Abstract
The regulatory systems underlying animal development must have evolved prior to the emergence of eumetazoans (cnidarians and bilaterians). Although representatives of earlier-branching animals – sponges ctenophores and placozoans – possess most of the developmental transcription factor families present in eumetazoans, the DNA regulatory elements that these transcription factors target remain uncharted. Here we characterise the core promoter sequences, U1 snRNP-binding sites (5′ splice sites; 5′SSs) and polyadenylation sites (PASs) in the sponge Amphimedon queenslandica. Similar to unicellular opisthokonts, Amphimedon’s genes are tightly packed in the genome and have small introns. In contrast, its genes possess metazoan-like core promoters populated with binding motifs previously deemed to be specific to vertebrates, including Nrf-1 and Krüppel-like elements. Also as in vertebrates, Amphimedon’s PASs and 5′SSs are depleted downstream and upstream of transcription start sites, respectively, consistent with non-elongating transcripts being short-lived; PASs and 5′SSs are more evenly distributed in bidirectional promoters in Amphimedon. The presence of bilaterian-like regulatory DNAs in sponges is consistent with these being early and essential innovations of the metazoan gene regulatory repertoire.
Similar content being viewed by others
Introduction
Spatiotemporal and cell type-specific gene regulation is prerequisite to multicellularity. Thus, any insights into the origin and evolution of animals requires a detailed understanding of gene structure – both regulatory and coding sequence – and the interplay between available transcription and regulatory factors with regulatory sequences that control gene expression1,2,3,4. Although the evolution of metazoan transcription factor families has been relatively well documented5,6,7,8,9, the evolution of regulatory sequences and gene regulatory networks remain restricted to a handful of case studies in bilaterian arthropods, chordates, nematodes and echinoderms10,11,12,13,14,15,16. It currently remains an open question as to what extent early-branching, morphologically simple animals, sponges (poriferans), ctenophores, placozoans and cnidarians, share regulatory elements with more complex bilaterians.
Although genomic regions that regulate gene expression often lie at a distance from the transcription start site (TSS)17, sequences overlapping with and in the vicinity of the TSS – core or basal promoter elements – are necessary for the integration of cis-regulatory inputs and the initiation of transcription18,19,20. These promoters also contribute directly to the regulation of gene expression in development and cell differentiation beyond simply the coordination of transcription21,22,23.
Some core promoter regulatory elements appear to be broadly conserved amongst eukaryotes (e.g. the TATA-box24,25), although there is marked variation in promoters between genes within a given species18,26,27,28. For instance, TATA-boxes and the initiator elements (Inr), which are considered the only core promoter motifs conserved from yeast to humans25, are present in only ~25% of human promoters25. Further, comparative analyses2,25,29,30 reveal that while there are a handful of other conserved elements that are present in some animal promoters [e.g., the transcription factor IIB recognition element (BRE), downstream core element (DCE), downstream promoter element (DPE), DNA recognition element (DRE) and motif ten element (MTE)], many regulatory sequence motifs appear to be restricted to specific species3,26,31,32. The diversity of promoter classes within and between metazoan species supports the view that this region contributes to the complex regulation of gene transcription in metazoans3,33.
The increase in the number and diversity of draft metazoan and eukaryotic genomes provides an opportunity to understand the relationship between the regulatory capacity of the genome and morphological evolution, including the origin of animal multicellularity. Leveraging on an increasingly detailed understanding of genome structure and function in model species, we use the reannotated draft genome of the demosponge Amphimedon queenslandica34, to begin to understand the relationship between gene regulation and animal evolution. Our findings are consistent with metazoan core promoter(s) originating in the pre-Cambrian before the divergence of eumetazoan and sponge lineages.
Results
The Amphimedon genome is compact
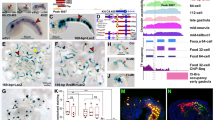
The reannotated Amphimedon genome contains 40,122 coding sequence gene models (excluding isoforms), covering nearly 65% of total genomic sequence34. These new gene models include better-annotated 5′ and 3′ untranslated regions (UTRs), allowing for the identification of transcript start and termination sites (Fig. 1). This new annotation has revealed the Amphimedon genome is phenomenally compact, having a median intergenic distance of a mere 587 bp34 and few gene deserts.
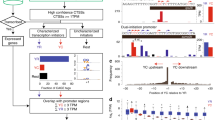
(a) A gene rich region with Aqu2 gene models shown in purple with thick lines depicting exons, mid size lines UTRs and thin lines introns. The wiggle tracks below show the RNA-seq expression of the Watson (top) and Crick (bottom) strand in precompetent larvae, competent larvae, juvenile and adult samples. The zoomed in region shows examples of genes orientated tail to tail, head to tail and head to head, with the direction of transcription shown by orange arrows and in introns as small arrow heads. (b) A gene depleted region. Color schema as in panel (a).
To place the gene density of the Amphimedon genome within a comparative framework, we surveyed the number of protein-coding genes in non-overlapping 50 kb windows of genomic DNA in a range of animals, two choanoflagellates, a filasterean and yeast (Fig. 2, Table 1). This survey of animal genomes is not exhaustive but includes representatives of ctenophores, placozoans and cnidarians, along with bilaterians with comparable genome sizes or known to have relatively compact genomes within the taxon they represent (e.g. the pufferfish, Takifugu rubripes). We also included the human genome in this analysis.
This analysis includes four non-metazoan opisthokonts (Saccharomyces cerevisiae, Capsaspora owczarzaki, and two choanoflagellates, Salpingoeca rosetta and Monosiga brevicollis), the sponge Amphimedon queenslandica, two ctenophores (Mnemiopsis leidyi and Pleurobrachia bachei), the placozoan Trichoplax adhaerens, the cnidarian Nematostella vectensis, the arthropod Drosophila melanogaster, and three chordates (Ciona intestinalis, Takifugu rubripes and Homo sapiens). Their relationships are shown to the left and the gene density distribution for each species is shown to the right. The percentage of genomic windows (y-axis) by the number of protein coding genes per 50 kb genomic window (x-axis) is shown, using data from all genomic scaffolds longer than 50 kb. Vertical blue lines in each panel mark the average gene density (see also Table 1). To diminish bias caused by multiple gene isoforms in well-characterized genomes (i.e. D. melanogaster), genes were counted as occurrences of a uniquely annotated genic 5′ ends (see Table 1). The broken axis for H. sapiens is to avoid scale distortion in other species.
We find that A. queenslandica has one of the most gene dense metazoan genome currently known, with a median of 9 genes per 50 kb. Only 2.7% of the Amphimedon genome is depleted of genes (Fig. 2, Table 1). The gene density of Amphimedon resembles that of the choanoflagellates Salpingoeca rosetta and Monosiga brevicollis, which have a median gene density of 10 and 11 per 50 kb, respectively, although it is less dense than the more evolutionary distant filasterean Capsaspora owczarzaki, with a median of 16 genes per 50 kb (Fig. 2, Table 1).
Gene density differences cannot be solely attributed to differences in genome size. The A. queenslandica genome (166.7 Mb) is over three and four times the size of that of S. rosetta (55.4 Mb) and M. brevicollis (41.6 Mb), respectively (Table 1). Although both the ctenophore genomes are similar in size to the Amphimedon genome (Mnemiopsis leidyi, 155.9 Mb; Pleurobrachia bachei, 156.1 Mb), they have a markedly lower median gene density (5 and 2 genes per 50 kb, respectively) and a higher percentage of gene depleted regions (5.8 and 17% in M. leidyi and P.bachei) (Table 1). These values for the ctenophores are closer to those found in Nematostella vectensis (356.6 Mb), although the cnidarian genome is nearly twice as large (Fig. 2, Table 1). The urochordate Ciona intestinalis has a smaller genome than A. queenslandica (115.2 Mb) and only 1.7% of its genome is composed of gene-depleted regions (Fig. 2), yet it is still less gene dense than that of A. queenslandica, with a median of seven genes per 50 kb (Fig. 2). Even the miniature genome of the holoplanktonic urochordate Oikopleura dioica (65 Mb) has a gene density similar to Amphimedon35. The genomes of Drosophila melanogaster, Takifugu rubripes and Homo sapiens are the least gene dense, and a mosaic of gene dense and depleted regions (Fig. 2, Table 1).
The A. queenslandica genome contains 18,054 (45%) genes that have the potential to be transcribed in opposite directions off the same core promoter. 11,379 of these head-to-head genes have transcription start sites (TSSs) 1 kb or less away from each other, and thus may be under the control of a bidirectional promoter (Table 2), as previously defined36,37. This makes Amphimedon currently the animal with the highest number of potential bidirectional promoters; 4,398 D. melanogaster genes (32.2%) are transcribed from bidirectional promoters36.
Amphimedon promoters are enriched in elements present in bilaterian promoters
Using deep stranded expression data from a variety of developmental stages34, we identified 3,309 gene models (8.2% of total) with both RNA-seq supported 5′ gene-ends (annotated as described in34) and a promoter that did not overlap with another gene promoter (Supplementary Table 1). This is comprised of 330 bidirectional (when both head-to-head promoters have 5′ ends), 645 putative bidirectional (when one head-to-head promoter pair has a single gene with an annotated 5′ end) and 2,334 unidirectional (when no evidence of bidirectional or divergent transcription is found within 1.0 kb of the identified promoter) promoters. Analysis of the nucleotide composition in the vicinity of the TSSs of these three promoter types (Supplementary Fig. 1) reveals no sequence differences, with all having an increase in C and G nucleotide frequency at and just after the TSS (Supplementary Fig. 1).
An unbiased survey of the DNA sequences most overrepresented in the vicinity of the TSSs of the 3,309 coding gene representatives revealed 15 motifs significantly enriched in unidirectional, bidirectional and putative bidirectional promoters (Fig. 3; Supplementary Table 2). The most abundant element is a specificity protein 1 (Sp1)-like GC–box, whose frequency is enriched within 50 bp upstream of the TSS peaking right at the TSS (Fig. 3). This GC-rich motif is more prevalent in the bidirectional and putative bidirectional promoters (~36%) compared to unidirectional promoters (29%) (Supplementary Table 2). The reverse compliment of this motif (Motif 2, Fig. 3) is also overrepresented in promoter regions (coding strand) and is most common at the TSS (Fig. 3). This motif is also more abundant in putative bidirectional and bidirectional promoters (Supplementary Table 2). E-box sequences are enriched around the A. queenslandica TSSs and practically identical to the known canonical sequence of a special type of E-box known as a G-box, characteristic of plants38. This motif is four times more common in bidirectional and putative bidirectional promoters than in unidirectional promoters (Supplementary Table 2). A motif containing the Kozak sequence (translation initiation site) is also overrepresented in A. queenslandica promoters, although only in a fraction of the genes (Fig. 3). This motif is AT rich, similar to previously reported Kozak consensus in cnidarians and yeast39. This motif also bears close resemblance to the YY1 active motif in immortalized human cell lines40. The YY1/Kozak motif is more prevalent ~10 bp after the transcription start site, although its low abundance suggests that some genes begin translation further than 100 bp downstream of the annotated TSS (Fig. 3). The post-TSS position of the identified Kozak sequence confirms the use of only genes with high confidence 5′ UTRs and a strict motif identification approach to identify promoter elements.
The sequence logo of DNA motifs significantly enriched in the promoters and the TATA-box are shown. The frequency per 10 bp bin from −400 bp to +100 bp relative to the TSS (grey vertical line) is displayed in column 4. The significance value is the e-value of the log likelihood ratio of each motif. The right column shows the number of promoters, amongst the 3,309 promoters surveyed, and the percentage of promoters (in parentheses) that possess the motif.
Additional motifs characteristic of bilaterian promoters include Nrf-1, EGR-2 and KLF family binding sites (Fig. 3). The Nrf-1 and EGR-2 binding sequences are distributed close to the TSS but without a clear positional association (Fig. 3). The KLF-family motif, when present, is found at a similar distance to the TSS as the Sp1/GC-box motif. This is because of the sequence similarity between the two motifs (Fig. 3). Poly-CT patches are enriched more downstream of the TSS site. Four additional motifs that have no clear similarity with previously identified binding motifs are also significantly enriched around the TSS; only one of them, “Unknown 2”, is more prevalent close to the TSS than the surrounding regions (Fig. 3).
Many of the promoter elements that we identified in the survey of 3,309 genes with well-defined TSSs and promoters are also enriched in the putative promoters in other protein-coding genes in the Amphimedon genome, despite these genes having less resolved TSSs (Supplementary Fig. 2). These enrichments include Sp1/GC-box and the Sp1rc motifs, which are the most abundant elements that precede or overlay TSSs across the genome.
Among the most highly enriched motifs in the Amphimedon promoters surveyed, we found only one A/T-rich motif, with similar nucleotide composition to the TATA-box. Nevertheless, this motif had no specific positional enrichment and was prevalent across the whole region upstream of the TSS (Fig. 3). Scanning the promoters using the TATA-box consensus from vertebrates, we find that the TATA-box frequency peaks approximately 30 bp upstream of the TSS as expected33. Similar to previous reports in other organisms25,41, we find that (865) 26% of all surveyed promoters contain an identifiable TATA-box, regardless of promoter type (Fig. 3). This is perhaps not surprising given the Amphimedon genome is also relatively AT-rich (64.2%) compared to many other animal genomes (Table 1).
Other animal core promoter elements, including Inr, BREu, BREd, DPE, MTE, CEBP, and XCPE1, were not identified as enriched in our analysis. A survey for these motifs shows the BREu is most abundant in the region corresponding to the TSS (Supplementary Fig. 3), and closely resembles the distribution of the Sp1/GC-box motif on the promoter region (Fig. 3). This observation, in addition to the similarity between BREu and the Sp1/GC-box motif, suggest that the latter element might be acting as BREu in A. queenslandica. The metazoan-specific ETS motif is infrequent in Amphimedon promoters but localizes to the region around the TSS when present (Supplementary Fig. 3). Other conserved motifs, including the human and Drosophila Inr, BREd, DPE, CEBP and XCPE1 motifs, were either absent, present in very low frequencies or appear to be randomly distribution in the core promote region, possibly reflecting the inherent low specificity of these motifs42 (Supplementary Fig. 3).
Differential enrichment of poly(A) signals upstream and U1 snRNP-binding sites (5′ splice sites) downstream of Amphimedon core promoters
Vertebrate transcriptional elongation is driven by pervasive bidirectional and divergent transcription from promoters, followed by quick destabilization and degradation of unstable transcripts from the non-coding strand43,44. The enrichment of U1 snRNP-binding sites (5′ splice sites; 5′SSs) and poly(A) signals (PASs) downstream and upstream of the TSS, respectively, correlate with transcript stability45. A survey of the abundance of PAS and 5′SS motifs in relation to all TSSs in the Amphimedon genome reveals PAS motifs accumulate more rapidly in the antisense (non-coding) than in the sense direction (coding) and 5′SS motifs are more prevalent on the coding strand, albeit at much lower frequency (Fig. 4a). A similar accumulation pattern is observed when the survey is restricted to genes with core promoters containing Sp1/GC box or Sp1rc motifs (Fig. 4b). 5′SS motifs accumulate nearly twice as rapidly downstream of these highly prevalent motifs compared to TSSs across the genome (Fig. 4a,b).
The polyadenylation signal (AWUAAA PAS) and the U1 snRNP-binding sites (5′ splice sites; 5′SS) frequencies are shown in top and bottom panels, respectively. Blue lines, cumulative frequency for 500 bp downstream of the TSS in the sense direction. Green lines, cumulative frequency for 500 bp upstream in the antisense direction. (a) All TSSs in the genome. (b) Promoters with an identifiable Sp1/GC-box motif (c) Bidirectional promoters with an overlapping pair of Sp1/GC-box and Sp1rc motifs.
Analysis of bidirectional promoters with co-occurring and/or overlapping Sp1/GC-box and Sp1rc motifs revealed PAS accumulated at almost identical rates in both sense and antisense direction (Fig. 4c). These motifs are over three times as likely to co-occur in a promoter than expected by chance (Supplementary Table 3, see supplementary note). 5′SS sequences tended to accumulate more rapidly in the sense than the antisense strand. Although the 5′SS cumulative frequency rises at almost similar rates between strands, they are not as similar as that of the PAS signal. This difference in cumulative frequency suggests polyadenylation is a more prominent transcriptional regulatory element than splicing in Amphimedon.
Discussion
Amphimedon queenslandica has one of the most gene dense genomes in the animal kingdom, with a median of 9 genes per 50 kb and a median intergenic distance of 0.59 kb. Amphimedon’s genome organization, average intron size and alternative splicing patterns34,46 are more similar to unicellular holozoans than to other animals. These may be symplesiomorphic traits that are shared between sponges and unicellular holozoans, with other metazoan lineages evolving larger introns and intergenic regions. Alternatively Amphimedon’s high gene density may be the result of a secondary reduction in intergenic and intron DNA size from a last common metazoan ancestor that had already had an increased amount of non-coding DNA. Secondary increases in gene density appear to have occurred elsewhere in the animal kingdom (e.g. urochordates).
The enlargement of intergenic space increases the availability of non-coding, and presumably sequences that are less constrained to evolve into transcription factor target sites47,48. Although most developmental transcription factor families evolved early in metazoan evolution, prior to the divergence of extant phyla5,8, a majority of these families underwent further expansion and diversification early in eumetazoan evolution after this lineage diverged from poriferan and ctenophore lineages. The increase in transcription factor family membership in eumetazoans is likely to have led to an expansion in regulatory potential and gene regulatory network complexity and may underlie the difference in morphological evolution in sponges and eumetazoans5. The sponge body plan has remained unchanged since before the Cambrian49, while the eumetazoan lineage consists of a huge diversity of body plans. Ctenophores, whose transcription factor repertoires are similar to sponges50,51, have larger intergenic regions than Amphimedon and are considered morphologically more complex.
The high gene density of the Amphimedon genome results in a high frequency of small intergenic regions, overlapping transcription start and termination sites, and bidirectional promoters. Almost one third of A. queenslandica genes appear to be transcribed from a bidirectional promoter. We did not detect any major differences in nucleotide composition or abundance or distribution of binding motifs between unidirectional and bidirectional core promoter regions. Despite the relative AT-richness of the Amphimedon genome, there is a marked increase in the frequency of CpG dinucleotides at the Amphimedon TSS, similar to mammalian promoters52. CpG dinucleotides are highly mutable and rapidly lost in organisms with active DNA methylation mechanisms, as previously proposed for Amphimedon52,53,54. The increase in CpG around the Amphimedon TSS suggests that the mechanism responsible for maintaining low methylation levels of CpG dinucleotides in core promoters may have evolved prior to the divergence of poriferans and eumetazoans, or independently in sponges and vertebrates.
Ancient origin of animal promoters
Several of the most prevalent sequence motifs are overrepresented in and around the Amphimedon promoter include sequences previously found in bilaterians or specifically vertebrates, including KLF, Nrf-1 and ERG binding sites. The most prevalent among these is the Sp1/GC-box like motif, present in about 29% of all promoters. Given its prevalence and similarity to bilaterian BREu elements3, the Sp1/GC-box may play a similar function in core promoters in this sponge. The increased presence of Sp1/GC-box and Sp1rc motifs in Amphimedon bidirectional promoters is consistent with these elements promoting transcription in both directions.
It is worth noting that no other conserved bilaterian transcription initiation motif is enriched in A. queenslandica promoters. However, a deliberate search for conserved promoter motifs did reveal a relatively well-preserved TATA-box motif 30 bp upstream to the TSS in about 26% of the Amphimedon genes. Its location relative to the TSS is in agreement with the known TATA distances in other metazoans33.
Members of the Sp/KLF family of proteins are known to bind to either GC-box55 or CACC-box motifs56, both which share similarity with Sp1rc, the second most abundant sequence enriched in Amphimedon promoters. This protein family, characterized by a three C2H2 zinc finger DNA-binding domain at the C-terminus55, arose and expanded early in metazoan evolution57,58,59. Amphimedon has about ten Sp/KLF family members (Fernandez-Valverde et al. unpublished59). The Sp transcription factor family is also known to help recruit TFIID (part of the pre-initiation complex) and GATA1 to the transcription complex60,61,62. Sp1 has also been shown to activate transcription without enhancing the DNA binding activity of the TATA box-factor63. The high abundance of the GC-box motif, which was also previously identified as enriched in the promoter region of A. queenslandica ribosomal proteins64, is consistent with Sp/KLF proteins being major recruiters of TFIID to the TSS in this sponge.
The Nrf-1 transcription factor gene family appears also to be restricted to metazoans. The Amphimedon genome has at least one member of this gene family. In addition to this study, the Nrf-1 binding motif has been found to be enriched in X. tropicalis promoters26 and in sponge ribosomal gene promoters64. Furthermore, a recent ultra-deep survey of transcribed enhancers in human immune cells displays almost a 100% similarity to the consensus sequence found in Amphimedon65. Together these observations suggest that Nrf-1 and its target binding site evolved before the divergence of sponge and eumetazoan lineages.
The poly-CT motif present in many A. queenslandica promoters has been previously identified in ribosomal gene promoters of this sponge, and in N. vectensis and Trichoplax adhaerens64. Although this motif has been proposed to be a metazoan innovation64, the presence of pyrimidine- and CTT-rich motifs in plant promoters66,67 suggests that the poly-CT motif present in sponge promoters may have a more ancient origin and possibly has been lost early in bilaterian evolution. Likewise, a plant-like E-box motif is also prevalent in Amphimedon promoters, suggesting it may be a conserved ancient feature of gene regulation.
A conserved metazoan U1-PAS axis
It has been proposed that the perceived directionality of transcription downstream of active promoters arises from the rapid degradation of transcripts produced in the other direction from the non-elongating strand43,44. The differential accumulation of PAS and 5′SS sites in both directions away from promoters appears to dictate transcript stability, with PAS and 5′SS sites depleted and enriched, respectively, in the direction that yields stable transcripts.
Amphimedon displays a depletion of PASs and accumulation of 5 ′SSs in the direction of stable transcription, suggesting these mechanisms also contribute to transcription directionality in this sponge. Amphimedon has a high abundance of bidirectional promoters. In contrast to unidirectional promoters, PASs accumulate at almost identical rates both upstream and downstream of bidirectional promoters, consistent with the differential accumulation of PAS sites contributing to transcript stability in Amphimedon. The differential presence of PAS sites upstream and downstream of unidirectional and bidirectional promoters with Sp1/GC-box and overlapping Sp1/GC-box and Sp1rc motifs suggests promoters harboring these motifs are capable of triggering stable bidirectional transcription in Amphimedon.
Our findings are consistent with pervasive bidirectional transcription followed by quick destabilization of the non-elongating strand via mechanisms associated with the “U1-PAS Axis” emerged prior to the divergence of sponges and eumetazoans, and antedates the expansion of non-coding DNA in eumetazoans. Interestingly, the “U1-PAS Axis” also has been proposed as a mechanism for the origin of new genes45 as well as a crucial contributor to the emergence of stable non-coding RNA transcripts68,69. The apparent existence of this axis in Amphimedon, along with a long non-coding RNA repertoire similar to that found in bilaterians70, suggests that this mode of gene origination may be also an ancient feature of metazoan genomes.
Summary
Overall, our observations are consistent with sponges having maintained genomic features that emerged early in metazoan evolution. The apparent dual unicellular-multicellular nature of the Amphimedon genome suggests that last common ancestor of sponges and eumetazoans had already evolved a regulatory core promoter to allow for context- and cell type-specific gene expression, yet had not evolved the ramified regulatory architecture observed in complex bilaterians. These evolved along the eumetazoan stem along with the expansion of key developmental transcription factor families5,8.
Material and Methods
Gene density analysis
The genomes and gene annotations of H. sapiens, T. rubripes, C. intestinalis, D. melanogaster and S. cerevisiae were downloaded through the ftp site of Ensembl (http://www.ensembl.org/info/data/ftp/index.html) on March 2014. Non-coding genes were filtered out using the “coding” biotype as specified by Ensembl. The genomes of M. brevicollis, S. rosetta and C. owczarzaki were downloaded from the Broad genomes “Origins of Multicellularity” website (http://www.broadinstitute.org/annotation/genome/multicellularity_project/Downloads.html) on December 2014. The genome of T. adhaerens was downloaded from the JGI website (http://genome.jgi.doe.gov/Triad1/Triad1.download.ftp.html) on December 2015. The NCBI repository was used to retrieve the genomes of M. leidyi (http://www.ncbi.nlm.nih.gov/nuccore/AGCP00000000.1/) P. bachei (http://www.ncbi.nlm.nih.gov/nuccore/AVPN00000000.1/) on May 2014. The A. queenslandica genome used in this study corresponds to that described in71.
All genomes were filtered to remove mitochondrial sequences and scaffolds shorter than 50 kb using awk. The gtf gene annotation files were converted to bed files using the UCSC utilities72. All TSSs were extracted and, if duplicated, collapsed into unique TSSs using a custom awk script. All genomes were divided onto non-overlapping windows of 50 kb and the number of TSSs overlapping each window were quantified using bedtools coverageBed73.
Promoter identification
Promoter regions were identified using a custom perl script and a combination of the UCSC utilities72 and bedtools73. Briefly, using the TSS defined as the start position of the 5′ UTR we extracted 150 bp upstream of the core promoter and 50 bp downstream. All sequences that fell outside of an annotated genome scaffold or overlapped a gap in the scaffold were excluded. Promoters that had any overlap with another promoter were also excluded. To further refine our promoter set we only used promoters that were found on scaffolds longer than 10 kb that did not have long CpG islands (longer than 10 kb) and might be due to bacterial contamination. To identify bidirectional promoters, all promoters that were within 1 kb of each other and had evidence of bidirectional transcription were retrieved via a MySQL query of the Amphimedon genome UCSC database. To classify promoters into unidirectional, bidirectional and putatively bidirectional all core promoters were intersected with bidirectional intergenic regions of 1 kb or less using overlapSelect from UCSC utilities72. When two divergent core promoters overlapped a bidirectional intergenic region these promoters were considered “bidirectional”, while if only a single promoter overlapped the region it was classified as “putatively bidirectional”, as there was evidence of bidirectional transcription but only one of the genes had an annotated 5′ UTR and thus an annotated promoter. Promoters that did not overlap with these regions were deemed “unidirectional”. From these groups, only promoters with annotated 5′ UTRs were used in the analysis described in the text.
Motif identification, co-occurrence and positional enrichment
All strict core promoter motif were identified using meme (parameters: -maxsize 20000000 -p 14 -dna -nmotifs 10 -minw 6 -maxw 15 -mod zoops)74. Each motif was searched against the Jaspar CORE and Jaspar POLII motif database75 using TomTom (http://meme.ebi.edu.au/meme/tools/tomtom), STAMP (http://www.benoslab.pitt.edu/stamp/) and manual searches and renamed according to their most similar motif in the database or literature, if any. Motif frequency matrixes were retrieved from MEME output, converted into Homer format and used to identify of frequency across the promoter region76.
Positional motif enrichment analysis was carried out using Homer76. All analyses on the same strand were done using the “-norevopp” parameter. For motif positional enrichment all motifs of interest were searched in the core promoter region of bidirectional, putative bidirectional and unidirectional promoters (see above) using the annotatePeaks.pl script from the Homer pipeline −400 and +100 nts around the TSS in adjacent windows of 10 bp each. All frequency metrics refer to the number of identified motifs per bin divided by the total number of peaks in region. For genome-wide promoter analysis the “tss” option in Homer was used and only sequences that had full regions around the identified motif (no gaps) were analyzed. The dinucleotide frequency of core promoter regions was calculated using the faCount tool from the UCSC utilities72, while the overall nucleotide and dinucleotide frequency across the promoter regions was quantified using the annotatePeaks.pl script included in Homer76. Core promoters were defined as the region 150 bp upstream and 50 bp downstream of the TSS for genes that had an annotated 5′UTR supported by RNA-seq and EST data34.
For the cumulative PAS and 5′SS motif identification, a position specific matrix for the vertebrate PAS consensus (AWUAAA) and the 5′SS consensus (Jaspar database – motif SD0001.1) were searched 500 bp downstream of all TSS or motifs of interest identified in the 400 bp around annotated TSS. The frequencies in adjacent windows of 10 bp were used to calculate the cumulative frequency in the corresponding strand. IUPAC consensus sequences were obtained by converting meme motifs to jaspar format using the meme-suite meme2jaspar followed by prediction of the consensus sequence using the web-version of rsa-tools convert-matrix77.
Additional Information
How to cite this article: Fernandez-Valverde, S. L. and Degnan, B. M. Bilaterian-like promoters in the highly compact Amphimedon queenslandica genome. Sci. Rep. 6, 22496; doi: 10.1038/srep22496 (2016).
References
Davidson, E. H. The Regulatory Genome. (Academic Press, 2010).
Carninci, P. et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nat Genet 38, 626–635 (2006).
Lenhard, B., Sandelin, A. & Carninci, P. Metazoan promoters: emerging characteristics and insights into transcriptional regulation. Nat Rev Genet 13, 233–245 (2012).
Levine, M. & Tjian, R. Transcription regulation and animal diversity. Nature 424, 147–151 (2003).
Degnan, B. M., Vervoort, M., Larroux, C. & Richards, G. S. Early evolution of metazoan transcription factors. Curr Opin Genet Dev 19, 591–599 (2009).
Sebé-Pedrós, A. et al. Early evolution of the T-box transcription factor family. Proc Natl Acad Sci USA 110, 16050–16055 (2013).
de Mendoza, A. et al. Transcription factor evolution in eukaryotes and the assembly of the regulatory toolkit in multicellular lineages. Proc Natl Acad Sci USA 110, E4858–66 (2013).
Larroux, C. et al. Genesis and expansion of metazoan transcription factor gene classes. Mol Biol Evol 25, 980–996 (2008).
Sebé-Pedrós, A. & de Mendoza, A. Transcription Factors and the Origin of Animal Multicellularity. Evolutionary Transitions to Multicellular Life (Springer, 2015).
Maduro, M. F. & Rothman, J. H. Making Worm Guts: The gene regulatory network of the Caenorhabditis elegans endoderm. Dev Biol 246, 68–85 (2002).
Smith, J., Theodoris, C. & Davidson, E. H. A gene regulatory network subcircuit drives a dynamic pattern of gene expression. Science 318, 794–797 (2007).
Rafiq, K., Shashikant, T., McManus, C. J. & Ettensohn, C. A. Genome-wide analysis of the skeletogenic gene regulatory network of sea urchins. Development 141, 950–961 (2014).
Wang, S., Sengel, C., Emerson, M. M. & Cepko, C. L. A gene regulatory network controls the binary fate decision of rod and bipolar cells in the vertebrate retina. Dev Cell 30, 513–527 (2014).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
modENCODE Consortium et al. Identification of functional elements and regulatory circuits by Drosophila modENCODE. Science 330, 1787–1797 (2010).
Negre, N. et al. A cis-regulatory map of the Drosophila genome. Nature 471, 527–531 (2011).
Plank, J. L. & Dean, A. Enhancer function: mechanistic and genome-wide insights come together. Mol Cell 55, 5–14 (2014).
Juven-Gershon, T. & Kadonaga, J. T. Regulation of gene expression via the core promoter and the basal transcriptional machinery. Dev Biol 339, 225–229 (2010).
Lee, D.-H. et al. Functional characterization of core promoter elements: the downstream core element is recognized by TAF1. Mol Cell Biol 25, 9674–9686 (2005).
Zabidi, M. A. et al. Enhancer-core-promoter specificity separates developmental and housekeeping gene regulation. Nature 518, 556–559 (2015).
Hendrix, D. A., Hong, J.-W., Zeitlinger, J., Rokhsar, D. S. & Levine, M. S. Promoter elements associated with RNA Pol II stalling in the Drosophila embryo. Proc Natl Acad Sci USA 105, 7762–7767 (2008).
Ohler, U. & Wassarman, D. A. Promoting developmental transcription. Development 137, 15–26 (2009).
Zehavi, Y., Sloutskin, A., Kuznetsov, O. & Juven-Gershon, T. The core promoter composition establishes a new dimension in developmental gene networks. Nucleus (Austin, Tex) 5, 298–303 (2014).
Wasylyk, B. et al. Specific in vitro transcription of conalbumin gene is drastically decreased by single-point mutation in T-A-T-A box homology sequence. Proc Natl Acad Sci USA 77, 7024–7028 (1980).
Yang, C., Bolotin, E., Jiang, T., Sladek, F. M. & Martinez, E. Prevalence of the initiator over the TATA box in human and yeast genes and identification of DNA motifs enriched in human TATA-less core promoters. Gene 389, 52–65 (2007).
van Heeringen, S. J. et al. Nucleotide composition-linked divergence of vertebrate core promoter architecture. Genome Res 21, 410–421 (2011).
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005).
Buratowski, S., Hahn, S., Guarente, L. & Sharp, P. A. Five intermediate complexes in transcription initiation by RNA polymerase II. Cell 56, 549–561 (1989).
Gekakis, N. Role of the CLOCK Protein in the Mammalian Circadian Mechanism. Science 280, 1564–1569 (1998).
FitzGerald, P. C., Sturgill, D., Shyakhtenko, A., Oliver, B. & Vinson, C. Comparative genomics of Drosophila and human core promoters. Genome Biol 7, R53 (2006).
Matsukage, A., Hirose, F., Hayashi, Y., Hamada, K. & Yamaguchi, M. The DRE sequence TATCGATA, a putative promoter-activating element for Drosophila melanogaster cell-proliferation-related genes. Gene 166, 233–236 (1995).
Grishkevich, V., Hashimshony, T. & Yanai, I. Core promoter T-blocks correlate with gene expression levels in C. elegans . Genome Res 21, 707–717 (2011).
Smale, S. T. & Kadonaga, J. T. The RNA polymerase II core promoter. Annu Rev Biochem 72, 449–479 (2003).
Fernandez-Valverde, S. L., Calcino, A. D. & Degnan, B. M. Deep developmental transcriptome sequencing uncovers numerous new genes and enhances gene annotation in the sponge Amphimedon queenslandica . BMC Genomics 16, 387 (2015).
Seo, H. C. et al. Miniature genome in the marine chordate Oikopleura dioica . Science 294, 2506 (2001).
Xu, C., Chen, J. & Shen, B. The preservation of bidirectional promoter architecture in eukaryotes: what is the driving force? BMC Syst Biol 6 Suppl 1, S21–S21 (2012).
Orekhova, A. S. & Rubtsov, P. M. Bidirectional promoters in the transcription of mammalian genomes. Biochemistry (Mosc) 78, 335–341 (2013).
Kong, Q. et al. Regulatory switch enforced by basic helix-loop-helix and ACT-domain mediated dimerizations of the maize transcription factor R. Proc Natl Acad Sci USA 109, E2091–7 (2012).
Mankad, R. V., Gimelbrant, A. A. & McClintock, T. S. Consensus translational initiation sites of marine invertebrate phyla. Biol. Bull. 195, 251–254 (1998).
Whitfield, T. W. et al. Functional analysis of transcription factor binding sites in human promoters. Genome Biol 13, R50 (2012).
Ohler, U., Liao, G.-C., Niemann, H. & Rubin, G. M. Computational analysis of core promoters in the Drosophila genome. Genome Biol 3, RESEARCH0087 (2002).
Siebert, M. & Söding, J. Universality of core promoter elements? Nature 511, E11–E12 (2014).
Almada, A. E., Wu, X., Kriz, A. J., Burge, C. B. & Sharp, P. A. Promoter directionality is controlled by U1 snRNP and polyadenylation signals. Nature 499, 360–363 (2013).
Ntini, E. et al. Polyadenylation site-induced decay of upstream transcripts enforces promoter directionality. Nat Struct Mol Biol 20, 923–928 (2013).
Wu, X. & Sharp, P. A. Divergent transcription: a driving force for new gene origination? Cell 155, 990–996 (2013).
de Mendoza, A., Suga, H., Permanyer, J., Irimia, M. & Ruiz-Trillo, I. Complex transcriptional regulation and independent evolution of fungal-like traits in a relative of animals. eLife 4, e08904 (2015).
Davidson, E. H. & Erwin, D. H. Gene regulatory networks and the evolution of animal body plans. Science 311, 796–800 (2006).
Nelson, C. E., Hersh, B. M. & Carroll, S. B. The regulatory content of intergenic DNA shapes genome architecture. Genome Biol 5, R25 (2004).
Li, C., Chen, J. & Hua, T. Precambrian sponges with cellular structures. Science 279, 879–882 (1998).
Moroz, L. L. et al. The ctenophore genome and the evolutionary origins of neural systems. Nature 510, 109–114 (2014).
Ryan, J. F. et al. The genome of the ctenophore Mnemiopsis leidyi and its implications for cell type evolution. Science 342, 1242592 (2013).
Long, H. K. et al. Epigenetic conservation at gene regulatory elements revealed by non-methylated. DNA profiling in seven vertebrates. eLife 2, e00348 (2013).
Cohen, N. M., Kenigsberg, E. & Tanay, A. Primate CpG islands are maintained by heterogeneous evolutionary regimes involving minimal selection. Cell 145, 773–786 (2011).
Deaton, A. M. & Bird, A. CpG islands and the regulation of transcription. Genes Dev 25, 1010–1022 (2011).
Philipsen, S. & Suske, G. A tale of three fingers: the family of mammalian Sp/XKLF transcription factors. Nucleic Acids Res 27, 2991–3000 (1999).
Miller, I. J. & Bieker, J. J. A novel, erythroid cell-specific murine transcription factor that binds to the CACCC element and is related to the Krüppel family of nuclear proteins. Mol Cell Biol 13, 2776–2786 (1993).
Schaeper, N. D., Prpic, N.-M. & Wimmer, E. A. A clustered set of three Sp-family genes is ancestral in the Metazoa: evidence from sequence analysis, protein domain structure, developmental expression patterns and chromosomal location. BMC Evol Biol 10, 88 (2010).
Shimeld, S. M. C2H2 zinc finger genes of the Gli, Zic, KLF, SP, Wilms’ tumour, Huckebein, Snail, Ovo, Spalt, Odd, Blimp-1, Fez and related gene families from Branchiostoma floridae . Dev Genes Evol 218, 639–649 (2008).
Presnell, J. S., Schnitzler, C. E. & Browne, W. E. KLF/SP transcription factor family evolution: expansion, diversification, and innovation in eukaryotes. Genome Biol Evol 7, 2289–2309 (2015).
Merika, M. & Orkin, S. H. Functional synergy and physical interactions of the erythroid transcription factor GATA-1 with the Krüppel family proteins Sp1 and EKLF. Mol Cell Biol 15, 2437–2447 (1995).
Pugh, B. F. & Tjian, R. Mechanism of transcriptional activation by Sp1: evidence for coactivators. Cell 61, 1187–1197 (1990).
Ryu, S., Zhou, S., Ladurner, A. G. & Tjian, R. The transcriptional cofactor complex CRSP is required for activity of the enhancer-binding protein Sp1. Nature 397, 446–450 (1999).
Schmidt, M. C., Zhou, Q. & Berk, A. J. Sp1 activates transcription without enhancing DNA-binding activity of the TATA box factor. Mol Cell Biol 9, 3299–3307 (1989).
Perina, D. et al. Over-represented localized sequence motifs in ribosomal protein gene promoters of basal metazoans. Genomics 98, 56–63 (2011).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Civán, P. & Svec, M. Genome-wide analysis of rice (Oryza sativa L. subsp. japonica) TATA box and Y Patch promoter elements. Genome 52, 294–297 (2009).
Choi, K. et al. Arabidopsis meiotic crossover hot spots overlap with H2A.Z nucleosomes at gene promoters. Nat Genet 45, 1327–1336 (2013).
Andersson, R. et al. Nuclear stability and transcriptional directionality separate functionally distinct RNA species. Nature Commun 5, 5336 (2014).
Uesaka, M. et al. Bidirectional promoters are the major source of gene activation-associated non-coding RNAs in mammals. BMC Genomics 15, 35 (2014).
Gaiti, F. et al. Dynamic and widespread lncRNA expression in a sponge and the origin of animal complexity. Mol Biol Evol 32, 2367–2382 (2015).
Srivastava, M. et al. The Amphimedon queenslandica genome and the evolution of animal complexity. Nature 466, 720–726 (2010).
Kuhn, R. M., Haussler, D. & Kent, W. J. The UCSC genome browser and associated tools. Brief Bioinform 14, 144–161 (2013).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol 2, 28–36 (1994).
Sandelin, A., Alkema, W., Engström, P., Wasserman, W. W. & Lenhard, B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res 32, D91–4 (2004).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell 38, 576–589 (2010).
Thomas-Chollier, M. et al. RSAT 2011: regulatory sequence analysis tools. Nucleic Acids Res 39, W86–W91 (2011).
Acknowledgements
This work was supported by an Australian Research Council grant FL110100044 to BMD.
Author information
Authors and Affiliations
Contributions
B.M.D. and S.L.F.V. conceived and designed the study. S.L.F.V. carried out all comparative and Amphimedon promoter characterization analyses. S.L.F.V. and B.M.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Fernandez-Valverde, S., Degnan, B. Bilaterian-like promoters in the highly compact Amphimedon queenslandica genome. Sci Rep 6, 22496 (2016). https://doi.org/10.1038/srep22496
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep22496
This article is cited by
-
Tracing animal genomic evolution with the chromosomal-level assembly of the freshwater sponge Ephydatia muelleri
Nature Communications (2020)
-
Convergent evolution of a vertebrate-like methylome in a marine sponge
Nature Ecology & Evolution (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.