Abstract
Prefoldin is a molecular chaperone complex that regulates tubulin function in mitosis. Here, we show that Prefoldin depletion results in disruption of neuroblast polarity, leading to neuroblast overgrowth in Drosophila larval brains. Interestingly, co-depletion of Prefoldin and Partner of Inscuteable (Pins) leads to the formation of gigantic brains with severe neuroblast overgrowth, despite that Pins depletion alone results in smaller brains with partially disrupted neuroblast polarity. We show that Prefoldin acts synergistically with Pins to regulate asymmetric division of both neuroblasts and Intermediate Neural Progenitors (INPs). Surprisingly, co-depletion of Prefoldin and Pins also induces dedifferentiation of INPs back into neuroblasts, while depletion either Prefoldin or Pins alone is insufficient to do so. Furthermore, knocking down either α-tubulin or β-tubulin in pins- mutant background results in INP dedifferentiation back into neuroblasts, leading to the formation of ectopic neuroblasts. Overexpression of α-tubulin suppresses neuroblast overgrowth observed in prefoldin pins double mutant brains. Our data elucidate an unexpected function of Prefoldin and Pins in synergistically suppressing dedifferentiation of INPs back into neural stem cells.
Similar content being viewed by others
Introduction
Control of tissue homeostasis is a central issue during development. The neural stem cells, or neuroblasts, of the Drosophila larval brain is an excellent model for studying stem cell homeostasis1,2,3,4,5. Asymmetric division of neuroblasts generates a self-renewing neuroblast and a different daughter cell that undergoes differentiation pathway to produce neurons or glia6. Following each asymmetric division, apical proteins such as aPKC are segregated into the neuroblast daughter and function as “proliferation factor”, while basal proteins are segregated into a smaller daughter cell to act as “differentiation factors”7,8,9,10. At the onset of mitosis, the Partitioning defective (Par) protein complex that is composed of Bazooka (Baz)/Par3, Par6 and atypical protein kinase C (aPKC) is asymmetrically localized at the apical cortex of the neuroblast11,12,13. Other apical proteins including Partner of Inscuteable (Pins), the heterotrimeric G protein Gαi and Mushroom body defect (Mud) also accumulate at the apical cortex through an interaction of Inscuteable (Insc) with Par protein complex14,15,16,17,18. Apical proteins control basal localization of cell fate determinants Numb, Prospero (Pros), Brain tumor (Brat) and their adaptor proteins Miranda (Mira) and Partner of Numb (Pon) that are segregated into the ganglion mother cell (GMC) following divisions1. Apical proteins and their regulators also control mitotic spindle orientation to ensure correct asymmetric protein segregation at telophase14,15,16,17,18,19,20,21,22,23. Several centrosomal proteins, Aurora A, Polo and Centrosomin, regulate mitotic spindle orientation24,25,26.
There are at least two different types of neuroblasts that undergo asymmetric division in the larval central brain27,28,29. Perturbation of asymmetric division in either type of neuroblast can trigger neuroblast overproliferation and/or the induction of brain tumors4,30. The majority of neuroblasts are type I neuroblasts that generate a neuroblast and a GMC in each division, while type II neuroblasts generate a neuroblast and an intermediate neural progenitor (INP), which undergoes three to five rounds of asymmetric division to produce GMCs27,28,29. Ets transcription factor Pointed (PntP1 isoform), exclusively expressed in type II neuroblast lineages, promotes the formation of INPs31. Failure to restrict the self-renewal potential of INPs can lead to dedifferentiation, allowing INPs to revert back into “ectopic neuroblasts”32. Notch antagonist Numb and Brat function cooperatively to promote the INP fate29. Loss of brat or numb leads to “ectopic type II neuroblasts” originating from uncommitted immature INPs that failed to undergo maturation29. A zinc-finger transcription factor Earmuff functions after Brat and Numb in immature INPs to prevent their dedifferentiation33. Earmuff also associates with Brahma and HDAC3, which are involved in chromatin remodeling, to prevent INP dedifferentiation34,35. However, the underlying mechanism by which INPs possess limited developmental potential is largely unknown.
Prefoldin (Pfdn) was first identified as a hetero-hexameric chaperone consisting of two α-like (PFDN3 and 5) and four β-like (PFDN 1, 2, 4 and 6) subunits, based on its ability to capture unfolded actin36. Prefoldin promotes folding of proteins such as tubulin and actin by binding specifically to cytosolic chaperonin containing TCP-1 (CCT) and by directing target proteins to it36. The yeast homologs of Prefoldin 2–6, named GIM1-5 (genes involved in microtubule biogenesis) are present in a complex that facilitates proper folding of α-tubulin and γ-tubulin37. All Prefoldin subunits are phylogenetically conserved from Archaea to Eukarya38. Structural study of the Prefoldin hexamer from the archaeum M. thermoautotrophicum showed that Prefoldin forms a jellyfish-like shape consisting of a double β barrel assembly with six long tentacle-like coiled coils that participate in substrate binding39. The function of Prefoldin as a chaperone has also been illustrated in lower eukaryotes like C. elegans, in which loss of prefoldin resulted in defects in cell division due to reduced microtubule growth rate40. Depletion of PFDN1 in mice displayed cytoskeleton-related defects, including neuronal loss and lymphocyte development defects41. The only Prefoldin subunit in Drosophila that has been characterized to date, Merry-go-round (Mgr), the Pfdn3 subunit, cooperates with the tumor suppressor Von Hippel Lindau (VHL) to regulate tubulin stability42. However, the functions of Prefoldin in the nervous system remain elusive.
Here, we describe the critical role of evolutionarily-conserved Prefoldin complex in regulating neuroblast and INP asymmetric division and suppressing INP dedifferentiation. Mutants for two Prefoldin subunits, Mgr and Pfdn2, displayed neuroblast overgrowth with defects in cortical polarity of Par proteins and microtubule-related abnormalities. Interestingly, co-depletion of Pins in mgr or pfdn2 mutants led to massive neuroblast overgrowth. Prefoldin and Pins synergistically regulate asymmetric division of both neuroblasts and INPs. Surprisingly, they also synergistically suppress dedifferentiation of INPs back into neuroblasts. Knocking down tubulins in pins mutant background resulted in severe neuroblasts overgrowth, mimicking that caused by co-depletion of Prefoldin and Pins. Our data provide a new mechanism by which Prefoldin and Pins regulates neural stem cell homeostasis through regulating tubulin stability in both neuroblasts and INPs.
Results
Pfdn2 depletion results in the formation of ectopic neuroblasts
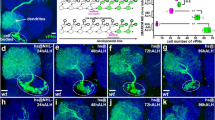
We identified pfdn2/CG6302, encoding a Prefoldin β-like subunit, from a RNA interference (RNAi) screen in larval brains (Zhang Y and Wang H, unpublished data). Ectopic neuroblasts labeled by a neuroblast marker, Deadpan (Dpn), were formed upon knocking down pfdn2 under a neuroblast driver insc-Gal4 (Fig. S1A). Only one neuroblast was observed in control type I neuroblast lineages using insc-Gal4 (Fig. S1B; 100%, n = 40) and type II neuroblast lineages using worniu-Gal4 with asense (ase)-Gal8043 (Fig. 1A; 100%, n = 40). In contrast, upon pfdn2 RNAi excess neuroblasts were observed in both type I neuroblast lineages (Fig. S1B; 53.1%, n = 32) and type II neuroblast lineages (Fig. 1A; 75.0%, n = 32), respectively. To verify the function of Pfdn2 in neuroblasts, we analyzed a putative hypomorphic allele of pfdn2, pfdn201239, which has a P element inserted at the 5′ untranslated region (UTR) of pfdn2. Hemizygous larval brains of pfdn201239 over Df(3L)BSC457 (referred to as pfdn2− thereafter) displayed 235.3 ± 31.7 neuroblasts per brain hemisphere (Fig. 1B–C, n = 25), suggesting that Pfdn2 inhibits the formation of ectopic neuroblasts in larval brains. Consistently, an increase of EdU (5-ethynyl-2′-deoxyuridine)-incorporation was also observed in pfdn2− mutants compared to the control (Fig. S1C). To generate pfdn2 null alleles, we mobilized a P element, EY06124. Its imprecise excision yielded two loss-of-function alleles, pfdn2Δ10 and pfdn2Δ17, both deleting the entire opening reading frame (ORF) of pfdn2 (Fig. 1D). pfdn2Δ10 and pfdn2Δ17 mutants survive to pupal stage and display strong phenotypes with ectopic neuroblasts labeled by Dpn (Fig. 1B–C; 335.0 ± 42.6 neuroblasts/lobe, n = 32 and 301.3 ± 22.7 neuroblasts/lobe, n = 25, respectively). These phenotypes in pfdn2Δ10 and pfdn2Δ17 mutant brains can be fully rescued by overexpression of wild-type pfdn2 or pfdn2-Venus transgene (Fig. S1D–F). Pfdn2 is abundantly expressed in neuroblasts, INPs and their immediate neural progeny- GMCs, detected by a specific antibody generated against Pfdn2 full length (Fig. S1G) and a transgenic Pfdn2 with a Venus tag at the C-terminus (Fig. S1J). In addition, Pfdn2 expression under the tubulin-Gal4 fully rescued the lethality of both pfdn2Δ10 and pfdn2Δ17 mutants. Pfdn2 protein was undetectable in pfdn2Δ10 zygotic mutants (Fig. S1G–H), further supporting that it is a null allele. Both type I and type II MARCM (Mosaic Analysis with Repressible Cell Marker)44 clones of pfdn2Δ10 generated excess neuroblasts (Fig. 1E–F; type I, 41.2%, n = 34; type II, 25.0%, n = 20). These phenotypes were slightly weaker than pfdn2Δ10 zygotic mutants, likely due to residual Pfdn2 protein in the clones (Fig. S1I). These data indicate that Pfdn2 is required in both type I and type II neuroblast lineages to prevent the formation of ectopic neuroblasts.
Pfdn2 suppresses neuroblast overproliferation in larval brains.
(A) Type II driver control (wor-Gal4 ase-Gal80 UAS-CD8-GFP) and pfdn2 RNAi were labeled with Dpn, Ase and CD8. (B) Dpn was labeled in wild-type, pfdn2− [pfdn201239/Df(3L)BSC457], pfdn2Δ10 and pfdn2Δ17 larval brains. The central brain (CB) is to the left of the white dotted line, which markers the border between the CB and the optic lobe (OL). (C) Quantification of larval brain neuroblasts. ***indicates p < 0.001. Error bars indicate mean standard deviation. NBs, neuroblasts. (D) Schematic representation of pfdn2 genomic locus and the pfdn2 deletion alleles used for this study. The extent of the deletion is indicated by the parentheses. (E,F) MARCM driver control and pfdn2Δ10 MARCM clones were labeled with Dpn, Ase and CD8. Neuroblasts (Dpn+ Ase+ in type I lineages and Dpn+ Ase− in type II lineages) in the clones are indicated by arrows. Individual neuroblast clones or lineages are marked by CD8::GFP and outlined by white dotted lines. Scale bars: 5 μm (A,E–F), 20 μm (B).
The Prefoldin complex suppresses the formation of ectopic neuroblasts
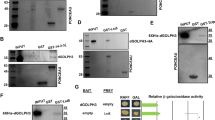
The full chaperone activity of the Prefoldin complex requires all six subunits39. Therefore, we ascertained the potential role of other Prefoldin subunits in neuroblasts. We generated a hemizygous mgr− mutant with mgrG5308, a putative mgr mutant with a P element inserted at the 5′UTR of mgr gene and a deficiency Df(3R)Exel6160 that deletes the entire mgr gene. This mgr− mutant accumulated ectopic neuroblasts in larval brains (Fig. 2A–B; 225.3 ± 25.0 neuroblasts/lobe, n = 35), suggesting that Mgr suppresses the formation of ectopic neuroblasts, similar to Pfdn2. Consistently, mgr RNAi knockdown led to ectopic neuroblasts in both type I (Fig. S2; 67.6%, n = 34) and type II neuroblast lineages (Fig. 2C; 83.3%, n = 30). Furthermore, RNAi knockdown of any of other four Prefoldin genes pfdn1/CG13993, pfdn4/CG10635, pfdn5/CG7048 or pfdn6/CG7770, resulted in ectopic neuroblasts in both type I (Fig. S2; pfdn1 RNAi, 43.2%, n = 37; pfdn4 RNAi, 26.3%, n = 38; pfdn5 RNAi, 38.2%, n = 34; pfdn6 RNAi, 68.4%, n = 38) and type II neuroblast lineages (Fig. 2C; pfdn1 RNAi, 41.7%, n = 36; pfdn4 RNAi, 28.6%, n = 35; pfdn5 RNAi, 46.0%, n = 50; pfdn6 RNAi, 80.0%, n = 40). To ascertain that these Prefoldin subunits form a protein complex as predicted, we carried out co-immunoprecipitation experiments in S2 cells. Indeed, Flag-tagged Pfdn2 physically associates with Myc-tagged Mgr in co-immunoprecipitation experiments (Fig. 2D). Likewise, Myc-tagged Pfdn5 physically interacts with Flag-tagged Pfdn2 (Fig. 2E). These data indicate that the Prefoldin complex is important for inhibition of the formation of ectopic neuroblasts.
Prefoldin is important for neuroblast homeostasis.
(A) Dpn was labeled in wild-type and mgr− [mgrG5308/Df(3R)Exel6160] larval brains. The central brain (CB) is to the left of the white dotted line, which markers the border between the CB and the optic lobe. (B) Quantification of larval brain neuroblasts. ***indicates p < 0.001. Error bars indicate mean standard deviation. NBs, neuroblasts. (C) Type II driver control (wor-Gal4 ase-Gal80 UAS-CD8-GFP), pfdn1 RNAi, mgr RNAi, pfdn4 RNAi, pfdn5 RNAi and pfdn6 RNAi were labeled with Dpn, Ase and CD8. Neuroblasts (Dpn+ Ase- in type II lineages) in the clones are indicated by arrows. Clones are marked by CD8::GFP and outlined by white dotted lines. (D,E) Co-IPs of S2 cells co-expressing Flag-Pfdn2, Myc-Mgr (D) or Flag-Pfdn2, Myc-Pfdn5 (E). Scale bars: 5 μm (C), 20 μm (A).
Mgr and Pfdn2 regulate Par polarity in neuroblasts
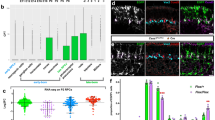
We wondered if Prefoldin is required for asymmetric division of neuroblasts to prevent the formation of ectopic neuroblasts. Indeed, mgr− mutant neuroblasts showed prominent defects of asymmetric division. In wild-type prometa/metaphase neuroblasts, aPKC and Par6 were always asymmetrically localized at the apical cortex while Mira and Pon were localized at the basal cortex (Fig. 3A, 100%, n = 50). Interestingly, aPKC was delocalized in 57.9% of the neuroblasts in mgr mutants during prometa/metaphase (Fig. 3A; n = 126). Similarly, Par6 was delocalized in 34.0% of prometa/metaphase in mgr− neuroblasts (Fig. 3A; n = 106). Basal proteins Mira (Fig. 3A; 10.8%, n = 102) and Pon (Fig. 3A; 10.2%, n = 108) were mildly disrupted in mgr− neuroblasts during prometa/metaphase. In contrast, neither Mgr nor Pfdn2 was important for Pins/Gαi cortical polarity (Fig. S3A–B). In most of previously reported mutants, asymmetric localization defects at prometa/metaphase were mostly restored during telophase by a poorly understood mechanism termed “telophase rescue”45. Surprisingly, all these proteins including aPKC (Fig. 3B; 56.0%, n = 25), Par6 (Fig. 3B; 25.0%, n = 32), Mira (Fig. 3B; 8.3%, n = 36) and Pon (Fig. 3B; 9.1%, n = 22) were still mis-segregated during telophase in mgr mutant neuroblasts, suggesting that telophase rescue did not happen in mgr− mutants. Therefore, Mgr is crucial for cortical polarity of Par proteins and asymmetric protein segregation in neuroblasts.
Pfdn2 and Mgr regulate neuroblast asymmetric division and mitotic spindle assembly.
(A,B) Larval brains of wild-type and mgr− [mgrG5308/Df(3R)Exel6160] hemizygote larvae were labeled with aPKC, Par6, Mira, Pon and DNA. (C) aPKC, Par6, Mira, Pon and DNA were labeled in wild-type and pfdn2− [pfdn201239/Df(3L)BSC457] hemizygote larvae. (D-F) Wild-type (D), pfdn2Δ10 (E) and pfdn2Δ10 mgrG5308 (F) larval brains were labeled with CNN, α-tubulin and DNA. Astral microtubules are indicated by white arrows and neuroblasts are outlined by white dotted lines. (G) Quantification of centrosome numbers. (H–K) Western blot of α-tubulin, β-tubulin and actin. Protein extracts from wild-type, pfdn2Δ10, mgr− [mgrG5308/Df(3R)Exel6160] hemizygote and pfdn2Δ10 mgrG5308 larval brains were probed by anti-α-tubulin, anti-β-tubulin and anti-actin. GAPDH is loading control. The expression levels of α-tubulin (I), β-tubulin (J) and actin (K) were quantified. ***indicates p < 0.001, *indicates p < 0.05, ns indicates p > 0.05. Error bars indicate mean standard deviation. Scale bars: 5 μm (A,D–F), 4 μm (B–C).
pfdn2 mutants also displayed defects of asymmetric protein localization of aPKC (Fig. S3C; pfdn2Δ10, 26.5%, n = 49; pfdn2−, 19.3%, n = 119) and Par6 (Fig. S3C; pfdn2Δ10, 15.2%, n = 46; pfdn2−, 13.7%, n = 95) during prometa/metaphase, albeit with weaker phenotypes. Interestingly, during telophase, mis-segregation of these proteins were evident in pfdn2− mutants (Fig. 3C; aPKC, 11.9%, n = 109; Par6, 7.5%, n = 107; Mira, 7.5%, n = 107; Pon, 10.2%, n = 118), suggesting that Pfdn2 is also important for asymmetric protein segregation at telophase. The defects in asymmetric protein segregation observed in pfdn2− and mgr− mutants were likely responsible for the neuroblast overgrowth.
In wild-type metaphase neuroblasts, the mitotic spindle marked by α-tubulin is aligned with the apicobasal axis (inferred by apical Baz) (Fig. S3D; 100%, n = 50), which is important for asymmetric protein segregation at telophase. In both mgr− (Fig. S3D; 23.7%, n = 76) and pfdn2Δ10 (Fig. S3D; 20.0%, n = 85) mutant neuroblasts, mitotic spindles were mis-oriented with apicobasal axis in cells that still retained asymmetric Baz localization. Taken together, both Mgr and Pfdn2 are required for Par polarity and mitotic spindle orientation in neuroblasts.
Pfdn2 is critical for centrosomal functions
Prefoldin is important for the assembly of αβ-tubulin dimers37,46 and mgr mutant cells have microtubule-based abnormalities42. Similarly, we observed various microtubule-associated abnormalities in pfdn2Δ10 mutant neuroblasts. Different from wild-type neuroblasts that always formed two centrosomes marked by Centrosomin (CNN) during mitosis (Fig. 3D,G; n = 60), in pfdn2Δ10 mutant, 20.5% of neuroblasts contained multiple centrosomes and 6.0% of neuroblasts contained only one centrosome and a monopolar spindle (Fig. 3E,G; n = 83). In addition, 54.2% of pfdn2Δ10 neuroblasts failed to assemble astral microtubules during metaphase (Fig. 3E; n = 83). These microtubule abnormalities in pfdn2Δ10 mutants were exacerbated by the presence of mgr mutation. In a pfdn2Δ10 mgrG5308 double mutant, 43.6% of neuroblasts had only one or multiple centrosome during mitosis (Fig. 3F,G; n = 78). In addition, vast majority of neuroblasts lacked obvious astral microtubules in the pfdn2Δ10 mgrG5308 double mutant (Fig. 3F; 89.7%, n = 78). It was shown previously that Mgr depletion led to reduction of both α-tubulin and β-tubulin levels42. Similarly, levels of both α-tubulin (Fig. 3H–I; n = 5) and β-tubulin (Fig. 3H,J; n = 5) were dramatically decreased in pfdn2Δ10 larval brains. They were further diminished to 10% and 26% of the wild-type levels in the pfdn2Δ10 mgrG5308 double mutant, respectively (Fig. 3H–J). In contrast, the levels of actin were not dramatically affected in pfdn2Δ10, mgr- single or pfdn2Δ10 mgrG5308 double mutants (Fig. 3H,K; n = 5). These data suggest that tubulin but not actin level is critically dependent on Prefoldin in the larval brain.
To examine microtubule growth, we depolymerized microtubules with cold treatment in pfdn2 mutants and monitored microtubule regrowth at 25 °C (Fig. S3E). In wild-type larval brains, we observed that prominent microtubule asters were formed 30 s after the recovery at 25 °C (Fig. S3E; 100%, n = 30) and a normal bipolar spindle was assembled at 120 s in all wild-type neuroblasts (Fig. S3E; 100%, n = 25). In contrast, pfdn2Δ10 mutant neuroblasts displayed much weaker asters after 30 s of recovery (Fig. S3E; 94.1%, n = 34) and only short and disorganized spindles were formed after 120 s of recovery (Fig. S3E; 100%, n = 53). These data suggest that Pfdn2 likely contributes to the microtubule growth through regulating tubulin levels.
Co-depletion of Pins and Prefoldin leads to massive neuroblast overgrowth
Given that depletion of Pfdn2 or Mgr only resulted in partial loss of Par polarity with intact Pins polarity, we wondered whether Prefoldin functions redundantly with Pins in regulating asymmetric division. Remarkably, simultaneous depletion of Pins and Pfdn2 resulted in severe neuroblast overgrowth in larval brains (Fig. 4A–B). Larval brains that were double mutant for pfdn2Δ10 pinsp89 formed 908.6 ± 127.1 neuroblasts per brain hemisphere (Fig. 4A–B; n = 35), while pfdn2Δ10 larval brains had 330.1 ± 43.9 neuroblasts per brain hemisphere (Fig. 4A–B; n = 35). In contrast, pinsp89 larval brains developed 43.8 ± 13.4 neuroblasts per brain hemisphere (Fig. 4A–B; n = 25), similar as the previous report7,47. Most of these ectopic neuroblasts were Dpn+ and Ase− (Fig. 4A) or Dpn+ PntP1+ (Fig. S4A), suggesting that type II neuroblasts are predominant in the population of ectopic neuroblasts. Interestingly, the population of immature INPs which were Dpn− PntP1+ appeared to be increased, too (Fig. S4A).
Co-depletion of Prefoldin and Pins results in supernumerary neuroblasts in larval brains.
(A) Dpn and Ase were labeled in wild-type, pfdn2Δ10, pinsp89 and pfdn2Δ10 pinsp89 larval brains. (B) Quantification of larval brain neuroblasts. (C) Dpn and Ase were labeled in wild-type, mgrG5308, pinsp89, mgrG5308 pinsp89 larval brains. (D) Quantification of larval brain neuroblasts. The central brain (CB) is to the left of the white dotted line, which markers the border between the central brain and the optic lobe. ***indicates p < 0.001. Error bars indicate mean standard deviation. Scale bar: 20 μm.
We next examined whether Mgr had a similar synergism with Pins in suppressing neuroblast overgrowth. Indeed, mgrG5308 pinsp89 larval brains formed 635.7 ± 238.0 neuroblasts per brain hemisphere (Fig. 4C–D; n = 30), while mgrG5308 homozygous larval brains had 190.0 ± 19.6 neuroblasts per brain lobe (Fig. 4C–D; n = 20). This data strongly suggest the synergism between Pins and Prefoldin in suppressing neuroblast overgrowth.
In contrast, co-depletion of Pfdn2 and aPKC or Par6 suppressed uncontrolled neuroblast proliferation in larval brains. pfdn2Δ10/01239 larval brains formed 283.7 ± 32.9 neuroblasts per brain hemisphere (Fig. S4B–C; n = 16), while knocking down aPKC under the insc-Gal4 formed 89.5 ± 9.0 neuroblasts per brain hemisphere (Fig. S4B–C; n = 20). However, aPKC RNAi knockdown partially suppressed neuroblast overgrowth in pfdn2Δ10/01239 larval brains with 174.8 ± 16.9 neuroblasts per brain hemisphere (Fig. S4B–C; n = 22). Likewise, RNAi-mediated knockdown of par6 in pfdn2Δ10/01239 larval brains resulted in 127.0 ± 22.8 neuroblasts per brain hemisphere (Fig. S4D–E; n = 18), compared to par6 RNAi knockdown alone (Fig. S4D–E; 83.2 ± 8.2 neuroblasts/lobe, n = 14). These data suggest that the synergism between Prefoldin and Pins in suppressing neuroblast overgrowth is specific.
Depletion of both Pins and Prefoldin results in symmetric division of neuroblasts
Given the synergism observed for Prefoldin and Pins in suppressing neuroblast overgrowth, we examined whether pfdn2 pins double mutants had more severe asymmetric division defects than pfdn2 or pins mutant alone. Indeed, asymmetric localization of aPKC and Mira was completely disrupted in pfdn2Δ10 pinsp89 neuroblasts at metaphase (Fig. 5A; 100%, n = 50). Moreover, all neuroblasts in pfdn2Δ10 pinsp89larval brains underwent equal-size division and segregated Mira into both daughter cells at telophase (Fig. 5B; 100%, n = 30), suggesting that neuroblast symmetric division indeed contributed to the dramatic neuroblast overgrowth. Very interestingly, co-depletion of Pfdn2 and Pins also led to symmetric division of Dpn+ Ase+ mature INPs. All INPs from wild-type (Fig. 5C; 100%, n = 15), pinsp89 (Fig. 5C; 100%, n = 12) or pfdn2Δ10 (Fig. 5C; 100%, n = 20) larval brains divided asymmetrically with Mira segregated into one of the daughter cells. However, all INPs in pfdn2Δ10 pinsp89 larval brains divided symmetrically with Mira segregated into both daughter cells (Fig. 5C; 100%, n = 12). These observations suggest that Pfdn2 and Pins function synergistically to regulate asymmetric division of both neuroblasts and INPs.
Prefoldin and Pins interact genetically to regulate asymmetric division of neuroblasts and INPs.
(A) aPKC, Mira and DNA were labeled in wild-type, pfdn2Δ10, pinsp89 and pfdn2Δ10 pinsp89 larval brains. (B–C) Dpn, Ase, Mira and DNA were labeled in wild-type, pfdn2Δ10, pinsp89 and pfdn2Δ10 pinsp89 larval brains. Open white dotted circles indicate INP daughters negative for Mira. (D) Wild-type, pfdn2Δ10, pinsp89 and pfdn2Δ10 pinsp89 larval brains were labeled with CNN, α-tubulin and DNA. Astral microtubules are indicated by white arrows and neuroblasts are outlined by white dotted lines. Scale bar: 5 μm.
Interestingly, the α-tubulin levels were further decreased in pfdn2Δ10 pinsp89 larval brains, compared to either pfdn2 or pins single mutant (Fig. S5A–B) suggesting that Pins may also have an unexpected function in regulating tubulin levels. To further probe the function of Pins in microtubules, we examined interphase microtubule aster in pfdn2 pins double mutant neuroblasts and compared them to pfdn2 or pins single mutant. In 23.7% of the pinsp89 neuroblasts, interphase microtubule aster was lost or strongly reduced (Fig. 5D, n = 38) and 66.7% of pfdn2Δ10 interphase neuroblasts failed to form interphase microtubule aster (Fig. 5D, n = 66). Strikingly, 96.6% of pfdn2Δ10 pinsp89 neuroblasts failed to form any interphase aster (Fig. 5D; n = 89). Thus, Pfdn2 and Pins function synergistically to regulate microtubule functions, control asymmetric cell division and to suppress the formation of ectopic neuroblasts.
Depletion of Pfdn2 or Mgr in pins mutants induces dedifferentiation of INPs back into neuroblasts
To explore whether Prefoldin is important for INP function, we knocked down pfdn2 or mgr in pins mutant background under an INP-specific driver erm-Gal4 (II), which is expressed in Ase− immature INPs. A wild-type INP clone consists of immature INPs (Ase− or Ase+), mature INPs (Dpn+ Ase+), their immediate progeny GMCs (Ase+) and neurons (Fig. 6A; 100%, n = 50). RNAi-mediated knockdown of pfdn2 (Fig. 6A; 100%, n = 50) under erm-Gal4 resembled wild-type INP clones. Surprisingly, pfdn2 RNAi knockdown in pinsp89 mutants resulted in numerous aberrant Dpn+ Ase- type II neuroblasts (Fig. 6A; 43.5%, n = 46), while INP clones in pinsp89 mutants alone did not contain any type II neuroblasts (Fig. 6A; 100%, n = 30). This observation suggests that upon depletion both Pfdn2 and Pins, INPs undergo dedifferentiation back into neuroblasts. Consistently, pinsp89 mutant with mgr RNAi knockdown resulted in similar dedifferentiation phenotype (Fig. 6A; 48.1%, n = 52; S6; 41.7%, n = 24), while mgr knockdown alone in INPs did not cause any phenotype (Fig. 6A; n = 42; S6; n = 25). These data suggest that co-depletion of Prefoldin and Pins induces dedifferentiation of INPs back into neuroblasts. The severe brain overgrowth observed in pfdn2Δ10 pinsp89 larval brains (Fig. 6B–C; 958.7 ± 107.0 neuroblasts/lobe, n = 15) can be partially reversed by overexpressing a wild-type pins under the INP specific driver erm-Gal4 (Fig. 6B–C; 302.7 ± 148.9 neuroblasts/lobe, n = 16), suggesting that Pins indeed functions to suppress INP dedifferentiation.
Knockdown of pfdn2 or mgr in pins mutant brains results in dedifferentiation of INPs back into neuroblasts.
(A) Driver control (erm-Gal4 UAS-CD8-GFP), pinsp89, pfdn2 RNAi UAS-Dicer2, pfdn2 RNAi UAS-Dicer2; pinsp89, mgr RNAi UAS-Dicer2 and mgr RNAi UAS-Dicer2; pinsp89 larval brains were labeled with Dpn, Ase and CD8. (B) pfdn2Δ10 pinsp89 and UAS-Pins pfdn2Δ10 pinsp89 under erm-Gal4 were labeled with Dpn and Ase. The central brain (CB) is to the left of the white dotted line, which markers the border between the CB and the optic lobe. (C) Quantification of larval brain neuroblasts. ***indicates p < 0.001. Error bars indicate mean standard deviation. Neuroblasts (NBs, Dpn+ Ase− in type II lineages) in the clones are indicated by arrows. Clones are marked by CD8::GFP and outlined by white dotted lines. Scale bars: 5 μm (A), 20 μm (B).
Knockdown of either α-tubulin or β-tubulin in pins mutants leads to massive neuroblast overgrowth
Given that Prefoldin and Pins synergistically regulate tubulin level in larval brains, we determined whether a decrease of microtubule function in pins mutant background could mimic the pins pfdn2 double mutants. First, we knocked down various α-tubulin and β-tubulin in neuroblasts using insc-Gal4. α-tubulin RNAi knockdown resulted in ectopic neuroblasts in both type I lineages (Fig. S7A; α-tub RNAi(I)/α-tub67C/CG8308, 45.5%, n = 66 and α-tub RNAi(II)/α-tub84B/CG1913, 53.3%, n = 60) and type II lineages (Fig. 7A; α-tub RNAi(I), 47.3%, n = 55 and α-tub RNAi(II), 51.7%, n = 60). Likewise, β-tubulin RNAi knockdown led to generation of ectopic neuroblasts in both type I neuroblast lineages (Fig. S7A; β-tub RNAi(I)/β-tub60D/CG3401, 47.5%, n = 40 and β-tub RNAi(II)/β-tub56D/CG9277, 52.1%, n = 48) and type II neuroblast lineages (Fig. 7A; β-tub RNAi(I), 52.5%, n = 40 and β-tub RNAi(II), 54.2%, n = 48). Next, we knocked down α-tubulin in pinsp89 mutant background. Surprisingly, this led to enormous overgrowth in neuroblast lineages (Fig. 7A; α-tub RNAi(I) pinsp89, 88.0%, n = 50; α-tub RNAi(II) pinsp89, 60.3%, n = 58), mimicking the phenotypes in pins pfdn2 double mutant. Consistently, β-tubulin knockdown in pinsp89 mutants also caused dramatic neuroblast overgrowth (Fig. 7A; β-tub RNAi(I) pinsp89, 84.0%, n = 50; β-tub RNAi(II) pinsp89, 68.3%, n = 41). Both α-tubulin and β-tubulin knockdown in these experiments were evident, as tubulin levels were dramatically reduced in these clones (Fig. S7B–E). Next, we examined whether this massive neuroblast overproliferation are attributed by asymmetric division defects. In wild-type prometa/metaphase neuroblasts, aPKC formed an apical crescent, while Mira accumulated at the basal cortex (Fig. S7F; 100%, n = 50). This asymmetric localization was also observed in most of the neuroblasts with α-tubulin knockdown alone (Fig. S7F; 98.4%, n = 63). However, pinsp89 mutant with α-tubulin RNAi further exacerbated delocalization of both aPKC (Fig. S7F; 96.7%, n = 30) and Mira (Fig. S7F; 86.7%, n = 30) compare to pinsp89 mutants alone (Fig. S7F; aPKC, 92.0%, n = 25; Mira, 65.2%, n = 23). In wild-type neuroblasts, apical aPKC and basal Mira were segregated exclusively into one of the daughter cells at telophase (Fig. 7B; 100%, n = 25). Asymmetric segregation of aPKC and Mira was also observed in pinsp89 mutant (Fig. 7B; 100%, n = 15) and α-tubulin RNAi knockdown neuroblasts (Fig. 7B; 97.7%, n = 44), presumably through “telophase rescue”. However, mis-segregation of both aPKC and Mira at telophase occurred in most of pinsp89 mutant neuroblasts with α-tubulin RNAi knockdown (Fig. 7B; 73.1%, n = 26). These observations suggest that co-depletion of tubulin and Pins results in severe defects of asymmetric division at both metaphase and telophase, which contributes to the severe overgrowth phenotype.
Reduction of tubulin levels in pins mutant brains results in neuroblast overgrowth and asymmetric division defects.
(A) Driver control (insc-Gal4 UAS-CD8-GFP), pinsp89, α-tub RNAi(I), α-tub RNAi(II), α-tub RNAi(I); pinsp89, α-tub RNAi(II); pinsp89, β-tub RNAi(I), β-tub RNAi(II), β-tub RNAi(I); pinsp89 and β-tub RNAi(II); pinsp89 were labeled with Dpn, Ase and CD8. (B) Driver control (insc-Gal4), pinsp89, α-tub RNAi(I) and α-tub RNAi(I); pinsp89 larval brains were labeled with aPKC, Mira and DNA. Type II neuroblasts (Dpn+ Ase-) in the clones are indicated by arrows. Clones are marked by CD8::GFP and outlined by white dotted lines. Scale bar: 5 μm.
Knockdown of either α-tubulin or β-tubulin in pins mutants leads to dedifferentiation of INPs back into neuroblasts
Next, we wondered whether tubulins also play a role in INPs to prevent dedifferentiation. Toward this end, we used erm-Gal4 to knock down α- or β-tubulin in INPs of pins mutants. Control brains formed 100.2 ± 8.2 neuroblasts per brain hemisphere (Fig. 8A, S8A; n = 15) and knockdown of α-tubulin under the erm-Gal4 gave rise to 110.3 ± 9.8 neuroblasts per brain hemisphere (Fig. 8A, S8A; n = 20), which was similar to that of the control brains. Remarkably, knockdown of α-tubulin under the control of erm-Gal4 in pinsp89 mutants caused severe neuroblast overproliferation (Fig. 8A, S8A; 438.7 ± 320.1 neuroblasts/lobe, n = 21), in contrast to pinsp89 larval brains that formed fewer neuroblasts (Fig. 8A, S8A; 45.5 ± 13.2 neuroblasts/lobe, n = 20). These data suggest that co-depletion of α-tubulin and Pins specifically in INPs results in ectopic neuroblasts due to INP dedifferentiation. Likewise, we observed the similar massive neuroblast overgrowth phenotype in pinsp89 larval brains with β-tubulin knockdown (Fig. 8A, S8A; 520.3 ± 405.2 neuroblasts/lobe, n = 18), while β-tubulin knockdown under erm-Gal4 alone did not result in obvious neuroblast overgrowth (Fig. 8A, S8A; 115.5 ± 10.6 neuroblasts/lobe, n = 25). Next, we examined single INP clones in these mutants. None of the INP clones in control (Fig. 8B; 100%, n = 30), pinsp89(Fig. 8B; 100%, n = 25), α-tubulin RNAi knockdown (Fig. 8B; 100%, n = 35), or β-tubulin RNAi knockdown (Fig. 8B; 100%, n = 35) larval brains contained any Dpn+ Ase- type II neuroblast. However, pinsp89 larval brains with α-tubulin knockdown produced ectopic type II neuroblasts in 85.4% of the INP clones (Fig. 8B; n = 41). Similarly, β-tubulin knockdown in pinsp89mutants displayed excess neuroblasts in 71.2% of the INP clones (Fig. 8B; n = 59). These data suggest that tubulins and Pins function synergistically in suppressing dedifferentiation of INPs back into neural stem cells.
Reduction of tubulin levels in pins mutant brains results in dedifferentiation of INPs back into neuroblasts.
(A) Dpn and Ase were labeled in driver control (erm-Gal4 UAS-Dicer2), pinsp89, α-tub RNAi(II), β-tub RNAi(II), α-tub RNAi(II); pinsp89 and β-tub RNAi(II); pinsp89 larval brains. The central brain (CB) is to the left of the white dotted line, which markers the border between the CB and the optic lobe. (B) Driver control (erm-Gal4 UAS-CD8-GFP), pinsp89, α-tub RNAi(II), β-tub RNAi(II), α-tub RNAi(II); pinsp89 and β-tub RNAi(II); pinsp89 larval brains were labeled with Dpn, Ase and CD8. Type II neuroblasts (Dpn+ Ase-) in the clones are indicated by arrows. Clones are marked by CD8::GFP and outlined by white dotted lines. (C) Driver control (erm-Gal4 UAS-Dicer2), pinsp89, β-tub RNAi(II) and β-tub RNAi(II); pinsp89 larval brains were labeled with Dpn, Ase, Mira and DNA. Open white dotted circles indicate INP daughters negative for Mira. Scale bars: 20 μm (A), 5 μm (B,C).
Knockdown of tubulin in pins mutant results in symmetric division of INPs
To assess whether tubulin knockdown in pins mutant affects asymmetric division of INPs, we examined the localization of Mira, which is asymmetrically localized in wild-type INPs at prometa/metaphase (Fig. S8B; 100%, n = 30). β-tubulin knockdown alone under erm-Gal4 did not influence the asymmetric localization of Mira (Fig. S8B; 100%, n = 26). In pinsp89 mutants, Mira was delocalized in 87.1% of the INPs (Fig. S8B; n = 31) and this phenotype was further enhanced with β-tubulin knockdown in INPs (Fig. S8B; 93.8%, n = 32). Next, we examined whether co-depletion of tubulin and Pins caused asymmetric division defects of INPs at telophase. Similar to wild-type INPs (Fig. 8C; 100%, n = 20), all INPs from pinsp89 (Fig. 8C; 100%, n = 10) and INP-specific knockdown of β-tubulin (Fig. 8C; 100%, n = 15) brains divided asymmetrically with Mira exclusively segregated to one of the daughter cells. However, 90.0% of the INPs in pinsp89 larval brains with β-tubulin knockdown in INPs divided symmetrically and mis-segregated Mira into both daughter cells (Fig. 8C; n = 20). These observations suggest that tubulins and Pins function synergistically to regulate asymmetric division of INPs.
Overexpression of α-tubulin suppresses various defects associated with prefoldin pins double mutants
Given that defects observed in pfdn2 pins double mutants can be phenocopied by co-depleting tubulin and Pins, we investigated whether restoring tubulin levels in pfdn2 pins double mutants is sufficient to prevent neuroblast overgrowth. Overexpression of GFP-α-tubulin under the ubiquitin promoter did not lead to any obvious changes to neuroblast number (Fig. S9A–C). In contrast to 866.1 ± 138.6 neuroblasts per brain hemisphere in pfdn2Δ10 pinsp89 mutant brains (Fig. S9A–B; n = 20), there were only 123.2 ± 49.4 neuroblasts per brain hemisphere in pfdn2Δ10 pinsp89double mutant brains expressing GFP-α-tubulin (Fig. S9A–B; n = 23). This observation suggests that the overexpression of GFP-α-tubulin dramatically suppresses the unrestrained neuroblast proliferation in pfdn2Δ10 pinsp89 larval brains. We ascertained whether this suppression was partially contributed by the restoration of asymmetric division. Indeed, the polarity of Baz (Fig. S9D; 33.3%, n = 21), aPKC (Fig. S9D; 18.8%, n = 32), Numb (Fig. S9D; 37.5%, n = 40) and Mira (Fig. S9D; 43.8%, n = 32) in prometa/metaphase neuroblasts were partially restored. Moreover, asymmetric segregation of aPKC and Mira was rescued in 66.7% of the neuroblasts at telophase with GFP-α-tubulin expression (Fig. S9E; n = 24), in contrast to a complete disruption in pfdn2Δ10 pinsp89 mutant brains (Fig. S9E; 100%, n = 20). These data suggest that Pfdn2 and Pins suppress neuroblast overgrowth through regulating tubulins levels.
Discussion
Here we have identified an unexpected synergism between Prefoldin and Pins in suppressing neuroblasts overgrowth (Fig. S9F). We show that various subunits of Prefoldin complex are implicated in asymmetric division of neuroblasts, especially during asymmetric protein segregation at telophase. It is known that depletion of Pins results in the formation of smaller larval brains, despite partial loss of neuroblasts polarity7,47. Interestingly, co-depletion of Pfdn2 and Pins results in severe neuroblasts overgrowth, while Pfdn2 depletion alone only causes mild brain overgrowth. This phenotype is contributed by a combination of loss of neuroblast polarity, defects of asymmetric division of INPs, as well as INP dedifferentiation. Knocking down tubulins in pins mutant background mimics the co-depletion of Prefoldin and Pins, suggesting that tubulin stability appears to be critical for the suppression of neuroblast overgrowth in the absence of Pins function. Our data also suggest that Prefoldin function and tubulin stability in INPs are important to suppress their dedifferentiation back into neuroblasts.
How microtubules induce cortical polarity is poorly understood in Drosophila neuroblasts. Previously, one report showed that kinesin Khc-73, which localized at the plus end of astral microtubules and Discs large (Dlg) induced cortical polarization of Pins/Gαi in neuroblasts48. However, microtubules are considered not essential for neuroblast polarity49,50. Here, we show that Drosophila Prefoldin regulates asymmetric division of both neuroblasts and INPs through tubulins, suggesting an important role of microtubules in neuroblast polarity. The essential role of microtubules directly regulating cell polarity is found in various systems. During C. elegans meiosis, a microtubule-organizing center is necessary and sufficient for the establishment of the anterior-posterior polarity51. In the fission yeast Schizosaccharomyces pombe, interphase microtubules directly regulate cell polarity through proteins such as tea1p52. In mammalian airway cilia, microtubules are required for asymmetric localization of planer cell polarity proteins53.
We show that the role of Drosophila Prefoldin complex in regulating asymmetric division is very likely dependent on microtubules. This is consistent with the known essential role of Prefoldin for maintaining tubulin levels in various organisms such as yeast, C. elegans, plants and mammals37,40,54,55. In yeast, Gim (Prefoldin) null mutants become super-sensitive to the microtubule-depolymerizing drug benomyl as a result of a reduced level of α-tubulin37. In the absence of Prefoldin, the function of the chaperone pathway is damaged and unable to fold sufficient amount of tubulins for normal yeast growth36. In C. elegans, reducing Prefoldin function causes defects in cell division presumably due to the reduction of tubulin levels and microtubule growth rate40. Genetic analysis of mammalian Prefoldin also suggests that cytoskeletal proteins like actin and tubulin make up the major substrate of Prefoldin in mammals41. These studies in different organisms together suggest that Prefoldin complex plays a conserved central role in tubulin folding.
“Telophase rescue”, a term refers to the phenomenon that protein mis-localization at metaphase is completely restored at telophase, is observed in many mutants that affect neuroblast asymmetric division45. However, both apical and basal proteins are still mis-segregated in pfdn2 and mgr mutants, suggesting that “telophase rescue” is defective in these mutants. Telophase rescue is regulated by TNF receptor-associated factor (DTRAF1), which binds to Baz and acts downstream of Egr/TNF56. Telophase rescue also depends on Worniu/Escargot/Snail family proteins and a microtubule-dependent Khc-73/Dlg pathway48,57. Pins did not form a protein complex with Mgr, α-tubulin or β-tubulin in co-immunoprecipitation assays (Fig. S10). Given that Dlg is a Pins-interacting protein58, Prefoldin appears to function in a different pathway with Dlg or Khc-73 during asymmetric division.
Recently, merry-go-round (mgr), encoding Prefoldin 3 (Pfdn3)/VBP1/Gim2 subunit, was reported to regulate spindle assembly42. Loss of mgr led to formation of monopolar mitotic spindles and loss of centrosomes because of improper folding and destabilization of tubulins42. Our analysis on Pfdn2 indicates that pfdn2 mutants displayed similar spindle and centrosome abnormalities. In addition, the incorrectly folded tubulin due to loss of mgr may be eliminated by Drosophila von Hippel Lindau protein (Vhl), an E3 ubiquitin-protein ligase42. Interestingly, our data suggest that Prefoldin has a tumor-suppressor like function in preventing neuroblast overgrowth. However, Drosophila Vhl is not important for brain tumor suppression, as its loss-of-function neither affects number of neuroblasts nor suppresses overgrowth observed in pfdn2 RNAi or mgr RNAi (data not shown).
We show a novel synergism between Prefoldin and Pins in suppressing dedifferentiation of INPs back into neuroblasts. Prefoldin and Pins apparently suppress dedifferentiation through regulating tubulin levels. It is likely that appropriate tubulin levels in INPs are important for their differentiation, while reducing tubulin levels can increase the risk of INP dedifferentiation. Currently, several cell fate determinants such as Brat, Numb and the SWI/SNF chromatin remodeling complex with its cofactors Erm and Hdac3 are critical to suppress INP dedifferentiation back into neuroblast29,32,33,34,35. It is currently unknown whether or how Prefoldin/Pins are linked to these known suppressors of dedifferentiation. It is possible that symmetric division of INPs causes reduced levels of Brat and Numb in these abnormal INP daughters, leading to their dedifferentiation. Alternatively, Prefoldin might regulate transcription of genes within INPs to suppress dedifferentiation. It was reported that the human homolog of Pfdn5, MM-1, has a role in transcriptional regulation by binding to the E-box domain of c-Myc and represses E-box-dependent transcriptional activity59. Interestingly, Prefoldin Subunit 5 gene is deleted in Canine mammary tumors, suggesting that it may be a tumor suppressor gene60. Our study has revealed a novel mechanism by which Prefoldin and Pins function through tubulin stability to suppress stem cell overgrowth. It is expected to contribute to the understanding of mammalian/human Prefoldin function in tumorigenesis.
Materials and Methods
Fly stocks and genetics
The fly strains used in this paper were: pfdn2Δ10, pfdn2Δ17, UAS-Pfdn2, UAS-Pfdn2-Venus, pinsp89(F. Yu), type II neuroblast driver (w; UAS-Dicer2, wor-Gal4, ase-Gal80/CyO; UAS-mCD8-GFP/TM3, Ser; J. Knoblich), neuroblast driver (insc-Gal4; J. Knoblich), INP driver (erm-Gal4/CyO; GM Rubin), pUbiquitin-α-tub-GFP (Gonzalez, C.). The following stocks were obtained from Bloomington Drosophila Stock Center (BDSC): pfdn201239(BDSC#11526), mgrG5308 (BDSC#30151), Df(3L)BSC457 (BDSC#24961), Df(3R)Exel6160 (BDSC#7639). The following RNAi stocks were obtained from Vienna Drosophila Resource Center (VDRC): pfdn2 RNAi (v28794/CG6302), pfdn1 RNAi (v18210/CG13993), mgr RNAi (v27727/CG6719), pfdn4 RNAi (v46220/CG10635), pfdn5 RNAi (v29812/CG7048) pfdn6 RNAi (v34204/CG7770), α-tubulin RNAi(I) (v24783/α-tub67C/CG8308), α-tubulin RNAi(II) (v52345/α-tub84B/CG1913), β-tubulin RNAi(I) (v104937/β-tub60D/CG3401), β-tubulin RNAi(II) (v24138/β-tub56D/CG9277), par6 RNAi (v19732/CG5884) and aPKC RNAi (v2907/CG42783).
Generation of pfdn2Δ10 and pfdn2Δ17
pfdn2Δ10 and pfdn2Δ17were generated by imprecise excision of P{EPgy2}l(3)01239EY06124 (BDSC#19918), a P element inserted at the 11th base pair from the pfdn2 transcription start site. Deletions were verified by PCR amplification and sequencing. Oligos used were: 5′-TGTGCAAGGCTGTTTCTCAC-3′ (forward) and 5′-TTATGTTTAAGTAACTGAAGTTGTGCT-3′ (reverse).
Molecular cloning
The full-length cDNAs of pfdn2, mgr and pfdn5 were amplified from cDNA clones obtained from Drosophila Genomics Resources Center (DGRC) and sub-cloned into Gateway® pENTRTM vector (pENTR™⁄D-TOPO® Cloning Kit, Invitrogen). Myc or Flag tags were added into the N-terminus of the gene sequences by LR recombination reactions (Gateway® LR Clonase® II Enzyme mix, Invitrogen) using pAMW or pAFW destination vectors respectively. The oligos that were used to amplify various DNA fragments were listed in Table 1.
Transgenic flies
To generate pUAS-Pfdn2-Venus or pUAS-Pfdn2 construct, the full-length coding region of Pfdn2 was tagged with (or without) Venus at the C-terminus by the LR recombination between pENTRY pfdn2 and pTWV (or pTW) vector. Briefly, using Gateway LR Clonase II enzyme mix, the full-length coding gene of Pfdn2 from the entry construct was transferred into the Gateway destination vector pTWV (or pTW). UAS-Pfdn2-Venus or UAS-Pfdn2 transgenic flies were generated by standard P element-mediated transformation of pUAS-Pfdn2-Venus or pUAS-Pfdn2 by BestGenes, Inc. Oligos used for sequencing were: 5′-TATAAATAGAGGCGCTTCGT-3′ (forward) and 5′-CTTCGGGCATGGCGGACTTG-3′ (reverse).
Clonal analysis
FRT2A pfdn2Δ10/TM6B and elav-Gal4 hsFlpase; UAS-nLacZ UAS-CD8:: GFP/CyO;FRT2A tubP-Gal80 (MARCM driver) flies were used to generate pfdn2 mutant clones. Larvae were heat-shocked twice at 37 °C for 2 hrs, at 24 h ALH and 40 h ALH respectively and were allowed to develop for another 3 days at 25 °C. RNAi knockdown was carried out in 29 °C for 4 days after egg-laying.
Immunohistochemisry
Larval brain dissection and immunostaining were carried out essentially as described previously25. Briefly, larval brains were dissected and fixed in 3.7% formaldehyde in PBS with 0.3% Triton X-100. Fixed brains were washed for three times (10 min each) before one hour-blocking with 3% BSA. Larval brains were then incubated with primary antibodies over night at 4 °C. Secondary antibodies were then added into the samples followed by three times washing with PBT. After incubated with ToPro-3 to stain DNA for 20 min, larval brains were mounted in Vectorshield (Vector Laboratory). Images were obtained using Zeiss LSM 710 confocal microscope and processed with Adobe Photoshop CS5.
Western blotting
Third-instar larval brains were dissected in PBS and homogenized in RIPA buffer (50 mM Tris HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.5% sodium deoxycholate, 0.1% SDS). Western blotting was carried out according to standard procedures.
Antibodies
The following primary antibodies were used: guinea pig anti-Dpn [J. Skeath, immunofluorescence (IF) 1:1000], rabbit anti-Ase (YN Jan, IF 1:500), mouse anti-Mira (F. Matsuzaki, IF 1:80), rat anti-CD8 (Life technologies, IF 1:200), rabbit anti-aPKCζ C20 (Santa Cruz Biotechnologies, IF 1:200, western blot (WB) 1:3000), guinea pig anti-Bazooka (F. Yu, IF 1:500, WB 1:1000), guinea pig anti-Gαi (F. Yu, IF 1:200), rabbit anti-Par6 (J. Knoblich, IF 1:200, WB 1:1000), rabbit anti-Pon (Y.N. Jan, IF 1:200), rabbit anti-Pins (F. Yu, IF 1:200, WB 1:3000), mouse anti-α-tubulin (Sigma, IF 1:100, WB 1:5000), mouse anti-β-tubulin (DSHB, IF 1:50, WB 1:25), rabbit anti-CNN (E. Schejter, IF 1:1000), rabbit anti-PntP1 (J. Skeath, IF 1:500), mouse anti-Actin (MP Biomedicals, WB 1:5000), rabbit anti-Pfdn2 (this study, IF 1:500, WB 1:1000), mouse anti-Myc (Abcam, WB 1:3000), mouse anti-Flag (Sigma, WB 1:3000).
Generation of anti-Pfdn2 antibody
The cDNA region encoding the full length of Pfdn2 was amplified by PCR and subsequently cloned into EcoRI and SalI sites of PAML-C2-X vector, using the In-Fusion HD cloning Kit (Clontech). The primers used were: 5′-AGGATTTCAGAATTCATGAGCACCGAATCGGCGAAG-3′ (forward), 5′-TTGCCTGCAGGTCGATCAGTTGAACACCAGGACATT-3′ (reverse). The expression of MBP-Pfdn2 was induced by isopropyl β-D-1-thiogalactopyranoside (IPTG) and purified using glutathione-Sepharose (GE Healthcare) and eluted with glutathione. MBP-Pfdn2 was injected into one rabbit and purified by GenScript (Hong Kong).
S2 cell culture, transient transfection and co-immunoprecipitation
Drosophila S2 cells were cultured in Shields and Sang m3 insect medium (Sigma-Aldich) with 10% fetal bovine serum (Hyclone) at 25 °C. Plasmids were transfected into S2 cells using Effectene Transfectin Reagent (QIAGEN). S2 cells were harvested 48 h after transfection and homogenized with lysis buffer (25 mM Tris pH8, 27.5 mM NaCl, 20 mM KCl, 25 mM sucrose, 10 mM EDTA, 10 Mm EGTA, 1 mM DTT, 10% (v/v) glycerol, 0.5% Nonidet P40) supplemented with Proteases inhibitors (Boeringher). Supernatants were collected and imunoprecipitated with mouse anti-Flag or anti-Myc for overnight at 4 °C, followed by incubation with Protein A/G beads (Pierces) for two hours at 4 °C. Bound proteins were analyzed by western blotting after washing for three times with cold PBS.
Quantification of spindle orientation
The apico-basal polarity of metaphase neuroblasts was indicated by a line perpendicular to the apical protein crescent, while the spindle axis was labeled by a second line parallel with the α-tubulin labeled mitotic spindle. The angles between these two lines were measured in metaphase neuroblasts.
EdU labeling
EdU labeling was processed with the Click-iT EdU Imaging Kits (Invetrogen). Briefly, third instar larval brains were dissected in PBS and incubated with 10 uM EdU working solution (Invetrogen) for 45 minutes. The brains were dissected and fixed in 3.7% formaldehyde with PBS for 15 mins at room temperature and permeabilized with 0.3% TritonX-100 in PBS. Incorporated EdU was detected after incubation with the Click-iT reaction cocktail (1 × Click-iT reaction buffer, 4% CuSO4, 0.24% Alexa Fluor azide, 10% Reaction buffer additive).
Microtubule regrowth assay
Larval brains were dissected in Shields and Sang m3 insect medium (Sigma-Aldich) with 10% fetal bovine serum (Hyclone) and incubated on ice for 30 min to depolymerize microtubules (MTs) completely. After that, brain samples were incubated in a water bath at 25 °C at various time points (0 s, 30 s and 120 s) to induce MTs regrowth. Samples were fixed in testis buffer (TB: 183 mM KCl, 47 mM NaCl, 10 mM Tris and 1 mM EDTA, pH6.8) supplemented with 37% formaldehyde.
Additional Information
How to cite this article: Zhang, Y. et al. Prefoldin and Pins synergistically regulate asymmetric division and suppress dedifferentiation. Sci. Rep. 6, 23735; doi: 10.1038/srep23735 (2016).
References
Doe, C. Q. Neural stem cells: balancing self-renewal with differentiation. Development 135, 1575–1587, doi: 10.1242/dev.014977 (2008).
Wu, P. S., Egger, B. & Brand, A. H. Asymmetric stem cell division: lessons from Drosophila. Semin Cell Dev Biol 19, 283–293, doi: 10.1016/j.semcdb.2008.01.007 (2008).
Gonczy, P. Mechanisms of asymmetric cell division: flies and worms pave the way. Nat Rev Mol Cell Biol 9, 355–366, doi: 10.1038/nrm2388 (2008).
Saini, N. & Reichert, H. Neural stem cells in Drosophila: molecular genetic mechanisms underlying normal neural proliferation and abnormal brain tumor formation. Stem Cells Int 2012, 486169, doi: 10.1155/2012/486169 (2012).
Gonzalez, C. Drosophila melanogaster: a model and a tool to investigate malignancy and identify new therapeutics. Nat Rev Cancer 13, 172–183, doi: 10.1038/nrc3461 (2013).
Ito, K. & Hotta, Y. Proliferation pattern of postembryonic neuroblasts in the brain of Drosophila melanogaster. Dev Biol 149, 134–148 (1992).
Lee, C. Y., Robinson, K. J. & Doe, C. Q. Lgl, Pins and aPKC regulate neuroblast self-renewal versus differentiation. Nature 439, 594–598, doi: 10.1038/nature04299 (2006).
Bello, B., Reichert, H. & Hirth, F. The brain tumor gene negatively regulates neural progenitor cell proliferation in the larval central brain of Drosophila. Development 133, 2639–2648, doi: 10.1242/dev.02429 (2006).
Betschinger, J., Mechtler, K. & Knoblich, J. A. Asymmetric segregation of the tumor suppressor brat regulates self-renewal in Drosophila neural stem cells. Cell 124, 1241–1253, doi: 10.1016/j.cell.2006.01.038 (2006).
Lee, C. Y., Wilkinson, B. D., Siegrist, S. E., Wharton, R. P. & Doe, C. Q. Brat is a Miranda cargo protein that promotes neuronal differentiation and inhibits neuroblast self-renewal. Dev Cell 10, 441–449, doi: 10.1016/j.devcel.2006.01.017 (2006).
Wodarz, A., Ramrath, A., Kuchinke, U. & Knust, E. Bazooka provides an apical cue for Inscuteable localization in Drosophila neuroblasts. Nature 402, 544–547, doi: 10.1038/990128 (1999).
Petronczki, M. & Knoblich, J. A. DmPAR-6 directs epithelial polarity and asymmetric cell division of neuroblasts in Drosophila. Nat Cell Biol 3, 43–49, doi: 10.1038/35050550 (2001).
Rolls, M. M., Albertson, R., Shih, H. P., Lee, C. Y. & Doe, C. Q. Drosophila aPKC regulates cell polarity and cell proliferation in neuroblasts and epithelia. J Cell Biol 163, 1089–1098, doi: 10.1083/jcb.200306079 (2003).
Schaefer, M., Shevchenko, A. & Knoblich, J. A. A protein complex containing Inscuteable and the Galpha-binding protein Pins orients asymmetric cell divisions in Drosophila. Curr Biol 10, 353–362 (2000).
Yu, F., Morin, X., Cai, Y., Yang, X. & Chia, W. Analysis of partner of inscuteable, a novel player of Drosophila asymmetric divisions, reveals two distinct steps in inscuteable apical localization. Cell 100, 399–409 (2000).
Fuse, N., Hisata, K., Katzen, A. L. & Matsuzaki, F. Heterotrimeric G proteins regulate daughter cell size asymmetry in Drosophila neuroblast divisions. Curr Biol 13, 947–954 (2003).
Izumi, Y., Ohta, N., Hisata, K., Raabe, T. & Matsuzaki, F. Drosophila Pins-binding protein Mud regulates spindle-polarity coupling and centrosome organization. Nat Cell Biol 8, 586–593, doi: 10.1038/ncb1409 (2006).
Siller, K. H., Cabernard, C. & Doe, C. Q. The NuMA-related Mud protein binds Pins and regulates spindle orientation in Drosophila neuroblasts. Nat Cell Biol 8, 594–600, doi: 10.1038/ncb1412 (2006).
Kraut, R., Chia, W., Jan, L. Y., Jan, Y. N. & Knoblich, J. A. Role of inscuteable in orienting asymmetric cell divisions in Drosophila. Nature 383, 50–55, doi: 10.1038/383050a0 (1996).
Schaefer, M., Petronczki, M., Dorner, D., Forte, M. & Knoblich, J. A. Heterotrimeric G proteins direct two modes of asymmetric cell division in the Drosophila nervous system. Cell 107, 183–194 (2001).
Hampoelz, B., Hoeller, O., Bowman, S. K., Dunican, D. & Knoblich, J. A. Drosophila Ric-8 is essential for plasma-membrane localization of heterotrimeric G proteins. Nat Cell Biol 7, 1099–1105, doi: 10.1038/ncb1318 (2005).
Wang, H. et al. Ric-8 controls Drosophila neural progenitor asymmetric division by regulating heterotrimeric G proteins. Nat Cell Biol 7, 1091–1098, doi: 10.1038/ncb1317 (2005).
Wang, C. et al. An ana2/ctp/mud complex regulates spindle orientation in Drosophila neuroblasts. Dev Cell 21, 520–533, doi: 10.1016/j.devcel.2011.08.002 (2011).
Lee, C. Y. et al. Drosophila Aurora-A kinase inhibits neuroblast self-renewal by Genes Dev 20, 3464–3474, doi: 10.1101/gad.1489406 (2006).
Wang, H. et al. Aurora-A acts as a tumor suppressor and regulates self-renewal of Drosophila neuroblasts. Genes Dev 20, 3453–3463, doi: 10.1101/gad.1487506 (2006).
Wang, H., Ouyang, Y., Somers, W. G., Chia, W. & Lu, B. Polo inhibits progenitor self-renewal and regulates Numb asymmetry by phosphorylating Pon. Nature 449, 96–100, doi: 10.1038/nature06056 (2007).
Bello, B. C., Izergina, N., Caussinus, E. & Reichert, H. Amplification of neural stem cell proliferation by intermediate progenitor cells in Drosophila brain development. Neural Dev 3, 5, doi: 10.1186/1749-8104-3-5 (2008).
Boone, J. Q. & Doe, C. Q. Identification of Drosophila type II neuroblast lineages containing transit amplifying ganglion mother cells. Dev Neurobiol 68, 1185–1195, doi: 10.1002/dneu.20648 (2008).
Bowman, S. K. et al. The tumor suppressors Brat and Numb regulate transit-amplifying neuroblast lineages in Drosophila. Dev Cell 14, 535–546, doi: 10.1016/j.devcel.2008.03.004 (2008).
Knoblich, J. A. Asymmetric cell division: recent developments and their implications for tumour biology. Nat Rev Mol Cell Biol 11, 849–860, doi: 10.1038/nrm3010 (2010).
Zhu, S., Barshow, S., Wildonger, J., Jan, L. Y. & Jan, Y. N. Ets transcription factor Pointed promotes the generation of intermediate neural progenitors in Drosophila larval brains. Proc Natl Acad Sci USA 108, 20615–20620, doi: 10.1073/pnas.1118595109 (2011).
Janssens, D. H. et al. Earmuff restricts progenitor cell potential by attenuating the competence to respond to self-renewal factors. Development 141, 1036–1046, doi: 10.1242/dev.106534 (2014).
Weng, M., Golden, K. L. & Lee, C. Y. dFezf/Earmuff maintains the restricted developmental potential of intermediate neural progenitors in Drosophila. Dev Cell 18, 126–135, doi: 10.1016/j.devcel.2009.12.007 (2010).
Eroglu, E. et al. SWI/SNF complex prevents lineage reversion and induces temporal patterning in neural stem cells. Cell 156, 1259–1273, doi: 10.1016/j.cell.2014.01.053 (2014).
Koe, C. T. et al. The Brm-HDAC3-Erm repressor complex suppresses dedifferentiation in Drosophila type II neuroblast lineages. Elife 3, e01906, doi: 10.7554/eLife.01906 (2014).
Vainberg, I. E. et al. Prefoldin, a chaperone that delivers unfolded proteins to cytosolic chaperonin. Cell 93, 863–873 (1998).
Geissler, S., Siegers, K. & Schiebel, E. A novel protein complex promoting formation of functional alpha- and gamma-tubulin. EMBO J 17, 952–966, doi: 10.1093/emboj/17.4.952 (1998).
Lupas, A. Coiled coils: new structures and new functions. Trends Biochem Sci 21, 375–382 (1996).
Siegert, R., Leroux, M. R., Scheufler, C., Hartl, F. U. & Moarefi, I. Structure of the molecular chaperone prefoldin: unique interaction of multiple coiled coil tentacles with unfolded proteins. Cell 103, 621–632 (2000).
Lundin, V. F., Srayko, M., Hyman, A. A. & Leroux, M. R. Efficient chaperone-mediated tubulin biogenesis is essential for cell division and cell migration in C. elegans. Dev Biol 313, 320–334, doi: 10.1016/j.ydbio.2007.10.022 (2008).
Cao, S. et al. Subunit 1 of the prefoldin chaperone complex is required for lymphocyte development and function. J Immunol 181, 476–484 (2008).
Delgehyr, N. et al. Drosophila Mgr, a Prefoldin subunit cooperating with von Hippel Lindau to regulate tubulin stability. Proc Natl Acad Sci USA 109, 5729–5734, doi: 10.1073/pnas.1108537109 (2012).
Neumuller, R. A. et al. Genome-wide analysis of self-renewal in Drosophila neural stem cells by transgenic RNAi. Cell Stem Cell 8, 580–593, doi: 10.1016/j.stem.2011.02.022 (2011).
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451–461 (1999).
Peng, C. Y., Manning, L., Albertson, R. & Doe, C. Q. The tumour-suppressor genes lgl and dlg regulate basal protein targeting in Drosophila neuroblasts. Nature 408, 596–600, doi: 10.1038/35046094 (2000).
Rommelaere, H. et al. Prefoldin recognition motifs in the nonhomologous proteins of the actin and tubulin families. J Biol Chem 276, 41023–41028, doi: 10.1074/jbc.M106591200 (2001).
Rossi, F. & Gonzalez, C. Synergism between altered cortical polarity and the PI3K/TOR pathway in the suppression of tumour growth. EMBO Rep 13, 157–162, doi: 10.1038/embor.2011.230 (2012).
Siegrist, S. E. & Doe, C. Q. Microtubule-induced Pins/Galphai cortical polarity in Drosophila neuroblasts. Cell 123, 1323–1335, doi: 10.1016/j.cell.2005.09.043 (2005).
Knoblich, J. A., Jan, L. Y. & Jan, Y. N. Asymmetric segregation of Numb and Prospero during cell division. Nature 377, 624–627, doi: 10.1038/377624a0 (1995).
Broadus, J. & Doe, C. Q. Extrinsic cues, intrinsic cues and microfilaments regulate asymmetric protein localization in Drosophila neuroblasts. Curr Biol 7, 827–835 (1997).
Wallenfang, M. R. & Seydoux, G. Polarization of the anterior-posterior axis of C. elegans is a microtubule-directed process. Nature 408, 89–92, doi: 10.1038/35040562 (2000).
Chang, F., Feierbach, B. & Martin, S. Regulation of actin assembly by microtubules in fission yeast cell polarity. Novartis Found Symp 269, 59–66; discussion 66–72, 223–230 (2005).
Vladar, E. K., Bayly, R. D., Sangoram, A. M., Scott, M. P. & Axelrod, J. D. Microtubules enable the planar cell polarity of airway cilia. Curr Biol 22, 2203–2212, doi: 10.1016/j.cub.2012.09.046 (2012).
Gu, Y. et al. Prefoldin 6 is required for normal microtubule dynamics and organization in Arabidopsis. Proc Natl Acad Sci USA 105, 18064–18069, doi: 10.1073/pnas.0808652105 (2008).
Lee, Y. et al. Prefoldin 5 is required for normal sensory and neuronal development in a murine model. J Biol Chem 286, 726–736, doi: 10.1074/jbc.M110.177352 (2011).
Wang, H., Cai, Y., Chia, W. & Yang, X. Drosophila homologs of mammalian TNF/TNFR-related molecules regulate segregation of Miranda/Prospero in neuroblasts. EMBO J 25, 5783–5793, doi: 10.1038/sj.emboj.7601461 (2006).
Cai, Y., Chia, W. & Yang, X. A family of snail-related zinc finger proteins regulates two distinct and parallel mechanisms that mediate Drosophila neuroblast asymmetric divisions. EMBO J 20, 1704–1714, doi: 10.1093/emboj/20.7.1704 (2001).
Bellaiche, Y. et al. The Partner of Inscuteable/Discs-large complex is required to establish planar polarity during asymmetric cell division in Drosophila. Cell 106, 355–366 (2001).
Fujioka, Y. et al. MM-1, a c-Myc-binding protein, is a candidate for a tumor suppressor in leukemia/lymphoma and tongue cancer. J Biol Chem 276, 45137–45144, doi: 10.1074/jbc.M106127200 (2001).
Hennecke, S. et al. Prevalence of the Prefoldin Subunit 5 Gene Deletion in Canine Mammary Tumors. PLos One 10, e0131280, doi: 10.1371/journal.pone.0131280 (2015).
Acknowledgements
We thank E Schejter, YN Jan, J. Skeath, C. Doe, F. Matsuzaki, J. Knoblich, C.Y. Lee, F. Yu, GM Rubin, the Bloomington Drosophila Stock Center and Vienna Drosophila RNAi Center for fly stocks and antibodies. This work is supported by Ministry of Education (MOE) Tier 2 funding MOE2014-T2-1-090 (H.W.), Duke-NUS Signature Research Program funded by Ministry of Health, Singapore (H.W.) and NUS Graduate School for Integrative Sciences and Engineering (NGS) Scholarship (Y.Z.). Work in C.G.´s laboratory is supported by AdG 2011 294603 advanced Grant from the European Research Council (ERC), BFU2015-66304 and BFU2014-52125-REDT-CellSYS from the Spanish MINECO and SGR Agaur 2014 100 from Generalitat de Catalunya. M.R. is supported by fellowship BES-2013-064543 from the MINECO, Spain.
Author information
Authors and Affiliations
Contributions
H.W., Y.Z. M.R. and C.G. designed the experiments and analyzed the data. Y.Z., M.R. and C.W. conducted the experiments. Y.Z. and H.W wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhang, Y., Rai, M., Wang, C. et al. Prefoldin and Pins synergistically regulate asymmetric division and suppress dedifferentiation. Sci Rep 6, 23735 (2016). https://doi.org/10.1038/srep23735
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep23735
This article is cited by
-
Mitochondria-enriched protrusions are associated with brain and intestinal stem cells in Drosophila
Communications Biology (2019)
-
An expandable embryonic stem cell-derived Purkinje neuron progenitor population that exhibits in vivo maturation in the adult mouse cerebellum
Scientific Reports (2017)
-
Identification of potential predictive markers of dexamethasone resistance in childhood acute lymphoblastic leukemia
Journal of Cell Communication and Signaling (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.