 antibodies was performed. Briefly, the twenty heart valve tissue samples of 20 cases patients with dilated cardiomyopathy were subjected to deparaffinization, rehydration, and antigen retrieval before the staining procedures were performed. The tissue slides were blocked with 2.5% normal horse serum for 10 minutes. Then, tissue slides were incubated with rabbit anti-human MAP-LC3, TGF-β R I, TGF-β R II, Fibronectin, MMP-2, MMP-9 antibody (dilution 1:50) overnight at 4 °C. After the tissue slides were washed, they were incubated with anti-mouse IgG HRP and anti-rabbit IgG HRP secondary antibody for 10 minutes. The slides were stained with 3, 3′-diaminobenzidine (DAB) (Vector Laboratories), counterstained with hematoxylin (Vector Laboratories), dehydrated, treated with xylene, and mounted. All slides were examined and representative pictures were taken using an Olympus BX41 microscope (Olympus America, Melville, NY).
Statistical analyses
All data were expressed as mean ± SEM and statistically compared by one-way ANOVA with Dunnett’s test and post-hoc tests were undertaken using the GraphPad Prism software. The details of each statistical analysis used were presented in the figure legends. Significance was indicated as *P < 0.05, #P < 0.05 and **P < 0.01, ##P < 0.01, $$P < 0.01.
Results
TGF-β1 simultaneously induces fibrosis and autophagy in human cardiac fibroblasts
Firstly, we investigated whether there was a correlation between TGF-β1-induced autophagy and fibrosis in human cardiac fibroblasts. Human cardiac fibroblasts were treated with TGF-β1 (10 ng/ml) for 0, 24, 48, 72, and 96 hours. Protein samples were harvested for western blotting assay. The autophagy hallmarks LC3-I, LC3-II, p62/SQSTM1, as well as indicator proteins of the fibrogenesis response Collagen I α2, Fibronectin and the Smad phosphorylation were analyzed. TGF-β1 induced LC3-II lipidation, with parallel increases in collagen I α2, fibronectin expression and Smad2 and Smad3 phosphorylation. Our results showed that TGF-β1 (10 ng/ml) significantly increased the synthesis of Collagen type I α2 and Fibronectin. Simultaneously, the conversion of water soluble MAP-LC3 (LC3- I) to the autophagosome-associated lipidated form (LC3-II) and p62 degradation were determined with the increase of Smad 2 and Smad 3 phosphorylation (Fig. 1a,b). The effect of TGF-β1 on cell cytotoxicity was determined by CCK-8 assay. As shown in Fig. 1c, we also showed that TGF-β1 treatment at a concentration of 10 ng/ml exhibited little effect on the viability of human cardiac fibroblasts. So, this dose was taken in the following experiments. To obtain more quantitative assessment of the induction of autophagy, we used flow cytometer analysis to count SNLYSO sensor (autolysosome fluorescent probe) labeled cells and assess the presence of autophagy flux. Figure 1d,e clearly show autophagosome and autophagolysosmes flux in human cardiac fibroblasts stimulated with TGF-β1.
(a) Human cardiac fibroblasts were treated with TGF-β1 (10 ng/ml) for 0, 24, 48, 72, and 96 hours. Protein samples were harvested for western blotting assay. The autophagy hallmarks (LC3-I, LC3-II, p62/SQSTM1), as well as indicator proteins of the fibrogenesis response in fibroblasts (collagen I α2, fibronectin and the Smad signaling pathway) were analyzed by western blotting assay. TGF-β1 induced LC3-II lipidation, with parallel increases in collagen Iα2, fibronectin expression and Smad2 and Smad3 phosphorylation. Data were normalized to GAPDH levels. Results are the means of three independent experiments from four different donors. (b) Representative quantitative data of densitometric analyses. Data are the means of three independent experiments from three different donors. For each experiment, LC3-I, LC3-II, p62/SQSTM1, collagen I α2, and fibronectin levels in HCF treated with TGF-β1 were compared with those from time-matched controls and normalized to GAPDH levels. *p < 0.05, **p < 0.01 vs. 0 hrs group. (c) TGF-β1 treatment does not affect cell viability of human cardiac fibroblasts. HCF cells were exposed to TGF-β1 (10 ng/ml) for the indicated time points (24, 48, 72, 96 hours), and cell viability and proliferation was measured by CCK-8 assay as described in the Materials and Methods section in three different culture experiments (n = 3). TGF-β1 treatment was not associated with any significant changes in cell viability. (d,e) HCF cells were treated with 10 ng/ml TGF-β1 for 0, 24, 48, 72, and 96 h. Cells were then probed by SNLYSO sensor (an autolysosome fluorescent probe) and autophagic cells were analyzed and quantitated by flow cytometer. Results from three independent experiments are shown as means ± SEM. **p < 0.01 vs. 0 hrs group.
TGF-β1-induced autophagy is required for fibrosis in human cardiac fibroblasts
To further determine whether autophagy induction contribute to TGF-β1-induced fibrosis, we examined the effects of TGF-β1 stimulation of human cardiac fibroblasts in the presence of autophagy inducer and inhibitor. Rapamycin and 3-methyladenine (3-MA) are known pharmacological inducer and inhibitor of autophagy, as reported by our group and others. Thus we co-treated human cardiac fibroblasts with rapamycin (1 μM) or 3-MA (5 mM) and TGF-β1 (10 ng/ml) for 48 and 72 hours, and collagen type I α2 and fibronectin expression levels were then compared with their corresponding controls (Fig. 2a,b). We found that rapamycin or 3-MA co-treatment significantly increased or decreased TGF-β1-induced pro-fibrotic effects in human cardiac fibroblasts (Fig. 2b). Densitometric analysis of LC3-II in co-treated human cardiac fibroblasts (TGF-β1 and autophagy inducer or inhibitor) versus controls revealed that TGF-β1 treatment is associated with an increase in LC3 -II at 48 hours versus control and an increase in LC3-II at 96 hours. We suggest that increased or decreased autophagy in the presence of autophagy inducer or inhibitor is linked to basal production of matrix component proteins and thus that autophagy is positively correlated to the synthesis of matrix components and human cardiac fibroblasts function.
(a) HCF were treated with TGF-β1 (10 ng/ml) in the presence of the rapamycin (1μM) and 3-MA (5 mM) for indicated durations. Western blotting revealed that inhibition of autophagy by 3-MA abrogated fibrogenic effects of TGF-β1, whereas rapamycin treatment enhanced fibrogenic effects of TGF-β1. (b,c) Densitometric analysis of fibronectin and collagen I α2 levels in HCF, which were stimulated with TGF-β1 or rapamycin (1μM) and 3-MA (5 mM). **p < 0.01 vs. TGF-β1-treated group. (d,e) Inhibition of autophagy by BECN1 siRNA decreased fibronection and collagen I α2 biosynthesis. BECN1 knocked down cells and their correspondence scramble infected cells were treated with TGF-β1 (10 ng/ml) for 48 and 72 hours. Protein loading was confirmed using GAPDH. Data are means of independent experiments of HCF cells from three different donors. For each experiment, LC3-I, LC3-II, p62/SQSTM1, fibronectin, and collagen I α2 levels were compared with those from time-matched controls and normalized to GAPDH levels. (f–h) Densitometry analysis showed that BECN1 knockdown was associated with a significant (**p < 0.01) decrease of TGF-β1-induced fibronectin and collagen I α2 biosynthesis in HCF. (I,j) HCF were pro-transfected with Atg 5 (pCMV-myc-Atg 5) and TGF-β1 (10 ng/ml) for another 48 hours. LC3-I, LC3-II, p62/SQSTM1, collagen I α2, and fibronectin levels were measured in whole-cell lysates. (k) Densitometry analysis revealed that autophagy induction by Atg 5 overexpression was associated with a significant (**p < 0.01) increase in TGF-β1-induced fibrogenic effects in HCF. (l) HCF were treated with TGF-β1 (10 ng/ml) in the presence of the rapamycin (1μM) and 3-MA (5 mM) for 48 hours. MMP-9, MMP-2, and Vimentin levels were measured in whole-cell lysates and normalized to GAPDH levels. (m,n) HCF were pre-transfected with BECN1 siRNA or Atg 5 (pCMV-myc-Atg 5) and TGF-β1 (10 ng/ml) for another 48 hours. MMP-9, MMP-2, and Vimentin levels were measured in whole-cell lysates and normalized to GAPDH levels.
Moreover, to utilize a parallel non-pharmacological approach to test the same hypothesis, beclin 1 (BECN1) gene expression was suppressed using siRNA interference in human cardiac fibroblasts (Fig. 2c) and later treated with TGF-β1 (10 ng/ml). These cells were compared with human cardiac fibroblasts, which were infected with scrambled siRNA. The expression of collagen type I α2 and fibronectin were then detected and compared between groups of scrambled siRNA and BECN1 siRNA transfection. The results showed that BECN1 knockdown significantly decreased TGF-β1-induced pro-fibrotic effects (Fig. 2d–g). We further used pCMV-myc-Atg5 plasmid to transfect into human cardiac fibroblasts and showed that TGF-β1 stimulation induced significantly more fibronectin synthesis in pCMV-myc-Atg5 overexpressed cells compared with corresponding control cells (Fig. 2h–k). Our results showed that rapamycin and pCMV-myc-Atg5 increased TGF-β1-induced expression of MMP2, MMP9, and Vimentin in human cardiac fibroblasts (Fig. 2l,m); 3-MA and BECN1 siRNA decreased TGF-β1-induced expression of MMP2, MMP9, and Vimentin in human cardiac fibroblasts (Fig. 2l,n), respectively. Our experiments showed that autophagy induction significantly increased the TGF-β1-induced fibrogenic effect.
Coincidence of elevated autophagy marker and fibronectin induction in human heart valve tissue
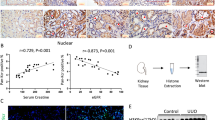
The twenty heart valve tissue samples of 20 cases patients with dilated cardiomyopathy were randomly selected for IHC assay. Representative photomicrographs of Fibronectin, MMP-2, MMP-9, MAP-LC3, TGF-β R I, and TGF-β R II expression in 20 samples of heart valve tissue from patients with HF were determined by IHC staining. Our results showed in Fig. 3 that heart valve tissue from patients with dilated cardiomyopathy was positive for Fibronectin, MMP-2, MMP-9, MAP-LC3, TGF-β R I, and TGF-β R II, except for sparse cytoplasmic staining in a few glia and neurons. Although these experiments were only conducted in specimens from a limited number of patients, their results are consistent with the preclinical evidence shown above and suggest that notably high expression of TGF-β R I and TGF-β R II may be associated with autophagy-mediated fibrosis in human heart diseases.
The twenty heart valve tissue samples of 20 cases patients with dilated cardiomyopathy were randomly selected for IHC assay. Representative photomicrographs of Fibronectin, MMP-2, MMP-9, MAP-LC3, TGF-β R I, and TGF-β R II expression in 20 samples of heart valve tissue from patients with dilated cardiomyopathy were determined by IHC staining. Image magnification: 100× (left); 200× (right). Scale bar of 10 mm (right) is equivalent to 50 μm.
MiR-19a-3p/19b-3p expresses with low levels in the plasma of HF patients
Studies of the last two years have shown that miRNA can be free outside the cells and stable in plasma or serum. Thus, these miRNAs can be used as disease diagnosis, therapeutic response and prognosis evaluation of effective and sensitive biomarker54. Thus, the differential expressions of 26 kinds of microRNAs from the literature were verified by Q-PCR assay and ath-156a was added as the external reference for corresponding control. Results show that expression of miRNA19a-3p/19b-3p is much low in HF patients (Fig. S1a,b). As shown in Fig. S1c,d, the expressions of miRNA 19a-3p/19b-3p were significantly down-regulated in DCM plasma pool samples. The expression of miRNA 19a-3p/19b-3p in the plasma samples of 38 patients with DCM was quantitatively tested by qRT-PCR. MiRNA 19a-3p/19b-3p decreased in DCM plasma samples. P < 0.001 (miRNA 19a); P = 0.001 (miRNA 19b). The expression of miRNA 19a-3p/19b-3p decreased more obviously in end stage of DCM plasma samples than in early stage. P = 0.009 (miRNA 19a); P = 0.008 (miRNA 19b) (Fig. S1e,f).
In addition, twenty plasma samples of 20 cases patients with heart failure were randomly selected as HF plasma samples; and another twenty plasma samples of 20 cases normal people were selected as normal control plasma samples. The characteristics of heart failure patients (HF group) and healthy controls (C group) were displayed in (Table 1). As shown in Fig. 4a,b, the expression of miR-19a-3p/19b-3p was significantly down-regulated in HF plasma samples, P < 0.001 compared to Control group.
MiR-19a-3p/19b-3p negatively regulates multiple players in fibrosis
To investigate whether the differential expression of miR-19a-3p/19b-3p was correlated with cell invasion, human cardiac fibroblasts were transfected with miR-19a-3p/19b-3p mimics and nonspecific miRNA Control. Transwell migration assay was used to investigate migration activity. As shown from the images in Fig. 5a,b, when the expression of miR-19a-3p/19b-3p was up-regulated by mimics in human cardiac fibroblasts, the cells demonstrated much low-migration potentiality compared to cells treated with miRNA Control. Q-PCR was used to analyze the expression of miR-19a-3p/19b-3p when treated with mimics and nonspecific miRNA Control. The result showed that the expression of miR-19a-3p/19b-3p was negative correlation with the migration ability of human cardiac fibroblasts (Fig. 5c,d).
(a) Photographs of the cell invasion through the polycarbonate membrane stain by crystal violet. The migratory cell numbers of human cardiac fibroblasts transfected with miR-19a-3p/19b-3p mimics were significantly more than that of cells transfected with miR Control respectively. (b) The inhibitory effect of miR-19a-3p/19b-3p on the invasion of the cells was quantified. **P < 0.01 compared with control. Data were presented as the mean ± SEM of three separate experiments. (c,d) The Q-PCR analyses of the expression of miR-19a-3p/19b-3p in human cardiac fibroblasts transfected with miR-19a-3p/19b-3p mimics, NC and untranfected one. −△△Ct = −(Ctx-CtU6x)-(CtNCx-CtU6NC). Data were present as mean ± SEM, n = 3, **P < 0.01 vs. NC group. (e) Human cardiac fibroblasts were transfected with miR-19a-3p/19b-3p mimics, miR Control. Expression of MMP-2, MMP-9, Vimentin and α-SMA after 72 hours were detected by western blotting. Protein loading was confirmed using GAPDH. (f) Quantitative data of densitometric analyses. The ratio of MMP-2, MMP-9, Vimentin and α-SMA to GAPDH were displayed as mean ± SEM, n = 3, *p < 0.05, **p < 0.01 vs. NC group.
It has been reported that MMP-2 and MMP-9 play a critical role in cell invasion by stimulating degradation of the ECM and cell migration. To probe the possible anti-invasion mechanism of miR-19a-3p/19b-3p, we tested the activity protein of MMP-2/9 in miR-19a-3p/19b-3p-transfected human cardiac fibroblasts. The protein of MMP-2 and MMP-9 in were decreased by miR-19a-3p/19b-3p, suggesting that miR-19a-3p/19b-3p suppresses the invasion ability of human cardiac fibroblasts through down-regulation of the expression of MMP-2/9 (Fig. 5e,f). Epithelial mesenchymal transition (EMT), characterized as loss of polarity and epithelial markers (including junctional and cell-cell adhesion proteins), has long been known to play a role in cellular metastasis and cell invasion. To further examine whether miR-19a-3p/19b-3p would inhibit EMT, consistent with metastasis ability, after transient transfection with miR-19a-3p/19b-3p mimics in human cardiac fibroblasts, both vimentin and α-SMA mRNA levels were decreased compared to miRNA Control group (Fig. 5e,f). In summary, miR-19a-3p/19b-3p was important in regulating reorganization of actin cytoskeleton and the maintenance of cell morphology.
MiR-19a-3p/19b-3p directly targeted TGF-β Receptor II
It is generally considered that miRNAs exert their function through regulating the expression of their downstream target genes. To investigate the target of miR-19a-3p/19b-3p in human cardiac fibroblasts, systemic bioinformatic publicly available algorithms were used to analyze and identify potential targets. We found that human TGFBR2 (TGF-β Receptor II) 3′-UTR contained putative miR-19a-3p/19b-3p complementary sites predicted using TargetScan 7.0 (http://www.targetscan.org) (Fig. 6a). To explore whether the TGF-β Receptor II was the target gene of the miR-19a-3p/19b-3p, we constructed the luciferase reporter vector with the putative TGF-β Receptor II 3′ UTR target site for the miR-19a-3p/19b-3p downstream of the luciferase gene (pGL3- TGF-β R II -3′-UTR). Luciferase reporter vector together with the miR-19a-3p/19b-3p mimics or the miRNA mimic control were transfected into HCF, respectively. In HCF, significant decrease in relative luciferase activity was noted when pGL3- TGF-β R II -3′-UTR was cotransfected with the miR-19a-3p/19b-3p mimics but not with the miRNA mimic control, respectively. These results showed that TGF-β R II was the target gene of the miR-19a-3p/19b-3p (Fig. 6b). To further identify the miR-19a-3p/19b-3p putative target gene TGF-β R II that might be involved in autophagy-related fibrosis, transfection and Q-PCR analysis was conducted to determine the mechanism by which miR-19a-3p/19b-3p inhibited TGF-β Receptor II mRNA. Figure 6c indicates that transfection of human cardiac fibroblasts from controls with miR-19a-3p/19b-3p mimics induced significant decrease in TGF-β Receptor II mRNA expression levels compared to miRNA Control transfection. Western blotting revealed that the protein level of TGF-β Receptor II but not TGF-β Receptor I was markedly reduced in the cells over-expressing miR-19a-3p/19b-3p compared to the non transfected cells (Fig. 6d,e).
(a) The miR-19a/b directly target on human TGF-β Receptor II were predicted using TargetScan 7.0. (b) Dual luciferase assay performed in HCF cells suggested that TGF-β Receptor II was the target gene of the miR-19a-3p/19b-3p. In HCF cells, significant decrease in relative luciferase activity was noted when pGL3- TGF-β R II -3′-UTR was cotransfected with the miR-19a-3p/19b-3p mimics but not with the miRNA mimic control, respectively. **p < 0.01. (c) TGF-β Receptor II mRNA levels were determined by Q-PCR and normalized to GAPDH. Human cardiac fibroblasts were transfected with miR-19a-3p/19b-3p mimics, miR Control. The gene expression of TGF-β Receptor II was detected after 48 hours. Data were present as the mean ± SEM, n = 3, **p < 0.01 vs. miR Control group. (d) Human cardiac fibroblasts were transfected with miR-19a-3p/19b-3p mimics, miR Control. TGF-β Receptor I/II protein and GAPDH after 72 hours were detected by western blotting. (e) Quantitative data of densitometric analyses. MiR-19a-3p/19b-3p mimic treatment was not associated with any significant changes in TGF-β Receptor I expression. The ratio of TGF-β Receptor II protein to GAPDH were displayed as mean ± SEM, n = 3, **p < 0.01 vs. miR Control group. (f) Western blotting analyses for total and phosphorylated forms of Smad 2 and Akt (p-Smad 2, p-Akt) expression in human cardiac fibroblasts. The experiments were normalized with GAPDH at baseline level and after pro-transfection with miR-19a-3p/19b-3p mimics, miR Control and stimulation with TGF-β1 (10 ng/ml) for 72 hours. Results are the means from three independent experiments using cells from three different donors. (g) Densitometric analysis of Smad2 and Akt phosphorylation levels in human cardiac fibroblasts. Each bar represents the mean ± SEM calculated from three independent experiments. *p < 0.05 compared with miR Control and TGF-β1 group; **p < 0.01 compared with miR Control and TGF-β1 group.
We further evaluated whether miR-19a-3p/19b-3p modulation altered the activation of Smad/Akt pathway proteins after TGF-β1 treatment. Western blotting analyses for total and phosphorylated forms of Smad 2 and Akt (p-Smad 2, p-Akt) expression in human cardiac fibroblasts were carried out. The experiments were normalized with GAPDH at baseline level and after pro-transfection with miR-19a-3p/19b-3p mimics, miR Control and stimulation with TGF-β1 (10 ng/ml) for 72 hours. As shown in Fig. 6f, the functional phosphorylation level of Smad 2 and Akt is up-regulated by TGF-β1 treatment in human cardiac fibroblasts. MiR-19a-3p/19b-3p modulation did not affect the expression of total Smad 2/3 and Akt. However, the over-expression of miR-19a-3p/19b-3p significantly reduced the level of the phosphorylated forms of Smad 2 and Akt (Fig. 6f,g).
MiR-19a-3p/19b-3p decreases TGF-β1 induced autophagy-related fibrosis
We validated that miR-19a-3p/19b-3p negatively regulates multiple players in autophagy and fibrosis and that TGF-β R II was one of miR-19a-3p/19b-3p targets was also validated in previous results. To further explore whether miR-19a-3p/19b-3p inhibited autophagy-mediated fibrosis induced by targeting TGF-β1, pharmacological induction of autophagy was applied. It was shown in Fig. 7 that we got an expected inhibition effect after transfected with miR-19a-3p/19b-3p for 72 h. The invasion potential of human cardiac fibroblasts was examined by transwell invasion assay with co-treatment of miR-19a-3p/19b-3p mimics and TGF-β1 (10 ng/ml). The treatment with miR-19a-3p/19b-3p mimics led to significantly decreased invasion cells compared to the treatment of Control group (Fig. 7a,b). Further we investigated the relationship between autophagy and invasion potential in human cardiac fibroblasts by SNLYSO autophagy detection assay with co-treatment of miR-19a-3p/19b-3p mimics and TGF-β1 (10 ng/ml). It was shown in Fig. 7c,d, the autophagy induction of TGF-β1 was significantly reduced by miR-19a-3p/19b-3p mimics (16.8% to 2.37% and 2.99%). Confocal fluorescence images of endogenous MAP-LC3 and SNLYSO sensor-labeled compartment in human cardiac fibroblasts also verified the above results. The results showed that miR-19a-3p/19b-3p reduced the accumulation of both the yellow and red puncta induced by TGF-β1 (Fig. 7e). The vacuoles assumed to be autophagosomes, would be expected to undergo acidification after maturation and finally, fuse with lysosomes so that their content is digested by lysosomal hydrolases. The large endosomes subsequently recruit multiple autophagosomes.
(a) Effects of miR-19a-3p/19b-3p and TGF-β1 on cell invasion in human cardiac fibroblasts in vitro. Photographs of the cell invasion through the polycarbonate membrane stained with crystal violet. (b) The inhibitory effect of miR-19a-3p/19b-3p and TGF-β1 on the invasion of the cells was quantified. Data were presented as the mean ± SEM of three separate experiments. *p < 0.05, **p < 0.01, group of pro-transfected with miR-19a-3p/19b-3p mimics and stimulation of TGF-β1 compared with control. (c,d) Human cardiac fibroblasts were pro-transfected with miR-19a-3p/19b-3p and then treated with 10 ng/ml TGF-β1 for 72 h. Cells were then probed by SNLYSO sensor and autolysosomes were analyzed and quantitated by flow cytometer. Results from three independent experiments are shown as means ± SEM. **p < 0.01, group of pro-transfected with miR-19a-3p/19b-3p mimics and stimulation of TGF-β1 compared with control. (e) Confocal fluorescence images of endogenous MAP-LC3 and SNLYSO sensor-labeled compartment in human cardiac fibroblasts. Human cardiac fibroblasts were pro-transfected with miR-19a-3p/19b-3p and then treated with 10 ng/ml TGF-β1. MAP-LC3 (green) was labeled with primary anti-MAP-LC3 polyclonal antibody. Goat anti-rabbit IgG/FITC were used as secondary antibody. Cells were simultaneously imaged in the presence of SNLYSO sensor (red) to visualize autophagy. Nuclei (blue) were labeled by DAPI. Confocal microscopy images were obtained. Bar = 30 μm.
We next determined whether the expression of autophagy and fibrosis related proteins would be modulated by miR-19a-3p/19b-3p. Human cardiac fibroblasts were pre-transfected with miR-19a-3p/19b-3p and miR Control. Then cells were treated with TGF-β1 (10 ng/ml) for the indicated duration. LC3-I, LC3-II, p62/SQSTM1, collagen Iα2, fibronectin, MMP-9, and MMP-2 levels were measured in whole-cell lysates. Protein loading was confirmed using GAPDH. The conversion from LC3-Ito LC3-II and degradation of p62/SQSTM1 induced by TGF-β1 were significantly restored by miR-19a-3p/19b-3p (Fig. 8a,b). Consistently, the increased synthesis of collagen type Iα2 and fibronectin and expression of MMP-2 and MMP-9 induced by TGF-β1 were also significantly restored by miR-19a-3p/19b-3p (Fig. 8a,c). To determine whether autophagy activation rescued the inhibition effect of miR-19a-3p/19b-3p on autophagy-related fibrosis, rapamycin was applied for autophagy inducer. As is shown in Fig. 8d–f, rapamycin could further increase LC3-II level, synthesis of collagen type I α2 and fibronectin and expression of MMP-2 and MMP-9 in human cardiac fibroblasts with miR-19a-3p/19b-3p and TGF-β1. Taken together, these results demonstrated that autophagy mediated the regulation of miR-19a-3p/19b-3p on fibrosis in human cardiac fibroblasts.
(a) Human cardiac fibroblasts were pre-transfected with miR-19a-3p/19b-3p and miR Control. Then cells were treated with TGF-β1 (10 ng/ml) for the indicated duration. LC3-I, LC3-II, p62/SQSTM1, collagen I α2, fibronectin, MMP-9, and MMP-2 levels were measured in whole-cell lysates. Protein loading was confirmed using GAPDH. (b,c) Quantitative data of densitometric analyses. The ratios of LC3-I, LC3-II, p62/SQSTM1, collagen I α2, fibronectin, MMP-9, and MMP-2 to GAPDH were present as mean ± SEM, n = 3, *p < 0.05, **p < 0.01 vs. group of miR Control and TGF-β1 co-treatment. (d) Rapamycin reverses the inhibition effect of miR-19a-3p/19b-3p on TGF-β1-induced autophagy-related fibrosis. Human cardiac fibroblasts were transfected with miR-19a-3p/19b-3p and miR Control. Then cells were pretreated with rapamycin (4 hours, 1 μM) and co-treated with TGF-β1 (10 ng/ml) for the indicated duration. LC3-I, LC3-II, p62/SQSTM1, collagen Iα2, fibronectin, MMP-9, and MMP-2 levels were measured in whole-cell lysates. Protein loading was confirmed using GAPDH. (e,f) Densitometry analysis showed that rapamycin (1 μM) significantly reversed the inhibition of miR-19a-3p/19b-3p on TGF-β1-induced autophagy-related fibrosis in human cardiac fibroblasts. *p < 0.05, **p < 0.01 vs group of miR Control and TGF-β1 co-treatment; ##p < 0.01 vs group of miR 19a-3p and TGF-β1 co-treatment; $$p < 0.001 vs group of miR 19b-3p and TGF-β1 co-treatment.
Discussion
HF is characterized by left ventricular (LV) remodeling and dilatation, with activation of a fetal gene program that triggers pathological changes in the myocardium associated with progressive dysfunction55. Tissue biopsy and cardiac radionuclide scan showed that there was no necrosis of cardiac muscle cells in the early stage of HF, but the degeneration, apoptosis and interstitial fibrosis of myocardial cells56,57,58,59,60,61. Therefore, most of the early HF patients firstly exhibit heart cavity expansion without obvious cardiac function insufficiency. With the continued existence of pathological matrix, necrosis of myocardial cells and extracellular matrix edema begin to appear, and then the heart function reduces till heart failure. Therefore, further study on the internal mechanism of HF pathology and explore the effective means to block the mechanism, will provide valuable basic research information for clinical treatment of HF. Indeed, certain mutations often induce HF with cardiac arrhythmia that is considered as the potential trigger of sudden cardiac death62,63. Thus, both the effective prognostic determination and appropriate cardiac care depend on accurate molecular and genetic diagnoses and therapy.
In this study, we have shown that TGF-β1 treatment simultaneously induced autophagy and fibrosis in human cardiac fibroblasts (Fig. 1). Either inhibition or induction of autophagy reduces and increases fibrotic characteristic in TGF-β1-induced fibrosis (Fig. 2). These findings highlighted a linking between autophagy and elevated matrix protein synthesis by human cardiac fibroblasts and strongly supported the hypothesis that TGF-β1-induced fibrosis was depending on its autophagy induction. In the present study we showed that the over-expressions of TGF-β Receptor I and TGF-β Receptor II are accompanied with hallmarks of autophagy, fibrosis, and ECM production in human cardiac tissues from heart valve (Fig. 3). Although miRNAs are essential to the regulation of cardiac remodeling, many of the mechanisms have not been well characterized45,46,47,64. The miR-19a-3p/19b-3p expression is specifically decreased in patients with DCM, especially in the end stage of DCM (Fig. 4). Herein we demonstrated that the cardiac-specific over-expression of miR-19a-3p/19b-3p resulted in inhibition of interstitial fibrosis, invasion potential, and epithelial mesenchymal transition (EMT) (Fig. 5). Furthermore, we confirmed and extended our previous discoveries, revealing that miR-19a-3p/19b-3p is a negative regulator of TGF-β1/Smad2 signaling via directly targeting TGF-β Receptor II (Fig. 6). To elucidate that miR-19a-3p/19b-3p is the important regulator of cardiac homeostasis and remodeling for potential therapeutic strategy for heart failure, we therefore demonstrated the miR-19a-3p/19b-3p inhibited the fibrosis through autophagy inhibition by targeting TGF-β R II in human cardiac fibroblasts (Figs 7 and 8).
Autophagy basically exists in most cells and can be rapidly activated as an adaptive response to renew intracellular substances and nutrition under cellular stimulation65,66. Autophagy (macro-autophagy) is a highly conserved process that is tightly regulated by various biological mechanisms and has important roles in many events, such as cellular remodeling during differentiation, development, adapt to environmental stress31,67. In contrast, autophagic cell death can drive cells towards type II programmed cell death, which is morphologically distinct from type I programmed cell death (apoptosis). However, a growing body of evidences points that autophagy is more important for cell survival. The accumulation of autophagosomes and autolysosomes exhibit survival response to deadly stress in order to rid the cell of harmful proteins or damaged organelles33,65. Autophagy is required to maintain cardiac homeostasis and function. Conversely, disrupted autophagy may contribute to cardiac remodeling37. TGF-β1 has been demonstrated to induce both autophagy and fibrosis in many tissues35,68. The coincidence of elevated autophagy marker and fibronectin induction in many diseases has been previously observed35. In the present study, our results demonstrate that autophagy is necessary for induction of fibrosis in human cardiac fibroblasts.
After TGF-β1 binding to the TGF-β R II, the phosphorylated TGF-β R I recruits and phosphorylates receptor-regulated Smad proteins (Smad2/3 complex). Thus, we have evaluated the functional phosphorylated form of Smad 2 and Smad 3 after TGF-β1 treatment and miR-19a-3p/19b-3p modulation. Our data show that the phosphorylated activated form of Smad 2 and Smad 3 after miR-19a-3p/19b-3p over-expression. Therefore the down-regulation of TGF-β R II and phosphorylated form of Smad 2 and Smad 3 are modulated by miR-19a-3p/19b-3p.
MiRNAs are considered as important regulators of gene expression, suppressing the expression of target genes through translational repression or degradation of a target transcript39,53. Because of the difficulty to obtain clinical HF heart tissue, we could not verify the miRNA expression in HF heart tissues. However, an important study on heart tissues of 25 patients with HF has shown the abnormal expression of various miRNA. Among them, the decreasing ratio of miRNA-19a/b is the most distinct48. Here, we focused on the regulation of miR-19a-3p/19b-3p on the autophagy-related fibrosis in human cardiac fibroblasts and the mechanisms of regulation of its target genes. We examined miR-19a-3p/19b-3p expression in human cardiac tissues and human cardiac fibroblasts by Q-PCR assay, as previously described. We integrated Targetscan 7.0 predictions and the resulting candidate functions and found that the TGF-β R II gene had the highest recurrence rate as a potential target gene of miR-19a-3p/19b-3p. MiRNAs are likely to bind partially to homologous sequence of a target gene in the 3′UTR53. We found that over-expression of miR-19a-3p/19b-3p could significantly reduce TGF-β R II gene expression. Furthermore, we detected the TGF-β/Smad2 signaling transduction undergoing the over-expression of miR-19a-3p/19b-3p. These results suggested that miR-19a-3p/19b-3p can directly and negatively regulate TGF-β R II gene expression and inhibit the TGF-β/Smad2 signaling transduction activated by TGF-β1.
Additional Information
How to cite this article: Zou, M. et al. Autophagy inhibition of hsa-miR-19a-3p/19b-3p by targeting TGF-β R II during TGF-β1-induced fibrogenesis in human cardiac fibroblasts. Sci. Rep. 6, 24747; doi: 10.1038/srep24747 (2016).
References
McKinsey, T. A. & Olson, E. N. Toward transcriptional therapies for the failing heart: chemical screens to modulate genes. J Clin Invest 115, 538–546, doi: 10.1172/JCI24144 (2005).
Zhou, J. et al. Engineering the heart: evaluation of conductive nanomaterials for improving implant integration and cardiac function. Sci Rep 4, 3733, doi: 10.1038/srep03733 (2014).
Yamada, T. et al. Relationship of myocardial fibrosis to left ventricular and mitochondrial function in nonischemic dilated cardiomyopathy–a comparison of focal and interstitial fibrosis. J Card Fail 19, 557–564, doi: 10.1016/j.cardfail.2013.05.018 (2013).
Dobaczewski, M., Chen, W. & Frangogiannis, N. G. Transforming growth factor (TGF)-beta signaling in cardiac remodeling. J Mol Cell Cardiol 51, 600–606, doi: 10.1016/j.yjmcc.2010.10.033 (2011).
Mascareno, E. et al. Cardiac lineage protein-1 (CLP-1) regulates cardiac remodeling via transcriptional modulation of diverse hypertrophic and fibrotic responses and angiotensin II-transforming growth factor beta (TGF-beta1) signaling axis. J Biol Chem 287, 13084–13093, doi: 10.1074/jbc.M111.288944 (2012).
Tarquini, R., Lazzeri, C., Pala, L., Rotella, C. M. & Gensini, G. F. The diabetic cardiomyopathy. Acta Diabetol 48, 173–181, doi: 10.1007/s00592-010-0180-x (2011).
Murarka, S. & Movahed, M. R. Diabetic cardiomyopathy. J Card Fail 16, 971–979, doi: 10.1016/j.cardfail.2010.07.249 (2010).
Gonzalez, A., Lopez, B., Querejeta, R. & Diez, J. Regulation of myocardial fibrillar collagen by angiotensin II. A role in hypertensive heart disease? J Mol Cell Cardiol 34, 1585–1593 (2002).
Puntmann, V. O. et al. Aortic stiffness and interstitial myocardial fibrosis by native T1 are independently associated with left ventricular remodeling in patients with dilated cardiomyopathy. Hypertension 64, 762–768, doi: 10.1161/HYPERTENSIONAHA.114.03928 (2014).
Brooks, W. W. & Conrad, C. H. Myocardial fibrosis in transforming growth factor beta(1)heterozygous mice. J Mol Cell Cardiol 32, 187–195, doi: 10.1006/jmcc.1999.1065 (2000).
Burchfield, J. S., Xie, M. & Hill, J. A. Pathological ventricular remodeling: mechanisms: part 1 of 2. Circulation 128, 388–400, doi: 10.1161/CIRCULATIONAHA.113.001878 (2013).
Kingsley, D. M. The TGF-beta superfamily: new members, new receptors, and new genetic tests of function in different organisms. Genes Dev 8, 133–146 (1994).
Brand, T. & Schneider, M. D. Transforming growth factor-beta signal transduction. Circ Res 78, 173–179 (1996).
del Pilar Alatorre-Carranza, M. et al. Liver fibrosis secondary to bile duct injury: correlation of Smad7 with TGF-beta and extracellular matrix proteins. BMC Gastroenterol 9, 81, doi: 10.1186/1471-230X-9-81 (2009).
Sui, X., Wei, H. & Wang, D. Novel mechanism of cardiac protection by valsartan: synergetic roles of TGF-beta1 and HIF-1alpha in Ang II-mediated fibrosis after myocardial infarction. J Cell Mol Med 19, 1773–1782, doi: 10.1111/jcmm.12551 (2015).
Montfort, I. & Perez-Tamayo, R. The distribution of collagenase in normal rat tissues. J Histochem Cytochem 23, 910–920 (1975).
Horn, M. A. et al. Age-related divergent remodeling of the cardiac extracellular matrix in heart failure: collagen accumulation in the young and loss in the aged. J Mol Cell Cardiol 53, 82–90, doi: 10.1016/j.yjmcc.2012.03.011 (2012).
Collier, P. et al. Getting to the heart of cardiac remodeling; how collagen subtypes may contribute to phenotype. J Mol Cell Cardiol 52, 148–153, doi: 10.1016/j.yjmcc.2011.10.002 (2012).
Levick, S. P. & Brower, G. L. Regulation of matrix metalloproteinases is at the heart of myocardial remodeling. Am J Physiol Heart Circ Physiol 295, H1375–1376, doi: 10.1152/ajpheart.907.2008 (2008).
Spinale, F. G. Myocardial matrix remodeling and the matrix metalloproteinases: influence on cardiac form and function. Physiol Rev 87, 1285–1342, doi: 10.1152/physrev.00012.2007 (2007).
Jugdutt, B. I. Matrix metalloproteinases as markers of adverse remodeling after myocardial infarction. J Card Fail 12, 73–76, doi: 10.1016/j.cardfail.2005.10.001 (2006).
Wieser, R., Wrana, J. L. & Massague, J. GS domain mutations that constitutively activate T beta R-I, the downstream signaling component in the TGF-beta receptor complex. EMBO J 14, 2199–2208 (1995).
Lee, S. C., Kim, S. H., Koh, H. J. & Kwon, O. W. TGF-betas synthesized by RPE cells have autocrine activity on mesenchymal transformation and cell proliferation. Yonsei Med J 42, 271–277 (2001).
Liu, X. et al. Negative interplay of retinoic acid and TGF-beta signaling mediated by TG-interacting factor to modulate mouse embryonic palate mesenchymal-cell proliferation. Birth Defects Res B Dev Reprod Toxicol 101, 403–409, doi: 10.1002/bdrb.21130 (2014).
Hsu, Y. C., Chen, M. J., Yu, Y. M., Ko, S. Y. & Chang, C. C. Suppression of TGF-beta1/SMAD pathway and extracellular matrix production in primary keloid fibroblasts by curcuminoids: its potential therapeutic use in the chemoprevention of keloid. Arch Dermatol Res 302, 717–724, doi: 10.1007/s00403-010-1075-y (2010).
Tseng, Z. H. et al. Association of TGFBR2 polymorphism with risk of sudden cardiac arrest in patients with coronary artery disease. Heart Rhythm 6, 1745–1750, doi: 10.1016/j.hrthm.2009.08.031 (2009).
McKnight, A. J., Savage, D. A., Patterson, C. C., Sadlier, D. & Maxwell, A. P. Resequencing of genes for transforming growth factor beta1 (TGFB1) type 1 and 2 receptors (TGFBR1, TGFBR2), and association analysis of variants with diabetic nephropathy. BMC Med Genet 8, 5, doi: 10.1186/1471-2350-8-5 (2007).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 8, 445–544 (2012).
Zou, M. et al. Beclin 1-mediated autophagy in hepatocellular carcinoma cells: implication in anticancer efficiency of oroxylin A via inhibition of mTOR signaling. Cell Signal 24, 1722–1732, doi: 10.1016/j.cellsig.2012.04.009 (2012).
Zou, M. et al. Oroxylin A induces autophagy in human malignant glioma cells via the mTOR-STAT3-Notch signaling pathway. Mol Carcinog 54, 1363–1375, doi: 10.1002/mc.22212 (2015).
Levine, B. & Klionsky, D. J. Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev Cell 6, 463–477 (2004).
He, C. & Klionsky, D. J. Regulation mechanisms and signaling pathways of autophagy. Annu Rev Genet 43, 67–93, doi: 10.1146/annurev-genet-102808-114910 (2009).
Berry, D. L. & Baehrecke, E. H. Autophagy functions in programmed cell death. Autophagy 4, 359–360 (2008).
Yeganeh, B. et al. Asthma and influenza virus infection:focusing on cell death and stress pathways in influenza virus replication. Iran J Allergy Asthma Immunol 12, 1–17, doi: 012.01/ijaai.117 (2013).
Ghavami, S. et al. Autophagy is a regulator of TGF-beta1-induced fibrogenesis in primary human atrial myofibroblasts. Cell Death Dis 6, e1696, doi: 10.1038/cddis.2015.36 (2015).
Nakai, A. et al. The role of autophagy in cardiomyocytes in the basal state and in response to hemodynamic stress. Nat Med 13, 619–624, doi: 10.1038/nm1574 (2007).
Taneike, M. et al. Inhibition of autophagy in the heart induces age-related cardiomyopathy. Autophagy 6, 600–606, doi: 10.4161/auto.6.5.11947 (2010).
Carthew, R. W. & Sontheimer, E. J. Origins and Mechanisms of miRNAs and siRNAs. Cell 136, 642–655, doi: 10.1016/j.cell.2009.01.035 (2009).
Ambros, V. The functions of animal microRNAs. Nature 431, 350–355, doi: 10.1038/nature02871 (2004).
Xu, J., Wang, Y., Tan, X. & Jing, H. MicroRNAs in autophagy and their emerging roles in crosstalk with apoptosis. Autophagy 8, 873–882, doi: 10.4161/auto.19629 (2012).
Vettori, S., Gay, S. & Distler, O. Role of MicroRNAs in Fibrosis. Open Rheumatol J 6, 130–139, doi: 10.2174/1874312901206010130 (2012).
Selcuklu, S. D. et al. MicroRNA-9 inhibition of cell proliferation and identification of novel miR-9 targets by transcriptome profiling in breast cancer cells. J Biol Chem 287, 29516–29528, doi: 10.1074/jbc.M111.335943 (2012).
Su, Z., Yang, Z., Xu, Y., Chen, Y. & Yu, Q. MicroRNAs in apoptosis, autophagy and necroptosis. Oncotarget 6, 8474–8490, doi: 10.18632/oncotarget.3523 (2015).
Zhao, Y. et al. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell 129, 303–317, doi: 10.1016/j.cell.2007.03.030 (2007).
Wang, J. et al. MicroRNA-24 regulates cardiac fibrosis after myocardial infarction. J Cell Mol Med 16, 2150–2160, doi: 10.1111/j.1582-4934.2012.01523.x (2012).
Ge, Y. et al. MicroRNA-350 induces pathological heart hypertrophy by repressing both p38 and JNK pathways. Biochim Biophys Acta 1832, 1–10, doi: 10.1016/j.bbadis.2012.09.004 (2013).
Ucar, A. et al. The miRNA-212/132 family regulates both cardiac hypertrophy and cardiomyocyte autophagy. Nat Commun 3, 1078, doi: 10.1038/ncomms2090 (2012).
Ikeda, S. et al. Altered microRNA expression in human heart disease. Physiol Genomics 31, 367–373, doi: 10.1152/physiolgenomics.00144.2007 (2007).
Yang, H. et al. Wogonin induces cell cycle arrest and erythroid differentiation in imatinib-resistant K562 cells and primary CML cells. Oncotarget 5, 8188–8201, doi: 2340/oncotarget2340 (2014).
Zhang, J. et al. MicroRNA-638 inhibits cell proliferation, invasion and regulates cell cycle by targeting tetraspanin 1 in human colorectal carcinoma. Oncotarget 5, 12083–12096, doi: 10.18632/oncotarget.2499 (2014).
Khan, S. et al. MicroRNA-145 targets MUC13 and suppresses growth and invasion of pancreatic cancer. Oncotarget 5, 7599–7609, doi: 10.18632/oncotarget.2281 (2014).
Konno, Y. et al. MicroRNA-101 targets EZH2, MCL-1 and FOS to suppress proliferation, invasion and stem cell-like phenotype of aggressive endometrial cancer cells. Oncotarget 5, 6049–6062, doi: 10.18632/oncotarget.2157 (2014).
Xu, X. M. et al. MicroRNA-19a and -19b regulate cervical carcinoma cell proliferation and invasion by targeting CUL5. Cancer Lett 322, 148–158, doi: 10.1016/j.canlet.2012.02.038 (2012).
Zile, M. R. et al. Relationship between the temporal profile of plasma microRNA and left ventricular remodeling in patients after myocardial infarction. Circ Cardiovasc Genet 4, 614–619, doi: 10.1161/CIRCGENETICS.111.959841.
Zhao, D. S. et al. Serum miR-210 and miR-30a expressions tend to revert to fetal levels in Chinese adult patients with chronic heart failure. Cardiovasc Pathol 22, 444–450, doi: 10.1016/j.carpath.2013.04.001 (2013).
Sanbe, A. Dilated cardiomyopathy: a disease of the myocardium. Biol Pharm Bull 36, 18–22 (2013).
Vrtovec, B. et al. Effects of intracoronary CD34+ stem cell transplantation in nonischemic dilated cardiomyopathy patients: 5-year follow-up. Circ Res 112, 165–173, doi: 10.1161/CIRCRESAHA.112.276519 (2013).
Chun, J. L., O’Brien, R., Song, M. H., Wondrasch, B. F. & Berry, S. E. Injection of vessel-derived stem cells prevents dilated cardiomyopathy and promotes angiogenesis and endogenous cardiac stem cell proliferation in mdx/utrn−/− but not aged mdx mouse models for duchenne muscular dystrophy. Stem Cells Transl Med 2, 68–80, doi: 10.5966/sctm.2012-0107 (2013).
Lelakowski, J. et al. Factors predisposing to ventricular tachyarrhythmia leading to appropriate ICD intervention in patients with coronary artery disease or non-ischaemic dilated cardiomyopathy. Kardiol Pol 70, 1264–1275 (2012).
Mozid, A. M., Arnous, S., Sammut, E. C. & Mathur, A. Stem cell therapy for heart diseases. Br Med Bull 98, 143–159, doi: 10.1093/bmb/ldr014 (2011).
Brunner, S. et al. Left ventricular functional assessment in murine models of ischemic and dilated cardiomyopathy using [18 F]FDG-PET: comparison with cardiac MRI and monitoring erythropoietin therapy. EJNMMI Res 2, 43, doi: 10.1186/2191-219X-2-43 (2012).
Gosselin-Badaroudine, P. et al. A proton leak current through the cardiac sodium channel is linked to mixed arrhythmia and the dilated cardiomyopathy phenotype. PLoS One 7, e38331, doi: 10.1371/journal.pone.0038331 (2012).
Tachi, M. et al. Relationship of postcontrast myocardial T1 value and delayed enhancement to reduced cardiac function and serious arrhythmia in dilated cardiomyopathy with left ventricular ejection fraction less than 35. Acta Radiol, doi: 10.1177/0284185115580840 (2015).
Callis, T. E. et al. MicroRNA-208a is a regulator of cardiac hypertrophy and conduction in mice. J Clin Invest 119, 2772–2786, doi: 10.1172/JCI36154 (2009).
Mizushima, N., Levine, B., Cuervo, A. M. & Klionsky, D. J. Autophagy fights disease through cellular self-digestion. Nature 451, 1069–1075, doi: 10.1038/nature06639 (2008).
Mehrpour, M., Esclatine, A., Beau, I. & Codogno, P. Autophagy in health and disease. 1. Regulation and significance of autophagy: an overview. Am J Physiol Cell Physiol 298, C776–785, doi: 10.1152/ajpcell.00507.2009 (2010).
Mancias, J. D. & Kimmelman, A. C. Targeting autophagy addiction in cancer. Oncotarget 2, 1302–1306, doi: 10.18632/oncotarget.384 (2011).
Xu, Y. et al. Tgf-beta1 induces autophagy and promotes apoptosis in renal tubular epithelial cells. Int J Mol Med 29, 781–790, doi: 10.3892/ijmm.2012.911 (2012).
Acknowledgements
The authors are grateful to the fund support by the National Natural Science Foundation of China (Grant number: 81201705; 81372035; 81571873; 81402959), the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD) and Science and Technology Development Foundation of Nanjing Medical University (2013NJMU011).
Author information
Authors and Affiliations
Contributions
M.Z., F.W., R.G., Y.O., X. C., T.J. and W.W. performed the experiments in cell line. M.Z., J.W., T.W., C.L., J.C. and Q.H. performed the experiments in human samples. M.Z., W.Z., P.L., L.Q. and X.Z. contributed reagents/materials/analysis tools. M.Z., F.W. and W.Z. collected and analyzed the data. M.Z., W.Z. and Y.C. conceived and designed the study. M.Z. and W.Z. wrote the main manuscript text. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zou, M., Wang, F., Gao, R. et al. Autophagy inhibition of hsa-miR-19a-3p/19b-3p by targeting TGF-β R II during TGF-β1-induced fibrogenesis in human cardiac fibroblasts. Sci Rep 6, 24747 (2016). https://doi.org/10.1038/srep24747
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24747
This article is cited by
-
MicroRNA delivery based on nanoparticles of cardiovascular diseases
Molecular and Cellular Biochemistry (2023)
-
Differential expression of aqueous humor microRNAs in central retinal vein occlusion and its association with matrix metalloproteinases: a pilot study
Scientific Reports (2022)
-
The potential regulatory role of hsa_circ_0004104 in the persistency of atrial fibrillation by promoting cardiac fibrosis via TGF-β pathway
BMC Cardiovascular Disorders (2021)
-
Nuclear miR-30b-5p suppresses TFEB-mediated lysosomal biogenesis and autophagy
Cell Death & Differentiation (2021)
-
Isoflurane and low-level carbon monoxide exposures increase expression of pro-survival miRNA in neonatal mouse heart
Cell Stress and Chaperones (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.